Abstract
The process of dog domestication is still somewhat unresolved. Earlier studies indicate that domestic dogs from all over the world have a common origin in Asia. So far, major histocompatibility complex (MHC) diversity has not been studied in detail in Asian dogs, although high levels of genetic diversity are expected at the domestication locality. We sequenced the second exon of the canine MHC gene DLA–DRB1 from 128 Asian dogs and compared our data with a previously published large data set of MHC alleles, mostly from European dogs. Our results show that Asian dogs have a higher MHC diversity than European dogs. We also estimated that there is only a small probability that new alleles have arisen by mutation since domestication. Based on the assumption that all of the currently known 102 DLA–DRB1 alleles come from the founding wolf population, we simulated the number of founding wolf individuals. Our simulations indicate an effective population size of at least 500 founding wolves, suggesting that the founding wolf population was large or that backcrossing has taken place.
Similar content being viewed by others
Introduction
The dog has a special position in human society. It was probably the first domesticated species (Clutton-Brock, 1995; Raisor, 2005; Pang et al., 2009) and it is currently the most widespread and diverse domestic animal. The questions of where and how dog domestication took place still remain somewhat unresolved. Recent genetic studies on the origin of domestic dogs have produced contrasting results, but all indicate that the geographical origin is somewhere in Asia. Studies of mitochondrial DNA (mtDNA) suggest that domestic dogs from all over the world have a common origin in East Asia (Leonard et al., 2002; Savolainen et al., 2002). Pang et al. (2009) narrowed the domestication locality to the southeastern part of Asia, south of the Yangtze River. On the other hand, based on genome-wide single-nucleotide polymorphism data in nuclear DNA, but less comprehensive sampling of Asian dogs, vonHoldt et al. (2010) suggested the Middle East as the main area of origin. Notably, all previous studies indicate an origin in Asia and, by contrast, show limited genetic diversity among the European dogs.
An analysis of mtDNA of dogs indicated that the global dog population originated from a minimum of 51 female wolves (Pang et al., 2009). Analysis based on maternally inherited mtDNA is problematic for estimating the number of founders because it does not depict the biparental evolutionary history. In addition, the effective population number for mtDNA is only one-fourth of that of the nuclear loci and mtDNA has a high mutation rate, which both complicate size estimates of founder populations. These problems can be overcome by using autosomal markers, which are retained in a population for a long time. The genes of the major histocompatibility complex (MHC) are suitable for this purpose, as suggested by Wayne and Ostrander (2007). MHC molecules have two pivotal roles in the immune system: (i) they are required for initiating immune response against pathogens (Germain, 1994), and (ii) they help in discriminating between self and non-self. High numbers of MHC alleles are essential in defeating the variety of pathogens. It is commonly accepted that MHC polymorphism is maintained by balancing selection, which hinders alleles from being lost or reaching fixation (Hedrick, 1999; Spurgin and Richardson, 2010). Instead, a great number of alleles fluctuate at intermediate frequencies over extended periods of time. This leads to a phenomenon known as trans-species polymorphism, in which similar alleles are conserved in different species (Klein, 1987). Retention of alleles over long periods of time make MHC genes suitable for population studies at a longer time scale than in other molecular markers (Prugnolle et al., 2005; Wayne and Ostrander, 2007). Therefore, MHC genes have been used in estimating the founder population sizes of, for example, Darwin’s finches (Vincek et al., 1997), humans (Klein et al., 1990; Ayala et al., 1994) and domestic mammals, including dogs (Vilà et al., 2005).
There are two major sources for new alleles in domestic animals: (i) mutation and (ii) gene flow from an ancestral or related species. In MHC genes, recombination also has a significant role in reorganizing sequences. In some cases, recombination may even outscore mutation in creating polymorphism (Schaschl et al., 2006). By estimating the number of novel alleles that have arisen since domestication, it is possible to estimate the number of alleles passed on from the founder population. Although the dog is the oldest domestic animal, from an evolutionary point of view it was domesticated relatively recently. Archeological as well as genetic evidence indicate that domestication took place approximately 15 000 years ago (Savolainen et al., 2002; Pang et al., 2009); an origin as early as 30 000 years ago has been suggested, but based on inconclusive archeological evidence (Germonpré et al., 2009). Both mtDNA and Y chromosomal studies suggest that introgression from wolves after domestication has been rare (Sundqvist, 2008; Pang et al., 2009).
Canine MHC (also referred to as dog leukocyte antigen, DLA) gene variation has been examined mostly in European dogs so far (Kennedy et al., 2007b), whereas mtDNA studies have shown that genetic variation is highest in East Asia and among the lowest in Europe. Therefore, it is probable that earlier studies of DLA diversity have revealed only a small part of the total diversity. In this study, we analyzed the highly polymorphic second exon of DLA–DRB1 gene from dogs across Asia. Combined with previously published DLA–DRB1 data (Kennedy et al., 2007b), we generated a more comprehensive picture of the global DLA diversity and used this information in simulations to estimate the minimum number of founding wolves for the current dog population.
Materials and methods
Samples
In total, 128 samples were collected from 69 dogs of North or East Asian (N/E Asia) origin and 59 dogs of South or West Asian (S/W Asia) origin (Supplementary Figure S1 and Supplementary Table S1). N/E Asia was defined as Asia east of the Ural Mountains and north of the Himalayas and S/W Asia was defined as Asia west and south of the Himalayas. These samples were combined with data from the largest study so far for DLA (Kennedy et al., 2007b). The 1484 samples that could be assigned to their geographic origins were used; 1051 of the samples were from Europe, 112 from South America, 110 from Africa, 90 from North America, 61 from S/W Asia and 60 from N/E Asia (Supplementary Table S2). A dog was assigned to a geographic region if (i) it belonged to a breed with a historic origin from the region or (ii) it was a non-breed dog from a remote part of the region that has presumably received little influx of foreign dogs. Most of the Asian breed dogs were sampled from Scandinavia, hence the environments of the Asian and European breed dogs were similar with respect to, for example, parasite load and mate choice.
DNA techniques
DNA was extracted from blood, hair or buccal cell swabs preserved on FTA-cards (Whatman International, Maidstone, UK). Amplification of the second exon of the DLA–DRB1 gene was carried out with a semi-nested PCR (Mullis and Faloona, 1987). DNA sequences were edited using Sequencing Analysis (Applied Biosystems, Carlsbad, CA, USA), assembled into contigs and further edited in Sequencher 4.1 (Gene Codes, Ann Arbor, MI, USA). To identify the alleles from heterozygous individuals, all heterozygotes were cloned using GeneJET PCR Cloning Kit (Fermentas, Burlington, Canada). All novel alleles were verified by a second independent analysis using another set of PCR primers (Kennedy et al., 2007a), to ensure that these alleles had not gone undetected in earlier studies because of primer mismatch (see Supplementary methods for detailed methodology).
Analyses of sequence data and tree construction
All available full-length sequences (270 bp) of the second exon of DLA–DRB1 gene were retrieved from NCBI GenBank database (Supplementary Table S3) and aligned in BioEdit 7 (Hall, 1999). The sequences included dogs mainly from Europe (Sarmiento and Storb, 1990; Murgia et al., 2006; Kennedy et al., 2007b), but also a lesser number of dogs from other regions (Runstadler et al., 2006) and wolves (Seddon and Ellegren, 2002; Kennedy et al., 2007a). To study phylogenetic relationships between the newly identified and previously published DLA alleles, a neighbor-joining tree was constructed for dog and wolf alleles with the software MEGA 4.1 (Tamura et al., 2007) using Jukes–Cantor distances.
Regional genetic diversity
Numbers of alleles (A) and numbers of unique alleles (AU) were compared between geographical regions. Alleles were categorized as unique if they were found only in one region. Genetic diversity in Asian dogs was compared with the diversity of the dogs from different regions from Kennedy et al. (2007b). Alleles obtained from GenBank from publications other than Kennedy et al. (2007b) could not be used in comparisons of diversity because the alleles could not be assigned to the breeds of origin.
Allelic richness (AR) was estimated using the program FSTAT v. 2.9.3.2 (Goudet, 2001) and observed heterozygosity (Ho) was estimated with Arlequin v. 3.11 (Excoffier et al., 2005). These parameters were estimated separately for the breed dogs and non-breed dogs in the N/E and S/W Asia. As a result it was not possible to assign genotypes for the individuals in Kennedy et al. (2007b), the data were not suitable for estimation of expected heterozygosity, allelic richness or other corrections for the sample size.
To compare how evenly each breed was sampled within a region, we calculated the evenness based on Shannon’s diversity index (Pielou, 1966):

where pi is the relative abundance of the breed i and S is the number of the breeds. Evenness or 'equality' is a measure of how similar the abundances of different breeds are in a given region. The measure assumes a value between 0 and 1. When there are similar proportions of all breeds, the evenness is one, and when the abundances are very dissimilar (some rare and some common) the value decreases.
To obtain a measure of evolutionary distinctiveness of different DLA–DRB1 alleles, we estimated Faith phylogenetic diversity (Crozier, 1997; Faith, 1992) for all regions using the program Conserve IV v.1.4 (Crozier et al., 1999). The program estimates phylogenetic diversity as a measure of the diversity of a subset of samples (the sum of the branch lengths in a phylogeny) relative to the length of the neighbor-joining tree of the complete set of samples. The neighbor-joining tree was constructed for dog alleles (270 bp) with the software MEGA 4.1 (Tamura et al., 2007) using Jukes–Cantor distances.
Estimation of number of founders
We assumed that mutation and recombination are the main sources for new alleles and calculated the number of alleles arisen after domestication with μtn (Vilà et al., 2005). Non-synonymous substitution rate (μ) was set to 5.9*10−9 per non-synonymous site per year (Klein et al., 1993), time since domestication (t) was set to 15 000 or to the more extreme 30 000 years (Savolainen et al., 2002; Germonpré et al., 2009) and the number of non-synonymous sites (n) within the second exon of the DLA–DRB1 gene was 200.39 (Vilà et al., 2005). Schaschl et al. (2006) showed that, on average, the effect of recombination to the DRB polymorphism in ungulates is eight times higher than the effect of mutation. Based on this result, we calculated the probability of novel alleles assuming that recombination increases the number of alleles at the same level as μ (in total equaling to μ=1.18*10–8), 10 times μ (μ=6.49*10–8) and 100 times μ (μ=5.96*10–7).
To estimate the minimum population size of wolf founders, we conducted simulations following the model of Vincek et al. (1997). In the model, 2Ne alleles are drawn and paired randomly in each generation. The probability of each allele pair to pass on to the next generation is one for heterozygotes and one-selection coefficient (s) for homozygotes. Dog has a single-locus DLA–DRB1 (Yuhki et al., 2007) and thus all heterozygosity is caused by variants of one locus. The effect of evolutionary forces and demography to the number of alleles was simulated with different population parameter values. These included founder population size (Ne=100–700), selection against homozygosity (s=0.01 (Satta et al., 1994) or 0.1), population growth rate (g=0.02, 0.05 or 0.1) and number of founder population alleles (Af=150 or 200). Initial allele frequencies in the founder population were set to be even. Mutation (infinite allele model) was included in the simulation with the mutation rate of μ=1.18*10–8. In addition, more extreme values were tested up to the magnitude of 10–7. Population was let to grow to 1000 individuals and each simulation was repeated 1000 times.
Results
Large number of novel DRB1 alleles among Asian dogs
Among the 128 dogs studied, 51 DLA–DRB1 alleles were identified. Five of these have previously not been submitted to GenBank and 17 are novel alleles. In all, 11 of the alleles were unique to N/E Asian dogs and 11 to S/W Asian dogs (Table 1). Allelic richness was at a similar level in the Asian regions for both breed and non-breed dogs. Observed heterozygosities were surprisingly low, between 0.313 and 0.455 among S/W Asian breed dogs and N/E Asian non-breed dogs, respectively. Considering the different environmental pressures and demographic factors for breed and non-breed dogs, the allelic richnesses and observed heterozygosities for the two groups were unexpectedly similar.
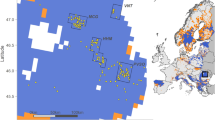
When the data of this study were combined with data of Kennedy et al. (2007b) for a global geographic comparison, the total number of alleles increased to 71 (Table 2). Other alleles obtained from GenBank could not be used in the diversity comparisons between regions, because they could not be assigned to the breeds of origin. The lowest number of alleles was found from North America (7) and the highest number from N/E Asia (36, Figure 1). The lowest number of unique alleles was also found among North American (1) and the highest among N/E Asian dogs (16). Phylogenetic diversity was similar in S/W Asia (0.585) and N/E Asia (0.584). Also other diversity values were very similar between these two regions. These values are strongly influenced by sampling schemes and sample sizes. Therefore, it is remarkable that the highest number of alleles and unique alleles were found among Asian dogs despite the small sample sizes.
The proportions of breed dogs differ among the regions (Table 2), which may bias comparisons of genetic diversity. However, the percentage of breed dogs was almost the same in the N/E Asian (91.5%) and European samples (91.3%), and still the genetic diversity was higher in the N/E Asian (A=36, AU=16) compared with the eight times larger European sample (A=30, AU=9). In addition, the numbers of individuals per breed were also uneven (Table 2), the African sample being the least even (J=0.09) and the N/E Asian sample the most even (J=0.83). The European and both Asian samples had similar evenness values (Table 2) suggesting that the sampling scheme did not have a large effect on the diversity values among these regions.
In the neighbor-joining tree, the newly identified full-length alleles from Asia were distributed among the previously published dog and wolf DLA–DRB1 sequences (Figure 2). There was no clustering of the new alleles that could be explained by population structure, selection or primers that amplify selectively.
Simulations indicate large number of founders for the dog population
The number of alleles descending from the time before dog domestication indicates the minimum number of wolf ancestors. Assuming a non-synonymous substitution rate of μ=5.9*10–9, the probability of new alleles arisen since domestication is 0.018 (15 000 years since domestication) or 0.035 (30 000 years since domestication). With the non-synonymous substitution rate of μ=1.18*10−8 and assuming that recombination increases the number of alleles at the same level as μ, the probability of new alleles is 0.035 or 0.071. If we assume that recombination increases the number of alleles 10 or, unlikely high, 100 times as much as μ, the probability of new alleles would be 0.195 or 0.390 and 1.791 or 3.583, respectively. Consequently, these results suggest that most or all of the alleles found in the current dog population were present already among the wolf ancestors; there is only a small chance that many new alleles have arisen by mutation since dog domestication. The number of new alleles would be considerable only if recombination rate is unlikely high.
In total, 102 DLA–DRB1 alleles (88 with non-synonymous differences) have been submitted to GenBank (Supplementary Table S3) suggesting a minimum of 51 wolf founders. However, this estimate assumes that all founding individuals were heterozygous and carried different alleles each. These assumptions are very unrealistic and the number of founding individuals must have been higher than this minimum estimate. In addition, many alleles may have been lost because of genetic drift during the domestication process. The effect of drift was probably strongest at the beginning of the process when the population size was small. We simulated the effect of drift to the number of alleles. A few possible scenarios are shown in Figure 3. According to these scenarios, the effective founder population size must have been at least 500 individuals to retain 100 alleles in the population (simulation A: Af=150, s=0.01, g=0.05 and μ=1.18*10–8). If a higher initial allele number (Af=200) is assumed, only 450 founder individuals are required. However, it is unlikely that a population of size 450 individuals would contain a higher number of alleles than a population of size 500. Therefore, we assumed the smaller number of alleles (Af=150) in the founder population.
The effect of drift to the number of retained alleles with different founder population sizes in simulations. The initial allele number (Af) was 150, mutation rate (μ) was 1.18*10–8 and each population was grown to 1000 individuals. Population growth rates (g) and selection against homozygotes (1-s) vary in different simulations in the following way: (a) g=0.05, s=0.01, (b) g=0.05, s=0.1, (c) g=0.02, s=0.01 and (d) g=0.1, s=0.01. Simulations A and B overlap. Each symbol represents the mean of 1000 simulations.
As mentioned above, the mutation rates of magnitude 10–9 or 10–8 are too low to essentially increase the number of alleles in the timescale of dog domestication, so it is likely that most of the current alleles were already present in the ancestral population. In the simulation timescale, even as high mutation rate as 10−7 did not increase the number of alleles considerably. The number of generations in a simulation is limited by the maximum number of individuals. The upper limit of 1000 individuals was reached in 49 and 15 generations (g=0.05), with 100 and 500 founder individuals, respectively. The magnitude of the selection coefficient did not have a considerable effect on the number of alleles retained (compare simulations A and B, Figure 3), which was also noticed by Vincek et al. (1997).
Discussion
DLA polymorphism and Asian origin of dogs
Albeit strong selection influencing MHC polymorphism, Prugnolle et al. (2005) found a strong negative correlation between MHC diversity and geographical distance of human populations from the assumed East African origin. The origin of dogs has previously been denoted to be somewhere in Asia (Pang et al., 2009; vonHoldt et al., 2010) and high levels of genetic diversity are expected at and nearby the domestication locality. Earlier studies of DLA diversity among dogs have been based almost exclusively on samples from European dogs (Kennedy et al., 2007b). It is remarkable that in the combined sample of 1612 dogs half (33/71) of the DLA–DRB1 alleles were found only among the 249 Asian dogs. Furthermore, with our sample of 128 Asian dogs we identified 22 full-length alleles, which had not been submitted to GenBank before. Our study confirms that until now a large proportion of the DRB1 diversity in Asia has remained unexplored. Our result also supports Asian origin of dogs.
Low levels of observed heterozygosity (0.444–0.455) among the Asian non-breed dogs in this study were surprising—even many dog breeds in Europe reach higher heterozygosity levels. In the earlier study of Kennedy et al. (2007b), the mean observed heterozygosity of over 80 mostly European dog breeds and some non-breed dogs was 0.665. In the wild, canids heterozygosities vary between 0.62 and 0.87 in grey wolf populations (Hedrick et al., 2000; Seddon and Ellegren, 2004) and 0.500–1.000 in coyotes (Hedrick et al., 2002). Our low estimate of heterozygosity could be caused by the presence of null alleles. If this was the case, the results would still be comparable between the Asian regions and would even emphasize more the high diversity of Asian dogs compared with European dogs. It is possible that the low heterozygosity observed here relates to sampling from small isolated villages and is caused by inbreeding. Adding to the effect of inbreeding, genetic drift may overcome balancing selection and reduce polymorphism in small populations unless if selection is very strong (Robertson, 1962).
The non-breed dogs in this study do not have similar backgrounds. Most of the European non-breed dogs have recent purebred ancestors, whereas the Asian non-breed dogs might never have been subject to systematic breeding. It is also more likely that Asian non-breed dogs are really descending from Asian dogs, whereas European non-breed dogs can have imported ancestors. To gain a more detailed picture of DLA polymorphism around the world and the domestication locality, a geographically and demographically more comprehensive sample would be needed. It would be especially useful to study non-breed dogs from remote places because they are least mixed with imported dogs. Such remote places are hard to find in Europe but a comprehensive study of non-breed dogs across most other parts of the world seems feasible.
Number of founders and the domestication process
Around 100 DLA–DRB1 alleles have been submitted to GenBank in full-length (270 bp). Our simulations suggest that retaining 100 alleles requires at least 500 wolf ancestors. If the allele frequencies in the founding wolf population were not even, as assumed in our simulations, the founder population must have been even larger. Our result is in line with earlier MHC simulation results (Vilà et al., 2005) based on less than half of the alleles included in our study. According to Vilà et al. (2005) possibly a few hundred wolf founders were involved in the domestication process. A large founding population is also supported by the finding that domestication has not considerably reduced nucleotide diversity in dogs (Gray et al., 2009).
According to Palstra and Ruzzante (2008), the ratio of effective population size (Ne) to census size averages 0.18 in stable populations. It is suggested that the Ne/N ratio is high for populations with small census size (Palstra and Ruzzante, 2008). For example, in the contemporary small Finnish wolf population, the Ne/N ratio was estimated to be 0.42 (Aspi et al., 2006). However, wolves have probably been widespread and numerous at the time of domestication (Boitani, 2003), so a low Ne/N ratio would be expected. Depending on the Ne/N estimate used, the 500 founding wolf individuals would translate into a census size of 1200–2800 individuals.
The contemporary wolf population is only a shadow of its past size and not well suited for estimations of ancient MHC allele numbers. Wolf is currently less polymorphic than, for example, coyote (Hedrick et al., 2002). This might be a consequence of wolf population fragmentation but it might also reflect a real biological difference in the ideal number of alleles. The wider continuous range of habitats occupied by coyotes (Arjo and Pletscher, 2004) might cause a higher pathogen load and drive higher polymorphism. Whichever the cause for the contemporary low allele number in wolves, the 150 alleles assumed in our simulations would be quite a high number in a single founding wolf population—especially when compared with the approximately 100 alleles so far found from domestic dogs all over the world. Taking into account also the large founding population needed to explain the present high dog DLA polymorphism, a reasonable conclusion is that domestication was not a very rapid or local process. This conclusion is supported by mtDNA diversity, which is markedly higher for dogs in Asia south of Yangtze River than all other parts of the world, but within this region diversity is relatively homogenous (Pang et al., 2009). Possibly, domestication of wolf was a cultural process, which was spread across a large geographical area and involved many wolf populations. It has also been suggested that the domestication process was started by the wolves taking advantage of the food waste around villages—rather than by deliberate human action (Coppinger and Coppinger, 2002). If so, this behavior probably developed over a period of time and in an extended region.
High DLA polymorphism in the dog could also be caused by backcrossing to wolves. vonHoldt et al. (2010) state that there has been ‘interbreeding with wolves early in the domestication process’. However, earlier studies suggest that the amount of hybridization has been small between domesticated dogs and wolves (Sundqvist, 2008; Pang et al., 2009). Instead, Sundqvist (2008) suggested that high recombination rate may explain the high MHC polymorphism. Our results show that even if recombination increases the number of alleles 10 times as much as mutation, an estimate close to the ratio observed in ungulates by Schaschl (2006), recombination has not added considerably to the polymorphism after domestication. Apparently, balancing selection and larger effective population size have maintained high MHC polymorphism compared with the uniparentally inherited neutral markers. After domestication dogs were exposed to a new set of pathogens, which could have resulted in co-evolution between the host and the pathogens. Also, denser populations of dogs compared with wolves might have favored pathogen invasion. This new kind of selection pressure would have maintained high polymorphism in the dog population—possibly even more efficiently than in the source wolf population. However, unless the recombination rate has been very high and there has been wide backcrossing, most of the polymorphism has been present in the founding population(s), which sets the lower limit to the number of domesticated individuals.
To conclude, we have shown that a large proportion of dog DLA–DRB1 diversity has remained unidentified because previous studies have primarily included European dogs. Accordingly, we found high DLA diversity in Asian dogs, which is in line with earlier studies of mtDNA and genome-wide single-nucleotide polymorphism diversity. Based on this more comprehensive DLA data set, we estimated that dogs originated from a large number, at least 500, founding wolves.
Data archiving
The novel allele sequences have been deposited at GenBank with accession numbers EU528627–EU528638, EU528641–EU528650 and EU528652–EU528656.
Accession codes
References
Arjo WM, Pletscher DH (2004). Coyote and wolf habitat use in northwestern Montana. Northwest Sci 78: 24–32.
Aspi J, Roininen E, Ruokonen M, Kojola I, VilÀ C (2006). Genetic diversity, population structure, effective population size and demographic history of the Finnish wolf population. Mol Ecol 15: 1561–1576.
Ayala FJ, Escalante A, O'Huigin C, Klein J (1994). Molecular genetics of speciation and human origins. Proc Nat Acad Sci USA 91: 6787–6794.
Boitani L (2003). Wolf conservation and recovery. In: Mech L, Boitani L, (eds) Wolves. Behavior, Ecology and Conservation. The University of Chicago Press: Chicago. pp 317–340.
Clutton-Brock J (1995). Origins of the dog: domestication and early history. In: Serpell J, (ed.) The Domestic Dog, its Evolution, Behavior and Interactions with People. Cambridge University Press: Cambridge. pp 7–20.
Coppinger R, Coppinger L (2002) Dogs: A New Understanding of Canine Origin, Behavior and Evolution. The University of Chicago Press: Chicago.
Crozier RH (1997). Preserving the information content of species: genetic diversity, phylogeny and conservation worth. Annu Rev Ecol Syst 28: 243–268.
Crozier RH, Agapow P-M, Pedersen K (1999). Towards complete biodiversity assessment: an evaluation of the subterranean bacterial communities in the Oklo region of the sole surviving natural nuclear reactor. FEMS Microbiol Ecol 28: 325–334.
Excoffier L, Laval LG, Schneider S (2005). Arlequin ver. 3.0: an integrated software package for population genetics data analysis. Evol Inform Online 1: 47–50.
Faith DP (1992). Conservation evaluation and phylogenetic diversity. Biol Conserv 61: 1–10.
Germain RN (1994). MHC-dependent antigen processing and peptide presentation: providing ligands for T lymphocyte activation. Cell 76: 287–299.
Germonpré M, Sablin MV, Stevens RE, Hedges REM, Hofreiter M, Stiller M et al (2009). Fossil dogs and wolves from Palaeolithic sites in Belgium, the Ukraine and Russia: osteometry, ancient DNA and stable isotopes. J Archaeol Sci 36: 473–490.
Goudet J (2001). FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.9.3). Available from http://www.unil.ch/izea/softwares/fstat.html . Updated from Goudet( (1995)).
Gray MM, Granka JM, Bustamante CD, Sutter NB, Boyko AR, Zhu L et al. (2009). Linkage disequilibrium and demographic history of wild and domestic canids. Genetics 181: 1493–1505.
Hall T (1999). BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41: 95–98.
Hedrick PW (1999). Balancing selection and MHC. Genetica 104: 207–214.
Hedrick PW, Lee RN, Garrigan D (2002). Major histocompatibility complex variation in red wolves: evidence for common ancestry with coyotes and balancing selection. Mol Ecol 11: 1905–1913.
Hedrick PW, Lee RN, Parker KM (2000). Major histocompatibility complex (MHC) variation in the endangered Mexican wolf and related canids. Heredity 85: 617–624.
Kennedy LJ, Angles JM, Barnes A, Carmichael LE, Radford AD, WER Ollier et al. (2007a). DLA-DRB1, DQA1, and DQB1 Alleles and haplotypes in North American gray wolves. J Hered 98: 491–499.
Kennedy LJ, Barnes A, Short A, Brown J, Lester S, Seddon J et al. (2007b). Canine DLA diversity: 1. New alleles and haplotypes. Tissue Antigen 69: 272–288.
Klein J (1987). Origin of major histocompatibility complex polymorphism: the trans-species hypothesis. Hum Immunol 19: 155–162.
Klein J, Gutknecht J, Fischer N (1990). The major histocompatibility complex and human evolution. Trends Genet 6: 7–11.
Klein J, Satta Y, O'Huigin C, Takahata N (1993). The molecular descent of the major histocompatibility complex. Annu Rev Immunol 11: 269–295.
Leonard JA, Wayne RK, Wheeler J, Valadez R, Guillen S, Vila C (2002). Ancient DNA evidence for old world origin of new world dogs. Science 298: 1613–1616.
Mullis KB, Faloona FA (1987). Specific synthesis of DNA in vitro via a polymerase catalyzed chain reaction. Methods Enzymol 155: 335–350.
Murgia C, Pritchard JK, Kim SY, Fassati A, Weiss RA (2006). Clonal origin and evolution of a transmissible cancer. Cell 126: 477–487.
Palstra FP, Ruzzante DE (2008). Genetic estimates of contemporary effective population size: what can they tell us about the importance of genetic stochasticity for wild population persistence? Mol Ecol 17: 3428–3447.
Pang J-F, Kluetsch C, Zou X-J, Zhang A-b, Luo L-Y, Angleby H et al. (2009). mtDNA data indicate a single origin for dogs south of Yangtze river, less than 16,300 years ago, from numerous wolves. Mol Biol Evol 26: 2849–2864.
Pielou EC (1966). The measurement of diversity in different types of biological collections. J Theor Biol 13: 131–144.
Prugnolle F, Manica A, Charpentier M, Guegan JF, Guernier V, Balloux F (2005). Pathogen-driven selection and worldwide HLA class I diversity. Curr Biol 15: 1022–1027.
Raisor M (2005) Determining the Antiquity of Dog Origins: Canine Domestication as a Model for the Consilience Between Molecular Genetics and Archaeology Vol. S1367, Archaeopress: Oxford.
Robertson A (1962). Selection for heterozygotes in small populations. Genetics 47: 1291–1300.
Runstadler JA, Angles JM, Pedersen NC (2006). Dog leucocyte antigen class II diversity and relationships among indigenous dogs of the island nations of Indonesia (Bali), Australia and New Guinea. Tissue Antigen 68: 418–426.
Sarmiento U, Storb R (1990). Nucleotide sequence of a dog DRB cDNA clone. Immunogenetics 31: 396–399.
Satta Y, O'Huigin C, Takahata N, Klein J (1994). Intensity of natural selection at the major histocompatibility complex loci. Proc Nat Acad Sci USA 91: 7184–7188.
Savolainen P, Zhang Y-p, Luo J, Lundeberg J, Leitner T (2002). Genetic evidence for an East Asian origin of domestic dogs. Science 298: 1610–1613.
Schaschl H, Wandeler P, Suchentrunk F, Obexer-Ruff G, Goodman SJ (2006). Selection and recombination drive the evolution of MHC class II DRB diversity in ungulates. Heredity 97: 427–437.
Seddon J, Ellegren H (2002). MHC class II genes in European wolves: a comparison with dogs. Immunogenetics 54: 490–500.
Seddon JM, Ellegren H (2004). A temporal analysis shows major histocompatibility complex loci in the Scandinavian wolf population are consistent with neutral evolution. Proc R Soc Lond Ser B 271: 2283–2291.
Spurgin LG, Richardson DS (2010). How pathogens drive genetic diversity: MHC, mechanisms and misunderstandings. Proc R Soc B Biol Sci 277: 979–988.
Sundqvist A (2008). Conservation genetics of wolves and their relationship with dogs. Doctor of Philosophy thesis, Uppsala University, Uppsala, Sweden.
Tamura K, Dudley J, Nei M, Kumar S (2007). MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24: 1596–1599.
Vilà C, Seddon J, Ellegren H (2005). Genes of domestic mammals augmented by backcrossing with wild ancestors. Trends Genet 21: 214–218.
Vincek V, O'Huigin C, Satta Y, Takahata N, Boag PT, Grant PR et al. (1997). How large was the founding population of Darwins Finches. Proc R Soc Lond B Biol Sci 264: 111–118.
vonHoldt BM, Pollinger JP, Lohmueller KE, Han E, Parker HG, Quignon P et al. (2010). Genome-wide SNP and haplotype analyses reveal a rich history underlying dog domestication. Nature 464: 898–902.
Wayne RK, Ostrander EA (2007). Lessons learned from the dog genome. Trends Genet 23: 557–567.
Yuhki N, Beck T, Stephens R, Neelam B, O'Brien S (2007). Comparative genomic structure of human, dog, and cat MHC: HLA, DLA, and FLA. J Hered 98: 390–399.
Acknowledgements
We thank Outi Savolainen, Lorna Kennedy and two anonymous reviewers for providing valuable comments on earlier versions of the manuscript. The study was financially supported by the Finnish Population Genetics Graduate School (AKN), Kone Foundation (AKN), the Academy of Finland (grant no. 116579 to JA, and grant no. 1118463 to HL), the Sigrid Juselius Foundation (HL), the Jane and Aatos Erkko Foundation (HL) and the Swedish Board of Agriculture (Jordbruksverket, PS). PS is a Royal Swedish Academy of Sciences Research Fellow supported by a grant from the Knut and Alice Wallenberg Foundation.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Heredity website
Supplementary information
Rights and permissions
About this article
Cite this article
Niskanen, A., Hagström, E., Lohi, H. et al. MHC variability supports dog domestication from a large number of wolves: high diversity in Asia. Heredity 110, 80–85 (2013). https://doi.org/10.1038/hdy.2012.67
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2012.67
Keywords
This article is cited by
-
Genetic diversity and selection signatures of the beef ‘Charolais de Cuba’ breed
Scientific Reports (2018)
-
Identification of novel polymorphisms and two distinct haplotype structures in dog leukocyte antigen class I genes: DLA-88, DLA-12 and DLA-64
Immunogenetics (2018)
-
Disease: A Hitherto Unexplored Constraint on the Spread of Dogs (Canis lupus familiaris) in Pre-Columbian South America
Journal of World Prehistory (2017)
-
No evidence of prenatal diversifying selection at locus or supertype levels in the dog MHC class II loci
Canine Genetics and Epidemiology (2016)
-
A comprehensive mapping of the structure and gene organisation in the sheep MHC class I region
BMC Genomics (2015)