Abstract
E3 protein ligases enhance transfer of ubiquitin-like (Ubl) proteins from E2 conjugating enzymes to substrates by stabilizing the thioester-charged E2~Ubl in a closed configuration optimally aligned for nucleophilic attack. Here, we report biochemical and structural data that define the N-terminal domain of the Homo sapiens ZNF451 as the catalytic module for SUMO E3 ligase activity. The ZNF451 catalytic module contains tandem SUMO-interaction motifs (SIMs) bridged by a Pro-Leu-Arg-Pro (PLRP) motif. The first SIM and PLRP motif engage thioester-charged E2~SUMO while the next SIM binds a second molecule of SUMO bound to the back side of E2. We show that ZNF451 is SUMO2 specific and that SUMO modification of ZNF451 may contribute to activity by providing a second molecule of SUMO that interacts with E2. Our results are consistent with ZNF451 functioning as a bona fide SUMO E3 ligase.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Kerscher, O., Felberbaum, R. & Hochstrasser, M. Modification of proteins by ubiquitin and ubiquitin-like proteins. Annu. Rev. Cell Dev. Biol. 22, 159–180 (2006).
Streich, F.C. Jr. & Lima, C.D. Structural and functional insights to ubiquitin-like protein conjugation. Annu. Rev. Biophys. 43, 357–379 (2014).
Hendriks, I.A. et al. Uncovering global SUMOylation signaling networks in a site-specific manner. Nat. Struct. Mol. Biol. 21, 927–936 (2014).
Sampson, D.A., Wang, M. & Matunis, M.J. The small ubiquitin-like modifier-1 (SUMO-1) consensus sequence mediates Ubc9 binding and is essential for SUMO-1 modification. J. Biol. Chem. 276, 21664–21669 (2001).
Tatham, M.H. et al. Polymeric chains of SUMO-2 and SUMO-3 are conjugated to protein substrates by SAE1/SAE2 and Ubc9. J. Biol. Chem. 276, 35368–35374 (2001).
Saitoh, H. & Hinchey, J. Functional heterogeneity of small ubiquitin-related protein modifiers SUMO-1 versus SUMO-2/3. J. Biol. Chem. 275, 6252–6258 (2000).
Reverter, D. & Lima, C.D. Insights into E3 ligase activity revealed by a SUMO–RanGAP1–Ubc9–Nup358 complex. Nature 435, 687–692 (2005).
Pruneda, J.N. et al. Structure of an E3:E2~Ub complex reveals an allosteric mechanism shared among RING/U-box ligases. Mol. Cell 47, 933–942 (2012).
Branigan, E., Plechanovova, A., Jaffray, E.G., Naismith, J.H. & Hay, R.T. Structural basis for the RING-catalyzed synthesis of K63-linked ubiquitin chains. Nat. Struct. Mol. Biol. 22, 597–602 (2015).
Buetow, L. et al. Activation of a primed RING E3–E2-ubiquitin complex by non-covalent ubiquitin. Mol. Cell 58, 297–310 (2015).
Dou, H., Buetow, L., Sibbet, G.J., Cameron, K. & Huang, D.T. BIRC7–E2 ubiquitin conjugate structure reveals the mechanism of ubiquitin transfer by a RING dimer. Nat. Struct. Mol. Biol. 19, 876–883 (2012).
Dou, H., Buetow, L., Sibbet, G.J., Cameron, K. & Huang, D.T. Essentiality of a non-RING element in priming donor ubiquitin for catalysis by a monomeric E3. Nat. Struct. Mol. Biol. 20, 982–986 (2013).
Gareau, J.R., Reverter, D. & Lima, C.D. Determinants of small ubiquitin-like modifier 1 (SUMO1) protein specificity, E3 ligase, and SUMO-RanGAP1 binding activities of nucleoporin RanBP2. J. Biol. Chem. 287, 4740–4751 (2012).
Plechanovová, A., Jaffray, E.G., Tatham, M.H., Naismith, J.H. & Hay, R.T. Structure of a RING E3 ligase and ubiquitin-loaded E2 primed for catalysis. Nature 489, 115–120 (2012).
Scott, D.C. et al. Structure of a RING E3 trapped in action reveals ligation mechanism for the ubiquitin-like protein NEDD8. Cell 157, 1671–1684 (2014).
Klug, H. et al. Ubc9 sumoylation controls SUMO chain formation and meiotic synapsis in Saccharomyces cerevisiae. Mol. Cell 50, 625–636 (2013).
Wickliffe, K.E., Lorenz, S., Wemmer, D.E., Kuriyan, J. & Rape, M. The mechanism of linkage-specific ubiquitin chain elongation by a single-subunit E2. Cell 144, 769–781 (2011).
Hochstrasser, M. SP-RING for SUMO: new functions bloom for a ubiquitin-like protein. Cell 107, 5–8 (2001).
Reindle, A. et al. Multiple domains in Siz SUMO ligases contribute to substrate selectivity. J. Cell Sci. 119, 4749–4757 (2006).
Pichler, A., Knipscheer, P., Saitoh, H., Sixma, T.K. & Melchior, F. The RanBP2 SUMO E3 ligase is neither HECT- nor RING-type. Nat. Struct. Mol. Biol. 11, 984–991 (2004).
Hecker, C.M., Rabiller, M., Haglund, K., Bayer, P. & Dikic, I. Specification of SUMO1- and SUMO2-interacting motifs. J. Biol. Chem. 281, 16117–16127 (2006).
Song, J., Durrin, L.K., Wilkinson, T.A., Krontiris, T.G. & Chen, Y. Identification of a SUMO-binding motif that recognizes SUMO-modified proteins. Proc. Natl. Acad. Sci. USA 101, 14373–14378 (2004).
Song, J., Zhang, Z., Hu, W. & Chen, Y. Small ubiquitin-like modifier (SUMO) recognition of a SUMO binding motif: a reversal of the bound orientation. J. Biol. Chem. 280, 40122–40129 (2005).
Guervilly, J.H. et al. The SLX4 complex is a SUMO E3 ligase that impacts on replication stress outcome and genome stability. Mol. Cell 57, 123–137 (2015).
Merrill, J.C. et al. A role for non-covalent SUMO interaction motifs in Pc2/CBX4 E3 activity. PLoS ONE 5, e8794 (2010).
Ouyang, J. et al. Noncovalent interactions with SUMO and ubiquitin orchestrate distinct functions of the SLX4 complex in genome maintenance. Mol. Cell 57, 108–122 (2015).
Yang, S.H. & Sharrocks, A.D. The SUMO E3 ligase activity of Pc2 is coordinated through a SUMO interaction motif. Mol. Cell. Biol. 30, 2193–2205 (2010).
Liu, Q. et al. The binding interface between an E2 (UBC9) and a ubiquitin homologue (UBL1). J. Biol. Chem. 274, 16979–16987 (1999).
Tatham, M.H. et al. Role of an N-terminal site of Ubc9 in SUMO-1, -2, and -3 binding and conjugation. Biochemistry 42, 9959–9969 (2003).
Capili, A.D. & Lima, C.D. Structure and analysis of a complex between SUMO and Ubc9 illustrates features of a conserved E2-Ubl interaction. J. Mol. Biol. 369, 608–618 (2007).
Duda, D.M. et al. Structure of a SUMO-binding-motif mimic bound to Smt3p-Ubc9p: conservation of a non-covalent ubiquitin-like protein-E2 complex as a platform for selective interactions within a SUMO pathway. J. Mol. Biol. 369, 619–630 (2007).
Knipscheer, P., van Dijk, W.J., Olsen, J.V., Mann, M. & Sixma, T.K. Noncovalent interaction between Ubc9 and SUMO promotes SUMO chain formation. EMBO J. 26, 2797–2807 (2007).
Brzovic, P.S., Lissounov, A., Christensen, D.E., Hoyt, D.W. & Klevit, R.E.A. UbcH5/ubiquitin noncovalent complex is required for processive BRCA1-directed ubiquitination. Mol. Cell 21, 873–880 (2006).
Eddins, M.J., Carlile, C.M., Gomez, K.M., Pickart, C.M. & Wolberger, C. Mms2–Ubc13 covalently bound to ubiquitin reveals the structural basis of linkage-specific polyubiquitin chain formation. Nat. Struct. Mol. Biol. 13, 915–920 (2006).
Hibbert, R.G., Huang, A., Boelens, R. & Sixma, T.K. E3 ligase Rad18 promotes monoubiquitination rather than ubiquitin chain formation by E2 enzyme Rad6. Proc. Natl. Acad. Sci. USA 108, 5590–5595 (2011).
Karvonen, U., Jaaskelainen, T., Rytinki, M., Kaikkonen, S. & Palvimo, J.J. ZNF451 is a novel PML body- and SUMO-associated transcriptional coregulator. J. Mol. Biol. 382, 585–600 (2008).
Hendriks, I.A., D'Souza, R.C., Chang, J.G., Mann, M. & Vertegaal, A.C. System-wide identification of wild-type SUMO-2 conjugation sites. Nat. Commun. 6, 7289 (2015).
Matic, I. et al. Site-specific identification of SUMO-2 targets in cells reveals an inverted SUMOylation motif and a hydrophobic cluster SUMOylation motif. Mol. Cell 39, 641–652 (2010).
Schimmel, J. et al. Uncovering SUMOylation dynamics during cell-cycle progression reveals FoxM1 as a key mitotic SUMO target protein. Mol. Cell 53, 1053–1066 (2014).
Tammsalu, T. et al. Proteome-wide identification of SUMO2 modification sites. Sci. Signal. 7, rs2 (2014).
Eisenhardt, N. et al. A novel vertebrate SUMO enzyme family reveals insights into SUMO-chain assembly. Nat. Struct. Mol. Biol. doi:10.1038/nsmb.3114 (2 November 2015).
Cappadocia, L. et al. Structural and functional characterization of the phosphorylation-dependent interaction between PML and SUMO1. Structure 23, 126–138 (2015).
Chang, C.C. et al. Structural and functional roles of Daxx SIM phosphorylation in SUMO paralog-selective binding and apoptosis modulation. Mol. Cell 42, 62–74 (2011).
Lima, C.D. & Schulman, B.A. Structural biology: a protein engagement RING. Nature 489, 43–44 (2012).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Bernier-Villamor, V., Sampson, D.A., Matunis, M.J. & Lima, C.D. Structural basis for E2-mediated SUMO conjugation revealed by a complex between ubiquitin-conjugating enzyme Ubc9 and RanGAP1. Cell 108, 345–356 (2002).
Pickart, C.M. & Rose, I.A. Functional heterogeneity of ubiquitin carrier proteins. J. Biol. Chem. 260, 1573–1581 (1985).
Parker, J.L. & Ulrich, H.D. SIM-dependent enhancement of substrate-specific SUMOylation by a ubiquitin ligase in vitro. Biochem. J. 457, 435–440 (2014).
Mascle, X.H. et al. Identification of a non-covalent ternary complex formed by PIAS1, SUMO1, and UBC9 proteins involved in transcriptional regulation. J. Biol. Chem. 288, 36312–36327 (2013).
Abascal, F., Tress, M.L. & Valencia, A. Alternative splicing and co-option of transposable elements: the case of TMPO/LAP2alpha and ZNF451 in mammals. Bioinformatics 31, 2257–2261 (2015).
Udeshi, N.D. et al. Refined preparation and use of anti-diglycine remnant (K-epsilon-GG) antibody enables routine quantification of 10,000s of ubiquitination sites in single proteomics experiments. Mol. Cell. Proteomics 12, 825–831 (2013).
Scaglioni, P.P. et al. A CK2-dependent mechanism for degradation of the PML tumor suppressor. Cell 126, 269–283 (2006).
Yunus, A.A. & Lima, C.D. in Methods in Molecular Biology: SUMO Protocols Vol. 497 (ed. Ulrich, H.D.) 167–186 (Springer, 2009).
Armstrong, A.A., Mohideen, F. & Lima, C.D. Recognition of SUMO-modified PCNA requires tandem receptor motifs in Srs2. Nature 483, 59–63 (2012).
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Strong, M. et al. Toward the structural genomics of complexes: crystal structure of a PE/PPE protein complex from Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 103, 8060–8065 (2006).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Acknowledgements
We would like to thank A.P.'s laboratory for providing the pLou3-ZNF4512–56 and pLou3-ZNF4512–247 plasmids, and A.P.'s and C.D.L.'s labs for discussion. Data for this study were measured at beamline X29 of the National Synchrotron Light Source, whose financial support comes principally from the Offices of Biological and Environmental Research and of Basic Energy Sciences of the US Department of Energy (DOE), the National Center for Research Resources (P41RR012408) and the National Institute of General Medical Sciences (NIGMS; P41GM103473) of the US National Institutes of Health (NIH). This work is also based on research conducted at the Northeastern Collaborative Access Team beamlines, which are funded by the NIGMS from the NIH (P41 GM103403). The Pilatus 6M detector on the 24-ID-C beamline is funded by a NIH Office of Research Infrastructure Programs High End Instrumentation grant (S10 RR029205). This research used resources of the Advanced Photon Source, a DOE Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under contract no. DE-AC02-06CH11357. Research reported in this publication was supported in part by the NIGMS of the NIH under award number GM065872 (C.D.L.) and the NIH–National Cancer Institute Cancer Center Support Grant P30 CA008748. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH. A.P. is supported by the Max Planck Society and the Deutsche Forschungsgemeinschaft (DFG-SPP1365 PI 917/2-1). L.C. is supported in part by a fellowship from the Fonds de Recherche du Québec–Santé. C.D.L. is supported as an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
A.P. initiated the project. L.C. and C.D.L. designed experiments that L.C. performed. L.C. and C.D.L. wrote the manuscript with input from A.P.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Noncovalent interaction between SUMO2 and UBC9
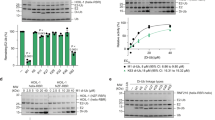
Fluorescence polarization assay performed using 50 nM Alexa488-labeled SUMO21–93 and serially diluted UBC9. Data is mean ± s.d. (n=3 technical replicates). Data was fitted to a single site binding model accounting for ligand depletion. The assay was performed in 20 mM HEPES pH 7.5, 50 mM NaCl and 0.1 % Tween-20.
Supplementary Figure 2 Electron density maps and analysis of the protein-protein contacts in the asymmetric unit.
(a) Stereo view of the electron density map around the ZNF451 PLRP motif. Protein is in stick representation with red spheres representing water molecules. ZNF451 is colored in green, SUMO2D in yellow and SUMO2B in orange. The simulated annealing composite omit 2mFo – DFc map is contoured at 1.5 σ and shown as a grey mesh. (b) Surface representation of the proteins in the asymmetric unit of the ZNF451–SUMO2~RANGAP1–UBC9 complex. Selected interface areas are presented and highlighted using arrows and filled circles. (c) Table generated with PISA (Krissinel, E. & Henrick, K. J Mol Biol 372, 774-97, 2007) showing the list of interactions in the crystal lattice. ΔiG is the solvation free energy gain upon formation of the interface. NHB and NSB are the numbers of hydrogen bonds and salt-bridges across the interface. Only interfaces burying more than 100 Å2 of accessible surface area are listed.
Supplementary Figure 3 Structural comparison of the SUMO-SIM interactions and the PLRP motif.
SUMO–SIM interactions at the (a) SUMO2D–ZNF451, (b) SUMO2D–RANBP2 (pdb 3UIN; Gareau, J.R et al., J Biol Chem 287, 4740-51, 2012), or (c) SUMO2B–ZNF451 interfaces. The left panels highlight the SUMO2 residues that interact with the SIMs. SUMO2 is in surface representation. The SUMO2 residues interacting with the SIMs are presented in stick representation. Residues belonging to ZNF451 and RANBP2 are also in stick representation. The right panels highlight the hydrogen bond and salt bridges interactions involved in the SUMO–SIM interface. For simplicity, only the side-chains establishing hydrogen bonds or salt bridges interactions are presented. Grey dashes and bold numbers represent hydrogen bonds and their length. The view was rotated by 45° along the x axis as compared to the left panels. (d) Structural similarity between the PLRP motif of ZNF451 and the one present in other proteins. Structures containing a PLRP sequence were selected and the PLRP sequence of each protein was aligned to the one of ZNF451. Only structural alignment resulting in a rmsd < 1 Å are presented. Also, only one representative structure per protein is presented. The pdb accession numbers are provided for each of the structures.
Supplementary Figure 4 Effect of the ZNF4512–55-SUMO215–89 fusion on the conjugation rates of SUMO2 to the p53 tetramerization domain.
Single-turnover conjugation rates of SUMO2 to p53 tetramerization domain (residues 320-393) in presence of 100 nM fusion ZNF4512–55SUMO215–89 or ZNF4512–56. Left, plot showing the initial rate of SUMO conjugation versus p53 concentration. Data show mean ± s.d. (n=3 technical replicates). Data for ZNF4512–56 is the same as in Figure 1b. Right, representative SDS-PAGE for SUMO2 conjugation to p53 in presence of 100 nM fusion ZNF4512–55SUMO215–89. Assays were done at 0°C in 20 mM HEPES pH 7.5, 50 mM NaCl, 0.1 % Tween-20 and 5 mM EDTA using 28 ± 5 nM thioester charged UBC9~Alexa488-labeled SUMO2, 100 nM fusion ZNF4512–55SUMO215–89 and serially diluted p53 at the indicated concentration.
Supplementary Figure 5 Purification of SUMO215–93~Znf4512–110
12 % NuPAGE gel run with MES buffer and stained with Coommassie Brillant Blue showing the purification of SUMO215–93-Znf4512–110. Conjugation reaction was performed for 1h and passed on a HiPrep Desalting column equilibrated with 20 mM Tris pH 8, 50 mM NaCl and 1 mM 2-mercaptoethanol. The desalted reaction was then applied on a MonoQ column and eluted with a 50 to 1,000 mM NaCl gradient. Fractions 32 and 33 were pooled and used in lysine discharge assays.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Tables 1 and 2 (PDF 1082 kb)
Supplementary Data Set 1
Uncropped gel and representative gels related to main figures (PDF 1941 kb)
Rights and permissions
About this article
Cite this article
Cappadocia, L., Pichler, A. & Lima, C. Structural basis for catalytic activation by the human ZNF451 SUMO E3 ligase. Nat Struct Mol Biol 22, 968–975 (2015). https://doi.org/10.1038/nsmb.3116
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3116
This article is cited by
-
Signalling mechanisms and cellular functions of SUMO
Nature Reviews Molecular Cell Biology (2022)
-
Structural basis for the E3 ligase activity enhancement of yeast Nse2 by SUMO-interacting motifs
Nature Communications (2021)
-
Overexpression of CpSIZ1, a SIZ/PIAS-Type SUMO E3 Ligase from Wintersweet (Chimonanthus praecox), Delays Flowering, Accelerates Leaf Senescence and Enhances Cold Tolerance in Arabidopsis
Plant Molecular Biology Reporter (2021)
-
Molecular mechanisms of topoisomerase 2 DNA–protein crosslink resolution
Cellular and Molecular Life Sciences (2020)
-
CRISPR-UMI: single-cell lineage tracing of pooled CRISPR–Cas9 screens
Nature Methods (2017)