Abstract
Mitochondrial disease can be attributed to both mitochondrial and nuclear gene mutations. It has a heterogeneous clinical and biochemical profile, which is compounded by the diversity of the genetic background. Disease-based epidemiological information has expanded significantly in recent decades, but little information is known that clarifies the aetiology in African patients. The aim of this study was to investigate mitochondrial DNA variation and pathogenic mutations in the muscle of diagnosed paediatric patients from South Africa. A cohort of 71 South African paediatric patients was included and a high-throughput nucleotide sequencing approach was used to sequence full-length muscle mtDNA. The average coverage of the mtDNA genome was 81±26 per position. After assigning haplogroups, it was determined that although the nature of non-haplogroup-defining variants was similar in African and non-African haplogroup patients, the number of substitutions were significantly higher in African patients. We describe previously reported disease-associated and novel variants in this cohort. We observed a general lack of commonly reported syndrome-associated mutations, which supports clinical observations and confirms general observations in African patients when using single mutation screening strategies based on (predominantly non-African) mtDNA disease-based information. It is finally concluded that this first extensive report on muscle mtDNA sequences in African paediatric patients highlights the need for a full-length mtDNA sequencing strategy, which applies to all populations where specific mutations is not present. This, in addition to nuclear DNA gene mutation and pathogenicity evaluations, will be required to better unravel the aetiology of these disorders in African patients.
Similar content being viewed by others
Introduction
Disorders of the mitochondrial oxidative phosphorylation (OXPHOS) system are among the most frequently inherited metabolic disorders in newborns.1 In all, 13 structural OXPHOS subunits, in addition to 22 tRNA and 2 rRNA molecules, are encoded by mitochondrial DNA (mtDNA), although the majority of the more than 100 proteins involved in its structure, import, assembly and control of expression are encoded by nuclear DNA (nDNA).2, 3 Depending on the position of a mutation, the mode of inheritance, if not de novo, can be maternal or Mendelian and present in a dominant or recessive manner. As a result, clinical and biochemical heterogeneity is a hallmark of these disorders and affects both adults and children. Although the aetiology of these disorders are mostly attributed to nDNA pathogenic mutations and even more so in paediatric cases,4 recent evidence have shown that the prevalence of pathogenic mtDNA variants are more common than estimated previously.5, 6, 7
More than 230 pathogenic mtDNA variants have already been reported and well-established mitochondrial syndromes, such as Leber's hereditary optic neuropathy (LHON), mitochondrial encephalopathy with lactic acidosis and stroke-like episodes, myoclonic epilepsy with ragged red fibres and neuropathy, ataxia and retinitis pigmentosa, have been associated with specific variants of mtDNA.6 However, the clinical heterogeneity with which paediatric patients present most often do not result in clear genotype–phenotype correlations and the diagnostic approach still requires an extensive, multi-disciplinary diagnostic approach, including the assessment of OXPHOS enzyme activities in a clinically relevant tissue (eg, muscle biopsy) to direct genetic testing. Screening for an array of nuclear and mtDNA variants, using disease-based epidemiological information, remains one of the principal approaches to identify a primary mitochondrial genetic defect with suspected mitochondrial disease.
We have recently shown in a cohort study that paediatric patients of African descent tend to have a predominantly muscle-associated phenotype and do not conform to well-defined, clinical syndromes.8 However, disease-based epidemiological genetic data are still generally lacking in African patients with primary mitochondrial disease. Although the variation between African and European mtDNA haplogroups is well documented, investigations of mtDNA disease are generally based on European mtDNA haplogroup disease information. We therefore hypothesized that a large number of unique mtDNA variants will be associated with mitochondrial disease in this mainly African cohort. In this investigation, we have used high-throughput sequencing technology to characterize the mtDNA variation in muscle samples from a cohort of 71 South African paediatric patients diagnosed with neuromuscular mitochondrial respiratory chain (RC) disease.
Patients and methods
Patients
Patients in this study originated from the northern provinces of South Africa and were all assessed at the Paediatric Neurology Unit at Steve Biko Academic Hospital (Pretoria, South Africa). In all, 71 paediatric patients presented with a neuromuscular phenotype and had a clinical mitochondrial disease criterion score indicative of a mitochondrial disorder as described by Wolf and Smeitink.9 On the basis of these criteria, the study group of 71 patients consisted of the following South African population groups: 48 (68%) African, 19 (27%) Caucasian, 3 Asian (4%) and 1 (1%) of mixed ancestry (see Supplementary Table S1). Ages of the subjects (36 males and 35 female) ranged between the neonatal periods to 10 years of age. Ethical approval was obtained from the University of Pretoria (number 91/98 and amendments). Informed consent and assent were obtained for all patients before initiation of this study, although a family history was not available for most cases. Muscle enzyme analyses were performed essentially as described elsewhere,10, 11, 12 and for 56/71 of the cases, an RC deficiency was confirmed. In addition, mtDNA relative copy number analysis on muscle DNA was performed by real-time polymerase chain reaction (PCR) using the ND1/GAPDH ratio using TaqMan chemistry and commercial probes (Applied Biosystems, Foster City, CA, USA).
DNA isolation and mtDNA amplification
Total DNA was isolated from muscle tissue samples using NucleoSpin Tissue kits (Macherey-Nagel, Düren, Germany). Total and amplified DNA was quantified using the Quant-iT PicoGreen dsDNA kit (Invitrogen, Carlsbad, CA, USA). The complete human mitochondrial genome (GenBank NC_012920.1) was amplified in two overlapping fragments by long template PCR (Long PCR Enzyme Mix, Fermentas, St Leon-Rot, Germany) as described previously.13 Fragment A (7546 bp) was amplified using a forward (nt 6115–6135) and reverse (nt 13 640–13 660) primer set and Fragment B (9250 bp) was amplified using a separate forward (nt 13 539–13 559) and reverse (nt 6200–6220) primer set. All the PCR amplifications were performed using conditions suggested by the supplier of the reagents at an annealing temperature of 58 °C. Amplified Fragments A and B for each patient were purified by gel extraction and combined at equimolar amounts to a final concentration of 62.5 ng/μl.
Next-generation sequencing and data analysis
Massively parallel DNA sequencing of the PCR fragments were performed in both strands on a Roche 454 GS-FLX platform at Inqaba Biotech (Pretoria, South Africa). Multiplex identifier adaptors, used during the GS-FLX Titanium Library preparation procedure, enabled multiple samples to be sequenced together in a single region of a PicoTiterPlate gasket and allowed for automated software identification of samples after multiplexing and sequencing. Primary data analysis was performed using the CLC genomics workbench (CLC bio, Aarhus, Denmark). Standard Flowgram Format files were imported and trimmed to remove low-quality sequences as well as the 454 sequence Primers A and B, using default settings. High-quality sequencing reads for each patient were mapped against the revised Cambridge Reference Sequence (rCRS) of human mtDNA (GenBank NC_012920.1), using default settings, to obtain a consensus sequence for each individual and to enable variation detection. Single-nucleotide polymorphisms (SNPs) were automatically detected using the High-throughput sequence SNP detection function, which also enables the estimation of variant allele frequency (%). For SNP detection, quality parameters were kept at default values and significance parameters were set as summarized in the Supplementary Table S2. Insertions and deletions (indels) were manually detected by visual inspection of consensus sequences, as recommended by the CLC genomics manual. A variation was classified as a high confidence variation (HCV) when detected in at least three sequences reads that included both forward and reverse strands, unless there were five reads with a quality score over 20 (or 30 if the variation is associated with a 5 mer or higher).14, 15 A variation was therefore classified as a low confidence variation when detected only in either forward or reverse strands, where the variation was an indel associated with homopolymer regions (6–8 bases) or when a heteroplasmic SNP may be due to homopolymer errors occurring at or adjacent to the nucleotide position.14, 15, 16 A control DNA sample with whole mtDNA sequence, which had been previously determined by conventional Sanger sequencing, was included during the process. Sanger sequencing was carried out on this control sample as well as those indicated in Tables 1 and 2, with allele frequencies higher than 10% using BigDye Terminator v.3.1 chemistries on an ABI 3130xl Genetic Analyzer (Applied Biosystems). For the control DNA sample, a 100% consistence was observed between the Roche 454 and Sanger consensus sequence.
Allele frequency (heteroplasmy) confirmation
For selected cases with low level of heteroplasmy, confirmation of the variation and levels was carried out using an alternative method. The Pyromark Assay Design Software v.2.0 (Qiagen, Crawley, UK) was used to design locus-specific amplification and pyrosequencing primers (for the m.4160T>C variation – forward primer, nt 4127–4160; reverse primer, nt 4219–4242; sequence primer, nt 4145–4159; and for the 14723T>C variation – forward primer, nt 14 688–14 710; reverse primer, nt 14 758–14 780; sequence primer, nt 14 704–14 721). Pyrosequencing was performed on a Pyromark Q24 platform (Qiagen) according to the manufacturer's instructions and data were analysed using the Pyromark Q24 software by comparing the data from a wild type and variant at the specific locus.
The mitochondrial genome consensus sequence of each patient was firstly analysed using Phylotree database (http://www.phylotree.org/, tree Build 7 February 2011) to assign mitochondrial haplogroups. All non-haplogroup-associated variants (ie, variants not reported in Phylotree) were then further analysed to classify the diversity present using the mtDNA-GeneSyn computer tool.17 Non-synonymous protein coding, RNA and regulatory region variants identified in patients were analysed using the SNP annotation using Blast function of the CLC genomic workbench to query the NCBI dbSNP database (http://www.ncbi.nlm.nih.gov/SNP/, accessed April 2011). The mitochondrial genome databases, MITOMAP (http://www.mitomap.org, accessed April 2011), mtDB (http://www.genpat.uu.se/mtDB/, accessed April 2011) and mtSNP (http://www.mtsnp.tmig.or.jp/mtsnp/index_e.shtml, accessed April 2011) were also consulted and Google searches were performed for rare variants to ultimately define variants as either previously reported or novel. Variants in protein-coding genes and in transfer RNA genes were further analysed to estimate the potential to be pathogenic, using the Alamut mutation interpretation software (Interactive Biosoftware, Rouen, France) and Mamit-tRNA database (http://www.mamit-trna.u-strasbg.fr/, accessed April 2011). Alamut reports the Align Grantham Variation Grantham Deviation (with a score of C65 most likely and C0 least likely to be deleterious), Polymorphism Phenotyping, version 2 and Sorting Intolerant From Tolerant predictions. Interspecies conservation indexes (CI) of variants were determined using the web-based bioinformatics platform MitoTool (http://www.mitotool.org/, accessed April 2011). All previously reported variants identified in this group, as well as several high confidence novel variants that are possibly pathogenic, were verified by conventional Sanger sequencing. Fisher's exact test (two tailed) was used for statistical analysis, where variants in the two groups (L-haplogroup and non-L-haplogroup patients) of the cohort were compared.
Results
mtDNA sequence data and haplogroup classification
We successfully sequenced the complete mitochondrial genome from the muscle of 71 paediatric patients diagnosed with a mitochondrial disorder and mapped >99% of all sequence fragments to the rCRS. The average amount of mapped reads for this patient group was 3941±1645, with the read length after trimming ranging between 249 and 392 bp, an average coverage of 81±26 and no zero coverage areas. The total amount of HCVs identified per individual, compared with the rCRS, ranged from 27 to 116 for patients with African haplogroups and 9 to 69 for patients with non-African haplogroups, most of which could be assigned to known polymorphic mtDNA variation. A total of 409 heteroplasmic positions were identified for the cohort, with an average of 9 heteroplasmic variants per patient (ranging from 0 to 68).
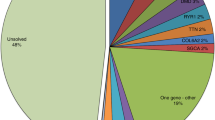
The mtDNA halpogroups of the patients, which were assigned according to Phylotree are as follows: 21 to haplogroup L0, 4 to L1, 10 to L2, 15 to L3 (total of L-haplogroups, which represent the African patients in this cohort: 50), 1 to M, 2 to N, 3 to J, 2 to T, 7 to H and 6 to U (total of non-L-haplogroup: 21). From long-range PCR and sequencing, no mtDNA rearrangements were detected. A separate muscle mtDNA copy number investigation on this cohort revealed two cases (P2, P59) with mtDNA depletion (mtDNA/nDNA <5). No candidate pathogenic mtDNA variants were detected in these two cases.
Non-haplogroup-associated mtDNA variants
After excluding all variants associated with any mtDNA haplogroup as reported in Phylotree, we firstly reviewed all high confidence substitutions in protein-coding genes (see Supplementary Figure S1A) and compared these in African patients with the non-African patients in the cohort. A total of 128 substitutions in 110 polymorphic positions were identified in patients with an African (L-) haplogroup. In 43/48 (86%) of African patients, one or more substitution were detected. Comparatively, substitutions occurred in a significantly lower percentage (13/23 or 57%) of non-African (non-L-haplogroup) patients, with a total of 59 substitutions found at 58 polymorphic positions (P=0.0037). No significant differences were observed between African and non-African haplogroups when comparing the percentages of variation among the three codon positions (results not shown). The variants were similar at the three codon positions for African and non-African haplogroups (results not shown graphically). Variants led to 49 synonymous and 61 non-synonymous substitutions in African haplogroup patients (ratio of 1:1.2). This was not significantly different when comparing the 28 synonymous versus 30 non-synonymous substitutions (ratio of 1:1.07) in non-African haplogroup patients (P=0.745, as illustrated in Supplementary Figure S1A).
Of the substitutions, 98/110 (89%) in the African haplogroups were transitions, which was similar (P=0.58) to the non-African haplogroups with 54/58 (93%) constituting transitions (Supplementary Figure S1B). This yields a transversion:transition ratio of 1:8 and 1:13.5, respectively, which is comparable to that reported by Pereira et al.17 In both patient groups, the majority of the variants occurred in neutral apolar amino acids at 78/110 and 38/58 for African and non-African patients, respectively (P=0.487), followed by 27/110 (African) and 12/58 (non-African) neutral polar changes (P=0.701). Only a small number occurred in basic polar (4/110 and 5/58, P=0.278) and acid polar (1/110 and 3/58, P=0.12) amino acids for the two groups, respectively (Supplementary Figure S1C).
When considering non-haplogroup substitutions in non-protein-coding positions, a total of 31 substitutions (in 28 polymorphic positions) were identified in 20 (40%) of the patients with an African haplogroup (see Supplementary Figure S2). In all, 20 substitutions (in 18 polymorphic positions) were identified in 10 (48%) patients with a non-African haplogroup. The majority of these variants were transversions in the African (22/28) and non-African (17/18) haplogroups (P=0.12; Supplementary Figure S2A). The distribution of these variants occurred generally in the same regions, that is, in the non-coding control region and rRNA genes with no significant differences in any of these regions owing to the relative small total number of variants observed (8 and 5, respectively).
For tRNA genes, most substitutions occurred in the stem regions for both groups, followed by the D-loop region and only African haplogroup patients showed variants in the variable and other regions (Supplementary Figure S2B). This is a significant observation as the acceptor and anticodon stem regions is considered hotspots for pathogenic mutations.18 The distribution of variants in the 12S and 16S ribosomal RNA genes indicates that a generally larger number of variants occurred in the non-stem regions (Supplementary Figure S2C). Although the total number of variants in the non-African patients in rRNA regions is higher (10/20) compared with African patients (10/31), it was statistically insignificant (P=0.249).
Disease-associated mtDNA variants
Table 1 summarizes the reported disease-associated variants found in this patient cohort. It shows, at varying allele frequencies, 10 different previously reported disease-associated variants in 12 of the 71 patients and included one variation in an rRNA, one in tRNA and eight in structural genes.
Non-coding regions
Firstly, a putative m.2756C>T mutation in the large mitochondrial ribosomal subunit was identified at a homoplasmic frequency in a female Caucasian patient, who also had a very low frequency and thus likely benign m.5958T>C variation in MTCO1. This case had a different phenotype (severe myopathy and cardiomyopathy) compared with the first case,19 at which time it was associated with two Thai LHON patients. More recently, this variation was described as a somatic mutation in pituitary adenoma, which leads to HIF1á destabilization.20 The second disease-associated variation occurring outside a non-protein region was an m.14723T>C variation in a conserved area of the genome in the TRNE gene and resides finally in the T-stem of the tRNA molecule. This variation has recently been described in a patient with chronic progressive external ophthalmoplegia, myopathy and a progressive increase in the proportion of COX-deficient fibres.32 We identified this variation also at a low frequency in an African female patient with a COX deficiency and a similar clinically profile.
Complex I
Five disease-associated variants were identified in complex I subunit encoding genes. Firstly, an m.3407G>A missense variation in a highly conserved region of the MTND1 gene, which results in the basic polar amino-acid substitution of p.Arg34His, was detected. This substitution was observed at an extremely low frequency and, considering that complex I is unaffected, it may not yet have a biological significant impact on the patient. This variation was initially associated with a rare variety of hypertrophic cardiomyopathy in a 65-year-old Indian patient,21 but subsequently suggested to be a polymorphism associated with the M5a haplogroup.22 The second complex I variation, m.4160T>C missense variation in the MTND1 gene, has been reported several times before23, 24, 25 and has been associated with LHON and the related neurological abnormalities involved. This variation substitutes a highly conserved amino-acid residue for another with the same polarity (p.Leu285Pro) and in silico analyses predicts it to have a detrimental impact. We identified this variation at a low frequency in one patient who had a clinical presentation indicative of LHON as well as a complex I enzyme deficiency. Two variants (m.10114T>C and m.10128C>A) in the MTND3 gene of complex I were observed in a female patient who presented with eye and muscular involvement (no enzyme data could be generated owing to a poor biopsy). The m.10114T>C missense variation was recently reported to be associated with aminoglycoside-induced ototoxicity in two African TB (Tuberculosis) patients31 and reported to have an impact on the OXPHOS capacity. This variation leads to the substitution of a poorly conserved amino-acid residue, neutral apolar isoleucine, to a neutral polar threonine at position 19 and have been predicted by in silico analyses to be a benign variation. The second variation in this patient, an m.10128C>A missense variation, was also described by Human et al31 for the same African TB patients with aminoglycoside-induced ototoxicity who harboured the m.10114T>C missense variation. This variation was predicted to be benign,31 but we found it to substitute a highly conserved amino-acid residue (p.Leu24Met) with a possible deleterious impact. Finally, a well-documented pathogenic m.14484T>C variation (p.Met64Val) was identified in one patient. This LHON mutation in the MTND6 gene of complex I was detected at a heteroplasmic level in an African male patient with a clinical and biochemical profile similar to what is commonly reported for this mutation.
Complex IV
Two disease-associated variants were identified in the MTCO1 encoding gene. Firstly, an m.5958T>C missense variation was identified at very low (∼3%, and thus likely not to contribute to the disease) allele frequency in a female patient (haplogroup U) who presented with a combined COX deficiency. This pathogenic variation substitutes a highly conserved neutral polar amino-acid residue to a basic polar residue (p.Tyr19His) and has been reported to cause a major defect in COX assembly.26 Secondly, the m.7080T>C missense variation, which changes a neutral apolar residue to one with similar polarity (p.Phe393Leu), has previously been reported as both a polymorphism in haplogroups U 27 and M12b,28 as well as a prostate cancer-associated point mutation.29 This substitution was identified at a homoplasmic frequency in a female patient who did not have a clear complex IV deficiency, but a combined deficiency of complexes I and III.
Complex III
Notably from the summary in Table 1, a high frequency (five cases) of the m.15735C>T variation in the MTCYB gene of complex III were found, which accounts for 7% in this cohort. The clinical profile among these five patients varies substantially, although a complex III deficiency was identified in four of these cases. However, some aspects of this putative pathogenic variation, which has recently been reported in a patient of European descent with muscle weakness, ptosis and cardiomyopathy,34 should be noted. The variation was initially observed in a European sequence,35 but later also identified in two African sequences,36 which resulted in its classification as a polymorphism belonging to the L2a1b haplogroup (Phylotree.org). In concurrence with this, of the five cases in our group where this variation was detected, four clustered to this haplogroup and the other to a subgroup of L0. It was also recently reported in two L-haplogroup South African TB patients, with aminoglycoside-induced ototoxicity, that it was predicted to be benign.31 Indeed, the CI for the resultant p.Ala330Val substitution is relatively low at 0.535 and further in silico evaluation for this variation indicates that it is probably benign and tolerated.
Novel variants of unknown significance
A large number of novel variants, which have not previously been reported to be either polymorphic or pathogenic, were unidentified at various allele frequencies in our group. To identify novel candidate pathogenic variants, we, however, limited the variants to an allele frequency equal or higher than 20% in this investigation. The impact of lower frequency (20%) novel variants can, however, not be disregarded as these levels may already have, or develop to, a significant biological impact. Using these limitations, 11 novel variants with possible, but unknown pathogenic significance were identified in 13 patients. These are summarized in Table 2, along with the in silico evaluation of their predicted impact.
Two of these variants occurred in non-structural genes. The first of these, an m.1835A>G heteroplasmic MTRNR2 variation, was found in a patient with a (likely unrelated) isolated complex II deficiency. Secondly, a homoplasmic m.4301A>T variation in the TRNI gene, which is next to the location for a pathogenic homoplasmic m.4300A>G mutation found in a patient with hypertrophic cardiomyopathy,37 was observed in a severely affected female who presented with a multi-systemic profile, isolated muscle complex III deficiency, but without cardiac involvement.
Of the five novel missense variants occurring in genes encoding complex I subunits, only one case presented with a (combined) complex I enzyme deficiency. For one case, harbouring a predicted damaging heteroplasmic m.4789G>A variation, maternal inheritance was documented. A clear complex I deficiency was not present in this case, albeit near the lower reference limit. Although these variants should be considered separately in the various cases, it was also interesting to note that eye involvement forms part of the clinical profile in all of these cases.
In the MTCO1 gene, one novel frameshift and one novel missense variation (in three cases) was observed. The heteroplasmic m.5935Adel, which results in an early frameshift and subsequent termination (p.Asn11ThrfsX19), occurs in a clinically severely affected patient, but without complex IV deficiency. Similarly, a homoplasmic (predicted benign) variation of m.6723G>A were detected in three cases without complex IV deficiency.
Finally, two missense variants in the MTCYB gene, m.14883C>T and m.15272A>G, occurred at almost homoplasmic levels in two separate patients. Although in silico predictions for both variants indicated that these are likely to be benign, muscle complex III deficiency was observed in both cases.
Discussion
We investigated the mtDNA variants and more specifically the occurrence of known and novel mtDNA variants in post-mitotic (muscle) tissue from a clinically and ethnically heterogeneous group of South African paediatric patients who were diagnosed with a mitochondrial disorder. A clinical evaluation of the greater section of this cohort recently revealed that among patients of African descent, a myopathic clinical presentation was more common, whereas Caucasian patients presented predominantly with central nervous system involvement.8 As reported here, next-generation sequencing technology of the entire mitochondrial genome on this cohort enabled the identification of a great number of mtDNA variants and at varied allele frequencies, which can in part be attributed to the post-mitotic tissue used in this study. The non-haplogroup-defining variants between the African and non-African patients in this cohort are clearly different in number, with a significantly higher number found in African patients. Although a more extensive investigation is required, our results may already indicate that in the African patient population, a greater number and diversity of pathogenic mtDNA mutations may be found.
The diversity of mtDNA variants found in this cohort is reflected in the varied disease-associated variants and novel variants of unknown significance, as well as the varied allele frequencies at which they occur. Several of these low-frequency, heteroplasmic variants in the muscle may indeed be due to somatic mtDNA mosaicism and not disease-causing variants,38, 39 which highlights the importance of follow-up investigations such as cybrid studies to establish pathogenicity for these variants. We have identified a number of previously documented disease-associated variants in this cohort, of which only one (m.14484T>C LHON mutation) can be considered as a frequently occurring syndrome-associated mutation. This correlates well with the absence of characteristic syndromes and difference in phenotypes as reported previously for the main part of this group.8 Using a minimum allele frequency of 20%, we also report a number of variants that was considered to have the potential, but at varied probabilities, to be pathogenic based on the biological data and predicted impact analysis. Although these predictions, along with clinical and biochemical data, could be an indication of which of these variants are likely to be pathogenic, the unavailability of a family history as well as additional tissues have greatly limited a better evaluation of the pathogenic potential of variants. Consequently, these variants have to be further investigated separately to determine pathogenicity, which is beyond the scope of this report.
In conclusion, in the absence of disease-based epidemiological data in African patients with mitochondrial RC disease, our strategy using next-generation sequence technology enabled the fairly rapid evaluation of full length mtDNA sequences of a relatively large cohort of patients. Furthermore, it allowed detection of low level of allele frequencies (heteroplasmy), which may have remained undetected if established sequencing technology was used. Although the cohort represents a small fraction of a mostly under-diagnosed, heterogeneous disease population, the data should nevertheless significantly contribute to expand our knowledge of the spectrum of causative mtDNA variants responsible for mitochondrial disease in South African paediatric patients. However, until the impact of some of the previously reported and novel variants has been fully resolved, it is not possible to determine accurately the prevalence of mtDNA mutations in this patient cohort or in the broader patient population. From a practical point of view, we finally conclude that molecular genetic investigations in African patients with RC disorders should follow a full-length mtDNA sequencing approach rather than single mutation detection strategies, which is most often based on clinical and genetic information from non-African patients. With the recent developments of next-generation sequence technology, this approach have become feasible and, with the inclusion of nDNA investigations, which constitutes the majority of pathogenic mutations, a better understanding of the aetiology of mitochondrial disease in the African population may soon be possible.
Accession codes
References
Skladal D, Halliday J, Thorburn DR : Minimum birth prevalence of mitochondrial respiratory chain disorders in children. Brain 2003; 126: 1905–1912.
Anderson S, Bankier AT, Barrell BG et al: Sequence and organization of the human mitochondrial genome. Nature 1981; 290: 457–465.
Reinecke F, Smeitink JAM, van der Westhuizen FH : OXPHOS gene expression and control in mitochondrial disorders. Biochim Biophys Acta 2009; 1792: 1113–1121.
Zeviani M, Di Donato S : Mitochondrial disorders. Brain 2004; 127: 2153–2172.
Swalwell H, Kirby DM, Blakely EL et al: Respiratory chain complex I deficiency caused by mitochondrial DNA mutations. Eur J Hum Genet 2011; 19: 769–775.
Tuppen HA, Blakely EL, Turnbull DM, Taylor RW : Mitochondrial DNA mutations and human disease. Biochim Biophys Acta 2010; 1797: 113–128.
Elliott HR, Samuels DC, Eden JA, Relton CL, Chinnery PF : Pathogenic mitochondrial DNA mutations are common in the general population. Am J Hum Genet 2008; 83: 254–260.
Smuts I, Louw R, du Toit H, Klopper B, Mienie LJ, van der Westhuizen FH : An overview of a cohort of South African patients with mitochondrial disorders. J Inherit Metab Dis 2010; e-pub ahead of print 5 February 2010; doi:10.1007/s10545-009-9031-8.
Wolf NI, Smeitink JAM : Mitochondrial disorders: a proposal for consensus diagnostic criteria in infants and children. Neurology 2002; 59: 1402–1405.
Rahman S, Blok RB, Dahl HH et al: Leigh syndrome: clinical features and biochemical and DNA abnormalities. Ann Neurol 1996; 39: 343–351.
Janssen AJ, Trijbels FJ, Sengers RC et al: Spectrophotometric assay for complex I of the respiratory chain in tissue samples and cultured fibroblasts. Clin Chem 2007; 53: 729–734.
Shepherd D, Garland PB : The kinetic properties of citrate synthase from rat liver mitochondria. Biochem J 1969; 114: 597–610.
Taylor RW, Taylor GA, Durham SE, Turnbull DM : The determination of complete human mitochondrial DNA sequences in single cells: implications for the study of somatic mitochondrial DNA point mutations. Nucleic Acids Res 2001; 29: 15e74.
Hedges D, Burges D, Powell E et al: Exome sequencing of a multigenerational human pedigree. PLoS One 2009; 4: e8232.
Zaragoza MV, Fass J, Diegoli M, Lin D, Arbustini E : Mitochondrial DNA variant discovery and evaluation in human cardiomyopathies through next-generation sequencing. PLoS One 2010; 5: e12295.
De Schrijver JM, De Leeneer K, Lefever S et al: Analysing 454 amplicon resequencing experiments using the modular and database oriented variant identification pipeline. BMC Bioinform 2010; 11: 269.
Pereira L, Freitas F, Fernandes V et al: The diversity present in 5140 human mitochondrial genomes. Am J Hum Genet 2009; 84: 628–640.
McFarland R, Elson JL, Taylor RW, Howell N, Turnbull DM : Assigning pathogenicity to mitochondrial tRNA mutations: when ‘definitely maybe’ is not good enough. Trends Genet 2004; 20: 591–596.
Joyjinda Y : The role of mitochondrial background in the expression of Leber hereditary optic neuropathy. Masters thesis, Mahidol University, Mahidol, Thailand, 2007.
Porcelli AM, Ghelli A, Ceccarelli C : The genetic and metabolic signature of oncocytic transformation implicates HIF1 destabilization. Hum Mol Genet 2010; 19: 1019–1032.
Prasad GN, Vanniarajan A, Emmanuel C, Cherian KM, Singh L, Thangaraj K : Novel mitochondrial DNA mutations in a rare variety of hypertrophic cardiomyopathy. Int J Cardiol 2006; 109: 432–433.
Bandelt H, Yao Y, Salas A : The search of ‘novel’ mtDNA mutations in hypertrophic cardiomyopathy: MITOMAPping as a risk factor. Int J Cardiol 2008; 126: 439–442.
Howell N, Kubacka I, Xu M, Mccullough DA : Leber hereditary optic neuropathy: involvement of the mitochondrial ND I gene and evidence for an intragenic suppressor mutation. Am J Hum Genet 1991; 48: 935–942.
Howell N, Kubacka I, Halvorson S, Howell B, Mccullough DA, Mackey D : Phylogenetic analysis of the mitochondrial genomes from Leber hereditary optic neuropathy pedigrees. Genetics 1995; 140: 285–302.
Mackey DA : Three subgroups of patients from the United Kingdom with Leber hereditary optic neuropathy. Eye 1994; 8: 431–436.
Pye D, Kyriakouli DS, Taylor GA et al: Production of transmitochondrial cybrids containing naturally occurring pathogenic mtDNA variants. Nucleic Acids Res 2006; 34: e95.
Hernstadt C, Elson JL, Fahy E et al: Reduced-median-network analysis of complete mitochondrial DNA coding-region sequences for the major African, Asian, and European haplogroups. Am J Hum Genet 2002; 70: 1152–1171.
Chandrasekar A, Kumar S, Sreenath J et al: Updating phylogeny of mitochondrial DNA macrohaplogroup M in India: dispersal of modern human in South Asian corridor. PLoS One 2009; 4: e7447.
Petros JA, Baumann AK, Ruiz-Pesini E et al: mtDNA mutations increase tumorigenicity in prostate cancer. Proc Natl Acad Sci USA 2005; 102: 719–724.
Gomez-Duran A, Pacheu-Grau D, Lopez-Gallardo E et al: Unmasking the causes of multifactorial disorders: OXPHOS differences between mitochondrial haplogroups. Hum Mol Genet 2010; 19: 3343–3353.
Human H, Hagen CM, De Jong G et al: Investigation of mitochondrial sequence variants associated with aminoglycoside-induced ototoxicity in South African TB patients on aminoglycosides. Biochem Biophys Res Commun 2010; 393: 751–756.
Alston CL, Lowe J, Turnbull DM, Maddison P, Taylor RW : A novel mitochondrial tRNA(Glu) (MTTE) gene mutation causing chronic progressive external ophthalmoplegia at low levels of heteroplasmy in muscle. J Neurol Sci 2010; 298: 140–144.
Johns DR, Neufeld MJ, Park RD : An ND-6 mitochondrial DNA mutation associated with Leber hereditary optic neuropathy. Biochem Biophys Res Commun 1992; 187: 1551–1557.
Bannwarth S, Procaccio V, Rouzier C et al: Rapid identification of mitochondrial DNA (mtDNA) mutations in neuromuscular disorders by using surveyor strategy. Mitochondrion 2008; 8: 136–145.
Coble MD, Just RS, O’Callaghan JE et al: Single nucleotide polymorphisms over the entire mtDNA genome that increase the power of forensic testing in Caucasians. Int J Legal Med 2004; 118: 137–146.
Kivisild T, Shen P, Wall DP et al: The role of selection in the evolution of human mitochondrial genomes. Genetics 2006; 172: 373–387.
Taylor RW, Giordano C, Davidson MM et al: A homoplasmic mitochondrial transfer ribonucleic acid mutation as a cause of maternally inherited hypertrophic cardiomyopathy. J Am Coll Cardiol 2003; 21: 1786–1796.
He Y, Wu J, Dressman DC et al: Heteroplasmic mitochondrial DNA mutations in normal and tumour cells. Nature 2010; 464: 610–614.
Li M, Schönberg A, Schaefer M et al: Detecting heteroplasmy from high-throughput sequencing of complete human mitochondrial DNA genomes. Am J Hum Genet 2010; 87: 237–249.
Acknowledgements
We acknowledge the supporting contribution of the clinical staff of the Department of Paediatrics and Child Health, Steve Biko Academic Hospital, University of Pretoria, South Africa, as well as Charlotte Alston, Emma Watson and John Yarham of the Mitochondrial Research Group, Institute for Ageing and Health, Newcastle University. We are grateful for the financial support of the Department of Science and Technology and Medical Research Council of South Africa. RWT and DMT are funded by a Wellcome Trust Programme Grant (074454/Z/04/Z) and the UK NHS Specialised Services ‘Rare Mitochondrial Disorders of Adults and Children’ Diagnostic Service (http://www.mitochondrialncg.nhs.uk). FHvdW is supported by a Newcastle University Research Committee Visiting Professorship bursary.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website
Rights and permissions
About this article
Cite this article
van der Walt, E., Smuts, I., Taylor, R. et al. Characterization of mtDNA variation in a cohort of South African paediatric patients with mitochondrial disease. Eur J Hum Genet 20, 650–656 (2012). https://doi.org/10.1038/ejhg.2011.262
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2011.262
Keywords
This article is cited by
-
Leber’s hereditary optic neuropathy plus dystonia caused by the mitochondrial ND1 gene m.4160 T > C mutation
Neurological Sciences (2022)
-
Mitochondrial disease patients with novel ND4 12058A > C and ND1 m.3911A > G variations: implications for a role in the phenotype following a bioinformatic investigation
Molecular Biology Reports (2021)
-
Human origins in a southern African palaeo-wetland and first migrations
Nature (2019)
-
Haplogroup Context is Less Important in the Penetrance of Mitochondrial DNA Complex I Mutations Compared to mt-tRNA Mutations
Journal of Molecular Evolution (2018)