Abstract
Retinoblastoma is an aggressive childhood cancer of the developing retina that is initiated by the biallelic loss of RB1. Tumours progress very quickly following RB1 inactivation but the underlying mechanism is not known. Here we show that the retinoblastoma genome is stable, but that multiple cancer pathways can be epigenetically deregulated. To identify the mutations that cooperate with RB1 loss, we performed whole-genome sequencing of retinoblastomas. The overall mutational rate was very low; RB1 was the only known cancer gene mutated. We then evaluated the role of RB1 in genome stability and considered non-genetic mechanisms of cancer pathway deregulation. For example, the proto-oncogene SYK is upregulated in retinoblastoma and is required for tumour cell survival. Targeting SYK with a small-molecule inhibitor induced retinoblastoma tumour cell death in vitro and in vivo. Thus, retinoblastomas may develop quickly as a result of the epigenetic deregulation of key cancer pathways as a direct or indirect result of RB1 loss.
Similar content being viewed by others
Main
Retinoblastoma is a rare childhood cancer of the retina that can develop in a sporadic or a heritable form and is fatal if untreated. When the RB1 gene was first cloned, it was found to undergo biallelic inactivation in virtually all retinoblastoma tumours1. Since then, hundreds of genetic lesions have been identified in human cancer. These genetic lesions can be grouped on the basis of the signalling pathways they affect that have direct or indirect mechanistic links to many of the common cellular properties, or hallmarks, of cancer. Thus, the rate of cancer progression is related to the kinetics of acquisition of multiple genetic lesions and/or epigenetic changes that ultimately lead to activation of growth-signalling pathways, evasion of cell death and senescence, acquisition of limitless replicative potential, sustained angiogenesis, and local tissue invasion and metastasis2.
RB1 inactivation confers limitless replicative potential to retinoblasts and is rate limiting for retinoblastoma tumorigenesis3. However, the mechanisms that enable retinoblastoma cells to acquire the additional hallmarks of cancer remain unknown. Evidence from molecular, cellular and cytogenetic studies suggest that RB1 is required for maintaining chromosomal stability4,5, and that its loss leads to chromosome instability in cells maintained in culture. These data raise the possibility that RB1 inactivation may underlie the rapid acquisition of cooperating mutations in key cancer pathways through chromosome instability. Alternatively, epigenetic changes may have a more dominant role in cooperating with the loss of RB1 retinoblastoma tumorigenesis. RB1 has been implicated in regulating most major epigenetic processes, including microRNA regulation, DNA methylation, histone modification and ATP-dependent chromatin reorganization6,7,8,9,10. Thus, inactivation of RB1 in retinoblasts may lead to the rapid epigenetic deregulation of cancer genes that contribute to the essential cellular properties of retinoblastoma.
In this study, the St Jude Children’s Research Hospital – Washington University Pediatric Cancer Genome Project characterized the genetic landscape of retinoblastoma. Whole-genome sequencing (WGS) of four retinoblastomas and their paired germline DNA samples showed no genetic lesions in known tumour suppressor genes or oncogenes, other than RB1 and MYCN. More importantly, an orthotopic xenograft derived from one of the primary tumours showed no evidence of clonal variation or new coding-region mutations. This finding suggests that the retinoblastoma genome is more stable than previously believed.
Unlike the genetic landscape of retinoblastoma, the epigenetic profile shows profound changes relative to that observed in normal retinoblasts. One of the most striking results was the induction of the expression of the proto-oncogene spleen tyrosine kinase (SYK) in human retinoblastoma. SYK is required for tumour cell survival, and inhibition of SYK with a small-molecule inhibitor caused the degradation of MCL1 and caspase-mediated cell death in retinoblastoma cells in culture and in vivo. These findings highlight how comprehensive genetic and epigenetic analyses of tumours can be integrated, leading to the discovery of promising new therapeutic approaches and shedding light on the mechanisms underlying the rapid progression of retinoblastoma following RB1 inactivation.
Retinoblastoma whole-genome sequencing
We performed WGS analysis on four primary human retinoblastoma samples (Supplementary Information, section 1, and Supplementary Table 1) and on matched normal tissue. Local tumour cell invasion, but not metastasis, was evident in each patient (Fig. 1a–c and Supplementary Fig. 1). We generated an orthotopic xenograft, SJRB001X, of the primary tumour SJRB001 by inoculating primary tumour cells into the vitreous humour of the eyes of immunocompromised mice (Supplementary Information, section 2). SJRB001X had molecular, genetic and histopathologic features similar to those of SJRB001 (Fig. 1d–f; Supplementary Figs 2–4; Supplementary Tables 2–4; and Supplementary Information, section 3).
a–c, Representative retinoblastoma tumour section (SJRB001) stained with haematoxylin and eosin (H&E), showing choroidal and optic nerve invasion (arrow). d–f, H&E-stained section of the SJRB001X orthotopic xenograft with choroidal (e) and optic nerve (f) invasion (arrows). AC, anterior chamber; ON, optic nerve; Sc, sclera. Scale bars, 25 μm.
Using a paired-end sequencing approach, we generated 1,040.9 gigabase pairs (Gb) of sequence data for the samples described; 956.8 Gb (92%) was successfully mapped to the NCBI 36.1 reference genome (Supplementary Information, section 4, and Supplementary Table 5). The average genome coverage was ×28.9, and the average exon coverage was ×23.8 with 98.4% of single nucleotide polymorphisms (SNPs) detected across all nine genomes showing concordance with their corresponding SNP array genotype calls at the same genomic positions (Supplementary Table 5). To provide additional sequence coverage, we performed transcriptome sequencing of all four primary tumours (Supplementary Information, section 5, and Supplementary Table 6).
We identified 668 validated somatic sequence mutations and 40 structural variations across the four cases (Table 1). These included 23 tier-1 mutations in genes, 35 tier-2 mutations in evolutionarily conserved regions of the genome (Supplementary Information, section 6), 309 tier-3 mutations in non-repetitive regions of the genome that are not part of tiers 1 and 2, and 301 tier-4 mutations in repetitive sequences in the genome (Table 1 and Supplementary Table 7). The average number of sequence mutations was 167 per case (range, 56–258), with only 3.25 mutations per case (range, 0–5) resulting in amino-acid changes (Table 1). The estimated mean mutation rate was 6.7 × 10−8 per base (range, 1.03 × 10−7–2.17 × 10−8), which is 15-fold less than that in adult tumours analysed by WGS, except for acute myeloid leukaemia11. The predominant changes were C → A and G → T transversions (Supplementary Fig. 5), which is consistent with the possibility that some of the transversions result from production of 8-oxoguanine during oxidative stress. Tumour SJRB002 had no somatic sequence variations that resulted in amino-acid changes; the only structural variations were the loss of heterozygosity (LOH) at the RB1 locus on chromosome 13 and a gain of chromosome 6p. This suggests that very few genetic lesions are required for retinoblastoma progression after RB1 inactivation.
RB1 inactivation in retinoblastoma
Both RB1 alleles were inactivated in each sample (Supplementary Figs 6 and 7). Tumours SJRB002 and SJRB003 had mutations in RB1 combined with copy-number-neutral LOH, and SJRB001 and SJRB004 had somatic sequence mutations combined with RB1-promoter hypermethylation (Supplementary Figs 6–8). Deep-sequence analysis of the germline sample from SJRB002 revealed that about 10% of reads contained the R445 nonsense mutation, suggesting the presence of germline chimaerism for the RB1 mutation (Supplementary Fig. 6). Combining the WGS data with SNP array data of an additional 42 samples, we found that tumours from patients with lower regional nucleotide diversity were much less likely to undergo LOH at the RB1 locus (Supplementary Tables 8 and 9 and Supplementary Information, section 7). These data show a significant association (P = 8 × 10−8, Fisher’s exact test) between a germline genetic variation and mechanism of biallelic RB1 inactivation in retinoblastoma.
Recurrent lesions in retinoblastoma
To determine whether any of the 11 genes with somatic mutations that caused amino-acid changes or a frameshift in the coding region (Table 1; Fig. 2a, b; Supplementary Figs 9 and 10; Supplementary Information, section 8; and Supplementary Table 10) were recurrently mutated in retinoblastoma, we sequenced all exons from the 11 genes in our recurrent screening cohort of 42 retinoblastomas (Supplementary Information, section 4). Only BCOR was recurrently mutated in retinoblastoma (6 of 46, or 13%). Five of the samples had BCOR mutations that resulted in truncation of the encoded protein, and one sample had a focal gene deletion (Fig. 2c, Supplementary Table 11 and Supplementary Fig. 11).
a, b, CIRCOS plots of genetic alterations in two retinoblastomas and the matched orthotopic xenograft. Loss of heterozygosity (orange), amplifications (red) and deletions (blue) are shown. Interchromosomal translocations (green lines) and intrachromosomal translocations (purple lines) are indicated. Sequence mutations in NCBI RefSeq genes included silent single nucleotide variants (SNVs; green), missense SNVs (brown), nonsense SNVs (dark blue), splice-site mutations (pink) and insertion/deletion mutations (indels, red). *Germline mosaic. c, BCOR mutations identified in the discovery and recurrency cohort.
We also used the WGS data to identify somatic structural variations including whole-chromosome gains and losses, focal deletions (DEL), insertions (INS), inversions (INV), intrachromosomal rearrangements (ITX), interchromosomal rearrangements (CTX) and regions of LOH (Fig. 2a–c; Supplementary Information, section 9; Supplementary Table 12; and Supplementary Fig. 12). The average number of structural variations was ten per case (range, 0–24) (Supplementary Table 1). Tumour SJRB001 had four structural variations (two DEL and ins INS) including a gain of a region of chromosome 2 spanning MYCN (Supplementary Table 12), and the only chromosomal lesion in SJRB002 was a gain of chromosome 6p, which occurs in about 40% of human retinoblastomas12 (Fig. 2b). Only a few genomic regions were affected by the structural variations in SJRB003 and SJRB004 (Supplementary Figs 10, 13 and 14 and Supplementary Table 12).
Orthotopic retinoblastoma xenograft
The genomic landscape of the orthotopic xenograft was remarkably similar to that of the primary tumour, despite continuous growth and multiple passages in vivo over nine months (Fig. 2 and Supplementary Table 1). All of the sequence mutations and structural variations detected in SJRB001 were retained in SJRB001X. Only 67 new SNVs and four structural variations were identified in the xenograft, and none affected annotated genes (Fig. 2 and Supplementary Table 1). Moreover, each mutation was identified at a subclonal level (range, 20–30%) and the mutant allele frequency for the lesions in tiers 1–4 in SJRB001 was retained in SJRB001X (Supplementary Fig. 15).
This result was surprising because several studies in mice and cell cultures have linked RB1 inactivation to defects in chromosome segregation that result in aneuploidy4,5,13,14,15 and chromosome instability4. We measured the distance between sister chromatids, the distance between kinetochores and the proportion of lagging chromatids in two RB1-deficient human retinoblastoma orthotopic xenografts16 (SJRB001X and SJRB002X). Consistent with results from RB1-deficient retinal pigmented epithelium cells4, the distances between sister chromatids and between kinetochores were increased, and there was evidence of lagging chromosomes (Supplementary Figs 16 and 17). However, less variation in ploidy was observed during spectral karyotype analysis of SJRB001X and SJRB002X, which was more consistent with the ploidy of wild-type cells (Fig. 3a, b and Supplementary Table 13). Moreover, copy number variations were much lower in our cohort of 46 retinoblastomas than in tumours with known genome instability such as ovarian cancer (Fig. 3c). Together, the cytogenetic data and WGS data suggest that the genome is stable and that newly acquired lesions do not provide a selective growth advantage and are thus probably passenger mutations (Supplementary Information, section 9).
a, Chromosomal mis-segregation of SJRB001X cells after at least 21 rounds of cell division is plotted in red. RPE, retinal pigment epithelium. b, Representative spectral karyotype image of SJRB001X after the third passage in mice. c, Alterations in the 46 retinoblastoma (Rb) cases relative to 153 high-grade serous ovarian cancer (Ov) cases from The Cancer Genome Atlas. The median percentage of the genome involved in copy number variations (CNVs) was 1.5% for retinoblastoma and 27.7% for ovarian cancer. Red circles, samples used for WGS.
Identifying deregulated cancer pathways
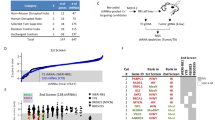
There are many examples over the past several decades of epigenomic changes such as DNA methylation contributing to tumorigenesis17,18,19. Indeed, a recent study demonstrated changes in DNA methylation in Wilms’ tumours20, which tend, like retinoblastomas, to have stable genomes. To explore whether epigenetic deregulation of genes or pathways promotes tumorigenesis in retinoblastoma, we carried out an integrative analysis of chromatin immunoprecipitation (ChIP) data, DNA methylation data and gene expression data using order statistics. The SJRB001X sample was used for ChIP assay (Supplementary Figs 18–20), and primary tumour and xenograft samples were used for both DNA methylation and gene expression assays. In all three analyses, experimental results in retinoblastoma tumours were compared with those from human fetal retinae. In this comparison, a total of 104 genes, including 15 known cancer genes (Fig. 4a and Supplementary Tables 14 and 15), were found to have significant differences, indicating that several key cancer genes were epigenetically deregulated.
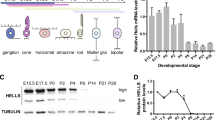
a, Whole-genome view of the gene ranks based on integrating ChIP-on-chip, methylation and gene expression results. The y axis shows −log(P), where P is the P value of the Q statistic corrected for multiple testing. Significantly (false discovery rate, ≤10%) downregulated (green) or upregulated (red) genes are shown. b, c, ChIP validation of histone markers of the SYK promoter including quantification by quantitative PCR with TaqMan probes. In c, each bar shows mean ± s.d. of triplicate samples. d, SYK expression measured by quantitative PCR in fetal-week-20 retina (fetal), primary retinoblastoma (tumour), orthotopic xenografts (SJRB001X and SJRB002X) and cell lines. Each bar is mean ± s.d. of duplicate samples normalized to GPI1 expression. e, Immunoblot of SYK (green) and actin (red) in orthotopic xenografts, human fetal retina and representative cell lines; black and white representation of the SYK immunoblot is in the lower panel. f, H&E (purple) and anti-SYK (brown) immunohistochemistry of retinoblastoma tissue. g, Immunoprecipitation analysis of SYK and pSYK Y525/526 from Weri1 retinoblastoma cells. h, Viability was measured in triplicate cultures 72 h after infection of retinoblastoma cells with a lentivirus vector expressing either a control lentivirus or an shRNA against SYK. Scale bars, 10 μm.
SYK is a novel therapeutic target
SYK is the fifth most significant gene identified by the integrative analysis and the only upregulated kinase gene (Supplementary Table 15 and Fig. 4a). SYK is expressed throughout the haematopoietic system, regulates immunomodulatory signalling and has been implicated in several haematologic malignancies21,22,23,24. Small-molecule inhibitors of SYK have been developed to treat autoimmune disorders25, and two of those agents, BAY 61-3606 and R406, have shown efficacy in preclinical leukaemia studies26,27,28.
ChIP-on-chip analysis showed increased activating histone modifications (H3K4me3 and K3K9/14Ac) at SYK’s promoter, and the repressive histone marker (H3k9me3) was unchanged. Binding of RNA polymerase II to the SYK promoter was also increased (Fig. 4b and Supplementary Tables 14 and 15). These ChIP-on-chip results were validated in independent samples by real-time PCR analysis with reverse transcription (Fig. 4c), and we confirmed the increase in SYK gene expression (Fig. 4d). SYK protein levels were higher in human retinoblastoma orthotopic xenografts and cell lines than in human fetal retinae (Fig. 4e). To determine whether SYK is expressed in primary human retinoblastomas, we performed immunohistochemistry on a retinoblastoma tissue microarray or whole-eye sections. In total, 100% (82 of 82) showed very strong expression (3+) of SYK in all tumour cells; SYK was not expressed in normal retina (Fig. 4f). SYK’s kinase activity is regulated through autophosphorylation at the Tyr 525 and Tyr 526 residues within its catalytic domain. These sites were phosphorylated in retinoblastoma cells, and this phosphorylation was reversed by exposure to BAY 61-3606 or R406 (Fig. 4g and data not shown).
To determine whether SYK expression is required for retinoblastoma growth, survival or both, we generated a short hairpin RNA (shRNA) against SYK and cloned it into the lentiviral vector Lenti-SYK-9. Lenti-SYK-9 efficiently knocked down SYK in retinoblastoma cell lines (Supplementary Fig. 21) and drastically increased apoptosis in retinoblastoma cells (Fig. 4h and Supplementary Fig. 21). Similar results were obtained in vivo using SJRB001X (data not shown). We used an empty lentiviral vector and a lentiviral vector encoding an SYK shRNA that less effectively reduced SYK expression (Lenti-SYK-6) as controls. Cell lines that do not express SYK (BJ, 293T and uninduced Jurkat cells) were used as controls and the Lenti-SYK-9 lentivirus had no effect on the growth or apoptosis of the control cells.
We exposed retinoblastoma cell lines that express high levels of SYK (Weri1 and RB355) to various concentrations of the SYK inhibitors BAY 61-3606 and R406 for 72 h and then measured cell viability. Jurkat (uninduced) and 293T cells were used as negative controls. Weri1 and RB355 cells were sensitive to both SYK inhibitors, but the Jurkat and 293T cells were unaffected (Fig. 5a). Transmission electron microscopy of retinoblastoma cells treated with the SYK inhibitors showed morphologic features consistent with cell death and mitochondrial defects (Supplementary Fig. 22); this was confirmed by scoring the proportion of activated caspase 3+ cells (Fig. 5b, c and Supplementary Fig. 22). Jurkat cells showed no increase in activated caspase 3+ cells after treatment with 10 μM R406 or BAY 61-3606 (data not shown).
a, Dose response for SYK inhibitors R406 (red) and BAY 61-3606 (black) in RB355 retinoblastoma cells and a negative control (Jurkat). Each data point is mean ± s.d. of triplicate samples. b–e, Immunofluorescence of activated caspase 3 (b, c) or EdU (d, e) (red) before and after treatment of RB355 cells with R406 or BAY 61-3606. A total of 250 cells were scored in duplicate for each sample and each treatment condition to derive the mean and s.d. Nuclei were stained with 4′,6-diamidino-2-phenylindole (DAPI, blue). f, Treatment of stimulated Jurkat or RB355 cells with 5 μM BAY 61-3606 for 24 h reduced MCL1 expression. g, Schematic of the treatment schedule for mice with SJRB001X tumours. h, Representative magnetic resonance images of a mouse whose tumour responded after four courses of treatment with BAY 61-3606 (left) and another whose disease progressed during treatment (right). i, Survival curves show that treatment with BAY 61-3606 plus TPT improved outcome. j, Immunostaining for activated caspase 3 (arrows) in eyes either untreated or treated with BAY 61-3606. k, Immunoblot showing reduction in MCL1 after treatment with BAY 61-3606 (BAY) or BAY 61-3606 plus TPT. C, control. Scale bars, 5 μm (b, d); 10 μm (j).
The proportions of cells from each line that incorporated EdU were similar, suggesting that retinoblastoma’s cell cycle is not affected by SYK inhibition (Fig. 5d, e and Supplementary Fig. 22). The effects of targeting SYK were partly rescued by the pan-caspase inhibitor Q-VD-OPH (Supplementary Fig. 22). Less MitoTracker Red accumulated in retinoblastoma cells exposed to the SYK inhibitors (Supplementary Fig. 22), yet treatment of Jurkat cells with either inhibitor had no effect on MitoTracker Red accumulation (data not shown). Inhibiting SYK in B-CLL cells reduced their autophosphorylation of SYK and destabilized MCL126,29, the only anti-apoptotic member of the BCL2 family that is upregulated in retinoblastoma (data not shown). Both inhibitors reduced autophosphorylation of SYK on Y525/526 (Fig. 4g) and reduced MCL1 expression coincident with apoptosis (Fig. 5f).
We tested the efficacy of BAY 61-3606 in vivo using our SJRB001X model30. The chemotherapy regimen consisted of a single subconjunctival dose of BAY 61-3606 on day 1 and daily doses of topotecan (TPT) on days 1–5 until either six courses (21 days per course) were administered or the tumour had progressed and surgical enucleation was required (Fig. 5g, h). BAY 61-3606 plus TPT significantly improved the outcome (P = 0.003; Fig. 5i), and its efficacy was correlated with an increase in activated caspase 3+ cells in the treated eyes (Fig. 5j). Previous studies using this model have shown that TPT combined with subconjunctival carboplatin had no effect on tumour response or outcome30, so the improvement seen here can be attributed to targeting SYK. MCL1 expression was also reduced in the treated eyes, which is consistent with increased apoptosis and targeting of SYK in vivo (Fig. 5k). Together, these results suggest that SYK is a promising new target for treating retinoblastoma.
Discussion
Genome sequencing reveals that retinoblastomas have a relatively stable genome. The mutational rate and number of structural variations per case that we assessed were among the lowest reported in human cancer to date. Moreover, in one example (SJRB002) the only non-silent mutation found was in RB1, and only two structural variations were detected. The minimal increase in passenger mutations in SJRB001X cells, despite prolonged passage, was also consistent with a relatively stable genome. These results are surprising because previous studies have shown that the functional inactivation of RB1 can cause genomic instability. Although retinoblastoma cells show the mitotic defects that can lead to errors in chromosome segregation, the ploidy of retinoblastoma cells is stable in vitro and in vivo31,32. We do not exclude the possibility that RB1 inactivation promotes change in other contexts, but our data show that genomic instability is not a hallmark of retinoblastoma and do not explain how retinoblastomas progress so rapidly.
We propose that epigenetic mechanisms contribute to retinoblastoma tumorigenesis. We identified several known oncogenes and tumour suppressor genes with histone modifications and altered DNA methylation that correlated with changes in gene expression. Our key discovery was that SYK is important in retinoblastoma. Retinal progenitor cells and retinal neurons express little or no SYK, and SYK has no known function in the developing visual system. Moreover, no recurrent genetic lesions in SYK were identified by WGS or SNP array analysis to suggest that this gene drives retinoblastoma tumorigenesis. Only by integrating epigenetic and gene expression analyses did we identify SYK as an important oncogene in retinoblastoma. This is important not only for expanding our understanding of the biology of retinoblastoma but also for advancing immediate therapeutic options that were not previously considered, such as the use of BAY 61-3606 or R406. This study highlights the value of integrating WGS analyses of the genetic and epigenetic features of tumour genomes to finding a cure for cancers such as retinoblastoma.
Methods Summary
Full details of sample acquisition, molecular and biochemical procedures, informatics and whole genome sequencing, and animal and drug studies are provided in Supplementary Information. The St Jude Children’s Research Hospital Institutional Review Board approved experiments involving human subjects and informed consent was obtained from all subjects. For animal studies, all experiments were performed in accordance with federal guidelines and regulations. The St Jude Children’s Research Hospital Institutional Animal Care and Use Committee approved all animal experiments. Lentiviral vectors (GIPZ with Lenti-SYK-9 #V3LHS-366147 and Lenti-SYK-6 #V3LHS-366143) encoding shRNAs against SYK were purchased from OpenBiosystems.
Accession codes
Data deposits
The whole-genome sequence data and SNP 6.0 data have been deposited in the NCBI dbGaP under accession number phs000352.v1.p1.
References
Friend, S. H. et al. A human DNA segment with properties of the gene that predisposes to retinoblastoma and osteosarcoma. Nature 323, 643–646 (1986)
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011)
Knudson, A. Mutation and cancer: statistical study of retinoblastoma. Proc. Natl Acad. Sci. USA 68, 820–823 (1971)
Manning, A. L., Longworth, M. S. & Dyson, N. J. Loss of pRB causes centromere dysfunction and chromosomal instability. Genes Dev. 24, 1364–1376 (2010)
Hernando, E. et al. Rb inactivation promotes genomic instability by uncoupling cell cycle progression from mitotic control. Nature 430, 797–802 (2004)
Chi, P., Allis, C. D. & Wang, G. G. Covalent histone modifications – miswritten, misinterpreted and mis-erased in human cancers. Nature Rev. Cancer 10, 457–469 (2010)
Lu, J., Ruhf, M. L., Perrimon, N. & Leder, P. A genome-wide RNA interference screen identifies putative chromatin regulators essential for E2F repression. Proc. Natl Acad. Sci. USA 104, 9381–9386 (2007)
Benetti, R. et al. A mammalian microRNA cluster controls DNA methylation and telomere recombination via Rbl2-dependent regulation of DNA methyltransferases. Nature Struct. Mol. Biol. 15, 998 (2008)
Wen, H., Andrejka, L., Ashton, J., Karess, R. & Lipsick, J. S. Epigenetic regulation of gene expression by Drosophila Myb and E2F2-RBF via the Myb-MuvB/dREAM complex. Genes Dev. 22, 601–614 (2008)
Bourgo, R. J. et al. SWI/SNF deficiency results in aberrant chromatin organization, mitotic failure, and diminished proliferative capacity. Mol. Biol. Cell 20, 3192–3199 (2009)
Ley, T. J. et al. DNA sequencing of a cytogenetically normal acute myeloid leukaemia genome. Nature 456, 66–72 (2008)
Corson, T. W. & Gallie, B. L. One hit, two hits, three hits, more? Genomic changes in the development of retinoblastoma. Genes Chromosom. Cancer 46, 617–634 (2007)
Ganem, N. J., Godinho, S. A. & Pellman, D. A mechanism linking extra centrosomes to chromosomal instability. Nature 460, 278–282 (2009)
Iovino, F., Lentini, L. & Amato, A. &. Di Leonardo, A. RB acute loss induces centrosome amplification and aneuploidy in murine primary fibroblasts. Mol. Cancer 5, 38 (2006)
Amato, A., Lentini, L., Schillaci, T., Iovino, F. & Di Leonardo, A. RNAi mediated acute depletion of retinoblastoma protein (pRb) promotes aneuploidy in human primary cells via micronuclei formation. BMC Cell Biol. 10, 79 (2009)
McEvoy, J. et al. Coexpression of normally incompatible developmental pathways in retinoblastoma genesis. Cancer Cell 16, 260–275 (2011)
Feinberg, A. P. & Tycko, B. The history of cancer epigenetics. Nature Rev. Cancer 4, 143–153 (2004)
Jones, P. A. & Laird, P. W. Cancer epigenetics comes of age. Nature Genet. 21, 163–167 (1999)
Laird, P. W. Cancer epigenetics. Hum. Mol. Genet. 14, (suppl. 1)R65–R76 (2005)
Hansen, K. D. et al. Increased methylation variation in epigenetic domains across cancer types. Nature Genet. 43, 768–775 (2011)
Hahn, C. K. et al. Proteomic and genetic approaches identify Syk as an AML target. Cancer Cell 16, 281–294 (2009)
Chen, L. et al. SYK-dependent tonic B-cell receptor signaling is a rational treatment target in diffuse large B-cell lymphoma. Blood 111, 2230–2237 (2008)
Feldman, A. L. et al. Overexpression of Syk tyrosine kinase in peripheral T-cell lymphomas. Leukemia 22, 1139–1143 (2008)
Young, R. M. et al. Mouse models of non-Hodgkin lymphoma reveal Syk as an important therapeutic target. Blood 113, 2508–2516 (2009)
Weinblatt, M. E. et al. An oral spleen tyrosine kinase (Syk) inhibitor for rheumatoid arthritis. N. Engl. J. Med. 363, 1303–1312 (2010)
Baudot, A. D. et al. The tyrosine kinase Syk regulates the survival of chronic lymphocytic leukemia B cells through PKCδ and proteasome-dependent regulation of Mcl-1 expression. Oncogene 28, 3261–3273 (2009)
Suljagic, M. et al. The Syk inhibitor fostamatinib disodium (R788) inhibits tumor growth in the Eμ-TCL1 transgenic mouse model of CLL by blocking antigen-dependent B-cell receptor signaling. Blood 116, 4894–4905 (2010)
Buchner, M. et al. Spleen tyrosine kinase inhibition prevents chemokine- and integrin-mediated stromal protective effects in chronic lymphocytic leukemia. Blood 115, 4497–4506 (2010)
Gobessi, S. et al. Inhibition of constitutive and BCR-induced Syk activation downregulates Mcl-1 and induces apoptosis in chronic lymphocytic leukemia B cells. Leukemia 23, 686–697 (2009)
Brennan, R. C. et al. Targeting the p53 pathway in retinoblastoma with subconjunctival Nutlin-3a. Cancer Res. 71, 4205–4213 (2011)
Yan, Y. et al. Engraftment and growth of patient-derived retinoblastoma tumour in severe combined immunodeficiency mice. Eur. J. Cancer 36, 221–228 (2000)
Squire, J., Gallie, B. L. & Phillips, R. A. A detailed analysis of chromosomal changes in heritable and non-heritable retinoblastoma. Hum. Genet. 70, 291–301 (1985)
Acknowledgements
The WGS was supported as part of the St Jude Children’s Research Hospital – Washington University Pediatric Cancer Genome Project. We thank J. Lahti, J. Dalton and M. Valentine for help with FISH analysis, L. Holmfeldt, J. Zhang and M. Barbato for help with sample preparation, and V. Valentine for spectral karyotype analysis. We thank I. Qaddoumi, C. Rodriguez-Galindo and B. Haik for the support of the St Jude Children’s Research Hospital retinoblastoma clinical research and W. Lei, D. McGoldrick, D. Alford, S. Espy, J. Obenauer and K. Johnson for assistance with data acquisition, handling and analysis. We thank A. McArthur and C. Guess for editing the manuscript, J. Temirov for help with sister chromatid analysis, J. Thurman for help with histology, F. Krafcik for help with cell culture, J. Wu and C. Billups for statistical analysis, J. Jeon for help with lentiviral preparations and C. Bradley for assistance with preclinical testing. This work was supported, in part, by Cancer Center Support (CA21765) from the NCI; grants to M.A.D. from the NIH (EY014867 and EY018599), the American Cancer Society and the Research to Prevent Blindness Foundation; and the American Lebanese Syrian Associated Charities. M.A.D. is a Howard Hughes Medical Institute Early Career Scientist. This work was also supported by an American Cancer Society Fellowship to A.L.M., the MGH Cancer Center Saltonstall Foundation Scholarship to N.J.D. and funding from AstraZeneca and NIH grants GM81607 and CA64402 to N.J.D.
Author information
Authors and Affiliations
Contributions
M.A.D., C.A.B., J. McEvoy, J.F.-O., R.B. and A.L.M. designed the experiments. L.D. and J.Z. led the data analysis. M.A.D., L.D., J.Z., X.C., A.U., G.W., J.W., M.R., J. Ma, S.P., S.M., P.G., G.N., D.Z., C.L., R.S.F., L.L.F., X.H., D.J.D. and K.O. performed the data analysis and provided bioinformatics support. M.A.D., J.Z., A.U., C.A.B., J. McEvoy, J.F.-O., R.B. and A.L.M. prepared the figures and tables. C.A.B, J.M., J.F.-O., R.B. and A.L.M. performed the laboratory experiments. J.F.-O. created the xenograft. M.W. provided samples. J.E., S.S. and C.M. provided assistance with sample processing and handling. A.B. and D.E. provided pathology support. M.A.D., C.N., E.R.M., R.K.W. and J.R.D. supervised the project. M.A.D. wrote the manuscript. J.Z., C.A.B., J. McEvoy, J.F.-O., L.D., M.W., E.R.M., S.S., C.M. and J.R.D. read and commented on the manuscript. J.Z., L.D., X.C., A.U., G.W., J.W., M.R., J. Ma, J.E., S.S., C.M., S.P., S.M., P.G., G.N., D.Z., C.L., R.S.F., L.L.F., X.H., D.J.D., K.O., C.N., E.R.M., A.B., D.E., R.K.W., J.R.D. and M.A.D. are part of the St Jude Children’s Research Hospital – Washington University Pediatric Cancer Genome Project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Materials and Methods, Supplementary Figures 1-22 with legends, Supplementary Tables 1-6, 8-9, 11 and 13 (see separate zipped file for Tables 7,10, 12, 14 and 15) and additional references. The zipped file containing Supplementary Tables 7, 10, 12, 14 and 15 was added on 3 February 2012. (PDF 13338 kb)
Supplementary Tables 7
This file contains Supplementary Tables 7, 10, 12, 14 and 15. (ZIP 22384 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This licence does not permit commercial exploitation, and derivative works must be licensed under the same or similar licence.
About this article
Cite this article
Zhang, J., Benavente, C., McEvoy, J. et al. A novel retinoblastoma therapy from genomic and epigenetic analyses. Nature 481, 329–334 (2012). https://doi.org/10.1038/nature10733
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10733
This article is cited by
-
Prox1 Suppresses Proliferation and Drug Resistance of Retinoblastoma Cells via Targeting Notch1
Current Medical Science (2024)
-
Retinoblastoma: present scenario and future challenges
Cell Communication and Signaling (2023)
-
The functional role of circular RNAs in the pathogenesis of retinoblastoma: a new potential biomarker and therapeutic target?
Clinical and Translational Oncology (2023)
-
A novel MYCN-YTHDF1 cascade contributes to retinoblastoma tumor growth by eliciting m6A -dependent activation of multiple oncogenes
Science China Life Sciences (2023)
-
LncRNA HOTAIR facilitates proliferation and represses apoptosis of retinoblastoma cells through the miR-20b-5p/RRM2/PI3K/AKT axis
Orphanet Journal of Rare Diseases (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.