Abstract
The two-step nitrification process is an integral part of the global nitrogen cycle, and it is accomplished by distinctly different nitrifiers. By combining DNA-based stable isotope probing (SIP) and high-throughput pyrosequencing, we present the molecular evidence for autotrophic growth of ammonia-oxidizing bacteria (AOB), ammonia-oxidizing archaea (AOA) and nitrite-oxidizing bacteria (NOB) in agricultural soil upon ammonium fertilization. Time-course incubation of SIP microcosms indicated that the amoA genes of AOB was increasingly labeled by 13CO2 after incubation for 3, 7 and 28 days during active nitrification, whereas labeling of the AOA amoA gene was detected to a much lesser extent only after a 28-day incubation. Phylogenetic analysis of the 13C-labeled amoA and 16S rRNA genes revealed that the Nitrosospira cluster 3-like sequences dominate the active AOB community and that active AOA is affiliated with the moderately thermophilic Nitrososphaera gargensis from a hot spring. The higher relative frequency of Nitrospira-like NOB in the 13C-labeled DNA suggests that it may be more actively involved in nitrite oxidation than Nitrobacter-like NOB. Furthermore, the acetylene inhibition technique showed that 13CO2 assimilation by AOB, AOA and NOB occurs only when ammonia oxidation is not blocked, which provides strong hints for the chemolithoautotrophy of nitrifying community in complex soil environments. These results show that the microbial community of AOB and NOB dominates the nitrification process in the agricultural soil tested.
Similar content being viewed by others
Introduction
Nitrification is the only oxidative process that links the reduced and oxidized pools of inorganic nitrogen to sustain the global nitrogen cycle (Gruber and Galloway, 2008). The rate-limiting step in microbial nitrification is the oxidation of ammonia to nitrite, which is then rapidly oxidized further to nitrate (Koops et al., 2006). Soil ammonia oxidation has long been considered to be performed by only a few phylogenetically restricted microorganisms belonging to Proteobacteria (Kowalchuk and Stephen, 2001). However, recent studies have expanded the known ammonia oxidizers from the domain Bacteria to Archaea in marine habitats (Könneke et al., 2005) and soil (Treusch et al., 2005). The widespread presence of archaeal amoA gene (Francis et al., 2005; Leininger et al., 2006; Erguder et al., 2009) and autotrophic lifestyle of cultured ammonia-oxidizing archea (AOA) (Könneke et al., 2005; Hatzenpichler et al., 2008) have suggested its important role in the global cycling of nitrogen and carbon. However, the relative contribution of AOA and ammonia-oxidizing bacteria (AOB) to ammonia oxidation in agricultural soil remains unclear. For instance, AOB has been shown to dominate the microbial ammonia oxidation in German agricultural soil (Jia and Conrad, 2009), whereas archaeal ammonia oxidation prevails in Scottish agricultural soil (Offre et al., 2009; Zhang et al., 2010).
Unlike AOB, the ecological study of nitrite-oxidizing bacteria (NOB) has been hampered because there is no universal 16S rRNA gene probe that can specifically target the highly diverse NOB community in complex soil environments (Freitag et al., 2005). In addition to the newly discovered Nitrotoga of Betaproteobacteria (Alawi et al., 2007), NOB is composed of four genera, namely Nitrobacter, Nitrococcus, Nitrospina and Nitrospira, which are phylogenetically affiliated to Alpha-, Gamma-, provisionally Deltaproteobacteria and the deep-branching bacterial phylum Nitrospirae, respectively (Bock and Wagner, 2006). For a long time, Nitrobacter has been studied most intensively and is believed to play a dominant role in nitrification, but an increasing line of evidence suggests the functional importance of Nitrospira. A recent study has shown that Nitrobacter-like NOB responds positively to high N availability and high nitrite oxidation potential, whereas Nitrospira-like NOB seems favored when soil nitrite oxidation activity is low (Attard et al., 2010). The autotrophic lifestyle of known NOB in culture makes it suitable for DNA-stable isotope probing (SIP) investigation to directly link NOB identity with nitrification activity in a way similar to that of AOB in agricultural soil (Jia and Conrad, 2009). Instead of focusing on the NOB-specific 16S rRNA gene, barcoded pyrosequencing of the total 16S rRNA in 13C-labeled DNA could provide useful insights into the relative role of Nitrospira and Nitrobacter in soil nitrification in agricultural ecosystems.
Agricultural ecosystems annually receive approximately 25% of global nitrogen input, mostly in the form of ammonium (Gruber and Galloway, 2008). The ammonium must be oxidized at least once to nitrate by a nitrifying community. A comprehensive understanding of microbial nitrification requires simultaneous characterization of AOB, AOA and NOB. However, no study appears to have been able to clearly link nitrite oxidation activity to the phylogenetic identity of Nitrobacter and/or Nitrospira in agricultural soil. In fact, soil nitrite concentration is generally below the detection limit, indirectly suggesting that some NOB may have high substrate affinity and are, therefore, extremely efficient for nitrite oxidation. As for ammonia oxidizers, direct evidence is still missing for ammonia oxidation by group I.1b AOA, which is commonly observed in soil (Schleper, 2010). Notably, active AOA in Scottish soil is phylogenetically most closely associated with group I.1a, which is typically dominant in marine habitats (Offre et al., 2009). Taking into account the recent conflicting observations (Gubry-Rangin et al., 2010), an investigation with regard to microbial nitrification in geographically broader regions is needed to assess accurately the global relevance of AOA and AOB in nitrogen cycling in agricultural ecosystem.
Therefore, by combining 13CO2-based DNA-SIP and high-throughput 454 pyrosequencing techniques, we aim to investigate the autotrophic growth of active nitrifying community upon ammonium fertilization in agricultural soil from a region typical of agricultural areas in the North China Plains. In addition, C2H2 inhibition technique was also exploited in an attempt to determine whether 13CO2 assimilation would occur in the absence of soil nitrification, which is thought to provide the sole energy source in support of the lithotrophic growth of the nitrifying community.
Materials and methods
DNA-SIP microcosm
Soil samples were collected in triplicate from the State Key Experimental Station for Ecological Agriculture (35°00′ N, 114°24′ E) of the Chinese Academy of Sciences; the field information is detailed in Supplementary Materials. DNA-SIP microcosm was constructed in triplicate to investigate the active soil-nitrifying community as previously described in detail (Jia and Conrad, 2009). Briefly, 6.58 g of fresh soil (equivalent to 6.0 g dry weight gram soil) was incubated at 60% maximum water-holding capacity in 100 ml serum bottles, which were tightly capped with black butyl stoppers at 27 °C in the dark. Incubation of the soil microcosm was performed with 5% of 13CO2 or 12CO2 with or without 100 Pa C2H2, a suicide substrate inhibitor of microbial ammonia oxidation. The treatment was renewed once a week, including the addition of 100 μg (NH4)2SO4-N g dry weight gram soil and of 5% 13CO2 or 12CO2. Thus, the soil microcosms received a total of 400 μg (NH4)2SO4-N g dry weight gram soil over an incubation period of 4 weeks. The headspace of the bottle was flushed weekly with pressurized synthetic air (20% O2, 80% N2) for 1 min to maintain oxic conditions, and, if stated, the addition of 100 Pa C2H2 was performed immediately after the headspace air was fully exchanged. The 13CO2 (99 atom% carbon) was purchased from Sigma-Aldrich Co. (St Louis, MO, USA) and 12CO2 was made by acidifying sodium carbonate. The headspace CO2 concentration was measured as described previously (Meng et al., 2005). Before incubation of the DNA-SIP microcosm, a 12-day pre-incubation of soil at 40% maximum water-holding capacity was carried out to reduce the amount of soil-respired CO2 so that the target concentration of 5% CO2 could be maintained at a constant level over the entire incubation period. The pre-incubation also increased the labeling efficacy of targeted microorganisms because the dilution of 13CO2 by soil-respired 12CO2 could be decreased significantly as reported previously (Jia and Conrad, 2009). In addition, a 3-day pre-incubation with 100 Pa C2H2 was executed to fully inactivate the AOB community for all the C2H2 treatments immediately before SIP incubation was started.
The destructive sampling was performed in triplicate at days 3, 7 and 28. About 2.0 g of fresh soil was removed from each replicate and immediately frozen at −20 °C for molecular analysis. The rest of the soil replicate was homogenized with 10 ml of 2 M KCl by shaking at 200 r.p.m. for 30 min, and then passed through filter paper for the determination of NH4+-N, NO2−-N and NO3−-N using a Skalar SAN Plus segmented flow analyzer (Skalar Inc., Breda, The Netherlands). The nitrite concentration was below the detection limit and nitrification was assessed by the changes in NH4+-N and NO3−-N content in the soil. These were then calculated based on oven-dried soil weight.
Nucleic acid extraction and SIP gradient fractionation
Soil DNA was extracted using a FastDNA spin kit for soil (MP Biomedicals, Cleveland, OH, USA), according to the manufacturer's instruction. Soil DNA quantity and purity were determined by a Nanodrop ND-1000 UV–Vis Spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA). SIP fractionation was performed as described previously (Neufeld et al., 2007). For each treatment, 2.0 μg of soil DNA extract was mixed well with CsCl stock solution to achieve an initial CsCl buoyant density of 1.725 g ml−1 before ultracentrifugation at 177 000 g for 44 h at 20 °C. DNA fractionation was carried out by displacing the gradient medium with sterile water from the top of the ultracentrifuge tube using an NE-1000 single syringe pump (New Era Pump Systems Inc., Farmingdale, NY, USA) with a precisely controlled flow rate of 0.38 ml min −1. Up to 14 or 15 DNA gradient fractions were generated with equal volumes of about 380 μl, and a 65 μl aliquot of each fraction was used for refractive index measurement using an AR200 digital hand-held refractometer (Reichert, Inc., Buffalo, NY, USA). The fractionated DNA was purified and dissolved in 30 μl TE buffer as discussed previously (Jia and Conrad, 2009).
Real-time quantitative PCR
Real-time quantitative PCR was performed in triplicate to verify the efficacy of 13C incorporation into the genomes of the autotrophic ammonia-oxidizing prokaryotes by analyzing the copy number of bacterial and archaeal amoA genes in the fractionated DNA across the entire buoyant density gradients from DNA-SIP microcosms as detailed previously (Jia and Conrad, 2009) on a CFX96 Optical Real-Time Detection System (Bio-Rad, Laboratories Inc., Hercules, CA, USA). Bacterial and archaeal 16S rRNA genes were also quantified using domain-specific primers. The primers and PCR conditions are detailed in Supplementary Table S6. The real-time PCR standard was generated using plasmid DNA from one representative clone containing amoA gene or 16S rRNA genes of Bacteria and Archaea, and a dilution series of standard template over six to eight orders of magnitude per assay was used to optimize the real-time PCR conditions. Blanks were always run with water as the template instead of soil DNA extract. Specific amplification was verified by melting curve analysis, which always results in a single peak, and by agarose gel electrophoresis.
Pyrosequencing, cloning and phylogenetic analysis
Pyrosequencing was carried out on a Roche 454 GS FLX Titanium sequencer (Roche Diagnostics Corporation, Branford, CT, USA) by analyzing the V4 regions of 16S rRNA genes in the fractionated DNA across the entire buoyant density gradients from the 13CO2-treated SIP microcosms with or without C2H2. Taq sequences were used to barcode the PCR amplicons in each DNA gradient fraction from labeled and control treatments (Supplementary Tables S1 and S2). Triplicate amplicons were pooled, purified and visualized on 1.2% agarose gels. The concentration of purified PCR amplicons was determined, and the purified PCR amplicons were then combined in equimolar ratios into a single tube in preparation for pyrosequencing analysis. In addition, pyrosequencing of the archaeal 16S rRNA genes was performed using the archaea-specific primer shown in Supplementary Table S6.
In total, 196 000 high-quality sequences reads were obtained after removing about 20% of the original reads by RDP online pyrosequencing analysis (Cole et al., 2009). Taxonomy of the high-quality sequence reads was assigned using the RDP classifier with a minimum support threshold of 60% and the RDP taxonomic nomenclature (Cole et al., 2009), and only sequence reads highly similar to the putative nitrifying microorganisms were further analyzed. From each sample of the 28 DNA gradient fractions (Supplementary Table S1) and 13 DNA gradient fractions (Supplementary Table S2), all sequences affiliated with AOB, NOB and crenarchaeota or unclassified archaea by the RDP classifier were extracted. The targeted sequence reads from the 13C-labeled ‘heavy’ DNA fractions were clustered into operational taxonomic unit (OTU) at 97% cutoff using the mothur software (Schloss et al., 2009), and the OTUs containing less than 10 reads were removed. A representative sequence was then used from each OTU for phylogenetic analysis.
Clone libraries of 16S rRNA and amoA genes were also constructed from the SIP microcosm after incubation for 4 weeks. Bacterial and archaeal 16S rRNA genes were amplified in the 13C-labeled ‘heavy’ DNA fractions for clone library construction using the universal primers 27f-1492r (Lane, 1991) and Arch21f-Arch958R (DeLong, 1992) as detailed in Supplementary Table S6. The bacterial amoA genes were amplified in the 13C-labeled ‘heavy’ DNA fractions and the ‘light’ DNA fractions from the 13CO2-labeled microcosm for clone library construction (each containing 25 clones) using amoA-1F/amoA-2R assay (Rotthauwe et al., 1997). PCR amplicons of archaeal amoA genes were obtained in both the 13C-labeled ‘heavy’ DNA fractions and the total DNA from the 13CO2-labeled microcosm for clone library construction (each containing 20 clones). Triplicate PCR products were pooled and purified before cloning. The Escherichia coli JM109-competent cells were used for transformation. Sequencing of the clones containing the correct insert was performed by the Invitrogen Sequencing Department (Invitrogen, Shanghai, China) and analyzed by DNAstar software package (DNASTAR, Inc., Madison, WI, USA).
Phylogenetic analysis was performed using the Molecular Evolutionary Genetics Analysis (MEGA 4.0) software package (Kumar et al., 2004). For the bacterial 16S rRNA sequence highly affiliated with AOB, the basic tree of sequences larger than 1.4 kb was constructed through a neighbor-joining algorithm. The closest relatives to the 13C-labeled AOB sequences in this study and those from literature were then selected to reconstruct a phylogenetic tree inferred from the distance matrix using the Fitch algorithm with global rearrangement and a randomized input order of sequences. The tree topology was checked by the neighbor-joining algorithm and the minimum evolution method (Kumar et al., 2004). Phylogenetic analysis of the bacterial and archaeal amoA genes were performed as described previously (Jia and Conrad, 2009).
Accession numbers of nucleotide sequences
The nucleotide sequences have been deposited at GenBank with accession numbers HQ678241–HQ678248 and HQ678202–HQ678208 for 16S rRNA genes of AOA and AOB, respectively, as well as HQ678187–HQ678201 and HQ678209–HQ678240 for the amoA genes of AOA and AOB, respectively, from the DNA-SIP experiment. The pyrosequencing reads have been deposited at GenBank with accession number DRA000318.
Results
SIP incubation was established by feeding the soil microcosm with 5% 13CO2 or 5% 12CO2 with or without 100 Pa C2H2. Ammonium fertilization with 100 μg NH4+-N g dry weight gram soil was performed once a week at days 0, 7, 14 and 21 as described previously (Jia and Conrad, 2009). In the absence of C2H2, active nitrification activity was observed through significant nitrate production, which was completely abolished in the C2H2-treated soil microcosms at days 3, 7 and 28 (Figure 1). At day 28, the recovered ammonium concentration in the C2H2-treated soil was largely consistent with the nitrate produced in the soil in the absence of C2H2, suggesting the effective inhibition of C2H2 on soil nitrification activity. The total DNA was extracted and subjected to isopycnic centrifugation for each of the four different treatments (13CO2, 12CO2, 13CO2+C2H2 and 12CO2+C2H2) at days 3, 7 and 28 in triplicates for molecular analysis.
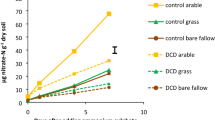
Change in concentrations of soil nitrate (a) and ammonium (b) in surface agricultural soil incubated with either 12CO2 or 13CO2 in the presence or absence of C2H2 after incubation for 3, 7 and 28 days. The error bars of soil nitrate and ammonium contents indicate standard deviations of triplicate microcosm incubations.
Real-time quantitative PCR indicated that the copy number of bacterial amoA genes peaked in the DNA fractions with buoyant densities with the 13CO2-labeled treatment having a slightly higher value than those from the 12CO2 control treatment. This implies that the genomes of AOB cells were partially labeled in the 13CO2-treated soils after incubation for 3 and 7 days (Figure 2). At day 28, the majority of bacterial amoA gene was observed in the heavier DNA fractions, indicating a much greater degree of labeling of the AOB cells during active nitrification (Figures 2 and 3a). Similarly, real-time quantitative PCR revealed that the abundance of archaeal amoA (Figure 3b) and 16S rRNA genes (Figure 3c) were significantly higher in the ‘heavy’ DNA fractions from the 13CO2-labeled treatment than those from the control treatments (that is, 12CO2, 12CO2+C2H2 and 13CO2+C2H2) at day 28. However, the labeling of archaeal amoA and 16S rRNA genes was not detected at days 3 and 7. In addition, the PCR signals of archaeal amoA and 16S rRNA genes in the ‘heavy’ DNA fractions were clearly visualized on agarose gel from the 13CO2-labeled treatment, but not from the 12CO2 control treatment (Figures 3b and c).
Quantitative distribution of bacterial amoA gene copy numbers across the entire buoyant density gradient of the fractionated DNA from the soil incubated with either 13CO2 or 12CO2 without C2H2 after incubation for 3, 7 and 28 days. The normalized data are the ratio of the gene copy number in each DNA gradient fraction to the maximum quantities from each treatment, as described previously (Lueders et al., 2004). For clarity, the standard error bar from the triplicate microcosm incubations was removed.
Quantitative distribution of the amoA gene copy numbers of Bacteria (a), Archaea (b) and of the archaeal 16S rRNA gene copy number (c) across the entire buoyant density gradient of the fractionated DNA from the soil incubated with either 12CO2 or 13CO2 in the presence or absence of C2H2 after incubation for 28 days. For clarity, the standard error bar from the triplicate microcosm incubations was removed, whereas it is shown in the insets. Strong signals of the PCR-amplified archaeal amoA (b) and 16S rRNA genes (c) were visualized by agarose gel electrophoresis in the 13C-labeled DNA fraction-6 from three biological replicates of the 13CO2-treated soil, but not from the 12CO2-treated soil. P, N and M denote positive control, negative control and DNA marker, respectively.
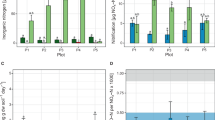
Pyrosequencing analysis of the total microbial community was performed in DNA gradient fractions from the labeled microcosm (13CO2) and the control treatment (13CO2+C2H2) at day 28 (Supplementary Tables S1 and S2). About 196 000 high-quality sequence reads were obtained with an average length of about 370 bp in the V4 region of the 16S rRNA gene. Across the entire DNA gradient, the highly enriched 16S rRNA gene sequence of AOB (Figure 4a) and of NOB (Figure 4b) were only observed in the labeled microcosm in which ammonia oxidation was not blocked by C2H2 (Figure 1). Up to 13.2% of bacterial 16S rRNA gene sequences in the 13C-labeled DNA were closely related to the Nitrosospira cluster 3 (Figure 5a), whereas sequences affiliated with the Nitrosomonas sp. comprised 4.5% of the total bacterial community in the 13C-labeled DNA fraction (Figure 5b). Similarly, NOB consisted of Nitrospira- and Nitrobacter-like 16S rRNA reads, accounting for 27% and 15% of total 16S rRNA sequences in the 13C-labeled DNA, respectively (Figures 5c and d). Phylogenetic analysis indicated that the majority of Nitrospira-like sequences could be grouped into five OTUs, accounting for 95% of the total NOB sequences at 97% sequence similarity (Supplementary Table S3). On the other hand, more than 98% of the Nitrobacter-like sequences were assigned to a single OTU group (Supplementary Table S4), which was most closely affiliated with the NOB enriched from wastewater treatment plant (Alawi et al., 2009). A similar result was observed for the uncultured crenarchaeal 16S rRNA reads, of which the relative frequency was apparently higher in the ‘heavy’ DNA fractions from the labeled microcosm than those in the control microcosms (Figure 4c). In addition, the crenarchaeal 16S rRNA reads in the labeled microcosm appeared to have a slightly higher buoyant density compared with those in the control microcosm (Figure 4c). This may imply an incremental labeling of uncultured crenarchaeota during active nitrification, despite the fact that the 13C label seems to be assimilated to a much lesser extent compared with the bacterial amoA gene (Figure 2).
Relative frequency of the 16S rRNA gene sequence reads affiliated with ammonia-oxidizing bacteria (a), nitrite-oxidizing bacteria (b) and uncultured crenarchaeota (c) across the entire buoyant density gradient of the fractionated DNA from the 13CO2-treated soil after incubation for 28 days with or without 100 Pa acetylene. The abundance is expressed as the percentage of the targeted sequence reads to the total 16S rRNA gene reads in each DNA gradient fraction.
Changes in the relative abundance of the 16S rRNA gene sequence reads affiliated with Nitrosospira (a) and Nitrosomonas (b) of the AOB community, and Nitrospira (c) and Nitrobacter (d) of the NOB community across the entire buoyant density gradient of the fractionated DNA from the 13CO2-treated soil after incubation for 28 days with or without 100 Pa acetylene. The relative abundance is expressed as the percentage of the targeted sequence reads to the total 16S rRNA gene sequence reads in each DNA gradient fraction.
Clone library construction of the bacterial amoA and 16S rRNA genes was performed in the 13C-labeled and native DNA fractions, as described previously (Jia and Conrad, 2009). Out of the 62 clones containing the near full-length bacterial 16S rRNA gene fragments, seven were identified as AOB sequences in the 13C-labeled DNA (Supplementary Table S5). Phylogenetic analysis revealed that the active AOB were exclusively related to the Nitrosospira cluster 3 (Supplementary Figure S1). However, a molecular survey of the amoA gene indicated that two out of the 25 cloned amoA sequences were affiliated with the Nitrosomonas sp. Nm 58, whereas the other 23 sequences were placed within the Nitrosospira cluster 3 (Supplementary Figure S2). This observation was largely consistent with the pyrosequencing results (Supplementary Figure S1).
Clone libraries of archaeal 16S rRNA and amoA genes were also constructed from the 13C-labeled DNA (Supplementary Figure S3). Phylogenetic analysis indicated that sequences affiliated with a moderately thermophilic AOA of N. gargensis accounted for the majority of archaeal amoA (33 out of 34 clones) and 16S rRNA genes (33 out of 64 clones). Pyrosequencing analysis of the archaeal 16S rRNA genes in the 13C-labeled DNA reveled that 2918 out of 5504 reads were closely related to N. gargensis (Supplementary Figure S4A and Supplementary Table S2). This was supported by the specific enrichment (517 out of 1080) of sequences that were highly similar with N. gargensis when the total 16S rRNA gene was analyzed (Supplementary Figure S4B and Supplementary Table S1).
Discussion
Microbial nitrification can serve as an ideal model for investigating community metabolism in complex soil environments by directly linking nitrification activity to the phylogenetic identity of physiologically distinct microorganisms. Using a SIP-targeted pyrosequencing approach, AOB and NOB were labeled to a much greater extent than AOA during active nitrification. Furthermore, the results in this study indicate that 13CO2 assimilation by AOB, AOA and NOB occurred only when nitrification was not inhibited, which suggests the chemolithoautotrophy of nitrifying community in the agricultural soil tested. These results imply that Bacteria dominated the nitrification activity in the agricultural soil tested, which is consistent with previous findings in German agricultural soil (Jia and Conrad, 2009).
The time-course incubation of SIP microcosms revealed that both AOB and AOA assimilated 13CO2 during active nitrification. The essential prerequisite for autotrophic ammonia oxidation is thought to be the amoA gene (Hyman and Arp, 1992). As shown in Supplementary Table S5, the amoA gene copy numbers of AOA and AOB in the ‘heavy’ DNA fractions from the labeled microcosm were generally one order of magnitude higher than those from the control microcosm. The 13C-labeled AOB amoA genes accounted for 56.4% of the total AOB amoA gene abundance, whereas a constant background of about 2.62% was observed in the control treatments. With respect to AOA, 9.41% of the total archaeal amoA genes were retrieved in the 13C-labeled DNA (Supplementary Table S5) and 0.40–0.55% was found in the heavy fractions as background from the control microcosms. Similar observations were obtained for the 16S rRNA gene quantification of Bacteria and Archaea (Supplementary Table S5). Pyrosequencing has further suggested the exceptionally high abundance of AOB and NOB, up to 51.2% of the total microbial community in the 13C-labeled DNA (Figure 4). These findings provide strong hints that nitrification was primarily catalyzed by AOB and NOB in the agricultural soil tested. Assuming the ecological significance of AOA and AOB could be better revealed by the relative abundance of bacterial versus archaeal amoA genes in the 13C-labeled DNA, the archaeal contribution to nitrification activity was then estimated by multiplying the number of 13C-labeled amoA-containing AOA cells in the soil with the maximum ammonia oxidation rates estimated from known AOA cultures in marine habitats (Könneke et al., 2005) and hot springs (de la Torre et al., 2008). Theoretical calculation suggested that AOA could contribute 1.51% to 23.4% of the net nitrification activity from three biological replicates of SIP microcosm (Supplementary Table S5).
The putative archaeal contribution in this study has to be cautiously interpreted and re-assessed in the future because the cell-specific rate of 13C-labeled AOA cells in the soil tested may be highly different with that of Nitrosopumilus maritimus and of the Candidatus Nitrosocaldus yellowstonii. In fact, no ammonia oxidation activity was observed after incubation of the soil tested for more than 5 months at 46 °C, which suggests that the labeled AOA was physiologically distinct from N. gargensis, despite the fact that they are the most phylogenetically related to one another. In addition, the carbon incorporation efficiency of AOA and AOB cannot adequately reflect nitrification activity because the relationship between CO2 fixation and ammonia oxidation is poorly understood. For instance, it remains unknown for AOA, whereas the mole ratio of NH3 oxidation to C incorporation is generally assumed to differ greatly among AOB species (Feliatra and Bianchi, 1993). Furthermore, archaeal ammonia oxidation activity may be underestimated by calculating AOA cells that are retrieved only in the ‘heavy’ DNA fractions containing the highly enriched 13C label (Supplementary Table S5). The overall distribution pattern of crenarchaeal 16S rRNA reads from the labeled microcosm showed a slightly drifting trend to the heavier DNA fractions when compared with that in the control microcosm (Figure 4c). Thus, the majority of AOA cells might have replicated at least once in the labeled soil microcosm even though only 9.41% of the total amoA-containing AOA cells actively replicated (Supplementary Table S5) and assimilated 13CO2 to such an extent that they were sufficiently heavy to be revolved in the ‘heavy’ DNA fractions. Nevertheless, the results in this study show the chemolithoautotrophic growth of AOA within the soil crenarchaeota group I.1b lineage, which so far eludes cultivation in soil (Schleper, 2010).
The time-course experiment indicated that bacterial amoA genes were incrementally labeled during active nitrification upon ammonium fertilization (Figure 2), implying that AOB is actively involved in soil nitrification. This was supported by the inferred cell-specific rate of ammonia oxidation. The AOB cell-specific rate fell well within the range reported in the literature (Okano et al., 2004; Jia and Conrad, 2009), being sufficiently large to account for the ammonium consumption in the labeled microcosm (Supplementary Table S5). In contrast, the cell-specific rate of the labeled AOA was estimated to be 4.06 fmol per NH3 oxidized cell per h, and it far exceeded the maximum capacity of the known AOA in cultures (0.59 fmol per NH3 oxidized cell per h). This indicates that the observed nitrification activity could not have solely resulted from the AOA unless the growth of the labeled AOA in the soil is at least six-fold faster than that of the known AOA under optimal culture conditions. Thus, AOB seemed to dominate microbial ammonia oxidation by contributing >76.7% of the nitrification activity in the soil tested (Supplementary Table S5). This result is consistent with recent findings from studies involving German agricultural soil (Jia and Conrad, 2009), zinc-contaminated soils (Mertens et al., 2009) and nitrogen-rich grassland soils (Di et al., 2009).
A meta-analysis of the 13C-labeled 16S rRNA genes in the literature was performed along with the active AOB retrieved in this study, showing physiologically distinct AOB ecotypes across various environments (Supplementary Figure S1). The Nitrosospira cluster 3 dominated active AOB population in this study, whereas the dominant sequence of active AOB in German agricultural soil (Jia and Conrad, 2009) was related to Nitrosospira sp. Nsp 65, of which the phylogenetic classification is still undefined (Koops et al., 2006). In the estuarine sediments, the labeled 16S rRNA genes were related to the Nitrosomonas sp. Nm143 lineage and obligatorily halophilic Nitrosomonas cryotolerans lineage (Freitag et al., 2006), whereas lake sediment AOB closely related to Nitrosomonas europaea or Nitrosomonas eutropha became metabolically active upon the addition of ammonium (Whitby et al., 2001). In aquatic environments, AOB members within the Nitrosospira cluster 0 and Nitrosomonas communis lineage are exclusively found in ammonium-rich stream biofilms, whereas sequences within the Nitrosomonas oligotroph lineage are found in both ammonium-poor and ammonium-rich stream biofilms, as well as the mat of a Romanian Movile Cave without light (Chen et al., 2009). The niche differentiation possibly selects the distinctly active nitrifiers in a wide variety of environments (Martens-Habbena et al., 2009), which implies that AOB plays a major role in nitrification process in these environments. Niche separation may indeed explain the failure of detecting active archaeal nitrification in a previous study (Jia and Conrad, 2009). The community of AOA highly similar to N. gargensis (Supplementary Figure S3) was not observed in the German agricultural soil study. Thus, AOA in the previous study might have had lower growth rates than those affiliated with N. gargensis in this study. The assimilation of 13CO2 might be too low to allow the detection of labeled AOA, as reported previously (Jia and Conrad, 2009). mRNA-SIP would be a powerful technique to elucidate the relative role of AOA and AOB in soil nitrification, with its much higher sensitivity than DNA-SIP (Huang et al., 2009).
DNA-SIP clearly indicated that the active replication of amoA-containing AOA cells depended on ammonia oxidation activity as verified by the C2H2 inhibition technique (Figures 3 and 4), providing strong evidence for the chemolithoautotrophic growth of AOA in complex soil environments. The widespread presence of archaeal amoA transcript suggests that AOA actively catalyzed ammonia oxidation under in situ conditions in soil (Treusch et al., 2005; Nicol et al., 2008) and marine habitats (Lam et al., 2007; Santoro et al., 2010). Archaeal ammonia oxidation is very likely a common feature that naturally occurs in a wide variety of environments on Earth, and its ecological importance varies greatly in accordance with the physicochemical characteristics of the habitats. For instance, recent studies showed that AOA within the marine crenarchaeota group I.1a lineage correlated well with nitrification activity in agricultural soil (Offre et al., 2009), whereas archaeal ammonia oxidation in agricultural alkaline soil was performed by AOA members within the soil crenarchaeota group I.1b lineage in this study. The expression of archaeal amoA genes at high temperatures of up to 94 °C is observed in hot springs (Jiang et al., 2010), implying that archaeal ammonia oxidation might have evolved under thermophilic conditions (de la Torre et al., 2008, Hatzenpichler et al., 2008). In fact, comparative genomic analysis has provided strong support for the taxonomic assignment of the mesophilic N. gargnesis and N. maritimus to the proposed Thaumarchaetoa phylum (Spang et al., 2010), a deeply branched group separate from, hyperthermophilic Crenarchaeota phylum. A considerable amount of crenarchaeota in moderate marine and terrestrial environments may not be ammonia oxidizers (Agogue et al., 2008). Thus, the newly proposed phylum Thaumarchaeota appears to represent the best niche difference between mesophilic and hyperthermophilic habitats in which crenarchaeota live (Brochier-Armanet et al., 2008), but not the putative primary physiology of archaeal ammonia oxidation (de la Torre et al., 2008; Reigstad et al., 2008).
The targeted pyrosequencing of the total 16S rRNA gene allows the fingerprinting of active NOB communities with an unprecedented level of detail. Phylogenetic analysis indicated that nitrite oxidation was primarily catalyzed by Nitrospira- and Nitrobacter-like NOB communities (Figure 5). Nitrite oxidation has long been speculated to be predominantly performed by some organisms other than the best-investigated Nitrobacter-like NOB in freshwater and aquarium biofilters (Hovanec and DeLong, 1996) and activated sludge (Wagner et al., 1996). This hypothesis was confirmed when Nitrospira-like NOB was detected as dominant nitrite oxidizers in the aquatic environment (Hovanec et al., 1998; Schramm et al., 1998). The results in this study show that the labeled community of Nitrospira-like NOB comprised a much higher proportion of the total microbial community in the 13C-labeled DNA than that of the Nitrobacter-like NOB (Figures 5c and d), suggesting that the former is involved in nitrification more actively than the latter. This may be explained by the high substrate affinity of Nitrospira (Koops and Pommerening-Röser, 2001). For instance, the best growth of Nitrospira moscoviensis was observed with 0.35 mM nitrite concentration (Ehrich et al., 1995), whereas 1.5 mM could result in complete growth inhibition of a novel marine Nitrospira-like bacterium (Off et al., 2010). The soil nitrite concentration was under detection limit in this study, implying the rapid conversion of nitrite into nitrate by the NOB. In the meantime, owing to the highly heterogeneous nature of soil aggregate, considerably higher concentrations of nitrite might occur in certain microhabitats, in which the labeling of Nitrobacter-like NOB is enhanced. Thus, a large and highly diverse group of Nitrospira-like NOB may have been functionally active, whereas a low diversity of Nitrobacter-like NOB was labeled. This was confirmed by a single OTU comprising 98% of Nitrobacter-like sequence reads (Supplementary Table S4), whereas Nitrospira-like NOB community was composed of phylogenetically more diverse groups of 16S rRNA genes in the 13C-labeled DNA (Supplementary Table S3). The result presented here is consistent with those previously reported in grassland soil (Freitag et al., 2005) and wastewater treatment plants (Daims et al., 2001). The low diversity of the labeled Nitrobacter-like NOB may also be attributed to the fact that putatively dominant Nitrobacter in this study might grow as heterotrophs (Daims et al., 2001) and could not be labeled with 13CO2. A recent study elegantly showed the functional importance of Nitrobacter-like NOB rather than Nitrospira-like NOB in soils and suggested the r- and K-strategy for the former and the latter, respectively (Attard et al., 2010). In combination with the functional gene assay (Poly et al., 2008), DNA-SIP would enable better assessment of the relative contribution of Nitrobacter and Nitrospira on soil nitrification.
The results in this study provide strong evidence that AOB, AOA, NOB and uncultured crenarchaeota assimilated 13CO2 as a result of active nitrification in the soil microcosm (Figures 3 and 4). The SIP-targeted pyrosequencing can, in principle, allow detailed analysis of the active microorganisms involved in the metabolism of labeled substrate in the soil with unprecedented coverage. This is most important for the detection of active microbial populations that have a slow rate of substrate turnover and growth due to low energy yield, or that comprise only a tiny fraction of the total microbial community. For instance, 20 out of 2744 AOB sequence reads were found to be closely related to the Nitrosomonas oligotroph lineage by pyrosequencing analysis (Supplementary Figure S1), and none of these sequences were detected by clone library analysis of 16S rRNA (Supplementary Figure S1) and bacterial amoA genes (Supplementary Figure S2). Similarly, AOB members affiliated with Nitrosomonas sp. accounted for a considerable proportion, up to 4.5%, of the total 16S rRNA genes in the 13C-labeled DNA (Figure 5), but these AOB sequences could not be retrieved by clone library analysis of the 16S rRNA genes (Supplementary Figure S1). Therefore, the active microorganisms with low abundance in the 13C-labeled DNA can be readily detected with tremendously improved resolution and sensitivity by pyrosequencing rather than by the conventional clone library technique. Interestingly, NOB was labeled to a much greater extent than AOB by pyrosequencing analysis (Figure 4). Sequences highly similar to Nitrospira and Nitrobacter sp. comprised 26.9% and 15.1% of the 16S rRNA genes in the 13C-labeled DNA fractions, respectively (Figure 5). The significantly simultaneous labeling of AOB and NOB has thus suggested the bacterial community dominate microbial nitrification in the soil studied. In addition, there might have been some other unknown organisms that provided trace amounts of nitrification activity and was overlooked in this study. However, the molecular survey showed that AOB highly similar to the Nitrosospira cluster 3 responded positively to in situ long-term fertilization and dominated the active nitrifying community under field conditions (Chu et al., 2007). Thus, the results presented here might largely reflect the predominant nitrification process occurring under field conditions, although the SIP microcosm did not entirely reproduce the physiochemical and biological conditions in the field.
Taken together, the results of this study show that Bacteria dominated nitrification in the soil tested. The labeling of AOB, AOA and NOB by 13CO2 indicates that nitrification is driven by a complex and multi-tiered autotrophic community. The acetylene inhibition technique suggests the chemolithoautotrophic metabolism of nitrifying community in complex soil environments. With the rapid advancement of next-generation sequencing techniques, the SIP-targeted pyrosequencing approach described herein will be a powerful tool in microbial ecology for investigating ecologically important processes in a wide variety of ecosystems with labeled substrates.
Accession codes
References
Agogue H, Brink M, Dinasquet J, Herndl GJ . (2008). Major gradients in putatively nitrifying and non-nitrifying Archaea in the deep North Atlantic. Nature 456: 788–791.
Alawi M, Lipski A, Sanders T, Eva Maria P, Spieck E . (2007). Cultivation of a novel cold-adapted nitrite oxidizing betaproteobacterium from the Siberian Arctic. ISME J 1: 256–264.
Alawi M, Off S, Kaya M, Spieck E . (2009). Temperature influences the population structure of nitrite-oxidizing bacteria in activated sludge. Environ Microbiol Rep 1: 184–190.
Attard E, Poly F, Commeaux C, Laurent F, Terada A, Smets BF et al. (2010). Shifts between nitrospira- and nitrobacter-like nitrite oxidizers underlie the response of soil potential nitrite oxidation to changes in tillage practices. Environ Microbiol 12: 315–326.
Bock E, Wagner M . (2006). Oxidation of inorganic nitrogen compounds as an energy source. In: Dworkin M, Falkow S (eds). The Prokaryotes. Springer Verlag: New York, pp 457–495.
Brochier-Armanet C, Boussau B, Gribaldo S, Forterre P . (2008). Mesophilic crenarchaeota: proposal for a third archaeal phylum, the Thaumarchaeota. Nat Rev Microbiol 6: 245–252.
Chen Y, Wu L, Boden R, Hillebrand A, Kumaresan D, Moussard H et al. (2009). Life without light: microbial diversity and evidence of sulfur- and ammonium-based chemolithotrophy in Movile Cave. ISME J 3: 1093–1104.
Chu H, Fujii T, Morimoto S, Lin X, Yagi K, Hu J et al. (2007). Community structure of ammonia-oxidizing bacteria under long-term application of mineral fertilizer and organic manure in a sandy loam soil. Appl Environ Microbiol 73: 485–491.
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ et al. (2009). The ribosomal database project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37: D141–D145.
Daims H, Nielsen JL, Nielsen PH, Schleifer K-H, Wagner M . (2001). In situ characterization of nitrospira-like nitrite-oxidizing bacteria active in wastewater treatment plants. Appl Environ Microbiol 67: 5273–5284.
de la Torre JR, Walker CB, Ingalls AE, Konneke M, Stahl DA . (2008). Cultivation of a thermophilic ammonia oxidizing archaeon synthesizing crenarchaeol. Environ Microbiol 10: 810–818.
DeLong E . (1992). Archaea in coastal marine environments. Proc Natl Acad Sci USA 89: 5685–5689.
Di HJ, Cameron KC, Shen JP, Winefield CS, O’ Callaghan M, BOWATTE S et al. (2009). Nitrification driven by bacteria and not archaea in nitrogen rich grassland soils. Nat Geosci 2: 621–624.
Ehrich S, Behrens D, Lebedeva E, Ludwig W, Bock E . (1995). A new obligately chemolithoautotrophic, nitrite-oxidizing bacterium, Nitrospira moscoviensis sp. nov. and its phylogenetic relationship. Arch Microbiol 164: 16–23.
Erguder TH, Boon N, Wittebolle L, Marzorati M, Verstraete W . (2009). Environmental factors shaping the ecological niches of ammonia-oxidizing archaea. FEMS Microbiol Rev 33: 855–869.
Feliatra F, Bianchi M . (1993). Rates of nitrification and carbon uptake in the Rhone River plume (northwestern Mediterranean Sea). Microb Ecol 26: 21–28.
Francis CA, Roberts KJ, Beman JM, Santoro AE, Oakley BB . (2005). Ubiquity and diversity of ammonia-oxidizing archaea in water columns and sediments of the ocean. Proc Natl Acad Sci USA 102: 14683–14688.
Freitag TE, Chang L, Clegg CD, Prosser JI . (2005). Influence of inorganic nitrogen management regime on the diversity of nitrite-oxidizing bacteria in Agricultural Grassland Soils. Appl Environ Microbiol 71: 8323–8334.
Freitag TE, Chang L, Prosser JI . (2006). Changes in the community structure and activity of betaproteobacterial ammonia-oxidizing sediment bacteria along a freshwater-marine gradient. Environ Microbiol 8: 684–696.
Gruber N, Galloway JN . (2008). An earth-system perspective of the global nitrogen cycle. Nature 451: 293–296.
Gubry-Rangin C, Nicol GW, Prosser JI . (2010). Archaea rather than bacteria control nitrification in two agricultural acidic soils. FEMS Microbiol Ecol 74: 566–574.
Hatzenpichler R, Lebedeva EV, Spieck E, Stoecker K, Richter A, Daims H et al. (2008). A moderately thermophilic ammonia-oxidizing crenarchaeote from a hot spring. Proc Natl Acad Sci USA 105: 2134–2139.
Hovanec T, DeLong E . (1996). Comparative analysis of nitrifying bacteria associated with freshwater and marine aquaria. Appl Environ Microbiol 62: 2888–2896.
Hovanec TA, Taylor LT, Blakis A, Delong EF . (1998). Nitrospira-like bacteria associated with nitrite oxidation in freshwater aquaria. Appl Environ Microbiol 64: 258–264.
Huang WE, Ferguson A, Singer AC, Lawson K, Thompson IP, Kalin RM et al. (2009). Resolving genetic functions within microbial populations: in situ analyses using rRNA and mRNA stable isotope probing coupled with single-cell Raman-fluorescence in situ hybridization. Appl Environ Microbiol 75: 234–241.
Hyman M, Arp D . (1992). 14C2H2- and 14CO2-labeling studies of the de novo synthesis of polypeptides by Nitrosomonas europaea during recovery from acetylene and light inactivation of ammonia monooxygenase. J Biol Chem 267: 1534–1545.
Jia ZJ, Conrad R . (2009). Bacteria rather than Archaea dominate microbial ammonia oxidation in an agricultural soil. Environ Microbiol 11: 1658–1671.
Jiang H, Huang Q, Dong H, Wang P, Wang F, Li W et al. (2010). RNA-based investigation of ammonia-oxidizing archaea in Hot Springs of Yunnan Province, China. Appl Environ Microbiol 76: 4538–4541.
Könneke M, Bernhard AE, de la Torre JR, Walker CB, Waterbury JB, Stahl DA . (2005). Isolation of an autotrophic ammonia-oxidizing marine archaeon. Nature 437: 543–546.
Koops HP, Pommerening-Röser A . (2001). Distribution and ecophysiology of the nitrifying bacteria emphasizing cultured species. FEMS Microbiol Ecol 37: 1–9.
Koops HP, Purkhold U, Pommerening-Roser A, Timmermann G, Wagner M . (2006). The lithoautotrophic ammonia-oxidizing bacteria. In: Dworkin M, Falkow S (eds). The Prokaryotes. Springer Verlag: New York, pp 778–811.
Kowalchuk GA, Stephen JR . (2001). Ammonia-oxidizing bacteria: a model for molecular microbial ecology. Annl Rev Microbiol 55: 485–529.
Kumar S, Tamura K, Nei M . (2004). MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5: 150–163.
Lam P, Jensen MM, Lavik G, McGinnis DF, Muller B, Schubert CJ et al. (2007). Linking crenarchaeal and bacterial nitrification to anammox in the Black Sea. Proc Natl Acad Sci USA 104: 7104–7109.
Lane DJ . (1991). 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds). Nucleic Acid Techniques in Bacterial Systematics. John Wiley and Sons: New York, USA, pp 177–203.
Leininger S, Urich T, Schloter M, Schwark L, Qi J, Nicol GW et al. (2006). Archaea predominate among ammonia-oxidizing prokaryotes in soils. Nature 442: 806–809.
Lueders T, Manefield M, Friedrich MW . (2004). Enhanced sensitivity of DNA- and rRNA-based stable isotope probing by fractionation and quantitative analysis of isopycnic centrifugation gradients. Environ Microbiol 6: 73–78.
Martens-Habbena W, Berube PM, Urakawa H, de la Torre JR, Stahl DA . (2009). Ammonia oxidation kinetics determine niche separation of nitrifying Archaea and Bacteria. Nature 461: 976–979.
Meng L, Ding W, Cai Z . (2005). Long-term application of organic manure and nitrogen fertilizer on N2O emissions, soil quality and crop production in a sandy loam soil. Soil Biol Biochem 37: 2037–2045.
Mertens J, Broos K, Wakelin SA, Kowalchuk GA, Springael D, Smolders E . (2009). Bacteria, not archaea, restore nitrification in a zinc-contaminated soil. ISME J 3: 916–923.
Neufeld JD, Vohra J, Dumont MG, Lueders T, Manefield M, Friedrich MW et al. (2007). DNA stable-isotope probing. Nat Protocols 2: 860–866.
Nicol GW, Leininger S, Schleper C, Prosser JI . (2008). The influence of soil pH on the diversity, abundance and transcriptional activity of ammonia oxidizing archaea and bacteria. Environ Microbiol 10: 2966–2978.
Off S, Alawi M, Spieck E . (2010). Enrichment and physiological characterization of a novel nitrospira-like bacterium obtained from a marine sponge. Appl Environ Microbiol 76: 4640–4646.
Offre P, Prosser JI, Nicol GW . (2009). Growth of ammonia-oxidizing archaea in soil microcosms is inhibited by acetylene. FEMS Microbiol Ecol 70: 99–108.
Okano Y, Hristova KR, Leutenegger CM, Jackson LE, Denison RF, Gebreyesus B et al. (2004). Application of real-time PCR to study effects of ammonium on population size of ammonia-oxidizing bacteria in soil. Appl Environ Microbiol 70: 1008–1016.
Poly F, Wertz S, Brothier E, Degrange V . (2008). First exploration of nitrobacter diversity in soils by a PCR cloning-sequencing approach targeting functional gene nxrA. FEMS Microbiol Ecol 63: 132–140.
Reigstad LJ, Richter A, Daims H, Urich T, Schwark L, Schleper C . (2008). Nitrification in terrestrial hot springs of Iceland and Kamchatka. FEMS Microbiol Ecol 64: 167–174.
Rotthauwe J, Witzel K, Liesack W . (1997). The ammonia monooxygenase structural gene amoA as a functional marker: molecular fine-scale analysis of natural ammonia-oxidizing populations. Appl Environ Microbiol 63: 4704–4712.
Santoro AE, Casciotti KL, Francis CA . (2010). Activity, abundance and diversity of nitrifying archaea and bacteria in the central California current. Environ Microbiol 12: 1989–2006.
Schleper C . (2010). Ammonia oxidation: different niches for bacteria and archaea? ISME J 4: 1092–1094.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB et al. (2009). Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75: 7537–7541.
Schramm A, de Beer D, Wagner M, Amann R . (1998). Identification and activities in situ of nitrosospira and nitrospira spp. as dominant populations in a nitrifying fluidized bed reactor. Appl Environ Microbiol 64: 3480–3485.
Spang A, Hatzenpichler R, Brochier-Armanet C, Rattei T, Tischler P, Spieck E et al. (2010). Distinct gene set in two different lineages of ammonia-oxidizing archaea supports the phylum thaumarchaeota. Trends Microbiol 18: 331–340.
Treusch AH, Leininger S, Kletzin A, Schuster SC, Klenk H-P, Schleper C . (2005). Novel genes for nitrite reductase and Amo-related proteins indicate a role of uncultivated mesophilic crenarchaeota in nitrogen cycling. Environ Microbiol 7: 1985–1995.
Wagner M, Rath G, Koops H-P, Flood J, Amann R . (1996). In situ analysis of nitrifying bacteria in sewage treatment plants. Water Sci Technol 34: 237–244.
Whitby CB, Hall G, Pickup R, Saunders JR, Ineson P, Parekh NR et al. (2001). 13C incorporation into DNA as a means of identifying the active components of ammonia-oxidizer populations. Lett Appl Microbiol 32: 398–401.
Zhang LM, Offre PR, He JZ, Verhamme DT, Nicol GW, Prosser JI . (2010). Autotrophic ammonia oxidation by soil thaumarchaea. Proc Natl Acad Sci USA 107: 17240–17245.
Acknowledgements
This work was financially supported by the National Science Foundation of China (40971153 to ZJ), the National High-Tech R&D Program (863) (2009AA02Z310 to JX), the Knowledge Innovation Programs of the Chinese Academy of Sciences (KZCX2-YW-BR-06 to ZJ) and the E-Science Program of the Chinese Academy of Sciences (INFO-115-D01-Z006 to JX and ZJ). We want to extend our gratitude to Professor Weixing Ding for sampling assistance at the Feng Qiu State Key Experimental Station for Ecological Agriculture.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Xia, W., Zhang, C., Zeng, X. et al. Autotrophic growth of nitrifying community in an agricultural soil. ISME J 5, 1226–1236 (2011). https://doi.org/10.1038/ismej.2011.5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2011.5
Keywords
This article is cited by
-
Soil pH and phosphorus drive the canonical nitrifiers and comammox Nitrospira communities in citrus orchards with different cultivation ages
Soil Ecology Letters (2024)
-
Continuous-flow membrane bioreactor enhances enrichment and culture of autotrophic nitrifying bacteria by removing extracellular free organic carbon
Environmental Science and Pollution Research (2023)
-
Soil organic matter contents modulate the effects of bacterial diversity on the carbon cycling processes
Journal of Soils and Sediments (2023)
-
The active role of comammox Nitrospira in nitrification in acidic orchard soils revealed by DNA-SIP
Biology and Fertility of Soils (2023)
-
Research advances of ammonia oxidation microorganisms in wastewater: metabolic characteristics, microbial community, influencing factors and process applications
Bioprocess and Biosystems Engineering (2023)