Abstract
Although recent work has shown that both deterministic and stochastic processes are important in structuring microbial communities, the factors that affect the relative contributions of niche and neutral processes are poorly understood. The macrobiological literature indicates that ecological disturbances can influence assembly processes. Thus, we sampled bacterial communities at 4 and 16 weeks following a wildfire and used null deviation analysis to examine the role that time since disturbance has in community assembly. Fire dramatically altered bacterial community structure and diversity as well as soil chemistry for both time-points. Community structure shifted between 4 and 16 weeks for both burned and unburned communities. Community assembly in burned sites 4 weeks after fire was significantly more stochastic than in unburned sites. After 16 weeks, however, burned communities were significantly less stochastic than unburned communities. Thus, we propose a three-phase model featuring shifts in the relative importance of niche and neutral processes as a function of time since disturbance. Because neutral processes are characterized by a decoupling between environmental parameters and community structure, we hypothesize that a better understanding of community assembly may be important in determining where and when detailed studies of community composition are valuable for predicting ecosystem function.
Similar content being viewed by others
Introduction
Over the last few decades, phylogenetic approaches have revealed that microbes exhibit biogeographic patterns in diversity and distribution (Martiny et al., 2006; Hanson et al., 2012) which often mirror those observed for macro-organisms (Langenheder and Prosser, 2008; Langenheder et al., 2010). For example, ample evidence suggests that different ecosystems host distinct types of microbes (Lozupone and Knight, 2007; Nemergut et al., 2011), and that these community differences likely reflect selection (sensu Vellend, 2010) acting on trait differences between suites of organisms. Indeed, evidence supports the role of environmental parameters, including pH, salinity and the abundance and quality of carbon (C) in structuring microbial communities (Fierer and Jackson, 2006; Lozupone and Knight, 2007; Logue and Lindström, 2010; Nemergut et al., 2010). Community assembly processes driven by environmental parameters acting on traits are often called ‘niche-based’ (the term we use here) but they have also been referred to as ‘habitat filters’ and ‘deterministic processes’ in the literature.
However, recent work suggests that ‘historical filters’ or stochastic processes also affect microbial biogeography (Martiny et al., 2006). In particular, there is increasing evidence that dispersal limitations may have a more important role in structuring microbial communities than previously thought (Telford and Vandvik, 2006; Peay et al., 2010; Chytrý et al., 2012). Indeed, the neutral theory of biodiversity (Bell, 2001; Hubbell, 2001) in which dispersal is a key determinant of community structure, has been shown to explain a significant portion of bacterial community variation in a variety of systems ranging from tree hole aquatic habitats to wastewater treatment facilities (Sloan et al., 2006; Woodcock et al., 2007; Ofiteru et al., 2010; Caruso et al., 2011; Langenheder and Székely, 2011). Although the relative contribution of niche and neutral processes in determining microbial community structure may vary across systems, evidence is mounting that both can be important (Östman et al., 2009; Ofiteru et al., 2010).
Results suggesting that both dispersal and selection can influence microbial community assembly raise an important question: what regulates the relative role of niche vs neutral processes in structuring microbial communities? Work from macrobial systems suggests that a suite of factors including ecosystem productivity, metacommunity (defined here as a set of communities linked by dispersing and interacting taxa) diversity, and dispersal rates are important in the relative balance of these assembly processes (Chase, 2003). Additionally, empirical studies have demonstrated that disturbance can cause an increase in the importance of niche-based processes in structuring communities (Chase, 2007; Jiang and Patel, 2008). Yet, other research suggests that disturbance may promote neutral processes (Didham et al., 2005; Didham and Norton, 2006). Although the specifics of community assembly in response to disturbance may vary with the type and intensity of disturbance, as well as the ecosystem examined, disturbance events frequently kill or severely impact many members of a community. This can ‘reset’ assembly processes and may create temporal gradients that provide excellent opportunities for examining general rules about community assembly.
Here, we examined bacterial community assembly processes in response to a wildfire. Fires are ecologically important disturbances (Bond et al., 2005) and their effects on plant and animal communities as well as soil biogeochemistry have been widely studied (Certini, 2005; Wang and Kemball, 2005; Ferrenberg et al., 2006). Recent work has also shown that fire induces microbial community shifts characterized by an increase in the relative abundance of Firmicutes and/or β-proteobacteria and an increase in the ratio of bacteria to fungi (Yeager et al., 2005; Smith et al., 2008; Waldrop and Harden, 2008; Bárcenas-Moreno et al., 2011). However, the relative role of niche vs neutral assembly processes in driving these community shifts is unknown. Fire-based disturbance can lead to major shifts in a variety of environmental parameters that are likely to have large direct and indirect effects on the soil microbial community through niche-based processes. For example, fires typically result in an ephemeral pulse of ammonium (NH4+), creation of a reactive charcoal layer, and subsequent changes in pH (Peitikäinen et al., 2000; DeLuca and Sala, 2006; Wardle et al., 1997, 1998). On the other hand, because fire causes large reductions in the standing soil microbial biomass (Hart et al., 2005; Wang et al., 2012), dispersal, which can be largely stochastic, may lead to an increase in the relative importance of neutral processes in early community assembly.
In this study, we used pyrosequencing of bacterial 16S rRNA genes and null deviation analysis (Chase and Myers, 2011) to examine changes in bacterial community assembly processes following a wildfire that killed all vegetation and consumed the surface litter layer of a conifer forest. We sampled the bacterial community and chemistry of soils from a burned site and from an adjacent unburned forest stand at 4 and 16 weeks after the fire. On the basis of previous studies, we expected that burned soils would demonstrate an increase in soil pH and ammonium pools, a decrease in soil organic matter and alpha (local) and gamma (regional) diversity, and shifts in bacterial community composition. Given the importance of dispersal in early recovery processes, we hypothesized that the burned communities at 4 weeks would show a greater relative importance of neutral processes than unburned communities. By 16 weeks, we hypothesized that niche-based processes, driven by the major shifts in soil chemistry, would become more important for microbial community assembly in the burned sites. Our data yielded insights into microbial community assembly following disturbance, which may be important for a better understanding of the relationships between assembly processes, microbial community structure and ecosystem function.
Materials and methods
Study site and sample collection
A total of 100 samples, 25 each from burned and unburned soils, collected at both 4 (October 2010) and 16 weeks postfire (January 2011) were analyzed in this study. Soils were sampled roughly 1 m from the base of living (unburned) and dead (burned) tree trunks near the southeastern edge of the Fourmile Fire (40.039N, 105.391W), that was ignited on 6 September 2010 on the eastern slope of the Colorado Front Range, Boulder County, CO, USA. Forests were dominated by ponderosa pines (Pinus ponderosa scopulorum) and Douglas firs (Pseudotsugae menziesii glauca) on similar northeastern aspects, between 2100–2285 m asl. The climate, fire history and soils of these forests were described by Schoennagel et al., (2011) and Veblen et al., (2000). Unburned and burned sites were ∼300 m apart with each site roughly 150 m from the dividing fire-line between burned and unburned forest. Trees in both treatments were located within a 650 m2 plot with a minimum of 3 m and a maximum of 25 m separating individual trees; soils were collected from under the same trees on both dates. Samples were a composite of three 130.5 cm3 cores from the top 5 cm of mineral soil with litter and visible organic material excluded, and were transported on ice, sieved through 2 mm mesh, and stored at −80 °C for DNA extraction or at +4 °C for biogeochemical analyses.
DNA extraction and pyrosequencing of partial 16 S rRNA genes
DNA was isolated using the MO BIO Power Soil DNA Extraction kit (MO BIO Laboratories, Carlsbad, CA, USA), and was processed as described in Nemergut et al., (2010) and Knelman et al., (2012). A fragment of the 16S rRNA gene encoding the V1-V2 region was amplified using modified primers of 27F and 338R adapted for Titanium chemistry (454 Life Sciences, Bradford, CT, USA). PCR reactions were performed in triplicate with 10 μl of sterile H2O, 10 μl of 5 PRIME hot master mix (5 PRIME, Gaithersburg, MD, USA), 2 μl (5 μM) of the reverse primer, 1 μl (10 μM) of the forward primer and 2 μl of the sample DNA. Samples were denatured for 3 min at 94 °C followed by 25 cycles at 94 °C for 45 s, 50 °C for 30 s, 72 °C for 90 s and a final elongation step at 70 °C for 10 min. Three replicate PCR products were quantified, pooled and cleaned using MO BIO UltraClean-htp PCR Clean-up kits and 16S rRNA gene amplicons were sent to the Environmental Genomics Core Facility (Engencore) at University of South Carolina for 454 Life Sciences GS FLX Titanium pyrosequencing.
Sequence Analysis
Pyrosequencing data were screened using the QIIME (version 1.2.1) toolkit (Caporaso et al., 2010) with the following parameters: quality score >25, sequence length >200 and <400, maximum homopolymer of 6, 0 maximum ambiguous bases and 0 mismatched bases in the primer. OTUs were denoised using Denoiser (Reeder and Knight, 2010) and were picked at the 97% identity level using UCLUST (Edgar, 2010) in QIIME. OTUs were randomly subsampled in QIIME so each library contained 1142 sequences (the fewest in a single sample). Quality data were not obtained from five samples which were excluded from analyses. The taxonomic identity of OTUs was assigned using RDP in QIIME, and QIIME was used to generate a weighted UniFrac distance matrix (Lozupone and Knight, 2005; Lozupone et al., 2006) and a Bray–Curtis distance matrix (Bray and Curtis, 1957). QIIME was also used to generate α and γ diversity metrics (OTU richness (unique OTUs), Shannon diversity, phylogenetic diversity, Pielou’s evenness and dominance (probability of randomly sampling two individuals of the same OTU, Caparaso et al., 2010)). All sequencing data have been deposited in the MG-RAST database (http://metagenomics.anl.gov/).
Soil analysis and microbial biomass
Soil moisture, pH and total C and nitrogen (N) were measured on samples collected on both dates. Soil moisture was determined with the gravimetric method after drying soils at 60 °C for 48 h. Soil pH was measured using a 1:5 ratio of soil to de-ionized H2O. Total C and N were determined by grinding and combustion in an elemental analyzer as described by (Knelman et al., 2012). NH4+, dissolved organic N (DON), dissolved organic C (DOC), and microbial biomass were measured for only the 16 week samples by adding 40 ml of 0.5 M K2SO4 to 10 g of soil, shaking the mix for 1 h, and filtering through Whatman no.1 paper (Whatman Incorporated, Florham Park, NJ, USA). NH4+ concentrations were determined using the sodium salicylate method and absorbance at 650 nm on a microplate reader (Mulvaney, 1996). DOC and DON were determined using a TIC/TOC analyzer. For DOC, biomass C=EC/kEC, where EC=extractable C from soil, and kEC (extractable C from microbial biomass) was estimated at 0.45 (Beck et al., 1997). DON was determined by Kjeldahl Digestion of 20 ml of extract; N=EN/kEN where kEN was estimated at 0.54 (Brookes et al., 1985). Microbial C and N pools were calculated as the difference between DOC and DON from non-fumigated and 5-day chloroform fumigated soils (Brookes et al., 1985; Beck et al., 1997).
Data Analysis
Burned and unburned soil chemistry, α diversity measures, and weighted UniFrac matrices were compared with SAS-JMP 9.0.0 (JMP 2011) using one-way analysis of variance followed by Tukey’s HSD means comparisons (Kruskal–Wallace tests followed by Steel-Dwass means comparisons when test assumptions were not met). To avoid violating assumptions of sample independence, sample OTU dissimilarities (β-diversity, calculated as mean group Bray–Curtis dissimilarity) were compared using ADONIS followed by the permutation method of ‘betadisper’ in the Vegan package for the R platform (Oksanen et al., 2011; Team RDC 2011). Because the hypotheses tested here focus on the difference between burned and unburned soils and less on differences between geographically collocated samples, and because samples were removed from the site, repeated measures analyses or paired tests were not used. Variables that were measured only in January (NH4+, DOC/DON, microbial biomass) were compared via t-tests or Mann–Whitney U-tests. When effective, log or log10 transformations were applied to meet test assumptions. Our figures and tables contain back-transformed values with statistical comparisons based on transformed data as noted.
PERMANOVA and non-metric multidimensional scaling were completed on the Bray–Curtis distance matrix in PC-ORD (McCune and Mefford, 2011) and used to compare community composition in burned and unburned samples. Mantel tests for correlations between environmental factors and the soil bacterial community of burned and unburned soils at 4 and 16 weeks were calculated in PC-ORD. Edaphic data are available in the MIMARKS database (Yilmaz et al., 2011).
We used the null deviation approach (Chase and Myers, 2011) to examine bacterial community assembly. This technique uses a null model to create stochastically assembled communities from the regional species pool to determine the degree to which observed β diversity patterns deviate from stochastic assembly. The null deviation approach disentangles variation in community compositional dissimilarity across sites from variation due to changes in α (local) and γ (regional) diversity (Chase and Myers, 2011). This approach assesses changes in β diversity that result from the relative influence of niche and neutral processes and not from changes in α diversity. We measured the null deviation as the relative difference of the observed β diversity from the null-model β diversity, (βobs-βnull)/βnull, where β diversity was measured as Sorenson-Czekanowski dissimilarity. For each sample, the expected β diversity under the null model was calculated from 10 000 stochastically assembled communities. Gamma diversity for each null model was calculated from each within-treatment (burned vs unburned at 4 and 16 weeks) species pool. As this analysis requires presence-absence data and does not weight species by their abundance (unlike the non-metric multidimensional scaling and Mantel tests described above) it is sensitive to noise from rare species. Therefore, taxa with very low abundances (<1% of sequences per community) were removed from pyrosequencing data before analyses (Ofiteru et al., 2010). To test for treatment differences in the null deviation we conducted permutation tests by first randomly permuting treatment labels, then resimulating null models and recalculating null deviations for each of 5000 permutations.
Results
Bacterial Community Diversity and Structure
After rarefaction to an equal sequencing depth, we found a total of 4760 unique OTUs across all samples (Supplementary Figures S1 and S2a). Unburned soils contained 2596 OTUs at 4 weeks and 2627 OTUs at 16 weeks postfire. Burned soils had 1889 OTUs at 4 weeks and 1656 at 16 weeks—28 and 37% lower γ diversity than was observed in unburned soils, respectively. Unburned soil had 850 and 1002 unique (that is, not found in samples from any other treatment/date) OTUs and burned soil had 357 and 273 unique OTUs at 4 and 16 weeks, respectively. A total of 698 ‘generalist’ OTUs were found in both burned and unburned soils on both dates.
Burning significantly reduced α diversity for both dates (Supplementary Figure S2 b–f) regardless of the diversity metric (richness, Shannon, phylogenetic diversity, evenness or dominance) applied. Alpha diversity within burned or unburned soils did not change significantly between sampling dates. Specifically, burned soils had an average of 31 and 50% lower OTU richness than unburned soil at 4 and 16 weeks, respectively (Supplementary Figure S2b; F3,84=37.49, P<0.0001). The Shannon diversity index of burned soil (four weeks=6.12, 16 weeks=5.46) was significantly lower than that of unburned soil (4 weeks=7.68, 16 weeks=7.92) (Supplementary Figure S2c, F3,84=26.69, P<0.0001). Finally, burning reduced phylogenetic diversity by 28% at 4 weeks (49.61 vs 68.76) and 42% at 16 weeks (41.99 vs 72.65) (Supplementary Figure S2d; F3,84=33.98, P<0.0001). OTU evenness was lower in burned soil than in unburned at 4 (0.74 vs 0.87) and 16 weeks (0.69 vs 0.89) (Supplementary Figure S2e; F3,84=22.23, P<0.0001), while dominance was higher in burned soils at 4 (0.10 vs 0.02) and 16 weeks (0.09 vs 0.01) (Supplementary Figure S2f; F3,84=10.17, P<0.0001). Average β diversity, in this case mean pairwise Bray-Curtis dissimilarity and UniFrac distance, was also altered by burning (Supplementary Figures S2g and h). Fire caused a significant increase in Bray-Curtis dissimilarity at 4 and 16 weeks (PERMANOVA, P=0.0002), and a significant increase in UniFrac for both dates (PERMANOVA, P=0.0002).
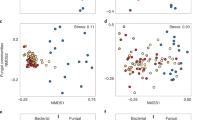
Burned and unburned soils harbored significantly different bacterial communities 4 and 16 weeks postfire (PERMANOVA, P<0.001) and within both treatments between dates (PERMANOVA, P<0.001). Non-metric multidimensional scaling clustered samples by treatment and date, with burned soils displaying a greater spread between samples than for unburned communities (Figure 1), consistent with the observed increases in β diversity. The relative abundances of seven dominant bacterial phyla (or subphyla in the case of Proteobacteria) differed significantly between burned and unburned soils at 4 (Supplementary Figure S3; χ26=58.07, P<0.0001) and 16 weeks (χ26=75.72, P<0.0001); and within the burned samples between dates (χ26=28.73, P<0.0001, Supplementary Figure S3). Proportional phyla and subphyla abundances in unburned soils were not different between dates (χ26=1.29, P>0.95). Firmicutes were in low relative abundance in unburned soils for both dates, but dominated burned soils. Betaproteobacteria were in low relative abundance in the 4 week burned samples, but increased at 16 weeks to become the second most abundant taxon.
Non-metric multidimensional scaling ordination based on Bray-Curtis distances showing the change in bacterial community composition and increase in β diversity in burned soil bacterial communities (4 week communities=gray squares; 16 week communities=black squares) compared with unburned soil bacterial communities (4 week=open triangles; 16 week=solid triangles) 4 and 16 weeks after the fire.
Soil chemistry
Burning reduced soil C and N (F3,96=36.61, P<0.0001), with 65% less C and 48% less N in burned than unburned samples from 4 weeks, and 69% less C and 60% less N in the burned soil from 16 weeks (Table 1). Burning reduced microbial biomass (microbial C; measured only at 16 weeks) by 77% compared with unburned samples (Supplementary Table S1). The C:N ratio of burned soils was 39% lower than that of unburned soils at 4 weeks, and 24% lower at 16 weeks (F3,96=107.94, P<0.0001; Table 1). Burning increased pH for both sample dates, while pH showed an overall decrease between 4 and 16 weeks regardless of treatment (F3,97=20.5, P<0.0001, Table 1). Soil moisture was lower in burned soils at 4 weeks (F3,93=10.38, P<0.0001), with no difference between treatments at 16 weeks (Table 1). Burned soils had greater than a 12-fold increase in mean NH4+ concentration (only measured at 16 weeks), averaging 3.12 μg −1g in unburned soils and 38.87 μg g−1 in burned soils (U=623, Z=-6.01, P<0.0001, Table 1). Burning did not significantly change DOC, but increased DON (both measured only at 16 weeks) by 60% over unburned soil (t24,24=−6.0, P<0.0001, Table 1).
Community Assembly Processes
The null deviation approach (Chase and Myers, 2011) created stochastically assembled communities from the regional species pool to determine the degree to which observed β diversity patterns deviate from stochastic assembly. A null deviation close to zero suggests that neutral processes are more important in structuring the community, whereas larger positive or negative null deviations suggest that niche-based processes are more important. After 4 weeks, burned communities deviated significantly less from the stochastic assembly model than unburned communities (permutation test, P=0.02; Figure 2). After 16 weeks, however, burned communities (relative null deviation=−0.17) deviated significantly more from the stochastic assembly model than unburned communities (P=0.001; Figure 2). Importantly, the unburned sites showed a moderate but consistent deviation from the stochastic assembly model (relative null deviation=−0.12; Figure 2), with no significant changes in the null deviation value between the 4 and 16 week samples (P=0.36). Burned sites, by contrast, were significantly more stochastic at 4 weeks than at 16 weeks (P<0.001).
Plot showing the null deviation (Chase and Myers, 2011) of burned and unburned communities 4 and 16 weeks after the fire. A null deviation close to zero suggests that neutral processes are more important in structuring the community, whereas larger positive or negative null deviations suggest that niche-based processes are more important. Different letters indicate significant differences between sample dates based on permutation tests (P<0.05).
Community structure and soil environmental characteristics
Our analysis revealed that the relationship between environmental characteristics and soil microbial community structure varied with sampling date and disturbance. We found no significant correlations between environmental characteristics and community structure for the 4 week samples (Table 2). For the 16 week burned samples we observed significant correlations between community composition and pH. By contrast, we observed significant correlations between soil C:N and community composition for the 16 week unburned soils. Tests of correlations between the bacterial community and environmental factors yielded similar results regardless of whether weighted or unweighted community metrics were considered (data not shown).
Discussion
A severe wildfire provided an opportunity to examine the relative roles of niche vs neutral assembly processes in recently disturbed soil bacterial communities. As expected, we found that burning caused substantial changes in soil bacterial diversity (Supplementary Figure S2), community structure (Figure 1, Supplementary Figures S1 and S3) and soil chemistry (Table 1). We also found that bacterial secondary succession proceeded very rapidly in the postdisturbance landscape, as communities from 4 and 16 weeks postburn were significantly different, not only in terms of the OTUs present, but also with respect to the phyla/subphyla proportional abundances (Figure 1, Supplementary Figures S1 and S3). Seasonal effects are known to influence soil microbial community abundances and activities (Monson et al., 2006; Schmidt et al., 2007) and, consistent with these observations, we observed a small but significant difference in the bacterial community structure (Figure 1) of unburned samples from 4 (fall) and 16 (winter) weeks. As well, pH and soil moisture were different between the two sampling time-points (Table 1). However, the magnitude of the differences in burned vs unburned soil community structure, along with significant reductions in γ and α diversity in both sample dates suggest that fire effects (direct or indirect) on microbial community assembly and secondary succession are much stronger than seasonal shifts over this time period.
A null deviation value close to zero suggests that community assembly is highly stochastic and neutral processes are more important in structuring the community. Larger positive or negative null deviations suggest that niche-based processes are more important, and environmental filters, for example, could have strong influences on community assembly. Regardless of disturbance or sampling time, null processes were important in structuring soil microbial communities (Figure 2). However, our analysis also revealed that fire caused a quantifiable change in assembly processes (that is, the relative importance of niche vs neutral processes) that shifted with time since disturbance. As with the observed changes in diversity and community structure, these shifts were evident over very short time frames: communities in the soils 4 weeks postburn were shaped by neutral processes (smaller null deviations) significantly more so than unburned communities, while burned communities at 16 weeks were shaped by niche processes (larger null deviations) more than unburned communities.
Interestingly, we also observed an increase in β diversity among postburn communities (Figure 1, Supplementary Figure S2e,f). Such increases in β diversity have been interpreted as support for neutral processes in community assembly (Kraft et al., 2007; Chase and Myers, 2011). However, our null deviation analysis supports that the observed increases in β diversity are due to both increases (4 weeks) and decreases (16 weeks) in neutral processes in burned soils relative to unburned sites (Figure 2). The initial increase in β diversity may thus reflect the stochastic nature of dispersal, and the fact that ‘seed’ microbes from air and precipitation vary in both space and time. As succession proceeds, however, our data suggest that this increase in β diversity reflects an increase in niche-based processes. Other work has demonstrated that fire tends to increase biogeochemical heterogeneity, likely due to variation in the severity of the burn across the landscape (for example, Turner et al., 2007; Hamman et al., 2008). Thus, these increases in environmental variation may be reflected, at least to some degree, in the changes in community composition observed in the burned soils, combined with an increase in niche-based processes in shaping communities. These shifts in community assembly may also be reflected in the trait differences of the dominant taxa. For instance, spore-forming Firmicutes —which may be more easily dispersed— are most abundant in the 4 week postburn soils. As well, Betaproteobacteria, which commonly dominate early successional landscapes (Nemergut et al., 2007; Sattin et al., 2009) are more abundant at 16 weeks, when the decrease in soil organic matter (Table 1) may select for more oligotrophic taxa.
We performed Mantel tests to examine correlations between environmental parameters and community structure. We measured a suite of standard soil chemical parameters (Table 1) but did not observe any significant relationships between these variables and community structure, (regardless of whether comparisons were completed with abundance based analyses or unweighted analyses), within treatments for the 4 week samples (Table 2). For the 16 week samples, β diversity in the burned samples was correlated with pH while unburned communities were correlated with soil C:N. Correlation coefficients for these relationships were roughly similar, and some have interpreted these as evidence for the amount of variation in community structure that is explained by niche-based processes. However, as noted by (Anderson et al., 2011), extreme caution should be taken in interpreting these relationships in terms of assembly mechanisms because of the potential for unmeasured environmental variation as well as the possibility of spatial structure in environmental parameters. Thus, these analyses provide hypotheses about the potential sources of local variation in bacterial community structure, but are not inconsistent with our null deviation analyses.
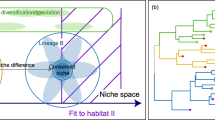
These observed differences in community assembly over very short time scales may reconcile the fact that different researchers have found support for increases in both niche (Chase, 2007; Jiang and Patel, 2008) and neutral (Didham et al., 2005; Didham and Norton, 2006; Leibold and McPeek, 2006) processes following disturbance. Indeed, our results suggest dynamic shifts in community assembly processes in postdisturbance landscapes, leading us to propose a conceptual model describing these changes. Specifically, we hypothesize that time since disturbance features at least three distinct phases in community assembly (Figure 3). Phase 1 immediately follows disturbance, and is characterized by a brief increase in the relative role of neutral processes in community assembly, perhaps because stochastic dispersal processes are strongly affecting community structure. This is supported by other work that suggests that ecological equivalence may be more likely immediately following a severe disturbance event or at the onset of primary succession when immigrants face less competition (Leibold and McPeek, 2006). During phase 2, organisms begin to grow and divide, and niche-based processes in the postdisturbance landscape act as strong filters on microbial community composition. This can be characterized by increases in β diversity if the disturbance was heterogeneous at the landscape level, or decreases if it was more homogenous. Similar results have been observed in other experimental systems as niche-based processes were shown to be important following drought as well as density-independent mortality disturbance events (Chase, 2007; Jiang and Patel, 2008). Finally, over longer time scales (phase 3), the environment becomes less harsh and neutral processes again become more important in shaping community structure. To some degree this supports other work that suggests that neutral processes may dominate community assembly within successional stages while niche processes may dominate during transition periods between successional stages (Denslow, 1980; Ellner and Fussmann, 2003; Cadotte, 2007).
An important caveat of our data is that we lack an understanding of community assembly from a functional level. It is possible that examining assembly processes using metagenomics or metatranscriptomics would reveal different patterns in the relative importance of niche vs neutral processes. For example, Burke et al. (2011) recently showed that microbial community succession on marine algae displayed functional convergence but lacked taxonomic coherence. This suggests a high degree of functional redundancy in microbial communities, which may decouple structure and function. If these same processes are at work in the postdisturbance landscape that we examined, analysis of the 16S rRNA gene data may suggest that communities are assembled by a larger predominance of neutral processes than is actually the case. Future studies should use both SSU rRNA and metagenomic approaches to examine community assembly processes, as trait-based approaches may yield deeper understandings of the mechanisms driving assembly.
As well, such trait-based approaches may be important for guiding approaches for how and where to sample microbial communities to understand ecosystem processes. We now possess the tools to reveal high-resolution details about temporal and structural changes in microbial community structure. As microbial community structure drives function, some argue that there is value in knowing ‘who does what’ to understand and predict ecosystem processes (Zak et al., 2003; Monson et al., 2006; Van Der Heijden et al., 2008). However, as mentioned above, many studies reveal that environmental factors are important determinants of microbial community structure (Fierer and Jackson, 2006; Lozupone and Knight, 2007; Logue and Lindstrom, 2010; Nemergut et al., 2010). Also, these same parameters are vital in regulating ecosystem processes (Bonan and Shugart 1989; Paul and Clark, 1996), raising the question: how much added value is provided by detailed investigations of microbial structure data? Indeed, a better understanding of microbial community assembly processes, and where and when they may change in response to disturbances, could be fundamental to understanding links between structure and function. We hypothesize that if communities are largely structured by neutral processes, then while environmental factors will still affect ecosystem processes in these communities by influencing the physiologies of individual microorganisms, soil communities should exhibit less of a direct link between edaphic factors and processes. In other words, the degree to which niche vs neutral processes guide microbial community assembly will affect the strength of the relationship between environmental factors and ecosystem processes.
References
Oliveros JC (2007), VENNY; An interactive tool for comparing lists with Venn Diagrams http://bioinfogp.cnb.csic.es/tools/venny/index.html.
Anderson MJ, Crist TO, Chase JM, Vellend M, Inouye BD, Freestone AL et al (2011). Navigating the multiple meanings of beta-diversity: a roadmap for the practicing ecologist. Ecol Lett 14: 19–28.
Beck T, Joergensen G, Kandeler E, Makeschin F, Nuss E, Oberholzer HR et al (1997). An inter-laboratory comparison of ten different ways of measuring soil microbial biomass C. Soil Biol Biochem 29: 1023–1032.
Bell G . (2001). Neutral macroecology. Science 293: 2413–2418.
Bonan GB, Shugart HH . (1989). Environmental factors and ecological processes in boreal forests. Annu Rev Ecol Syst 20: 1–28.
Bond WJ, Woodward FI, Midgley GF . (2005). The global distribution of ecosystems in a world without fire. New Phytol 165: 525–538.
Bray JR, Curtis JT . (1957). An ordination of the upland forest communities of southern Wisconsin. Ecol Monogr 27: 325–349.
Brookes PC, Landman A, Pruden G, Jenkinson DS . (1985). Chloroform fumigation and the release of soil nitrogen: a rapid direct extraction method to measure microbial biomass nitrogen in soil. Soil Biol Biochem 17: 837–842.
Burke C, Thomas T, Lewis M, Steinberg P, Kjelleberg S . (2011). Composition, uniqueness and variability of the epiphytic bacterial community of the green alga Ulva australis. ISME J 5: 590–600.
Bárcenas-Moreno G, García-Orenes F, Mataix-Solera J, Mataix-Beneyto J, Bååth E . (2011). Soil microbial recolonisation after a fire in a Mediterranean forest. Biol Fert Soils 47: 261–272.
Cadotte MW . (2007). Concurrent niche and neutral processes in the competition–colonization model of species coexistence. Proc R Soc B 274: 2739–2744.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger J, Bushman FD, Costello EK et al (2010). QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7: 335–336.
Caruso T, Chan Y, Lau MCY, McKay CP, Pointing SB . (2011). Stochastic and deterministic processes interact in the assembly of desert microbial communities on a global scale. ISME J 5: 1406–1413.
Certini G . (2005). Effects of fire on properties of forest soils: a review. Oecologia 143: 1–10.
Chase JM, Myers JA . (2011). Disentangling the importance of ecological niches from stochastic processes across scales. Philos Trans R Soc Lond B Biol Sci 366: 2351–2363.
Chase JM . (2003). Community assembly: when should history matter? Oecologia 136: 489–498.
Chase JM . (2007). Drought mediates the importance of stochastic community assembly. Proc Natl Acad Sci USA 104: 17430–17434.
Chytrý M, Lososová Z, Horsák M, Uher B, Čejka T, Danihelka J et al (2012). Dispersal limitation is stronger in communities of microorganisms than macroorganisms across Central European cities. J Biogeography 39: 1101–1111.
DeLuca TH, Sala A . (2006). Frequent fire alters nitrogen transformations in ponderosa pine stands of the Inland Northwest. Ecology 87: 2511–2522.
Denslow JS . (1980). Gap portioning among tropical rainforest trees. Biotropica 12: 47–55.
Didham RK, Norton DA . (2006). When are alternative stable states more likely to occur? Oikos 113: 357–362.
Didham RK, Watts CH, Norton DA . (2005). Are systems with strong underlying abiotic regimes more likely to exhibit alternative stable states? Oikos 110: 409–416.
Edgar RC . (2010). Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26: 2460–2461.
Ellner SP, Fussmann G . (2003). Effects of successional dynamics on metapopulation persistence. Ecology 84: 882–889.
Ferrenberg SM, Schwilk DW, Knapp EE, Groth E, Keeley JE . (2006). fire decreases arthropod abundance but increases diversity: early and late season prescribed fire effects in a sierra nevada mixed-conifer forest. Fire Ecol 2: 79–102.
Fierer N, Jackson R . (2006). The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci USA 103: 626–631.
Hamman ST, Burke IC, Knapp EE . (2008). Soil nutrients and microbial activity after early and late season prescribed burns in a Sierra Nevada mixed conifer forest. Forest Ecol Manag 256: 367–374.
Hanson CA, Fuhrman J, Horner-Devine MC, Martiny JBH . (2012). Beyond biogeographic patterns: processes shaping the microbial landscape. Nat Rev Microbiol 10: 497–506.
Hart SC, DeLuca TH, Newman GS, MacKenzie MD, Boyle SI . (2005). Post-fire vegetative dynamics as drivers of microbial community structure and function in forest soils. For Ecol Manage 220: 166–184.
Hubbell SP . (2001) The Unified Neutral Theory Of Biodiversity And Biogeography. Princeton University Press: Princeton, NJ.
Jiang L, Patel SN . (2008). Community assembly in the presence of disturbance: a microcosm experiment. Ecology 89: 1931–1940.
JMP Version 10 (2011). SAS Institute Inc.: Cary, NC.
Knelman JE, Legg TM, O’Neill SP, Washenberger CL, González A, Cleveland CC et al (2012). Bacterial community structure and function change in association with colonizer plants during early primary succession in a glacier forefield. Soil Biol Biochem 46: 172–180.
Kraft NJB, Cornwell WK, Webb CO, Ackerly DD . (2007). Trait evolution, community assembly, and the phylogenetic structure of ecological communities. Am Nat 170: 271–283.
Langenheder S, Bulling MT, Solan M, Prosser JI . (2010). Bacterial biodiversity-ecosystem functioning relations are modified by environmental complexity. PloS one 5: 10834.
Langenheder S, Prosser JI . (2008). Resource availability influences the diversity of a functional group of heterotrophic soil bacteria. Environ Microbiol 10: 2245–2256.
Langenheder S, Székely AJ . (2011). Species sorting and neutral processes are both important during the initial assembly of bacterial communities. ISME J 5: 1086–1094.
Leibold MA, McPeek MA . (2006). Coexistence of the niche and neutral perspectives in community ecology. Ecology 8: 1399–1410.
Logue JB, Lindström ES . (2010). Species sorting affects bacterioplankton community composition as determined by 16S rDNA and 16S rRNA fingerprints. ISME J 4: 729–738.
Lozupone C, Knight R . (2005). ‘UniFrac: a new phylogenetic method for comparing microbial communities’. Appl Environ Microbiol 71: 8228–8235.
Lozupone C, Hamady M, Knight R . (2006). UniFrac – an online tool for comparing microbial community diversity in a phylogenetic context. BMC bioinformatics 7: 371.
Lozupone CA, Knight R . (2007). Global patterns in bacterial diversity. Proc Natl Acad Sci USA 104: 11436–11440.
Martiny JBH, Bohannan BJM, Brown JH, Colwell RK, Fuhrman JA, Green JL et al (2006). Microbial biogeography: putting microorganisms on the map. Nat Rev Microbiol 4: 102–112.
McCune B, Mefford MJ . (2011) PC-ORD. Multivariate Analysis of Ecological Data. Version 6.0. MjM Software: Gleneden Beach, OR.
Monson RK, Lipson DL, Burns SP, Turnipseed AA, Delany AC, Williams MK et al (2006). Winter forest soil respiration controlled by climate and microbial community composition. Nature 439: 711–714.
Mulvaney RL . (1996). Nitrogen - Inorganic forms. In: Sparks, DL, et al. (ed) Methods of soil analysis, Part 3, SSSA Book Series 5. SSSA: Madison, WI, pp 1123–1184.
Nemergut DR, Anderson SP, Cleveland CC, Martin AP, Miller AE, Seimon A et al (2007). Microbial community succession in an unvegetated, recently-deglaciated soil. Microb Ecol 53: 110–122.
Nemergut DR, Cleveland CC, Wieder WR, Washenberger CL, Townsend AR . (2010). Plot-scale manipulations of organic matter inputs to soils correlate with shifts in microbial community composition in a lowland tropical rain forest. Soil Biol Biochem 42: 2153–2160.
Nemergut DR, Costello EK, Hamady M, Lozupone C, Jiang L, Schmidt SK et al (2011). Global patterns in the biogeography of bacterial taxa. Environ Microbiol 13: 135–144.
Ofiteru ID, Lunn M, Curtis TP, Wells GF, Criddle CS, Francis CA et al (2010). Combined niche and neutral effects in a microbial wastewater treatment community. Proc Natl Acad Sci USA 107: 15345–15350.
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’Hara RB et al (2011). Vegan: Community Ecology Package. R package version 2: 0–1.
Östman Ö, Drakare S, Kritzberg ES, Langenheder S, Logue JB, Lindström ES . (2009). Regional invariance among microbial communities. Ecology letters 13: 118–127.
Paul EA, Clark FE . (1996). Soil Microbiology and Biochemistry, 2nd Edition. Academic Press: New York, NY.
Peay KG, Garbelotto M, Bruns TD . (2010). Evidence of dispersal limitation in soil microorganisms: isolation reduces species richness on mycorrhizal tree islands. Ecology 91: 3631–3640.
Peitikäinen J, Hiukka R, Fritze H . (2000). Does short-term heating of forest humus change its properties as a substrate for microbes? Soil Biol Biochem 32: 277–288.
Reeder J, Knight R . (2010). Rapidly denoising pyrosequencing amplicon reads by exploiting rank-abundance distributions. Nat Methods 7: 688–689.
Sattin SR, Cleveland CC, Hood E, Reed SC, King AJ, Schmidt SK et al (2009). Functional shifts in unvegetated, perhumid, recently deglaciated soils do not correlate with shifts in soil bacterial community composition. J Microbiol 47: 673–681.
Schmidt SK, Costello EK, Nemergut DR, Cleveland CC, Reed SC, Weintraub MN et al (2007). Biogeochemical consequences of rapid microbial turnover and seasonal succession in soil. Ecology 88: 1379–1385.
Schoennagel T, Sherriff RL, Veblen TT . (2011). Fire history and tree recruitment in the Colorado Front Range upper montane zone: implications for forest restoration. Ecol Appl 21: 2210–2222.
Sloan WT, Mary Lunn M, Woodcock S, Head IM, Nee S, Curtis TP . (2006). Quantifying the roles of immigration and chance in shaping prokaryote community structure. Env Microbiol 8: 732–740.
Smith NR, Kishchuk BE, Mohn WW . (2008). Effects of wildfire and harvest disturbances on forest soil bacterial communities. Appl Environ Microbiol 74: 216–224.
Team RDC . (2011) R: A Language And Environment For Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria.
Telford RJ, Vandvik V Birks HJB (2006). Dispersal limitation matters for microbial morphospecies. Science 312: 1015.
Turner MG, Smithwick EAH, Metzger KL, Tinker DB, Romme WH . (2007). Inorganic nitrogen availability following severe stand-replacing fire in the Greater Yellowstone Ecosystem. Proc Natl Acad Sci USA 104: 4782–4789.
Van Der Heijden MGA, Bardgett RD, Van Straalen NM . (2008). The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett 11: 296–310.
Veblen TT, Kitzberger T, Donnegan J . (2000). Climatic and human influences on fire regimes in ponderosa pine forests in the Colorado Front Range. Ecol Appl 10: 1178–1195.
Vellend M . (2010). Conceptual synthesis in community ecology. Q Rev Biol 85: 183–206.
Waldrop MP, Harden JW . (2008). Interactive effects of wildfire and permafrost on microbial communities and soil processes in an Alaskan black spruce forest. Global Change Biol 14: 2591–2602.
Wang GG, Kemball KJ . (2005). Effects of fire severity on early development of understory vegetation. Can J Forest Res 35: 254–262.
Wang Q, Zhong M, Wang S . (2012). A meta-analysis on the response of microbial biomass, dissolved organic matter, respiration, and N mineralization in mineral soil to fire in forest ecosystems. Forest Ecol Manag 271: 91–97.
Wardle DA, Zackrisson O, Hörnberg G, Gallet C . (1997). The influence of island area on ecosystem properties. Science 277: 1296–1299.
Wardle DA, Zackrisson O, Nilsson MC . (1998). The charcoal effect in Boreal forests: mechanisms and ecological consequences. Oecologia 115: 419–426.
Woodcock S, Van Der Gast CJ, Bell T, Lunn M, Curtis TP, Head IM et al (2007). Neutral assembly of bacterial communities. FEMS Microbiol Ecol 62: 171–180.
Yeager CM, Northrup DE, Grow CC, Barns SM, Kuske CR . (2005). Changes in nitrogen-fixing and ammonia-oxidizing bacterial communities in soil of a mixed conifer forest after wildfire. Appl Environ Microbiol 71: 2713–2722.
Yilmaz P, Kottmann R, Field D, Knight R, Cole JR, Amaral-Zettler L et al (2011). Minimum information about a marker gene sequence (MIMARKS) and minimum information about any (x) sequence (MIxS) specifications. Nat Biotechnol 29: 415–420.
Zak DR, Holmes WE, White DC, Peacock AD, Tilman D . (2003). Plant diversity, soil microbial communities, and ecosystem function: are there any links? Ecology 84: 2042–2050.
Acknowledgements
This research was supported, in part, through grants from the National Science Foundation to DN, SS and CC (DEB1545913), DEB1120281 to LJ and the Niwot Ridge LTER grant (DEB-1027341) to MW, AT, and SS. This research used the Janus supercomputer, which is supported by the National Science Foundation (award number CNS-0821794) and the University of Colorado Boulder. The Janus supercomputer is a joint effort of the University of Colorado Boulder, the University of Colorado Denver and the National Center for Atmospheric Research. We thank Timothy Seastedt for comments on our manuscript and the University of Colorado’s Department of Ecology and Evolutionary Biology for generous support. We acknowledge Bob Head and Bill Nemergut (WFN) for assisting with sample collection and processing. Our manuscript was greatly improved by comments and suggestions from three anonymous reviewers and the editor.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
Ferrenberg, S., O'Neill, S., Knelman, J. et al. Changes in assembly processes in soil bacterial communities following a wildfire disturbance. ISME J 7, 1102–1111 (2013). https://doi.org/10.1038/ismej.2013.11
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2013.11
Keywords
This article is cited by
-
Elevated methane flux in a tropical peatland post-fire is linked to depth-dependent changes in peat microbiome assembly
npj Biofilms and Microbiomes (2024)
-
Bacterial community network complexity and role of stochasticity decrease during primary succession
Soil Ecology Letters (2024)
-
Spatial distribution pattern across multiple microbial groups along an environmental stress gradient in tobacco soil
Annals of Microbiology (2023)
-
Implications of Soil Microbial Community Assembly for Ecosystem Restoration: Patterns, Process, and Potential
Microbial Ecology (2023)
-
Assembly Processes and Biogeographical Characteristics of Soil Bacterial Sub-communities of Different Habitats in Urban Green Spaces
Current Microbiology (2023)