Abstract
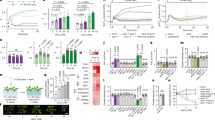
Innate immunity represents the first line of inducible defence against microbial infection in plants and animals1,2,3. In both kingdoms, recognition of pathogen- or microbe-associated molecular patterns (PAMPs or MAMPs, respectively), such as flagellin, initiates convergent signalling pathways involving mitogen-activated protein kinase (MAPK) cascades and global transcriptional changes to boost immunity1,2,3,4. Although Ca2+ has long been recognized as an essential and conserved primary mediator in plant defence responses, how Ca2+ signals are sensed and relayed into early MAMP signalling is unknown5,6. Using a functional genomic screen and genome-wide gene expression profiling, here we show that four calcium-dependent protein kinases (CDPKs) are Ca2+-sensor protein kinases critical for transcriptional reprogramming in plant innate immune signalling. Unexpectedly, CDPKs and MAPK cascades act differentially in four MAMP-mediated regulatory programs to control early genes involved in the synthesis of defence peptides and metabolites, cell wall modifications and redox signalling. Transcriptome profile comparison suggests that CDPKs are the convergence point of signalling triggered by most MAMPs. Double, triple and quadruple cpk mutant plants display progressively diminished oxidative burst and gene activation induced by the 22-amino-acid peptide flg22, as well as compromised pathogen defence. In contrast to negative roles of calmodulin and a calmodulin-activated transcription factor in plant defence7,8, the present study reveals Ca2+ signalling complexity and demonstrates key positive roles of specific CDPKs in initial MAMP signalling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Data deposits
All microarray data are available at the Gene Expression Omnibus (GEO; http://www.ncbi.nlm.nih.gov/geo/) under accession number GSE16557.
References
Nürnberger, T., Brunner, F., Kemmerling, B. & Piater, L. Innate immunity in plants and animals: striking similarities and obvious differences. Immunol. Rev. 198, 249–266 (2004)
Akira, S., Uematsu, S. & Takeuchi, O. Pathogen recognition and innate immunity. Cell 124, 783–801 (2006)
Boller, T. & Felix, G. A renaissance of elicitors: perception of microbe-associated molecular patterns and danger signals by pattern-recognition receptors. Annu. Rev. Plant Biol. 60, 379–406 (2009)
Asai, T. et al. MAP kinase signalling cascade in Arabidopsis innate immunity. Nature 415, 977–983 (2002)
Lecourieux, D., Ranjeva, R. & Pugin, A. Calcium in plant defence-signalling pathways. New Phytol. 171, 249–269 (2006)
Ma, W. & Berkowitz, G. A. The grateful dead: calcium and cell death in plant innate immunity. Cell. Microbiol. 9, 2571–2585 (2007)
Kim, M. C. et al. Calmodulin interacts with MLO protein to regulate defence against mildew in barley. Nature 416, 447–451 (2002)
Du, L. et al. Ca2+/calmodulin regulates salicylic-acid-mediated plant immunity. Nature 457, 1154–1158 (2009)
Lecourieux, D. et al. Proteinaceous and oligosaccharidic elicitors induce different calcium signatures in the nucleus of tobacco cells. Cell Calcium 38, 527–538 (2005)
Gust, A. A. et al. Bacteria-derived peptidoglycans constitute pathogen-associated molecular patterns triggering innate immunity in Arabidopsis. J. Biol. Chem. 282, 32338–32348 (2007)
Ma, W., Smigel, A., Tsai, Y. C., Braam, J. & Berkowitz, G. A. Innate immunity signaling: cytosolic Ca2+ elevation is linked to downstream nitric oxide generation through the action of calmodulin or a calmodulin-like protein. Plant Physiol. 148, 818–828 (2008)
Harper, J. F. & Harmon, A. Plants, symbiosis and parasites: a calcium signalling connection. Nature Rev. Mol. Cell Biol. 6, 555–566 (2005)
Boudsocq, M. & Sheen, J. in Abiotic Stress Adaptation in Plants: Physiological, Molecular and Genomic Foundation (eds Pareek, A., Sopory, S. K., Bohnert, H. J. & Govindjee) Ch. 4 (Springer, 2009)
Luan, S. The CBL-CIPK network in plant calcium signaling. Trends Plant Sci. 14, 37–42 (2009)
Cheng, S. H., Willmann, M. R., Chen, H. C. & Sheen, J. Calcium signaling through protein kinases. The Arabidopsis calcium-dependent protein kinase gene family. Plant Physiol. 129, 469–485 (2002)
Romeis, T., Piedras, P. & Jones, J. D. G. Resistance gene-dependent activation of a calcium-dependent protein kinase in the plant defense response. Plant Cell 12, 803–815 (2000)
Romeis, T., Ludwig, A. A., Martin, R. & Jones, J. D. G. Calcium-dependent protein kinases play an essential role in a plant defence response. EMBO J. 20, 5556–5567 (2001)
Kobayashi, M. et al. Calcium-dependent protein kinases regulate the production of reactive oxygen species by potato NADPH oxidase. Plant Cell 19, 1065–1080 (2007)
Zipfel, C. et al. Bacterial disease resistance in Arabidopsis through flagellin perception. Nature 428, 764–767 (2004)
Shan, L. et al. Bacterial effectors target the common signaling partner BAK1 to disrupt multiple MAMP receptor-signaling complexes and impede plant immunity. Cell Host Microbe 4, 17–27 (2008)
Zimmermann, P., Hennig, L. & Gruissem, W. Gene-expression analysis and network discovery using Genevestigator. Trends Plant Sci. 10, 407–409 (2005)
Sheen, J. Ca2+-dependent protein kinases and stress signal transduction in plants. Science 274, 1900–1902 (1996)
Uno, Y., Rodriguez Milla, M. A., Maher, E. & Cushman, J. C. Identification of proteins that interact with catalytically active calcium-dependent protein kinases from Arabidopsis. Mol. Genet. Genomics 281, 375–390 (2009)
Baena-González, E., Rolland, F., Thevelein, J. M. & Sheen, J. A central integrator of transcription networks in plant stress and energy signalling. Nature 448, 938–942 (2007)
Bednarek, P. et al. A glucosinolate metabolism pathway in living plant cells mediates broad-spectrum antifungal defense. Science 323, 101–106 (2009)
Clay, N. K., Adio, A. M., Denoux, C., Jander, G. & Ausubel, F. M. Glucosinolate metabolites required for an Arabidopsis innate immune response. Science 323, 95–101 (2009)
Huffaker, A. & Ryan, C. A. Endogenous peptide defense signals in Arabidopsis differentially amplify signaling for the innate immune response. Proc. Natl Acad. Sci. USA 104, 10732–10736 (2007)
Nemhauser, J. L., Hong, F. & Chory, J. Different plant hormones regulate similar processes through largely nonoverlapping transcriptional responses. Cell 126, 467–475 (2006)
Zhu, S. Y. et al. Two calcium-dependent protein kinases, CPK4 and CPK11, regulate abscisic acid signal transduction in Arabidopsis. Plant Cell 19, 3019–3036 (2007)
Sheen, J. et al. in Biology of Plant Microbe Interactions Vol. 6, Ch. 86 (IS-MPMI Symposium Proceedings, 2008)
Alonso, J. M. & Stepanova, A. N. T-DNA mutagenesis in Arabidopsis. Methods Mol. Biol. 236, 177–188 (2003)
He, P. et al. Specific bacterial suppressors of MAMP signaling upstream of MAPKKK in Arabidopsis innate immunity. Cell 125, 563–575 (2006)
Schreiber, K., Ckurshumova, W., Peek, J. & Desveaux, D. A high-throughput chemical screen for resistance to Pseudomonas syringae in Arabidopsis. Plant J. 54, 522–531 (2008)
Yoo, S. D., Cho, Y. H. & Sheen, J. Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nature Protocols 2, 1565–1572 (2007)
Boudsocq, M., Barbier-Brygoo, H. & Laurière, C. Identification of nine sucrose nonfermenting 1-related protein kinases 2 activated by hyperosmotic and saline stresses in Arabidopsis thaliana. J. Biol. Chem. 279, 41758–41766 (2004)
Burch-Smith, T. M., Schiff, M., Liu, Y. & Dinesh-Kumar, S. P. Efficient virus-induced gene silencing in Arabidopsis. Plant Physiol. 142, 21–27 (2006)
Felix, G., Duran, J. D., Volko, S. & Boller, T. Plants have a sensitive perception system for the most conserved domain of bacterial flagellin. Plant J. 18, 265–276 (1999)
Chinchilla, D. et al. A flagellin-induced complex of the receptor FLS2 and BAK1 initiates plant defence. Nature 448, 497–500 (2007)
Acknowledgements
We thank K. H. Liu and E. Baena-González for contributing to the CPK clone collection, O. R. Patharkar for initial microarray data analysis, B. Mueller and G. Tena for offering qPCR primers, M. N. Soler and S. Bolte for assistance on confocal microscopy, C. Laurière for discussions, S. P. Dinesh-Kumar for VIGS vectors, K. Schreiber and D. Desveaux for the seedling pathogen assay, and A. Gust and T. Nürnberger for the original PGN microarray data. We thank the Salk Institute, Syngenta Biotechnology and A. Sessions for sharing the Arabidopsis T-DNA collections, and the Arabidopsis Biological Resource Center for providing Arabidopsis mutant seeds. This research has been supported by a Marie Curie International fellowship within the 6th European Community Framework Program to M.B., an NSF predoctoral fellowship to M.R.W., and grants from the National Science Foundation and the National Institute of Health, and the MGH CCIB fund to J.S.
Author Contributions J.S. and M.R.W. initiated the project; M.B., M.R.W. and J.S. designed the experiments; M.R.W., M.B. and S.-H.C. built the CPK clone collection; M.B., M.R.W., H.L., L.S., P.H. and J.S. conducted the experiments and analysed the data; M.M., J.S. and M.B. analysed microarray data; J.B. managed plants; M.B., J.S. and M.M. prepared the manuscript with inputs from all co-authors.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-14 with Legends, Supplementary Tables 1and 9-14 (see separate files for Supplementary Tables 2-8), Supplementary Methods and Supplementary References. (PDF 12933 kb)
Supplementary Table 2
This table shows the total target gene list of CPK5 and CPK11, with the induction level (log2 ratio) by CPK and flg22 treatments in protoplasts, leaves and seedlings. For each gene, the Affymetrix ID, AGI number and TAIR annotations are provided. (XLS 205 kb)
Supplementary Table 3
This table shows shared CPK5 and CPK11 target genes overlapping with early flg22 responsive genes. For each gene, the induction level (log2 ratio) by CPK, the Affymetrix ID, AGI number, MapMan function category and annotations, Sheen lab annotations and TAIR annotations are provided. (XLS 126 kb)
Supplementary Table 4
This table shows CPK5-specific target genes overlapping with early flg22 responsive genes. For each gene, the induction level (log2 ratio) by CPK, the Affymetrix ID, AGI number, MapMan function category and annotations, Sheen lab annotations and TAIR annotations are provided. (XLS 106 kb)
Supplementary Table 5
This table shows CPK11-specific target genes overlapping with early flg22 responsive genes. For each gene, the induction level (log2 ratio) by CPK, the Affymetrix ID, AGI number, MapMan function category and annotations, Sheen lab annotations and TAIR annotations are provided. (XLS 76 kb)
Supplementary Table 6
This table shows CPK5 and CPK11 target genes (shared and specific) overlapping with early flg22 responsive genes. For each gene, the induction level (log2 ratio) by CPK and flg22 treatments in protoplasts, leaves and seedlings, the Affymetrix ID, AGI number and TAIR annotations are provided. (XLS 57 kb)
Supplementary Table 7
This table shows CPK5 and CPK11 target genes (shared and specific) overlapping with early MAMP responsive genes. For each gene, the induction level (log2 ratio) by CPK, flg22 treatments in leaves and seedlings and ABA treatment, the Affymetrix ID, AGI number and TAIR annotations are provided. (XLS 52 kb)
Supplementary Table 8
This table shows CPK5 and CPK11 target genes (shared and specific) overlapping with microbe responsive genes. For each gene, the induction level (log2 ratio) by CPK and microbes, the Affymetrix ID, AGI number and TAIR annotations are provided. (XLS 44 kb)
Rights and permissions
About this article
Cite this article
Boudsocq, M., Willmann, M., McCormack, M. et al. Differential innate immune signalling via Ca2+ sensor protein kinases. Nature 464, 418–422 (2010). https://doi.org/10.1038/nature08794
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08794
This article is cited by
-
Comparative RNA-seq analysis of resistant and susceptible banana genotypes reveals molecular mechanisms in response to banana bunchy top virus (BBTV) infection
Scientific Reports (2023)
-
NET4 and RabG3 link actin to the tonoplast and facilitate cytoskeletal remodelling during stomatal immunity
Nature Communications (2023)
-
Comparative transcriptome profiling of near isogenic lines PBW343 and FLW29 to unravel defense related genes and pathways contributing to stripe rust resistance in wheat
Functional & Integrative Genomics (2023)
-
Uncovering molecular mechanisms involved in microbial volatile compounds-induced stomatal closure in Arabidopsis thaliana
Plant Molecular Biology (2023)
-
Ca2+-responsive phospholipid-binding BONZAI genes confer a novel role for cotton resistance to Verticillium wilt
Plant Molecular Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.