Abstract
Interpretation of variants of unknown significance (VUS) is a major challenge for laboratories performing molecular diagnosis of hereditary breast and ovarian cancer (HBOC), especially considering that many genes are now known to be involved in this syndrome. One important way these VUS can have a functional impact is through their effects on RNA splicing. Here we present a custom RNA-Seq assay plus bioinformatics and biostatistics pipeline to analyse specifically alternative and abnormal splicing junctions in 11 targeted HBOC genes. Our pipeline identified 14 new alternative splices in BRCA1 and BRCA2 in addition to detecting the majority of known alternative spliced transcripts therein. We provide here the first global splicing pattern analysis for the other nine genes, which will enable a comprehensive interpretation of splicing defects caused by VUS in HBOC. Previously known splicing alterations were consistently detected, occasionally with a more complex splicing pattern than expected. We also found that splicing in the 11 genes is similar in blood and breast tissue, supporting the utility and simplicity of blood splicing assays. Our pipeline is ready to be integrated into standard molecular diagnosis for HBOC, but it could equally be adapted for an integrative analysis of any multigene disorder.
Similar content being viewed by others
Introduction
Since the identification of germline inactivating variants within the BRCA1 and BRCA2 genes that are implicated in hereditary breast and ovarian cancer (HBOC),1, 2 numerous other genes have also been found to be involved in this syndrome. Pathogenic variants within genes like TP53, PTEN, STK11 and CDH1 are associated with an increased risk of breast cancer, as well as other cancers and/or pathologies, defining the Li-Fraumeni syndrome, Cowden syndrome, Peutz-Jeghers syndrome and hereditary diffuse gastric cancer, respectively.3, 4, 5, 6 In the case of these syndromes, the clinical phenotype can target more easily the gene involved in the disease. Pathogenic variants in the PALB2 gene have a similar risk spectrum to BRCA2 for developing breast cancer. Pathogenic variants in RAD51C and in BRIP1 confer an increased risk of ovarian cancer,7, 8 whereas pathogenic variants of CHEK2 are associated with moderate risks of breast cancer.9 Finally, variants of other genes, including BARD1, RAD51B, RAD51D, XRCC2 and XRCC3, are associated with a low risk of breast and/or ovarian cancer, but their contribution and their penetrance remain to be characterized.10, 11, 12, 13, 14
Analysis of gene panels by next-generation sequencing (NGS) has resulted in the detection of a large number of new variants of unknown significance (VUS).15 However, current genetic counselling practice only considers variants if they directly affect protein structure; VUS are mostly ignored simply because their potentially deleterious character has not yet been confirmed. One way VUS can be deleterious is if they modify RNA splicing.16 Splicing depends on highly redundant but specific sequences in the gene’s pre-mRNA, such as 3′ and 5′ splice site (3′ss and 5′ss) consensus sequences and splicing regulatory elements.17, 18 VUS can directly impact these important sequences.19, 20 Most exons are constitutively included in transcripts but a few are alternatively included in a regulated manner and both alternative and constitutive splicing can be disturbed by splicing variants.19 The aberrant transcript that results can then contribute to carcinogenesis, as can mis-regulation of expression levels between transcripts.21, 22 Often the effect of the splicing variant is partial and normal transcript continues to be expressed by the mutated allele at varying levels.23, 24 In HBOC genes, it is estimated that one-third of potential deleterious variants impact mRNA splicing.15 Therefore study of how VUS impact gene expression at the RNA level in HBOC-related genes is a major means of identifying new molecular alterations that could be used for genetic counselling. However, the complexity of splicing regulation, mentioned above, makes the interpretation of VUS effects difficult. Moreover, while the splicing pattern of BRCA1 and BRCA2 has been extensively characterized,25, 26 the splicing pattern of the other HBOC-related genes has been less studied and the lack of a reference set of common splicing junctions currently hinders the interpretation of splicing VUS.
Targeted high-throughput sequencing of mRNA (targeted RNA-Seq) is a powerful method for detection of any alternative or abnormal transcripts.27 Here we chose a method based on the capture of exons28 without designing specific baits of known exon–exon junctions, to target a selected panel of transcripts of interest. In parallel, we developed a specific quantitative and qualitative bioinformatics and biostatistics pipeline to analyse transcripts and splicing variants in targeted RNA-Seq data. This pipeline enabled the identification of new alternative and abnormal junctions. For the first time, we describe, using this targeted RNA-Seq method, the splicing pattern of 11 HBOC-related genes: BARD1, BRCA1, BRCA2, BRIP1, CHEK2, PALB2, RAD51B, RAD51C, RAD51D, XRCC2 and XRCC3.
Patients and methods
Gene nomenclature
Nucleotide numbering of the transcripts was based on the cDNA sequences denoting c.1 as the first nucleotide of the translation initiation codon, according to the Human Genome Variation Society recommendations. Descriptions containing intronic positions were based on a genomic reference sequence. The NCBI accession numbers of the sequences used in this study are listed as follows: BARD1 (NG_012047.2 and NM_000465.3), BRCA1 (NG_005905.2 and NM_007294.3), BRCA2 (NG_012772.3 and NM_000059.3), BRIP1 (NG_007409.2 and NM_032043.2), CHEK2 (NG_008150.1 and NM_001005735.1), PALB2 (NG_007406.1 and NM_024675.3), RAD51B (NG_023267.2 and NM_133510.3), RAD51C (NG_023199.1 and NM_058216.2), RAD51D (NG_031858.1 and NM_002878.3), XRCC2 (NG_027988.1 and NM_005431.1) and XRCC3 (NG_011516.1 and NM_005432.3).
Biological material
Total RNA from eight patient lymphoblastoid cell lines (LCLs) were used as positive controls for the validation of the assay. These patients were carriers of variants causing abnormal splicing in BRCA1, BRCA2 or RAD51C: BRCA1 c.4675G>C (https://www.ncbi.nlm.nih.gov/clinvar/ SCV000538190); BRCA2 c.39−1G>A (SCV000536676); BRCA2 c.156_157insAlu (SCV000538191); BRCA2 c.475+3A>G (SCV000538192); BRCA2 c.7975A>G (SCV000538193); BRCA2 c.9501+3A>T (SCV000538194); RAD51C c.706−2A>G (SCV000536677); RAD51C c.1026+5_1026+7del (SCV000536678). All patients were carriers of splicing variant in only one gene, so they were used as negative controls for the other 10 non-mutated genes. In addition, total RNA from one voluntary healthy donor was also used as an independent negative control, as well as total RNA from one healthy breast tissue obtained after cosmetic contralateral surgery in a breast cancer patient with no HBOC familial history. Total RNA from 15 patient LCLs were also studied. These patients with suspicion of HBOC syndrome, who had previously tested negative for a BRCA1/2 pathogenic variant, were selected based on a predisposition probability higher than 90% according to the Claus model.29 All subjects gave informed consent for genetic analysis and were approved by the French Biomedicine Agency.
RNA extraction
LCLs were established for patients and controls. Total RNA was extracted using either TRIzol reagent (Invitrogen, Carlsbad, CA, USA), the AllPrep DNA/RNA Kit (Qiagen, Courtaboeuf, France) or the NucleoSpin RNA Kit (Macherey-Nagel, Hoerdt, France), according to the manufacturer’s instructions. Quality of all RNAs was assessed on the 2200 TapeStation (Agilent, Santa Clara, CA, USA) by the RNA integrity number. For all samples RNA integrity number was >7.5.
Sample preparation and targeted enrichment for NGS
We used Agilent eArray (SureDesign; Agilent) to design 120 nucleotide SureSelect solution library baits that target all known exons of the 11 genes of interest (see Supplementary Data). Fifty nucleotides of intron surrounding exons were also covered by baits, to allow bait design for all small exons up to 60 nucleotides. There were no baits spanning exon–exon junctions, to avoid bias for the enrichment of already known exon–exon junctions. The enrichment for targets of interest was performed on 1 μg of total RNA using the SureSelect RNA Reagent Kit, ILM (Agilent) according to the manufacturer’s instructions. Libraries were then sequenced on a NextSeq500 (Illumina, San Diego, CA, USA) using the high-output paired-end 2 × 101 bps program, with 16 to 18 samples per run.
Bioinformatics and biostatistics pipeline for the analysis of RNA-Seq data
We developed a specific bioinformatics pipeline for quantitative and qualitative analysis of targeted RNA-Seq data (Supplementary Figure S1). RNA-Seq reads were aligned to the human reference genome (hg19) using STAR v.2.3.0e with a splice junction database consisting of junctions from UCSC known genes and RefSeq.30 Splice junction read coverages were obtained from the output file from STAR, SAMtools v.1.3 and BEDTools v.2.17.0 to obtain counts for known or unknown splice junctions.31, 32 The HTseq-count v.0.6.1 program was used to count RNA-Seq reads by gene, with the ‘union’ mode option.33 Gene expression was normalized by DESeq2 v.2.1.6.0.34 We used BEDTools and homemade scripts (available upon request from GenoSplice technology) for read counts and junction annotations. Only junctions with 100 reads or more in at least one sample per run were analysed; those whose coverage was below this threshold were considered as background sequencing noise or related to very weakly expressed spliced transcripts. However, with this method, every aberrant junction, even if detected in only one sample, was selected for analysis. The Δ6q, 7 transcript in BRCA2, which contains an exon 6-derived TG dinucleotide between flanking exons 5 and 8, was detected and analysed using a homemade pattern research script (available upon request from GenoSplice technology). We classified detected junctions into different types of splicing events: cryptic exon inclusion, exon skipping and use of intronic/exonic cryptic splice site. For each junction and each individual, we considered the ratio of alternative/constitutive junction counts. Thereafter, these ratios were expressed as percentages. We considered only spliced junctions that were expressed over 0.1%.
We established the splicing pattern of each captured gene (see formulae in Figure 1) with a calculation method described in Supplementary Information. First, for each junction, we applied a procedure to eliminate outliers that might be abnormal or overexpressed junctions. Then we modelled the distribution of percentages of junction reads according to a gamma distribution or a negative binomial distribution (see Supplementary Information for further details). Given the modelling of the values of the percentages of junction reads, we computed for each percentage the probability that the value was in the distribution. Given the ith event, considering Ri a random variable distributed according to the modelling (gamma or negative binomial distributions), then the probability that a percentage (p) to be in the distribution is defined as P(Ri>p). A P-value <5% was then considered significant.
Splicing junctions and calculation of the percentage of junction reads. We classified each detected junction and calculated the percentage of junction reads using the formulae on the right. Exons are represented by black boxes, constitutive junctions are represented by thick black lines and alternative junctions are represented by thin grey lines. The calculation method is described in Supplementary Information. Types of splicing modifications considered are: (a) cryptic exon inclusion; (b) exon skipping; (c) multiple exon skipping; (d) splice intronic donor shift; (e) splice intronic acceptor shift; (f) splice exonic donor shift and (g) splice exonic acceptor shift.
Alternative spliced transcripts nomenclature
The nomenclature of alternative spliced transcripts in this study follows the convention used by Colombo et al25 and Fackenthal et al,26 and the Human Genome Variation Society guidelines. The letter delta (Δ) indicates alternative event resulting from single exon skipping. Commas or dashes indicate events resulting in skipping of two or more contiguous exons, respectively. Events involving a shifting of 5′ss (distal) or 3′ss (proximal) sites are indicated with p or q, respectively. Cryptic alternative 5′ss or 3′ss uses within introns are indicated as ▾Xp or ▾Xq, where X represents the exon number. Cryptic exon inclusion is indicated as ▾XA.
Validation of new alternative spliced transcripts
Reverse transcriptase PCRs (RT-PCRs) were performed from 200 ng of total RNA using the Onestep RT-PCR Kit (Qiagen) as described previously.35 Two strategies were used depending on the frequency of the alternative spliced transcript tested. For highly expressed alternative spliced transcripts, RT-PCRs were performed with primers flanking the spliced region. To detect very weakly expressed alternative spliced transcripts, we chose primers overlapping the splice junction, with 1–4 nucleotides' overlap (sequences available upon request).
Results
Description of the splicing patterns of the 11 captured HBOC genes
We used a tailored exon capture enrichment strategy to focus on the RNA splicing junctions of 11 HBOC genes: BARD1, BRCA1, BRCA2, BRIP1, CHEK2, PALB2, RAD51B, RAD51C, RAD51D, XRCC2 and XRCC3. On average, for each run, we generated 900 million sequence reads, with at least 50 million reads per sample. Coverage of all the small and large exons of interest was 100%, with a sequencing depth between 700x and 305 000x (data not shown). First, the global splicing pattern of these genes was determined using all LCL samples, by considering all alternative spliced transcripts with a percentage of alternative/normal junction reads more than 0.1% and excluding outliers events (see Patients and Methods) (Supplementary Table S1).
To verify if our method could reliably detect alternative splicing in more detail, we compared BRCA1 junctions detected by our method, in controls, with those described previously by Colombo et al.25 All the 10 alternative spliced transcripts previously classified as predominant were detected, as well as the majority of the very weakly expressed alternative spliced transcripts (Supplementary Table S2). Transcripts containing alternative terminal exons (eg, BRCA1-IRIS),36 resulting from the use of alternative transcription termination sites, could not be detected in our analyses because they do not create additional junction. We confirmed the presence of the newly discovered transcripts by RT-PCR. These include three inclusions of cryptic exons (▾2Aa (0.95%), ▾2Ab (2.1%) and ▾2B (0.28%)) and two splice donor shifts (Δ15q (0.16%) and ▾16q (0.24%)) (Supplementary Figure S2). Of the 38 spliced transcripts detected in the BRCA1 gene by RNA-Seq, 13 (34%) generate premature stop codons (Supplementary Table S2). The 17 most frequent BRCA1 spliced transcripts defined according to our method are represented in the Figure 2a.
Schematic representation of the main ubiquitously expressed alternative splicing events in 11 HBOC-related genes, BRCA1, BRCA2, RAD51C, PALB2, BARD1, RAD51B, BRIP1, XRCC2, CHEK2, XRCC3, RAD51D (a-k respectively). Summary of splicing events detected by targeted RNA-Seq in the current study. Genes are represented in grey, boxes correspond to exons and horizontal lines correspond to introns. Exons and introns are not drawn to scale. For each alternative junction, we calculated the percentage of junction reads and established the splicing pattern of each captured gene (see Figure 1 and Supplementary Information for the calculation method). All the splicing events shown here were detected with a percentage of junction reads >1%. Junctions expressed more than 10% are represented in bold, junctions expressed between 5 and 10% are represented by dotted lines and junctions expressed <5% are represented with thin lines. The exhaustive list of the splicing events detected in all captured genes is available in the Supplementary Table S1.
For BRCA2, we performed the same comparison of detected junctions between our method and those recently described by Fackenthal et al.26 As for BRCA1, all of the alternative spliced transcripts previously classified as predominant were detected, as well as the majority of the very weakly expressed alternative spliced transcripts (Supplementary Table S3). We confirmed nine newly discovered splicing events in BRCA2 by RT-PCR: five cryptic exon inclusions (▾18 A (0.31%), ▾20B (0.22%), ▾24 A (0.7%), ▾24B (3.1%) and ▾25 A (0.81%)), three exon skipping (Δ7 (0.09%), Δ(9,10) (0.14%) and Δ17_19 (0.04%)) and one splice acceptor shift (Δ4p (1.2%)) (Supplementary Figures S3 and S4). The new exon skipping Δ17_19 was the most weakly expressed event to be detected by RNA-Seq and confirmed by RT-PCR. Of the 27 spliced transcripts detected in the BRCA2 gene by RNA-Seq, 15 (56%) generate premature stop codons (Supplementary Table S3). The most frequent BRCA2 spliced transcripts defined according to our method are represented in Figure 2b.
These results confirmed that our approach was able to identify known and new alternative spliced transcripts in BRCA1 and BRCA2. In the same way, we described the main splicing pattern of the other nine genes studied (Figure 2). Our targeted RNA-Seq method could detect all splicing species (cryptic exon inclusion, single or multiple exon(s) skipping, splice donor/acceptor shifts). All splicing junctions detected by RNA-Seq for these 11 genes are listed in the Supplementary Table S1. Next, we performed a comprehensive screen of alternative spliced transcripts in one healthy breast tissue sample. Interestingly, according to statistical analysis, we qualitatively and quantitatively detected the same well-represented spliced transcripts previously identified in these 11 genes in LCLs (Supplementary Table S4).
Detection of splicing alterations in captured HBOC genes
Here, we analysed RNA extracted from eight LCLs with known splicing variants in BRCA1, BRCA2 and RAD51C (Table 1). All specific transcript anomalies were detected in these specific LCLs (see final column in Table 1), thus validating our technique.
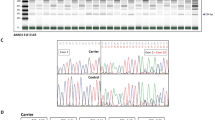
For the BRCA1 c.4675G>C pathogenic variant (Table 1), located in the last position of exon 15, a total effect on splicing has been shown by RT-PCR (Supplementary Figure S5). Analysing the junction data, we observed, as expected, a slight strengthening of the out-of-frame skipping of exon 15 (Δ15) (1.3% against 0.15±0.11%, corrected P-value <0.01) and a strong reinforcement of the out-of-frame deletion of the last 11 nucleotides of BRCA1 exon 15 (Δ15q) (13.4% against 0.16±0.13%, corrected P-value <0.001) (Figure 3a and Supplementary Table S5). When visualizing the mapped RNA-Seq sequences at position c.4675, only the nucleotide G corresponding to the wild-type allele was observed, confirming the drastic splicing defect (Figure 3b).
Impact of the BRCA1 c.4675G>C pathogenic variant on the splicing pattern of BRCA1 exon 15 in lymphoblastoid cell lines of a heterozygous HBOC patient. (a) Cartography of the percentage of junction reads observed between BRCA1 exons 14 and 16. Bold lines represent the junctions 14–15 and 15–16. Thin lines represent the alternative junctions 14–16, corresponding to exon 15 skipping. The abnormally enhanced junction Δ15q detected in the cell line carrying the variant is shown below and represents the out-of-frame deletion of the last 11 nucleotides of BRCA1 exon 15. (b) Coverage, sequencing depth and sequence observed at the variant positions.
For the BRCA2 c.39−1G>A pathogenic variant (Table 1), located in intron 1, a total effect on splicing has been also identified by RT-PCR (Supplementary Figure S5). Analysing the junction data, we observed, as expected, the aberrant skipping of exon 2 (Δ2) (20.9%) and aberrant combined skipping of exons 2 and 3 (Δ2, 3) (4.8%). These two spliced transcripts were not detected in the controls (Supplementary Table S1). Both these events cause loss of the translation initiation codon of BRCA2, which is naturally located in exon 2. The LCL carrying this variant also carried the heterozygous single-nucleotide polymorphism c.−26G>A. When visualizing the mapped RNA-Seq sequences at position c.−26, only the nucleotide G was observed, confirming the drastic splicing defect (data not shown).
In the same manner, our junction data confirmed that the BRCA2 c.156_157insAlu, which is a Portuguese founder pathogenic variant,37 causes the strong reinforcement of the in-frame skipping of exon 3 (Δ3) (46% against 1.2±0.9%, corrected P-value <0.001) (Table 1 and Supplementary Table S5).
Another total effect on splicing was observed for the BRCA2 c.475+3A>G pathogenic variant, which causes the out-of-frame skipping of exon 5, as shown in our RT-PCR experiments (Table 1 and Supplementary Figure S5). Our targeted RNA-Seq data confirmed the strong exon 5 skipping (Δ5) (94% against 2.5±2%, corrected P-value <0.001) (Supplementary Table S5).
The BRCA2 c.7975 A>G unclassified variation, located in the penultimate position of exon 17, caused the moderate in-frame skipping of exon 17 (Δ17) (3% against undetected here), as expected38 (Table 1 and Supplementary Table S5). When visualizing the mapped RNA-Seq sequences at position c.7975, the two alleles were observed, which confirms the partial effect of this variant on splicing (data not shown).
Another partial effect on splicing was observed for the BRCA2 c.9501+3 A>T unclassified variation, located in intron 25, which causes moderate (2.6%) frameshift skipping of exon 25 (Table 1).23 This spliced transcript was not detected in the controls (Supplementary Table S1).
The RAD51C c.1026+5_1026+7del (intron 8) and the RAD51C c.706−2A>G (intron 4) pathogenic variants caused the total frameshift skipping of exon 8 and in-frame skipping of exon 5, respectively11 (Table 1). These two exon skips were detected as very weak spliced transcripts in controls (Supplementary Table S1). For the first pathogenic variant, we correctly detected the exon 8 skipping (Δ8) (17.9% against 0.08±0.05%, corrected P-value <0.001). For the second pathogenic variant, we found that spliced transcript without exon 5 was overexpressed (Δ5) (29.2% against 0.03±0.03%, corrected P-value <0.001), but there were two other spliced transcripts, including overexpression of the frameshifting combined skipping of exons 4 and 5 (Δ4, 5) (3.6% against 0.04±0.05%, corrected P-value <0.001), as well as the expression of an abnormal frameshifting combined skipping of exons 5, 6 and 7 (Δ5_7) (0.48%) (Figure 4a and Supplementary Table S5). We confirmed these two aberrant spliced transcripts by RT-PCR with primers located in exons 2 and 8 (Figure 4b). We also noticed the presence of a weakly expressed additional transcript corresponding to the combination of the skipping of exons 5 and 7 (Δ5, Δ7) caused by the mutated allele. Moreover, we detected a new splice junction, corresponding to an alternative acceptor splice site, created de novo by the variant and recognized by the splicing machinery (Δ5p) (Figure 4a). Overall, these results confirm the detection and quantification of known and new mRNA transcripts in HBOC-relevant genes by our targeted RNA-Seq approach.
Impact of the RAD51C c.706−2 A>G pathogenic variant on the splicing of RAD51C exon 5 in LCLs of an HBOC patient, heterozygous for this variation. (a) RAD51C exon 5 cartography of the abnormal splicing junctions detected in LCLs from a patient with the splicing variant c.706−2 A>G (Table 1). The gene is represented by boxes corresponding to exons and horizontal lines corresponding to introns. Exons and introns are not drawn to scale. Junctions that are used at over 10% are represented in bold, junctions expressed between 1 and 10% are shown as dotted lines and junctions <1% are thin lines. (b) RT-PCR analysis of the RAD51C variation’s effect on splicing, using primers located in exons 2 and 8. Identities of the different transcripts are indicated on the right of the gel. The Δ7 splice transcript is detected in both control and patient samples, whereas the Δ5, Δ4, 5 and combination of Δ5 and Δ7 (Δ5, Δ7) (in bold) are only detected in the patient.
Discussion
Interpretation of VUS is a major challenge for the laboratories performing the molecular diagnosis of HBOC, especially considering that many genes are now known to be involved in the syndrome. One of the main areas contributing to understanding the functional impact of these variants is an investigation of their effects on RNA splicing to find out if they could lead to aberrant RNAs and consequently to potential loss of function of the proteins. Theoretically, any detected variation can affect RNA splicing.20 Here we developed a targeted RNA-Seq approach with bioinformatics and biostatistics analyses, which allows detection and quantification of splicing junctions in many genes simultaneously, with excellent sensitivity. Our ‘exon-restricted’ capture set designed for 11 HBOC-related genes enables the efficient capture of all exons and the efficient detection of known and new alternative splicing junctions, as well as cryptic exon inclusions. This strategy is applicable to any RNA-Seq platform capable of sequencing at least 5 Gb per sample after targeted capture.
All presented analyses were performed on LCL samples. However, the PAXgene system, which provides a snapshot of the transcripts at the time of sampling, is widely used in laboratories for molecular diagnosis and for studying the effect of a variant on splicing.24 Today this system is even used for RNA collection in the ‘100 000 Genomes Project’39 led by Genetics England for the National Health Service (http://www.genomicsengland.co.uk). Indeed, we observed, in a preliminary step, that the PAXgene system does not seem to be adapted to RNA-Seq, for very low expressed HBOC-associated genes, such as BRCA2 (Supplementary Figure S6). Analysis of this sample type would require a technical adjustment, with a decrease in the number of samples sequenced per run and an increase in sequencing capacity. However, these changes would have a significant impact on sequencing costs. Analysis of LCLs from patients, albeit time consuming, overcomes this limitation because in this type of sample the expression levels of all the genes were compatible with splicing analysis by RNA-Seq. However, Epstein–Barr virus transformation, modifying the global gene expression,40 might modify the splicing pattern. These results were obtained by sequencing mRNA from LCLs that we did not treat with a nonsense-mediated mRNA decay inhibitor, such as puromycin. In this way, we detected the most representative splicing pattern of these 11 genes (Supplementary Table S1). Moreover, we detected all expected abnormal spliced transcripts (Table 1). Of note, no outlier event was detected in the 15 patient LCLs (Supplementary Table S5). We assumed that they did not have splicing anomalies that could explain the familial presentation.
For BRCA1 and BRCA2, our method detected all most frequent spliced transcripts and most of the minor alternative spliced transcripts described in the literature by fragment analysis25, 26 (Supplementary Tables S1–S3). All the events that we did not detect were known to be very weakly expressed. This difference in sensitivity can be explained by the fact that we do not use a targeted RT-PCR method but a capture of exons to study simultaneously the set of captured transcripts. Targeted RNA-Seq has the advantage of simultaneously analysing the alternative spliced transcripts in many genes with high sensitivity. Our targeted RNA-Seq strategy does not attempt to describe rare splicing junctions since we established a detection limit of 100 reads (Supplementary Tables S1–S5). Importantly, our targeted RNA-Seq method allowed the detection of 14 novel alternative spliced transcripts in BRCA1 and BRCA2, with a percentage of junction reads between 0.04 and 3.1%.
We present here the characterization of the splicing pattern of 11 genes of interest in HBOC. This is the first time that the near exhaustive alternative splicing junctions sets have been described in HBOC-related genes other than BRCA1 and BRCA2 by RNA-Seq, and for these two genes we have increased the described repertoire of alternative spliced transcripts. Altogether, these new results provide an important resource for interpretation of VUS’ splicing impact for these genes. Furthermore, our data suggest that predominant alternative splicing in these 11 genes is similar in blood and breast tissues. The conclusion is that splicing analyses performed on, easy-to-obtain, blood samples are relevant for diagnosis.
In addition, our bioinformatics and biostatistics pipeline detected all the qualitatively and quantitatively abnormal splice junctions caused by variants that were previously detected by conventional methods in BRCA1, BRCA2 and RAD51C (Table 1). Indeed, we detected both partial and total effects on splicing without clearly distinguishing them in all cases, as our approach gives globally indicative rather than definitive exact quantification. Interestingly, we observed that a variant may cause a series of complex splicing anomalies (Figure 4a). Here we suggest a method for detection of the most representative splicing pattern and aberrant spliced transcripts in 11 HBOC-related genes. The method allows detection of abnormal splicing junctions caused by variants, including potential variants located in deep intronic regions that are far away from those that can be detected by routine exon-centric DNA sequencing. Our method could be included in a global strategy to classify variants for their pathogenicity. Indeed, we could consider our method as a first line method to detect abnormal splicing in patients. These events have to be confirmed with specific targeted approaches such as minigene assay or quantitative RT-PCR, to characterize the total or partial effect on splicing.23, 41 If the interpretation of RNA studies remain tricky, these results may be included in a multifactorial likelihood model calculating the posterior probability that the variant is pathogenic.42, 43, 44
The proposed strategy for the data analysis could be applied for studying splicing by targeted RNA-Seq for any complex genetic disease.
References
Miki Y, Swensen J, Shattuck-Eidens D et al: A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science 1994; 266: 66–71.
Wooster R, Bignell G, Lancaster J et al: Identification of the breast cancer susceptibility gene BRCA2. Nature 1995; 378: 789–792.
Walsh T, Casadei S, Coats KH et al: Spectrum of mutations in BRCA1, BRCA2, CHEK2, and TP53 in families at high risk of breast cancer. JAMA 2006; 295: 1379–1388.
Bubien V, Bonnet F, Brouste V et al: High cumulative risks of cancer in patients with PTEN hamartoma tumour syndrome. J Med Genet 2013; 50: 255–263.
Hearle N, Schumacher V, Menko FH et al: Frequency and Spectrum of Cancers in the Peutz-Jeghers Syndrome. Clin Cancer Res 2006; 12: 3209–3215.
Pharoah PDP, Guilford P, Caldas C : Incidence of gastric cancer and breast cancer in CDH1 (E-cadherin) mutation carriers from hereditary diffuse gastric cancer families. Gastroenterology 2001; 121: 1348–1353.
Loveday C, Turnbull C, Ruark E et al: Germline RAD51C mutations confer susceptibility to ovarian cancer. Nat Genet 2012; 44: 475–476.
Rafnar T, Gudbjartsson DF, Sulem P et al: Mutations in BRIP1 confer high risk of ovarian cancer. Nat Genet 2011; 43: 1104–1107.
Southey MC, Goldgar DE, Winqvist R et al: PALB2, CHEK2 and ATM rare variants and cancer risk: data from COGS. J Med Genet 2016; 53: 800–811.
Ratajska M, Antoszewska E, Piskorz A et al: Cancer predisposing BARD1 mutations in breast-ovarian cancer families. Breast Cancer Res Treat 2012; 131: 89–97.
Golmard L, Caux-Moncoutier V, Davy G et al: Germline mutation in the RAD51B gene confers predisposition to breast cancer. BMC Cancer 2013; 13: 484.
Loveday C, Turnbull C, Ramsay E et al: Germline mutations in RAD51D confer susceptibility to ovarian cancer. Nat Genet 2011; 43: 879–882.
Hilbers FS, Wijnen JT, Hoogerbrugge N et al: Rare variants in XRCC2 as breast cancer susceptibility alleles. J Med Genet 2012; 49: 618–620.
Park DJ, Lesueur F, Nguyen-Dumont T et al: Rare mutations in XRCC2 increase the risk of breast cancer. Am J Hum Genet 2012; 90: 734–739.
Castéra L, Krieger S, Rousselin A et al: Next-generation sequencing for the diagnosis of hereditary breast and ovarian cancer using genomic capture targeting multiple candidate genes. Eur J Hum Genet 2014; 22: 1305–1313.
Gaildrat P, Krieger S, Di Giacomo D et al: Multiple sequence variants of BRCA2 exon 7 alter splicing regulation. J Med Genet 2012; 49: 609–617.
Cartegni L, Chew SL, Krainer AR : Listening to silence and understanding nonsense: exonic mutations that affect splicing. Nat Rev Genet 2002; 3: 285–298.
Sterne-Weiler T, Howard J, Mort M, Cooper DN, Sanford JR : Loss of exon identity is a common mechanism of human inherited disease. Genome Res 2011; 21: 1563–1571.
Venables JP : Aberrant and alternative splicing in cancer. Cancer Res 2004; 64: 7647–7654.
Soukarieh O, Gaildrat P, Hamieh M et al: Exonic splicing mutations are more prevalent than currently estimated and can be predicted by using in silico tools. PLoS Genet 2016; 12: e1005756.
Chen J, Weiss WA : Alternative splicing in cancer: implications for biology and therapy. Oncogene 2015; 34: 1–14.
Oltean S, Bates DO : Hallmarks of alternative splicing in cancer. Oncogene 2014; 33: 5311–5318.
Bonnet C, Krieger S, Vezain M et al: Screening BRCA1 and BRCA2 unclassified variants for splicing mutations using reverse transcription PCR on patient RNA and an ex vivo assay based on a splicing reporter minigene. J Med Genet 2008; 45: 438–446.
Houdayer C, Caux-Moncoutier V, Krieger S et al: Guidelines for splicing analysis in molecular diagnosis derived from a set of 327 combined in silico/in vitro studies on BRCA1 and BRCA2 variants. Hum Mutat 2012; 33: 1228–1238.
Colombo M, Blok MJ, Whiley P et al: Comprehensive annotation of splice junctions supports pervasive alternative splicing at the BRCA1 locus: a report from the ENIGMA consortium. Hum Mol Genet 2014; 23: 3666–3680.
Fackenthal JD, Yoshimatsu T, Zhang B et al: Naturally occurring BRCA2 alternative mRNA splicing events in clinically relevant samples. J Med Genet 2016; 53: 548–558.
Levin JZ, Berger MF, Adiconis X et al: Targeted next-generation sequencing of a cancer transcriptome enhances detection of sequence variants and novel fusion transcripts. Genome Biol 2009; 10: R115.
Mercer TR, Clark MB, Crawford J et al: Targeted sequencing for gene discovery and quantification using RNA CaptureSeq. Nat Protoc 2014; 9: 989–1009.
Claus EB, Risch N, Thompson WD : Genetic analysis of breast cancer in the cancer and steroid hormone study. Am J Hum Genet 1991; 48: 232–242.
Dobin A, Gingeras TR : Mapping RNA-seq reads with STAR. Curr Protoc Bioinformatics 2015; 51: 11.14.1-19.
Li H, Handsaker B, Wysoker A et al: The Sequence Alignment/Map format and SAMtools. Bioinformatics (Oxf, Engl) 2009; 25: 2078–2079.
Quinlan AR, Hall IM : BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics (Oxf, Engl) 2010; 26: 841–842.
Anders S, Pyl PT, Huber W : HTSeq – a Python framework to work with high-throughput sequencing data. Bioinformatics (Oxf, Engl) 2015; 31: 166–169.
Love MI, Huber W, Anders S : Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 2014; 15: 550.
Gaildrat P, Krieger S, Théry J-C et al: The BRCA1 c.5434C->G (p.Pro1812Ala) variant induces a deleterious exon 23 skipping by affecting exonic splicing regulatory elements. J Med Genet 2010; 47: 398–403.
ElShamy WM, Livingston DM : Identification of BRCA1-IRIS, a BRCA1 locus product. Nat Cell Biol 2004; 6: 954–967.
Machado PM, Brandão RD, Cavaco BM et al: Screening for a BRCA2 rearrangement in high-risk breast/ovarian cancer families: evidence for a founder effect and analysis of the associated phenotypes. J Clin Oncol 2007; 25: 2027–2034.
Caux-Moncoutier V, Castéra L, Tirapo C et al: EMMA, a cost- and time-effective diagnostic method for simultaneous detection of point mutations and large-scale genomic rearrangements: application to BRCA1 and BRCA2 in 1525 patients. Hum Mutat 2011; 32: 325–334.
Siva N : UK gears up to decode 100000 genomes from NHS patients. Lancet (Lond, Engl 2015; 385: 103–104.
Maia A-T, Spiteri I, Lee AJ et al: Extent of differential allelic expression of candidate breast cancer genes is similar in blood and breast. Breast Cancer Res 2009; 11: R88.
Muller D, Rouleau E, Schultz I et al: An entire exon 3 germ-line rearrangement in the BRCA2 gene: pathogenic relevance of exon 3 deletion in breast cancer predisposition. BMC Med Genet 2011; 12: 121.
Plon SE, Eccles DM, Easton D et al: Sequence variant classification and reporting: recommendations for improving the interpretation of cancer susceptibility genetic test results. Hum Mutat 2008; 29: 1282–1291.
Thomassen M, Blanco A, Montagna M et al: Characterization of BRCA1 and BRCA2 splicing variants: a collaborative report by ENIGMA consortium members. Breast Cancer Res Treat 2012; 132: 1009–1023.
Vallée MP, Di Sera TL, Nix DA et al: Adding in silico assessment of potential splice aberration to the integrated evaluation of BRCA gene unclassified variants. Hum Mutat 2016; 37: 627–639.
Acknowledgements
We thank the members of our department of biopathology for their input, the high-throughput sequencing platform of Basse-Normandie SéSAME (Sequencing for Health, Agronomy, the Sea and the Environment) and the Clinical Research of the Centre François Baclesse. We also thank the members of the Evidence-Based Network for the Interpretation of Germ-Line Mutant Alleles consortium (ENIGMA) for their interactions during this study. This work was supported by grants from the Institut National du Cancer (INCa); the French Cancéropôle Nord-Ouest (CNO); and the Ligue Régionale du Calvados Contre le Cancer. We thank the Brazilian agency Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for providing a postdoctoral fellowship for ASS.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Davy, G., Rousselin, A., Goardon, N. et al. Detecting splicing patterns in genes involved in hereditary breast and ovarian cancer. Eur J Hum Genet 25, 1147–1154 (2017). https://doi.org/10.1038/ejhg.2017.116
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2017.116
This article is cited by
-
Identification of prognostic alternative splicing signature in gastric cancer
Archives of Public Health (2022)
-
Mutational and splicing landscape in a cohort of 43,000 patients tested for hereditary cancer
npj Genomic Medicine (2022)
-
Interpretation of BRCA2 Splicing Variants: A Case Series of Challenging Variant Interpretations and the Importance of Functional RNA Analysis
Familial Cancer (2022)
-
The intronic BRCA1 c.5407-25T>A variant causing partly skipping of exon 23—a likely pathogenic variant with reduced penetrance?
European Journal of Human Genetics (2020)
-
Assessment of branch point prediction tools to predict physiological branch points and their alteration by variants
BMC Genomics (2020)