Abstract
In higher eukaryotes, the 5S rRNA genes occur in tandem units and are arranged either separately (S-type arrangement) or linked to other repeated genes, in most cases to rDNA locus encoding 18S–5.8S–26S genes (L-type arrangement). Here we used Southern blot hybridisation, PCR and sequencing approaches to analyse genomic organisation of rRNA genes in all large gymnosperm groups, including Coniferales, Ginkgoales, Gnetales and Cycadales. The data are provided for 27 species (21 genera). The 5S units linked to the 35S rDNA units occur in some but not all Gnetales, Coniferales and in Ginkgo (∼30% of the species analysed), while the remaining exhibit separate organisation. The linked 5S rRNA genes may occur as single-copy insertions or as short tandems embedded in the 26S–18S rDNA intergenic spacer (IGS). The 5S transcript may be encoded by the same (Ginkgo, Ephedra) or opposite (Podocarpus) DNA strand as the 18S–5.8S–26S genes. In addition, pseudogenised 5S copies were also found in some IGS types. Both L- and S-type units have been largely homogenised across the genomes. Phylogenetic relationships based on the comparison of 5S coding sequences suggest that the 5S genes independently inserted IGS at least three times in the course of gymnosperm evolution. Frequent transpositions and rearrangements of basic units indicate relatively relaxed selection pressures imposed on genomic organisation of 5S genes in plants.
Similar content being viewed by others
Introduction
Organisation of ribosomal DNA in plants
The four types of ribosomal RNA (18S, 5.8S, 26S and 5S) are essential constituents of the ribosomes of all eukaryotes, encoded by variable numbers of copies of rRNA genes, called rDNA. Plants are known to bear an extraordinarily high number of these genes (from 1000 to more than 50 000) arranged in long tandem arrays on chromosomes (Hemleben and Zentgraf, 1994). The 18S–5.8S–26S genes are usually organised, in that order, in a single operon, in one or several chromosomal loci. In cell nucleoli, the genes are cotranscribed by RNA polymerase I into a primary transcript, which is shorter in plants (35S rRNA) than in animals (45S rRNA) (Seitz and Seitz, 1979). We will therefore refer to 35S rDNA throughout the work. The 5S is the only rRNA gene that is transcribed separately, by RNA polymerase III in nucleoplasm. The 5S rDNA typically occurs at positions separate from 35S rDNA in multicellular eukaryotic organisms, sometimes forming long arrays of tandemly arranged genes spaced by intergenic spacers (IGS) of variable lengths. Nevertheless, there are several exceptions to this rule, for example, 5S gene linkage to other repetitive gene families including 35S, small nuclear RNAs, histone genes or the trans-spliced leader (Drouin and de Sa, 1995), which were formerly believed to represent transition states between linked (typically prokaryotic) and unlinked (typically eukaryotic) arrangement. Consistent with this assumption, 5S rRNA genes in plants were thought to be mostly arranged separately from 35S rDNA (S-type arrangement), organised in tandem repeats whose number varied from less than 1000 to over 75 000 (Campell et al., 1992; Sastri et al., 1992).
Regardless, several lines of evidence indicate that the transition hypothesis may not work for green plants, at least. Earlier independent studies showed 5S gene integration into the 26S–18S spacers in the moss Funaria hygrometrica and the liverwort Marchantia polymorpha (Sone et al., 1999). In the recent genome-wide assessment of rDNA organisation in land plants done by Wicke et al. (2011), it was concluded that a linked 35S–5S rDNA (L-type arrangement) should be regarded as the ancestral state as this type has been observed in streptophyte algae and early diverging green plants such as liverworts, mosses, hornworts, lycophytes and monilophytes. However, linkage between 5S and 35S genes was also found in several angiosperm genera such as the phylogenetically derived genus Artemisia (Garcia et al., 2009). Later, deeper molecular and cytogenetic studies revealed that the L-type arrangement could be present in ∼25% of the Asteraceae family (Garcia et al., 2010; Mazzella et al., 2010).
Gymnosperms: a rather unexplored field for rDNA research
Although showing a wide diversity in morphology and ecology, gymnosperms consist of only 14 families with roughly 1000 species, a derisory number when compared with the estimated number of angiosperms or even with some of its largest families (such as Asteraceae and Orchidaceae, believed to hold more than 20 000 species each). There are four classically considered gymnosperm orders (Figure 1): (i) Coniferales are the largest, with around 650 species (division Pinophyta, including Pinaceae and the non-Pinaceae Pinophyta, frequently considered as a division on its own and named cupressophytes); (ii) Cycadales, with three extant families and about 130 species; (iii) Gnetales with three monogeneric families, including 70 species; and lastly (iv) Ginkgoales, with a single extant species, Ginkgo biloba. Phylogenetic relationships between gymnosperm families remain questionable (see Supplementary Figure S1 for a summary on available hypotheses on gymnosperm phylogeny), to the extent that even its monophyly and if a common ancestor is shared with flowering plants is still debated. Ecological, genetic and epigenetic factors might have acted distinctly to shape angiosperm and gymnosperm genomes (Leitch and Leitch, 2012).
A proposed organisation of 5S–35S genes in gymnosperm phylogeny. The support for the hypothesis comes from the works listed in Table 1. Grey lines—L-type arrangement; black lines—S-type arrangement. Grey arrows show putative integration events of 5S genes into the 26S–18S IGS. Dashed lines—putative ancestors having either L- of S-type units. The phylogeny was adapted from the currently most-accepted gnepine hypothesis—see Mathews (2009). Phylogeny of cupressophytes (non-Pinaceae conifers) adapted from Chaw et al. (1997).
Cytogenetic studies carried out mostly in Pinaceae (Pinophyta) revealed separate arrangement of 5S and 35S genes (Table 1). The situation outside this family is less clear. Cytogenetic studies using rDNA fluorescence in situ hybridisation revealed colocalisation of 35S and 5S signals in Podocarpus (Podocarpaceae, Murray et al., 2002) and some Cupressaceae (Hizume et al., 1999), both non-Pinaceae conifers from the division Pinophyta and also in G. biloba (Nakao et al., 2005). In most cases, however, because of low resolution of cytogenetic methods, it is not certain whether the 35S and 5S genes are physically linked or whether they form distinct arrays close to each other. There have been several molecular studies of 5S rDNA in gymnosperms with the aim of determining rDNA organisation at the unit level. Out of these, independent 5S clusters were confirmed in Pinus (Mashkova et al., 1990; Gorman et al., 1992; Moran et al., 1992; Liu et al., 2003a), Abies (Besendorfer et al., 2005), Larix (Trontin et al., 1999), Picea (Brown and Carlson, 1997) and Pseudotsuga (Amarasinghe and Carlson, 1998). PCR screens and sequence analysis revealed S-type rDNAs in Cycas, Ginkgo and Gnetum (Wicke et al., 2011). On the basis of these numerous studies, it has been proposed that in gymnosperms the 5S repeats also hold the separated configuration typical for vascular plants (Wicke et al., 2011). Conversely, Galián et al. (2012) undertaking a PCR-based strategy promptly isolated clones from G. biloba with 5S genes integrated in the 26S–18S spacer. The potentially functional 5S gene was located ∼2 kb downstream of the 26S gene. However, there were also clones without 5S genes pointing to some heterogeneity of rDNA arrays. Although in situ hybridisation results favour a dominant L-type arrangement, it remains unclear as to what is the proportion of S- and L-type units in the Gingko's genome.
Taking into account the relative paucity of knowledge about rDNA organisation in most gymnosperms despite the rather reduced size of the group (1000 gymnosperm versus 250 000 angiosperm species), we aim to tackle the global picture on rDNA arrangement in all major gymnosperm clades by studying its organisation in selected representatives through the use of molecular methods. This will enable an update of the state-of-art of rDNA linkages across land plants and the comparison of the different kinds of 5S insertions in all the groups studied up to now.
Materials and methods
Plant sampling
Fresh leaves from 27 species (21 genera), representing 9 families and all major gymnosperm groups, were obtained either from trees at the Botanical Garden of Barcelona, through plants grown at the greenhouse of the Botanical Institute of Barcelona or from trees planted in public gardens (Garden ‘Palau de Pedralbes’ and Pius XII Square gardening, Barcelona). Table 2 summarises the details of the materials collection.
DNA extraction
We extracted the total genomic DNA using either the CTAB method of Doyle and Doyle (1987) or the Nucleospin Plant II kit (Macherey-Nagel, GmbH et Co, Düren, Germany), depending on the quality of the vegetal material, either from silica gel-dried leaves or directly from fresh leaves. The final product was dissolved in Tris EDTA buffer. Optical density readings of the DNA concentration were estimated at an optical density of 260 nm using a Nanodrop spectrophotometer (Thermo Scientific, Wilmington, DE, USA).
Southern blot hybridisation
Purified genomic DNAs were digested with restriction enzymes (BamHI, BglII, EcoRI and EcoRV depending on the experiment) and separated by gel electrophoresis on a 0.9% (w/v) agarose gel. After electrophoresis, the gels were alkali blotted onto Hybond-XL membranes (GE Healthcare, Little Chalfont, Buckinghamshire, UK) and hybridised with 32P-labelled DNA probes (DekaLabel kit, MBI, Fermentas, Vilnius, Lithuania) as described in Garcia et al. (2009). After washing under high-stringency conditions, the hybridisation bands were visualised with a PhosphorImager (Typhoon 9410, GE Healthcare, Piscataway, NJ, USA) and the data were processed by ImageQuant software (GE Healthcare). The probes were a 220-bp PCR product derived from the 3′ end of the 26S rRNA gene of tobacco (Lim et al., 2000), a cloned ∼120 bp of the 5S genic region, also from tobacco (Fulnecek et al., 2002) and an 18S probe from tomato (Kiss et al., 1989). The IGS1-5S probe was a 580-bp fragment from Ginkgo prepared by cutting the insert of a pGIN2 plasmid construct with EcoRV.
PCR and cloning experiments
Six PCR primers were used to amplify products, assuming some linkage of 35S and 5S rRNA genes in either direct (head-to-tail) or inverted (tail-to-tail) orientations for several candidate species. The positions of the PCR primers are depicted in Supplementary Figure S2a. The primers used were: 5SgF (5′-GGTGCGATCATACCAGCACT-3′) and 5SgR (5′-GGTGCAACACGAGGACTTC-3′) (Garcia et al., 2012a); 26SF (5′-AGACGACTTTAAATACGCGAC-3′) and 18SR (5′-GGCTTAATCTTTGAGACAA-3′) are slightly modified versions of the Pr1 and Pr2 primers, respectively (Komarova et al., 2004); 26SR (5′-CTTTTCCTCCGCTTATTGATATGC-3′) and 18SF (5′-GCGCTACACTGATGTATTCAACGA-3′) primers are from (Kovarik et al., 2005). The variants of 5S genic primers were designed based on Ginkgo sequence: 5SgGINF (5′-CAATGGGTGCGATCATACC-3′) and 5SgGINR (5′-GTGCAACACTGGGGACTTC-3′). Most reactions were conducted with DyNAzymeII DNA polymerase (Finnzymes, Espoo, Finland) or the PfuUltraTM II Fusion HS DNA Polymerase (Stratagene, La Jolla, CA, USA), designed for amplifying long-length DNA targets; each reaction contained ∼500 ng of genomic DNA. The PCR profile used for amplification was: initial denaturation at 94 °C for 3 min followed by 35 cycles of 20 s at 94 °C, 1 : 30–3 : 00 (depending on experiment) min at 57 °C and 30 s at 72 °C; and a final extension step at 72 °C, 10 min. All reactions were carried out in a MJ MINI Personal thermal cycler (Bio-Rad Laboratories Inc., Hercules, CA, USA) and the PCR products were separated on a 1.2% agarose gel, stained with ethidium bromide and photo documented (Ultralum, Claremont, CA, USA).
To isolate IGS sequences we used PCR products obtained from reactions involving different combinations of primers. Multiple bands of variable sizes (from several hundred bp up to more than 2 kb) were usually obtained. The long fragments (>2 kb) were isolated using the QIAquick PCR purification kit (Hilden, Germany) and cloned into the pDrive vector (Qiagen PCR Cloning kit, Hilden, Germany). The PCR products obtained from PfuUltraTM II Fusion HS DNA Polymerase were treated with DyNAzymeII DNA Polymerase in the presence of dATP to add A overhangs. Several positive clones were recovered from each transformation reaction and analysed for insert length by gel electrophoresis. The IGS constructs were obtained for G. biloba (clone pGIN2, containing linked 26S–IGS1–5S sequences, GenBank JX402064), Podocarpus elongatus (p3g, containing the 5S–IGS2–18S sequences, GenBank JX402063) and Ephedra nebrodensis (pEPHN4, containing the 5S–IGS2–18S sequences, GenBank JX843794). The clones were sequenced by the Sanger method (Eurofins, MWG Operon, Eberberg, Germany) using primer walking and subcloning strategies. Highly repetitive regions were generally difficult to sequence by primer walking. In these cases, several subclones were prepared, sequenced and assembled.
Bioinformatic analysis
The contigs of IGS clones were assembled in BioEdit Sequence Alignment Editor 7.0.9.0 (Hall, 1999) and blasted against the GenBank database. The rRNA coding sequences were identified by pairwise comparisons using dot plot graphical outputs (http://www.vivo.colostate.edu/molkit/dnadot/ and http://bioinfo.lifl.fr/yass/) (Noe and Kucherov, 2005). Sliding Windows were set to 9 with stringencies corresponding to 0–1 mismatches. Annotation of regulatory motifs was carried out based on analogy with A. thaliana gene (Cloix et al., 2003). Restriction mapping was done using NEBcutter at the NEB server (http://tools.neb.com/NEBcutter2; Vincze et al., 2003). Searches for repeats were carried out by the tandem repeat finder program (Benson, 1999). Subrepeated structure of IGS was analysed by matrix dot plot comparisons of using similarity search tools box at the Colorado State University (http://www.vivo.colostate.edu/molkit/dnadot/). Sliding Windows were set to 9 with stringencies corresponding to 0–2 mismatches. Sequence divergences were calculated by a DNADIST algorithm (Felsenstein, 1989). Phylogenetic trees were constructed using a SEAVIEW software (Gouy et al., 2010).
Results
The data on fundamental rDNA organisation are provided here for 27 species, 21 genera, 4 families and 1 division of gymnosperms previously unstudied from this point of view. This provides coverage of basic rDNA organisation for at least one representative of all gymnosperm orders, as well as a 15% increase in coverage of genera and an almost 30% increase in coverage of families of gymnosperms. The newly assessed taxa are indicated in Table 2.
Southern blot hybridisation reveals both S- and L-type arrangements in gymnosperms
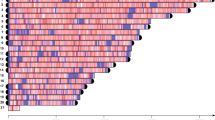
The BamHI hybridisation profiles obtained for 25 species are shown in Figures 2 and 3. Basically, two hybridisation profiles could be distinguished:
Southern blot hybridisation analysis of species with L-type arrangement of 5S and 35S units. Restriction map showing conserved BamHI sites and regions of probe hybridisation (thin lines above boxes) are schematically drawn in (a). Genomic DNAs were digested with BamHI and hybridised on blots with the 5S, 26S and GIN2 (containing IGS1 from Ginkgo) DNA probes (b).
The first group (Figure 2) included species with profiles typical for a dominant linked arrangement (L-type) of both genes as revealed by extensive cohybridisation of 26S and 5S probes on blots. The four Podocarpus and the single Afrocarpus (both Coniferales) species showed multiple 5S and 26S cohybridising bands of 5–20 kb in size. Podocarpus henkelii, P. elongatus, P. neriifiolius and Afrocarpus falcatus (Podocarpaceae) had a similar restriction profile with respect to position and number of hybridisation bands (data not shown for P. neriifolius).
Both Ephedra viridis and E. nebrodensis (Gnetales) had similar restriction profiles with all fragments hybridising to both 5S and 26S probes although in this case significant smearing of signals occurred. There was a significant fraction or unresolved signals in >20 kb regions, which may indicate mutation of heavy methylation of units, as BamHI is sensitive to cytosine methylation (McClelland and Nelson, 1985). We therefore used EcoRI, which is less sensitive to methylation, to confirm the results (Supplementary Figure S3). The 26S and 5S probes cohybridised to one or two fragments in all species. Cohybridisation of fragments with all three rDNA (5S, 18S and 26S) probes was visible in A. falcatus, P. latifolius and both Ephedra species.
The BamHI hybridisation profile of Ginkgo (Figure 2) was somewhat different from those of Podocarpus, Afrocarpus and Ephedra. In Ginkgo, only high (>15 kb) molecular weight BamHI fragments cohybridised with the 26S and 5S probes while the low (6 kb) molecular weight fragment hybridised with the 26S probe only; however, the spacer probe, isolated from the pGIN2 clone (described further below) hybridised to both 6- and >15-kb fragments.
The second group (Figure 3) comprised species with a separated organisation of 5S and 35S repeats (S-type). In this group, the 5S probe hybridised to ladders of regularly spaced fragments of variable lengths considered to be indicative of tandem arrangement of units (Campell et al., 1992). The 26S probe usually hybridised to one or several high-molecular-weight bands. The 5S hybridisation bands had no significant overlaps with the 26S signals except for a faint 4-kb fragment in Araucaria bidwillii (Araucariaceae, Coniferales). The periodicity of BamHI bands varied significantly from species to species ranging from 0.5 kb (Abies pinsapo) up to 1.8 kb in A. bidwillii. Relatively irregular ladders in Macrozamia moorei and Encephalartos msinganus (both from Zamiaceae, Cycadales) may be explained by the presence of multiple BamHI sites in the unit monomer and/or more complex organisation of genes.
Southern blot hybridisation analysis of species with S-type arrangement of 5S and 35S units. The 5S tandem arrays with conserved BamHI sites and probe hybridisation regions (lines above boxes) are schematically drawn in (a). Genomic DNAs were digested with BamHI and hybridised on blots with the 5S and 26S rDNA probes (b). The sizes of 5S monomeric units are indicated.
rDNA repeat size
In most gymnosperms analysed the rDNA probes hybridised to much larger restriction fragments than in angiosperms. We therefore wished to determine an approximate size of rDNA units of studied species. GenBank searches of 26S genes revealed that there is a single BglII site in the coding region (26S gene) that is highly conserved in all land plants. Consequently, digestion of genomic DNA with BglII would liberate monomeric units that could be revealed by subsequent hybridisation with the 5S, 18S and 26S probes (Supplementary Figure S3). Indeed, in all L-type species analysed (Podocarpus, Afrocarpus, Ephedra and Ginkgo), the probes hybridised to the high molecular weight fragments of >20 kb in length indicating that the size of basic 35S–5S units was within that range. The differences in the mobility of major BglII bands between species were only marginal. Fragments of smaller size (around 12 kb) seen in P. latifolius could be explained by the presence of additional BglII site(s) in some units. The sequence information on the Ginkgo rDNA unit allowed a more accurate estimation of unit size in this species. The genomic DNA was digested with the methylation-insensitive EcoRV, for which there are conserved target sites in the IGS1, IGS2 and 5.8S genes (Supplementary Figure S4). The subsequent hybridisation with the 5S, 18S and 26S probes revealed distinct fragments of 12, 3 and 6 kb, respectively, indicating that the size of the Ginkgo unit is 21 kb, at least.
Cloning of 35S–5S units from G. biloba
In G. biloba, the 26SF and 5SgR primers yielded several >2-kb PCR bands while the 26SF and 5SgF yielded a low molecular weight <1 kb band (Supplementary Figure S2). Cloning and sequencing of the 26S–5S PCR products revealed that only high molecular weight 26SF/5SgR bands contained both 26S and 5S sequences while the low molecular weight band amplified by the 26SF/5SgF primer set lacked either one or both genes and appeared to be non-specific. The inserts of five plasmid clones carrying the putative 26S–IGS1–5S sequences were slightly heterogeneous in size (1.6–1.9 kb). One clone carrying the 1827-bp insert (pGIN2 construct) was fully sequenced and submitted to GenBank (JX402064). BLAST searches revealed that there was a 5S gene ∼1.6-kb downstream from the 26S gene (Figure 4). The 5S coding strand was the same as the one coding for the 26S gene (direct orientation). In the 120-bp coding region, the BamHI site was conserved, as were the regulatory motifs, Boxes A and C and the internal element. The upstream TATA sequence in the non-coding part at -33 was equally conserved. The pyrimidine-rich motif was found at the 5′ end of the IGS1 (spacer between the 26S and 5S genes) as in other plant IGS sequences. The self-to-self alignment revealed the presence of two repetitive subregions on a dot plot (Supplementary Figure S5). The A1 subregion located 85–330-bp downstream from the 26S gene was composed of short 39-bp long units. The second subregion (A2) corresponding to the A subregion reported by Galián et al. (2012) was located within 671–1228 bp and contained 2.3 copies of a 240 bp repeat. The A2 monomeric unit was internally repetitive composed of several subrepeats of 39–57 bp in size. One subrepeat type was partially homologous to the A1 subrepeat. Sequence comparison with the 6-kb IGS clone (accession no. JQ279501.1) isolated by Galián et al. (2012) showed high identity over the 26S–IGS1–5S region, and differences were caused mostly by variation in the A1 and A2 tandem repeats. The sequences downstream from the 5S gene (IGS2 region) could not be compared because the pGIN2 clone lacked these sequences due to the cloning strategy. The pGIN2 clone also displayed significant similarities to non-5S bearing rDNA clones from Ginkgo at GenBank (Supplementary Figure S6).
Schematic representation 35S–5S units in G. biloba, Podocarpus elongatus and E. nebrodensis. Arrows above boxes indicate direction of transcription. The lines with arrows represent clones carrying 5S insertions. Subrepeated portions of IGS are indicated and labelled as (a–c). The 5S coding region sequences are shown below each graph, with conserved regulatory elements underlined. The positions of BamHI sites are indicated; in Podocarpus the site is mutated (asterisk).
Cloning of 35S–5S units from Podocarpus elongatus
In Podocarpus elongatus, the PCR using 5SgR/18SR and 5SgF/26SF yielded one prominent plus several minor bands (Supplementary Figure S2). The major 1.2-kb band from the 5SgF/26SF PCR reaction appeared to be non-specific as the DNA did not hybridise with the labelled 5S probe whereas the the 3-kb band obtained from the 5SgR/18SR reaction strongly hybridised with the probe (not shown). The 3-kb band was cloned and sequenced. The insert of clone p3g (GenBank JX402063) was 2612-bp long and contained part of the 5S and 18S genes on both the ends (Figure 4). The 5S was encoded by the opposite strand to the 18S gene (inverse orientation). The regulatory boxes A and C as well as the upstream regulatory elements were fully conserved. However, no internal element was found and there were also several point mutations, including the one in the BamHI site at the 5′ end. The TATA regulatory element in the upstream non-coding region was not conserved. The IGS between the 5S and 18S genes (IGS2) was highly repetitive. The self-to-self alignment revealed the presence of two prominent repetitive subregions on a dot plot graph (Supplementary Figure S5). The B subregion at position 490–745 (numbers are from the first 5 S nucleotide) was composed of short 28-bp subrepeats. The C subregion at position 1669–2391 was composed of 2.8 copies of a 257-bp repeat. The 18S coding sequence started at position 2580. A TATAGGG motif resembling the TATA(R)TA(n)GGG transcription initiation site (Perry and Palukaitis, 1990) was located immediately upstream of the C subregion.
Cloning of 35S–5S units from E. nebrodensis
In E. nebrodensis, the PCR using 5SgGINF and 18SR yielded a single strong ∼4 kb band plus several minor bands of larger size. The ∼4-kb fragment was cloned and one clone (pEPHN4) was sequenced (GenBank JX843794). The insert was 3358 bp long and contained three 5S copies on one end and a 5′ terminus of the 18S gene on the other end. The 5S insertion formed a short tandem of closely (9 bp) spaced genes (Figure 4). The 5S rRNA was encoded by the same DNA strand as the 35S genes (direct orientation). The genic region of the first copy (5S1) was 100% identical to the sequence of 5S rRNA from E. kokanica (GenBank X06996 from Melekhovets et al. (1988)) with conserved regulatory elements. The second (Ψ5S2) and third (Ψ5S3) genes had a 40 bp deletion at the 5′end and contained several mutations in regulatory boxes. Hence, these copies probably represent pseudogenes. The homology between pseudogenes and apparently functional 5S1 was 80–85% while that between both pseudogenes was 70% (Supplementary Figure S7). The terminators following coding region were, however, conserved in all the three units. Sequence downstream from the 5S genes was highly repetitive composed from short subrepeats (Supplementary Figure S5). Except of rRNA coding regions there was no significant homology between IGS clones from Ginkgo, Podocarpus, Ephedra and Gnetum (EMBL FR695703 from Wicke et al. (2011)).
Discussion
Genomic organisation of vital rRNA genes has been a topic of numerous investigations in a wide range of biological taxa. Gymnosperms are considered as early diverging seed plants, and hence, represent an important transition between seedless plants and angiosperms. Here, we report on arrangement of 5S and 35S rDNA that may be linked (L-type) or separate (S-type) across the main gymnosperm orders.
Uniform S-type rDNA arrangement in Pinaceae (Coniferales) and Cycadales
A good deal of previous molecular cytogenetic studies had been focused on family Pinaceae, particularly on economically and ecologically important genera such as Pinus and Picea, among others. According to the cytogenetic and molecular data compiled (Table 1), the most commonly found rDNA arrangement was the unlinked (S-type) configuration. Nevertheless, partially overlapping 5S and 35S fluorescence in situ hybridisation signals were observed on chromosomes of Abies alba (Puizina et al., 2008), some Picea (Lubaretz et al., 1996; Brown and Carlson, 1997; Siljak-Yakovlev et al., 2002; Shibata and Hizume, 2008) or certain Pinus (Hizume et al., 2002; Cai et al., 2006; Islam-Faridi et al., 2007) species. However, the Southern blot hybridisation did not reveal significant cohybridisation of 26S (18S) and 5S probes in any of these, indicating that the most likely interpretation of colocalised signals is the juxtaposition of independent 5S and 35S arrays. The Pinaceae karyotypes typically display many more 35S than 5S loci (more than twice in most cases; Siljak-Yakovlev et al., 2002; Islam-Faridi et al., 2007). Yet, Southern hybridisations did not reveal extraordinary heterogeneity of signals (Figure 3b), which could be explained by interlocus homogenisation and/or recent origin of rDNA loci. On the other hand, length heterogeneity of 5S units is indicated in Abies pinsapo, consistent with previous studies in this genus (Besendorfer et al., 2005).
Species from the family Zamiaceae (Cycadales) investigated—genera Ceratozamia, Zamia and Encephalartos—clearly displayed an S-type rDNA pattern as in situ hybridisation data had yet pointed out for both Ceratozamia and Zamia (Tagashira and Kondo, 2001), the only genera of this group investigated to date. The fact that recent molecular phylogenetic approaches (Supplementary Figure S1) point towards cycadophytes being more ancient than Ginkgo (Mathews, 2009) hinders even more the interpretation about the ancestral condition of rDNA organisation in seed plants.
Both S- and L-type rDNAs in non-Pinaceae Coniferales
In this group, we observed certain variations in rDNA organisation. Most members of Araucariaceae, Cupressaceae and Taxaceae showed separation of both 5S and 35S loci. The 5S genes were organised in long tandems as separate arrays. Araucaria bidwillii was interesting in having an extraordinarily long 5S–5S spacer (1.8 kb), which is much longer than that of any gymnosperm 5S spacers, whose units range between 0.4–0.87 kb (Gorman et al., 1992; Moran et al., 1992; Trontin et al., 1999; Besendorfer et al., 2005); known angiosperm 5S units range between 0.2–0.9 kb (Sastri et al., 1992; Fulnecek et al., 2006). It is possible that the large size of IGS was caused by the expansion of repeated elements as apparently happened in Larix species (Trontin et al., 1999) or in A. alba (Besendorfer et al., 2005). It will be interesting to analyse positions of 5S genes on Araucaria chromosomes that appear to bear a single 35S locus (Miranda et al., 2007).
Of note is that the genera Podocarpus and the closely related Afrocarpus (Podocarpaceae) appear to be exceptional among non-Pinaceae conifers in showing clear linkage of 26S and 5S units. The 35S–5S arrays are highly homogeneous, without any evidence of abundant separate loci. The 5S insertions occurred between 6–9 kb downstream from the 26S gene (estimates are based on Southern blot mapping) and >2-kb upstream of 18S gene (based on cloning). The cloned 5S–18S fragment could potentially transcribe both 5S and 35S genes as regulatory motifs for both RNA polymerases were identified in the sequence. The direction of transcription would be opposite as the rRNAs are encoded from complementary DNA strands. In a cytogenetic study, five Podocarpus species of Australian and New Zealand origin showed colocalisation of 5S and 35S signals, mostly on a single chromosome (Murray et al., 2002). In our study, four Podocarpus species of South African origin and one of the closely related genus Afrocarpus, showed linkage between 5S and 35S genes. Thus, it seems that the linked character of 5S and 35S genes is a characteristic feature for both genera that may comprise about 100 (Podocarpus) and up to 6 (Afrocarpus) species, respectively.
L- and S-type rDNAs in Gnetales
In this restricted group of plants, which in our sample consists only on both Ephedra viridis and E. nebrodensis (Ephedraceae) we found cohybridisation of 26S, 18S and 5S probes. Sequencing of the 5S–18S E. nebrodensis clone showed the presence of a functional 5S gene in a direct orientation as in Ginkgo. Contrast to Ginkgo, the E. nebrodensis clone contained additional pseudogenised 5S copies, which were directly linked to the functional copy. The phylogenetic relationships between 5 S sequences showed that Ephedra genes are closely related to those of Pinaceae (in accord with the gnepine hypothesis—further below) while both of them are clearly separated from Ginkgo genes (Supplementary Figure S8). These observations suggest that L-type units in Ginkgo and Ephedra arose after divergence of these species as a result of independent insertions and subsequent homogenisation steps. The rDNA units with short 5S tandems embedded in the IGS seem to be homogenised as MboI digestion liberated the 5S monomers in E. nebrodensis genomic DNA after the Southern blot hybridisation (Supplementary Figure S4). However, divergence between three 5S copies present in the IGS of E. nebrodensis and mutation patterns (Supplementary Figure S7) indicate considerable lack of homogeneity. Thus concerted evolution does not seem to operate efficiently between the individual 5S repeats embedded in the 26 S–18 S spacer while it is highly effective at the higher order repeats comprising whole 35S–5S–Ψ5S–Ψ5S unit (see further below). Duplicated 5S copies were occasionally found in other L-type species (Garcia et al., 2009; Wicke et al., 2011). It is an open question whether all members of Gnetales family display L-type arrangement as the IGS clone from Gnetum gnemon (EMBL FR695703) appears to lack a 5S insertion (Wicke et al., 2011). Our results are consistent with S-type organisation of 5S units in this species supporting the original finding of Wicke et al. (2011).
The origin of Gnetales has often been disputed in the literature. The gnepine hypothesis places Gnetales as a sister to Pinaceae (Burleigh and Mathews, 2004; Wu et al., 2011) while the anthopyte hypothesis (Rydin et al., 2002) considers close relationship of Gnetales to angiosperms. According to Southern blot mapping and sequencing of Ephedra species, their rDNA units are similar to those of Pinaceae (pinophytes) in many aspects, both being clearly different from the angiosperm pattern. First, the lengths of Ephedra and Gnetum units (>20 kb) closely match those of Pinus (>27 kb) and Picea (>37 kb) (Bobola et al., 1992) while the typical length of angiosperm units ranges 10–14 kb (Hemleben and Zentgraf, 1994). Secondly, the 5S insertions in both Ephedra species analysed appear to be located distally from the 26S gene, as are those of Podocarpus and Afrocarpus (non-Pinaceae conifers). These observations would partially support the gnepine hypothesis although preliminary comparisons of 5S sequence divergence suggest an even more unique position of Gnetales in seed plant phylogeny (Garcia et al., unpublished results).
Homogenised L-type rDNAs in Ginkgoales
Opinions split on the organisation of 35S and 5S genes in G. biloba, the only representative of its order. While fluorescence in situ hybridisation showed colocalisation of 35S and 5S signals on chromosomes (Nakao et al., 2005; Galián et al., 2012), the PCR-cloning analysis of Wicke et al. (2011) failed to demonstrate 5S linkage to the large rDNA cluster. The pGIN2 clone isolated by us harboured a 5S insertion in direct orientation located downstream from the 26S gene. The different orientation of 5S insertions in Ginkgo and Podocarpus suggests their independent origins. The 5S coding region was 100% identical to the directly sequenced 5S rRNA from Ginkgo (GenBank M10433.1 from Hori et al., 1985) indicating that we cloned an apparently functional gene that is likely to be expressed. On the other hand, there were two mutations in the 5S coding region of the clone obtained by Galián et al. (2012) (GenBank JQ279501.1) although these may not influence gene functionality as both locate outside of the internal controlling region.
Despite these observations some uncertainty remains over the organisation of rDNA units in Ginkgo. This is because there have been six IGS clones obtained in different laboratories out of which only two (GenBank JQ279501 and JX402064, from the University of Valencia, ES and Academy of Sciences, CZ, respectively) contained 5S insertions, while these were absent in the remaining four clones (GenBank FR695705-7 and JQ279502, from the University of Vienna, AU and University of Valencia, ES, respectively). Furthermore, there is apparent inconsistency between the unit size calculated from cloning experiments (<12 kb, in Galián et al. 2012) and Southern blot mapping (>20 kb). So, we are left with several possible explanations. The clone to clone differences can be explained by unhomogenised rDNA pools containing multiple gene families and pseudogenised copies, as suggested by Galián et al. (2012). Next generation sequencing of ITS pools revealed that the intragenomic diversity of rDNA units is higher in species with multiple chromosomal loci than in species with a single locus in Nicotiana (Matyasek et al., 2012). In this context, sex chromosomes in G. biloba (Newcomer, 1954) distinguished by number of satellites that likely represent rDNA loci, may generate some unit diversity. However, the Southern hybridisations do not support significant heterogeneity in the spectrum of rDNA families, and >90% of rDNAs seem to form a single or possibly two families. Such pattern is consistent with clustered rDNA-fluorescence in situ hybridisation signals on Ginkgo chromosomes (Nakao et al., 2005; Galián et al., 2012). We favour the explanation that the differences between individual IGS clones (with and without 5S insertions) isolated in different laboratories could be caused by biased amplification of rare rDNA variants (pseudogenes) and/or a PCR failure to accurately amplify DNA templates. The latter hypothesis is supported by the observation that different thermostable polymerases appear to produce products of different lengths on Ginkgo’s DNA (not shown). The longest products were obtained with polymerases bearing a proofreading activity (3′→5′exonuclease activity), and we recommend preferential usage of these polymerases when amplifying difficult IGS templates, in general.
Together, the data in this and others work (Galián et al., 2012) support the hypothesis that the predominant organisation of Ginkgo rDNAs is L-type.
Comparison of rDNA organisation in gymnosperms and angiosperms
The S-type arrangement seems to predominate in most angiosperm plant lineages (Campell et al., 1992; Sastri et al., 1992; Wicke et al., 2011) while in some families the L-type arrangement is quite frequent (Garcia et al., 2010). In gymnosperms the situation may be similar, in that the species from the group with the greatest diversity (Pinaceae conifers, accounting for almost one quarter of gymnosperm diversity) have the separate arrangement; in other lineages, however, such as Podocarpaceae, Ephedraceae and Ginkgoaceae, the L-type arrangement seems to be the dominant. The phylogenetic relationships (Figure 1) indicate that the 5S rDNA has changed its fundamental genomic organisation at least three times during the evolution of gymnosperms. Hence, conclusions based on single species observations should not be generalised. As in the angiosperms, parallel existence of comparable number of L-type and S-type units in the same genome has not been observed, suggesting that homogenisation pressures acting on rDNA repeats are strong. Yet coexistence of minor S-type loci along with major L-type arrangement (and vice versa) cannot be excluded as in some angiosperm species (Garcia et al., 2010, 2012a). Sequence analysis of 5S insertions, however, did reveal differences between angiosperms and gymnosperms. First, while in angiosperms the 5S insertions occur in a non-repetitive part of the IGS (within <1 kb downstream from the 26S gene), in gymnosperms the 5S gene is located distally to the 26S and 18S genes, embedded in a highly repetitive DNA region as in bryophytes (Sone et al., 1999). Second, to our knowledge the 5S genes of angiosperms are exclusively encoded by the opposite DNA strand to the 35S genes while in gymnosperms both direct and inverse orientations occur. Third, many angiosperm species bear retroelement signatures in regions flanking the 5S insertions resembling Cassandra TRIM elements (Kalendar et al., 2008). No such features have been found in sequences of gymnosperm units. Perhaps, retroelement motifs might have already been eroded and/or replaced by tandem repeats in gymnosperms.
Unit arrangement and chromosomal position of rDNA arrays
Contrast to angiosperms, relatively little data exists on the chromosomal organisation of rDNA in gymnosperms. In most angiosperm species the 35S rDNA genes are located at (sub)telomeric positions of short arms (Roa and Guerra, 2012) while this trend is not so obvious in gymnosperms, in which interstitial positions were reported, particularly in Pinaceae (Islam-Faridi et al., 2007). However, in both gymnosperms and angiosperms analysed to date the L-type rDNAs occur at terminal position (Garcia et al., 2012b and Garcia et al., unpublished). It will be interesting to determine chromosomal position of rDNA in other species in order to see whether the (sub)telomeric locations are favoured by the L-type arrays. It is also an open question of whether inversions, fusions (Olson and Gorelick, 2011) and other fundamental changes in chromosome structure (Leitch and Leitch, 2012) that accompany the long (300 Myrs) history of gymnosperm evolution contribute to switches in rDNA arrangements.
Conclusions
Our data show that in G. biloba the 5S genes integrated into the 26S–18S spacer and are nearly homogenised across the genome (L-type arrangement). Similar arrangement has been found in genera Podocarpus, Afrocarpus (non-Pinaceae Coniferales or cupressophytes) and Ephedra (Gnetales) while other gymnosperm lineages, including the cycads evolved a mostly separated arrangement of rDNAs. As both cycads and Ginkgo are considered to be early diverging 250–300 Myrs-old species (Zhou and Zheng, 2003) and display contrasting L- and S-type arrangements, respectively, it is likely that the two rDNA organisations are evolutionary neutral as both are also present in modern plants. Rapid switches between fundamental organisations of these vital rRNA genes are unexpected, yet occurred frequently in both gymnosperm and angiosperm lineages. The overwhelming prevalence of S-type arrangement in seed plants, however, still needs to be explained.
Data archiving
Sequence data have been submitted to GenBank: accession numbers JX402063, JX402064 and JX843794.
IGS and 5S sequences from G. biloba deposited in the Dryad repository: doi:10.5061/dryad.fq228
References
Amarasinghe V, Carlson JE (1998). Physical mapping and characterization of 5S rRNA genes in Douglas-fir. J Hered 89: 495–500.
Benson G (1999). Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res 27: 573–580.
Besendorfer V, Krajacic-Sokol I, Jelenic S, Puizina J, Mlinarec J, Sviben T et al (2005). Two classes of 5S rDNA unit arrays of the silver fir, Abies alba Mill.: structure, localization and evolution. Theor Appl Genet 110: 730–741.
Bobola MS, Smith DE, Klein AS (1992). Five major nuclear ribosomal repeats represent a large and variable fraction of the genomic DNA of Picea rubens and P. mariana. Mol Biol Evol 9: 125–137.
Bogunic F, Siljak-Yakovlev S, Muratovic E, Ballian D (2011). Different karyotype patterns among allopatric Pinus nigra (Pinaceae) populations revealed by molecular cytogenetics. Plant Biol 13: 194–200.
Brown GR, Carlson JE (1997). Molecular cytogenetics of the genes encoding 18s-5.8s-26s rRNA and 5s rRNA in two species of spruce (Picea). Theor Appl Genet 95: 1–9.
Burleigh JG, Mathews S (2004). Phylogenetic signal in nucleotide data from seed plants: Implications for resolving the seed plant tree of life. Am J Bot 91: 1599–1613.
Cai Q, Zhang DM, Liu ZL, Wang XR (2006). Chromosomal localization of 5S and 18S rDNA in five species of subgenus Strobus and their implications for genome evolution of Pinus. Ann Bot 97: 715–722.
Campell BR, Song Y, Posch TE, Cullis CA, Town CD (1992). Sequence and organization of 5S ribosomal RNA-encoding genes of Arabidopsis thaliana. Gene 112: 225–228.
Chaw SM, Aharkikh A, Sung HM, Lau TC, Li WH (1997). Molecular phylogeny of extant gymnosperms and seed plant evolution: analysis of nuclear 18SrRNA sequences. Mol Biol Evol 14: 56–68.
Cloix C, Yukawa Y, Tutois S, Sugiura M, Tourmente S (2003). In vitro analysis of the sequences required for transcription of the Arabidopsis thaliana 5S rRNA genes. Plant J 35: 251–261.
Cullis CA, Creissen GP, Gorman SW, Tiasdale RD (1988). The 25S, 18S, and 5S ribosomal RNA genes from Pinus radiata D. DON. In: Cheliak WM, Yapa AA, (eds) Molecular Genetics of Forest Tree Vol. PI-X-80, Canadian Forest Service: Petawa National Forest Institute. pp 34–40.
Doudrick RL, Heslopharrison JS, Nelson CD, Schmidt T, Nance WL, Schwarzacher T (1995). Karyotype of slash pine (Pinus-Elliottii Var Elliottii) using patterns of fluorescence in-situ hybridization and fluorochrome banding. J Hered 86: 289–296.
Doyle JJ, Doyle JL (1987). A rapid DNA isolation procedure from smallquantities of fresh leaf tissue. Phytochem Bull 19: 11–15.
Drouin G, de Sa MM (1995). The concerted evolution of 5S ribosomal genes linked to the repeat units of other multigene families. Mol Biol Evol 12: 481–493.
Felsenstein J (1989). PHYLIP — phylogeny inference package (version 3.2). Cladistics 5: 164–166.
Fulnecek J, Lim KY, Leitch AR, Kovarik A, Matyasek R (2002). Evolution and structure of 5S rDNA loci in allotetraploid Nicotiana tabacum and its putative parental species. Heredity 88: 19–25.
Fulnecek J, Matyasek R, Kovarik A (2006). Plant 5S rDNA has multiple alternative nucleosome positions. Genome 49: 840–850.
Galián JA, Rosato M, Rosselló JA (2012). Early evolutionary colocalization of the nuclear ribosomal 5S and 45S gene families in seed plants: evidence from the living fossil gymnosperm Ginkgo biloba. Heredity 108: 640–646.
Garcia S, Crhak Khaitova L, Kovarik A (2012a). Expression of 5S rRNA genes linked to 35S rDNA in plants, their epigenetic modification and regulatory element divergence. BMC Plant Biol 12: 95.
Garcia S, Garnatje T, Hidalgo O, McArthur ED, Siljak-Yakovlev S, Valles J (2007). Extensive ribosomal DNA (18S-5.8S-26S and 5S) colocalization in the North American endemic sagebrushes (subgenus Tridentatae, Artemisia, Asteraceae) revealed by FISH. Plant Syst Evol 267: 79–92.
Garcia S, Garnatje T, Kovarik A (2012b). Plant rDNA database: ribosomal DNA loci information goes online. Chromosoma 121: 389–394.
Garcia S, Lim KY, Chester M, Garnatje T, Pellicer J, Valles J et al (2009). Linkage of 35S and 5S rRNA genes in Artemisia (family Asteraceae): first evidence from angiosperms. Chromosoma 118: 85–97.
Garcia S, Panero JL, Siroky J, Kovarik A (2010). Repeated reunions and splits feature the highly dynamic evolution of 5S and 35S ribosomal RNA genes (rDNA) in the Asteraceae family. BMC Plant Biology 10: 176.
Gorman SW, Teasdale RD, Cullis CA (1992). Structure and organization of the 5S ribosomal-RNA genes (5S-DNA) in Pinus-radiata (pinaceae). Plant Syst Evol 183: 223–234.
Gouy M, Guindon S, Gascuel O (2010). SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol Biol Evol 27: 221–224.
Hall TA (1999). BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acids Symp Ser 41: 95–98.
Hemleben V, Zentgraf U (1994). Structural organisation and regulation of transcription by RNA polymerase I of plant nuclear ribosomal genes. In: L N, (eds). Results and Problems in Cell Differentiation 20: Plant Promoters and Transcription Factors. Springer-Verlag: Berlin/Heidelberg. pp 3–24.
Hizume M, Akiyama M (1992). Size variation of chromomycin a(3)-band in chromosomes of Douglas-fir, Pseudotsuga-menziesii. Jpn J Genet 67: 425–435.
Hizume M, Shibata F, Matsusaki Y, Garajova Z (2002). Chromosome identification and comparative karyotypic analyses of four Pinus species. Theor Appl Genet 105: 491–497.
Hizume M, Shibata F, Matsusaki Y, Kondo T (1999). Mapping of 45S rRNA and 5S rRNA loci in Cryptomeria japonica, Cunninghamia lanceolata and Ginkgo biloba. Chromosome Sci 3: 142.
Hori H, Lim BL, Osawa S (1985). Evolution of green plants as deduced from 5S ribosomal-RNA sequences. Proc Natl Acad Sci USA 82: 820–823.
Islam-Faridi MN, Nelson CD, Kubisiak TL (2007). Reference karyotype and cytomolecular map for loblolly pine (Pinus taeda L.). Genome 50: 241–251.
Kalendar R, Tanskanen J, Chang W, Antonius K, Sela H, Peleg O et al (2008). Cassandra retrotransposons carry independently transcribed 5S RNA. Proc Natl Acad Sci USA 105: 5833–5838.
Karvonen P, Savolainen O (1993). Variation and inheritance of ribosomal DNA in Pinus-Sylvestris L (Scots Pine). Heredity 71: 614–622.
Kiss T, Szkukalek A, Solymosy F (1989). Nucleotide sequence of a 17S (18S) rRNA gene from tomato. Nucleic Acids Res 17: 2127.
Komarova NY, Grabe T, Huigen DJ, Hemleben V, Volkov RA (2004). Organization, differential expression and methylation of rDNA in artificial Solanum allopolyploids. Plant Mol Biol 56: 439–463.
Kovarik A, Pires JC, Leitch AR, Lim KY, Sherwood AM, Matyasek R et al (2005). Rapid concerted evolution of nuclear ribosomal DNA in two Tragopogon allopolyploids of recent and recurrent origin. Genetics 169: 931–944.
Leitch AR, Leitch IJ (2012). Ecological and genetic factors linked to contrasting genome dynamics in seed plants. New Phytol 194: 629–646.
Lim KY, Kovarik A, Matyasek R, Bezdek M, Lichtenstein CP, Leitch AR (2000). Gene conversion of ribosomal DNA in Nicotiana tabacum is associated with undermethylated, decondensed and probably active gene units. Chromosoma 109: 161–172.
Liu B, Zhang SG, Zhang Y, Lan TY, Qi LW, Song WQ (2006). Molecular cytogenetic analysis of four Larix species by bicolor fluorescence in situ hybridization and DAPI banding. Int J Plant Sci 167: 367–372.
Liu ZL, Zhang D, Hong DY, Wang XR (2003b). Chromosomal localization of 5S and 18S-5.8S-25S ribosomal DNA sites in five Asian pines using fluorescence in situ hybridization. Theor Appl Genet 106: 198–204.
Liu ZL, Zhang D, Wang XQ, Ma XF, Wang XR (2003a). Intragenomic and interspecific 5S rDNA sequence variation in five Asian pines. Am J Bot 90: 17–24.
Lubaretz O, Fuchs J, Ahne R, Meister A, Schubert I (1996). Karyotyping of three Pinaceae species via fluorescent in situ hybridization and computer-aided chromosome analysis. Theor Appl Genet 92: 411–416.
Mashkova TD, Barciszewska MZ, Joachimiak A, Nalaskowska M, Barciszewski J (1990). Molecular evolution of plants as deduced from changes in free energy of 5S ribosomal RNAs. Int J Biol Macromol 12: 247–250.
Mathews S (2009). Phylogenetic relationships among seed plants: persistent questions and the limits of molecular data. Am J Bot 96: 228–236.
Matyasek R, Renny-Byfield S, Fulnecek J, Macas J, Grandbastien MA, Nichols RA et al (2012). Next generation sequencing analysis reveals a relationship between rDNA unit diversity and locus number in Nicotiana diploids. BMC Genomics 13: 722.
Mazzella C, Rodriguez M, Vaio M, Gaiero P, Lopez-Carro B, Santinaque FF et al (2010). Karyological features of Achyrocline (Asteraceae, Gnaphalieae): stable karyotypes, low DNA content variation and linkage of rRNA genes. Cytogenet Genome Res 128: 169–176.
McClelland M, Nelson M (1985). The effect of site specific methylation on restriction endonuclease digestion. Nucleic Acids Res 13 (Suppl): r201–r207.
Melekhovets YF, Troitsky AV, Valiejo-Roman KM, Bobrova VK, Antonov AS (1988). Nucleotide sequences of cytosolic 5S ribosomal RNAs from two gymnosperms, Gnetum gnemon and Ephedra kokanica. Nucleic Acids Res 16: 4155.
Miranda M, Almeida CCD, Guerra M (2007). Karyotype of Araucaria angustifolia and the decondensation/activation mode of its nucleolus organiser region. Aust J Bot 55: 165–170.
Moran GF, Smith D, Bell JC, Appels R (1992). The 5S-RNA genes in pinus-radiata and the spacer region as a probe for relationships between Pinus species. Plant Syst Evol 183: 209–221.
Murray BG, Friesen N, Heslop-Harrison JS (2002). Molecular cytogenetic analysis of Podocarpus and comparison with other gymnosperm species. Ann Bot 89: 483–489.
Nagano K, Matoba H, Yonemura K, Matsuda Y, Murata T, Hoshi Y (2007). Karyotype analysis of three Juniperus species using fluorescence in situ hybridization (FISH) with two ribosomal RNA genes. Cytologia 72: 37–42.
Nakao Y, Taira T, Horiuchi S, Kenji K, Mukai Y (2005). Chromosomal difference between male and female trees of Ginkgo biloba examined by karyotype analysis and mapping of rDNA on the chromosomes by fluorescence in situ hybridization. J Jpn Soc Hortic Sci 74: 275–280.
Newcomer EH (1954). The karyotype and possible sex chromosomes of Ginkgo-Biloba. Am J Bot 41: 542–545.
Noe L, Kucherov G (2005). YASS: enhancing the sensitivity of DNA similarity search. Nucleic Acids Res 33 (Web Server issue): W540–W543.
Olson K, Gorelick R (2011). Chromosomal fission accounts for small-scale radiations in Zamia (Zamiaceae; Cycadales). Bot J Linn Soc 165: 168–185.
Perry KL, Palukaitis P (1990). Transcription of tomato ribosomal DNA and the organization of the intergenic spacer. Mol Gen Genet 221: 103–112.
Puizina J, Sviben T, Krajacic-Sokol I, Zoldos-Pecnik V, Siljak-Yakovlev S, Papes D et al (2008). Cytogenetic and molecular characterization of the Abies alba genome and its relationship with other members of the Pinaceae. Plant Biol 10: 256–267.
Roa F, Guerra M (2012). Distribution of 45S rDNA sites in chromosomes of plants: structural and evolutionary implications. BMC Evol Biol 12: 225.
Rydin C, Kallersjo M, Friist EM (2002). Seed plant relationships and the systematic position of Gnetales based on nuclear and chloroplast DNA: conflicting data, rooting problems, and the monophyly of conifers. Int J Plant Sci 163: 197–214.
Sastri DC, Hilu K, Appels R, Lagudah ES, Playford J, Baum BR (1992). An overview of evolution on plant 5S-DNA. Plant Syst Evol 183: 169–181.
Seitz U, Seitz U (1979). Molecular-weight of ribosomal-RNA precursor molecules and their processing in higher-plant cells. Z Naturforsch C 34: 253–258.
Shibata F, Hizume M (2008). Comparative FISH karyotype analysis of 11 Picea species. Cytologia 73: 203–211.
Siljak-Yakovlev S, Cerbah M, Coulaud J, Stoian V, Brown SC, Zoldos V et al (2002). Nuclear DNA content, base composition, heterochromatin and rDNA in Picea omorika and Picea abies. Theor Appl Genet 104: 505–512.
Sone T, Fujisawa M, Takenaka M, Nakagawa S, Yamaoka S, Sakaida M et al (1999). Bryophyte 5S rDNA was inserted into 45S rDNA repeat units after the divergence from higher land plants. Plant Mol Biol 41: 679–685.
Tagashira N, Kondo K (2001). Chromosome phylogeny of Zamia and Ceratozamia by means of Robertsonian changes detected by fluorescence in situ hybridization (FISH) technique of rDNA. Plant Syst Evol 227: 145–155.
Trontin JF, Grandemange C, Favre JM (1999). Two highly divergent 5S rDNA unit size classes occur in composite tandem array in European larch (Larix decidua Mill.) and Japanese larch (Larix kaempferi (Lamb.) Carr.). Genome 42: 837–848.
Vincze T, Posfai J, Roberts RJ (2003). NEBcutter: a program to cleave DNA with restriction enzymes. Nucleic Acids Res 31: 3688–3691.
Wicke S, Costa A, Munoz J, Quandt D (2011). Restless 5S: the re-arrangement(s) and evolution of the nuclear ribosomal DNA in land plants. Mol Phylogenet Evol 61: 321–332.
Wu CS, Wang YN, Hsu CY, Lin CP, Chaw SM (2011). Loss of different inverted repeat copies from the chloroplast genomes of Pinaceae and Cupressophytes and influence of heterotachy on the evaluation of gymnosperm phylogeny. Genome Biol Evol 3: 1284–1295.
Zhang SG, Yang WH, Han SY, Han BT, Li MX, Qi LW (2010). Cytogenetic analysis of reciprocal hybrids and their parents between Larix leptolepis and Larix gmelinii: implications for identifying hybrids. Tree Genet Genomes 6: 405–412.
Zhou ZY, Zheng SL (2003). Palaeobiology: the missing link in Ginkgo evolution—The modern maidenhair tree has barely changed since the days of the dinosaurs. Nature 423: 821–822.
Acknowledgements
The authors are grateful to personnel from the Botanical Garden of Barcelona and particularly to Samuel Pyke and Miquel Veny (Institut Botànic de Barcelona-CSIC) who assessed at the identification and supervised the collection of plant materials from the garden. The comments and suggestions on the work from Dr Teresa Garnatje (Institut Botànic de Barcelona-CSIC-ICUB), Dr Josep Rosselló (Universitat de València) and Professor Joan Vallès (Universitat de Barcelona), have been much appreciated. We thank Dr Ilia Leitch (Royal Botanic Gardens, Kew) for providing Gnetum gnemon leaf material. Alastair Plant (Ohio University) is thanked for reading and correcting the English of the manuscript. This work has been supported by the Spanish and Catalan governments (projects CGL2010-22234-C02-01 and 02/BOS, and project 2009SGR00439, respectively), Academy of Sciences of the Czech Republic (RVO68081707) and the Czech Science Foundation (P501/13/10057S; P501/12/G090). SG benefited from a Beatriu de Pinós postdoctoral contract with the support of the Comissionat per a Universitats i Recerca (CUR) del Departament d’Economia i Coneixement (Catalan government).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Heredity website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Garcia, S., Kovařík, A. Dancing together and separate again: gymnosperms exhibit frequent changes of fundamental 5S and 35S rRNA gene (rDNA) organisation. Heredity 111, 23–33 (2013). https://doi.org/10.1038/hdy.2013.11
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2013.11
Keywords
This article is cited by
-
Intragenomic rDNA variation - the product of concerted evolution, mutation, or something in between?
Heredity (2023)
-
Chromosomal mapping of 5S and 18S-5.8S-25S rRNA genes in Saccharina japonica (Phaeophyceae) as visualized by dual-color fluorescence in situ hybridization
Journal of Oceanology and Limnology (2021)
-
Heterochromatin Bands and rDNA Sites Evolution in Polyploidization Events in Cynodon Rich. (Poaceae)
Plant Molecular Biology Reporter (2019)
-
Rapid and Efficient FISH using Pre-Labeled Oligomer Probes
Scientific Reports (2018)
-
Retroelements and DNA Methylation Could Contribute to Diversity of 5S rDNA in Agave L.
Journal of Molecular Evolution (2018)