Abstract
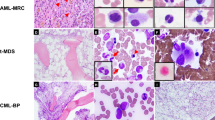
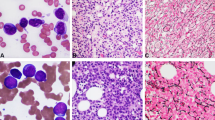
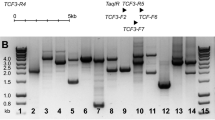
Six patients with de novo acute myeloid leukemia (AML) and a t(2;3)(p15–21;q26–27) were identified among approximately 1000 cases enrolled in the GIMEMA trial. The t(2;3) was the sole anomaly in three patients, whereas in three cases monosomy 7, trisomy 15 and 22, and trisomy 14 represented additional aberrations. No cryptic chromosome deletions at 5q, 7q, 12p, and 20q were observed. One patient carried a FLT3 D835 mutation; FLT3 internal tandem duplication (ITD) was not detected in three patients tested. Characterization of the translocation breakpoints using a 3q26 BAC contig specific for the PRDM3 locus showed that the breakpoints were located 5′ to EVI1 as follows: within myelodysplatic syndrome (MDS) intron 1 (# 3), between MDS1 exons 2 and 3 in three patients (# 1, 2, 4) with a 170 bp cryptic deletion distal to the breakpoint in one (# 2), and in a more centromeric position spanning from intron 2 to the 5′ region of EVI1 (# 6, 5). A set of 2p16–21 BAC probes showed that the breakpoints on chromosome 2p were located within BCL11A in two separate regions (# 1, 4 and # 2–5), within the thyroid adenoma-associated (THADA) gene (# 6) or distal to the ZFP36L2 locus (# 3). Regulatory elements were present in proximity of these breakpoints. RACE PCR studies revealed a chimeric transcript in 1/6 patient analyzed, but no fusion protein. Quantitative PCR showed a 21–58-fold overexpression of the EVI1 gene in all cases analyzed. The patients showed dysplasia of at least two myeloid cell lineages in all cases; they had a low-to-normal platelet count and displayed an immature CD34+ CD117+ immunophenotype. Despite intensive chemotherapy and a median age of 43 years (range 36–59), only two patients attained a short-lived response; one patient is alive with active disease at 12 months, five died at 4–14 months. We arrived at the following conclusions: (a) the t(2;3) is a recurrent translocation having an approximate 0.5% incidence in adult AML; (b) breakpoints involve the 5′ region of EVI1 at 3q26, and the BCL11A, the THADA gene or other regions at 2p16.1–21; (c) cryptic deletions distal to the 3q26 breakpoint may occur in some cases; (d) the juxtaposition of the 5′ region of EVI1 with regulatory elements normally located on chromosome 2 brings about EVI1 overexpression; (e) clinical outcome in these cases is severe.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Mrozek K, Heerema NA, Bloomfield CD . Cytogenetics in acute leukemia. Blood Rev 2004; 18: 115–136.
Alcalay M, Meani N, Gelmetti V, Fantozzi A, Fagioli M, Orleth A et al. Acute myeloid leukemia fusion proteins deregulate genes involved in stem cell maintenance and DNA repair. J Clin Invest 2003; 112: 1751–1761.
Grimwade D, Walker H, Oliver F, Wheatley K, Harrison C, Harrison G et al. The importance of diagnostic cytogenetics on outcome in AML: analysis of 1,612 patients entered into the MRC AML 10 trial. The Medical Research Council Adult and Children's Leukaemia Working Parties. Blood 1998; 92: 2322–2333.
Byrd JC, Mrozek K, Dodge RK, Carroll AJ, Edwards CG, Arthur DC, et al., Cancer and Leukemia Group B (CALGB 8461). Pretreatment cytogenetic abnormalities are predictive of induction success, cumulative incidence of relapse, and overall survival in adult patients with de novo acute myeloid leukemia: results from Cancer and Leukemia Group B (CALGB 8461). Blood 2002; 100: 4325–4336.
Secker-Walker LM, Mehta A, Bain B . Abnormalities of 3q21 and 3q26 in myeloid malignancy: a United Kingdom Cancer Cytogenetic Group study. Br J Haematol 1995; 91: 490–501.
Cuneo A, Mecucci C, Hagemeijer A . Cytogenetic and molecular abnormalities in myeloid malignancies. In: Degos LLD, Lowenberg B (eds). Textbook of Malignant Hematology. Martin Dunitz Publ.: London, 1999.
Grimwade D . The clinical significance of cytogenetic abnormalities in acute myeloid leukaemia. Best Pract Res Clin Haematol 2001; 14: 497–529.
Nucifora G . The EVI1 gene in myeloid leukemia. Leukemia 1997; 11: 2022–2031.
Morishita K, Parganas E, William CL, Whittaker MH, Drabkin H, Oval J et al. Activation of EVI1 gene expression in human acute myelogenous leukemias by translocations spanning 300–400 kilobases on chromosome band 3q26. Proc Natl Acad Sci USA 1992; 89: 3937–3941.
Barjesteh van Waalwijk van Doorn-Khosrovani S, Erpelinck C, van Putten WL, Valk PJ, van der Poel-van de Luytgaarde S, Hack R et al. High EVI1 expression predicts poor survival in acute myeloid leukemia: a study of 319 de novo AML patients. Blood 2003; 101: 837–845.
Barjesteh van Waalwijk van Doorn-Khosrovani S, Erpelinck C, Lowenberg B, Delwel R . Low expression of MDS1-EVI1-like-1 (MEL1) and EVI1-like-1 (EL1) genes in favorable-risk acute myeloid leukemia. Exp Hematol 2003; 31: 1066–1072.
Martinelli G, Ottaviani E, Buonamici S, Isidori A, Borsaru G, Visani G et al. Association of 3q21q26 syndrome with different RPN1/EVI1 fusion transcripts. Haematologica 2003; 88: 1221–1228.
Buonamici S, Chakraborty S, Senyuk V, Nucifora G . The role of EVI1 in normal and leukemic cells. Blood Cells Mol Dis 2003; 31: 206–212.
Voutsadakis IA, Maillard N . Acute myelogenous leukemia with the t(3;12)(q26;p13) translocation: case report and review of the literature. Am J Hematol 2003; 72: 135–137.
Storlazzi CT, Anelli L, Albano F, Zagaria A, Ventura M, Rocchi M et al. A novel chromosomal translocation t(3;7)(q26;q21) in myeloid leukemia resulting in overexpression of EVI1. Ann Hematol 2004; 83: 78–83.
Stevens-Kroef M, Poppe B, van Zelderen-Bhola S, van den Berg E, van der Blij-Philipsen M, Geurts van Kessel A et al. Translocation t(2;3)(p15–23;q26–27) in myeloid malignancies: report of 21 new cases, clinical, cytogenetic and molecular genetic features. Leukemia 2004; 18: 1108–1114.
Cuneo A, Bigoni R, Cavazzini F, Bardi A, Roberti MG, Agostini P et al. Incidence and significance of cryptic chromosome aberrations detected by fluorescence in situ hybridization in acute myeloid leukemia with normal karyotype. Leukemia 2002; 16: 1745–1751.
Noguera NI, Breccia M, Divona M, Diverio D, Costa V, De Santis S et al. Alterations of the FLT3 gene in acute promyelocytic leukemia: association with diagnostic characteristics and analysis of clinical outcome in patients treated with the Italian AIDA protocol. Leukemia 2002; 16: 2185–2189.
van Dongen JJ, Macintyre EA, Gabert JA, Delabesse E, Rossi V, Saglio G et al. Standardized RT-PCR analysis of fusion gene transcripts from chromosome aberrations in acute leukemia for detection of minimal residual disease. Report of the BIOMED-1 Concerted Action: investigation of minimal residual disease in acute leukemia. Leukemia 1999; 13: 1901–1928.
Lichter P, Tang Chang CJ, Call K, Hermanson G, Evans GA, Housman D et al. High resolution mapping of human chromosomes 11 by in situ hybridization with cosmid clones. Science 1990; 247: 64–69.
Davuluri RV GI, Zhang MQ . Computational identification of promoters and first exons in the human genome. Nat Genet 2001; 29: 412–417.
Bennett JM, Catovsky D, Daniel MT, Flandrin G, Galton DA, Gralnick HR et al. Proposals for the classification of the acute leukaemias. French–American–British (FAB) co-operative group. Br J Haematol 1976; 33: 451–458.
Vardiman JW, Harris NL, Brunning RD . The World Health Organization (WHO) classification of the myeloid neoplasms. Blood 2002; 100: 2292–2302.
Satterwhite E, Sonoki T, Willis TG, Harder L, Nowak R, Arriola EL et al. The BCL11 gene family: involvement of BCL11A in lymphoid malignancies. Blood 2001; 98: 3413–3420.
Zent C, Rowley JD, Nucifora G . Rearrangements of the AML1/CBFA2 gene in myeloid leukemia with the 3;21 translocation: in vitro and in vivo studies. Leukemia 1997; 11: 273–278.
Charrin C, Belhabri A, Treille-Ritouet D, Theuil G, Magaud JP, Fiere D et al. Structural rearrangements of chromosome 3 in 57 patients with acute myeloid leukemia: clinical, hematological and cytogenetic features. Hematol J 2002; 3: 21–31.
Kolomietz E, Al-Maghrabi J, Brennan S, Karaskova J, Minkin S, Lipton J et al. Primary chromosomal rearrangements of leukemia are frequently accompanied by extensive submicroscopic deletions and may lead to altered prognosis. Blood 2001; 97: 3581–3588.
Buonamici S, Li D, Chi Y, Zhao R, Wang X, Brace L et al. EVI1 induces myelodysplastic syndrome in mice. J Clin Invest 2004; 114: 713–719.
Acknowledgements
Work in our laboratories was supported by FIRB- MIUR, PRIN-MIUR, fondi 60%, AIRC (Associazione Italiana Ricerca sul Cancro), CNR, Rome.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Leukemia website (http://www.nature.com/leu).
Supplementary information
Rights and permissions
About this article
Cite this article
Trubia, M., Albano, F., Cavazzini, F. et al. Characterization of a recurrent translocation t(2;3)(p15–22;q26) occurring in acute myeloid leukaemia. Leukemia 20, 48–54 (2006). https://doi.org/10.1038/sj.leu.2404020
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2404020
Keywords
This article is cited by
-
Marked thrombocytosis and dysmegakaryopoiesis in acute myeloid leukemia with t(2;3)(p22;q26.2) and EVI1 rearrangement
Annals of Hematology (2013)
-
Genomic instability and myelodysplasia with monosomy 7 consequent to EVI1 activation after gene therapy for chronic granulomatous disease
Nature Medicine (2010)
-
Contributions of the Raf/MEK/ERK, PI3K/PTEN/Akt/mTOR and Jak/STAT pathways to leukemia
Leukemia (2008)