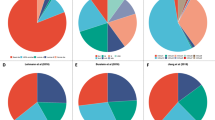
Abstract
The explosion of knowledge in cancer biology in the past two decades has led to the identification of specific molecular circuits in solid tumors. These pathways reflect specific abnormalities thought to drive malignant progression. This knowledge has also generated a vast panel of cancer biomarkers although many of these biomarkers lack sufficient research and validation to be used in the clinic. This Review discusses relevant molecular prognostic and/or predictive biomarkers in the six leading tumors with the highest contribution to cancer mortality: breast, lung, colorectal, prostate, pancreatic and ovarian cancer. Each biomarker is described according to its associated clinicopathological presentation and specific associated molecular interactions. Despite only few biomarkers being currently implemented in clinical practice, a new generation of predictors is emerging that could modify the classic organ-based cancer classification (for example, defects in DNA repair, aberrant MAPK signaling and aberrant PI3K/Akt/mTOR signaling). The advent of high-throughput strategies will also probably substitute monobiomarker strategies.
Key Points
-
Key oncogenic events and dysfunctional pathways have been identified over the past few decades in solid tumors
-
Many molecular-targeted agents have recently been approved or are in advanced clinical development, and these therapeutic tools target specific molecular abnormalities in the tumor allowing 'tailored treatment strategies'
-
Despite huge efforts, very few biomarkers are used in daily practice; examples of biomarkers in clinical use include ER, PR, HER2, Oncotype Dx®, MammaPrint® (breast cancer), EGFR mutations (lung cancer) and KRAS mutations (colorectal cancer)
-
Nevertheless a new generation of novel biomarkers that will modify the organ-based classification of cancer is emerging—such as DNA repair dysfunctionality, MAPK signaling and the PI3K/Akt/mTOR axis
-
Researchers and clinical investigators should be particularly aware of methodological issues when they consider biomarkers for bedside use
-
High-throughput analyses are being implemented in most tumor types, and given the huge number of possible effectors and the complex intricate pathways, high-throughput-based analysis will probably supercede monobiomarker strategies
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Jemal, A. et al. Cancer statistics, CA. Cancer J. Clin. 59, 225–249 (2009).
Early Breast Cancer Trialists' Collaborative Group (EBCTCG). Effects of chemotherapy and hormonal therapy for early breast cancer on recurrence and 15-year survival: an overview of the randomised trials. Lancet 365, 1687–1717 (2005).
Harris, L. et al. American Society of Clinical Oncology update of recommendations for the use of tumor markers in breast cancer. J. Clin. Oncol. 25, 5287–5312 (2007).
Grann, V. R. et al. Hormone receptor status and survival in a population-based cohort of patients with breast carcinoma. Cancer 103, 2241–2251 (2005).
Dowsett, M. & Dunbier, A. K. Emerging biomarkers and new understanding of traditional markers in personalized therapy for breast cancer. Clin. Cancer Res. 14, 8019–8026 (2008).
Penault-Llorca, F. et al. Ki67 expression and docetaxel efficacy in patients with estrogen receptor-positive breast cancer. J. Clin. Oncol. 27, 2809–2815 (2009).
Urruticoechea, A., Smith, I. E. & Dowsett, M. Proliferation marker Ki-67 in early breast cancer. J. Clin. Oncol. 23, 7212–7220 (2005).
Look, M. P. et al. Pooled analysis of prognostic impact of urokinase-type plasminogen activator and its inhibitor PAI-1 in 8377 breast cancer patients. J. Natl Cancer Inst. 94, 116–128 (2002).
Jänicke, F. et al. Randomized adjuvant chemotherapy trial in high-risk, lymph node-negative breast cancer patients identified by urokinase-type plasminogen activator and plasminogen activator inhibitor type 1. J. Natl Cancer Inst. 93, 913–920 (2001).
O'Malley, F. P. et al. Topoisomerase II alpha and responsiveness of breast cancer to adjuvant chemotherapy. J. Natl Cancer Inst. 101, 644–650 (2009).
Colozza, M. et al. Proliferative markers as prognostic and predictive tools in early breast cancer: where are we now? Ann. Oncol. 16, 1723–1739 (2005).
Harris, L. N. et al. Topoisomerase IIα amplification does not predict benefit from dose-intense cyclophosphamide, doxorubicin, and fluorouracil therapy in HER2-amplified early breast cancer: results of CALGB 8541/150013. J. Clin. Oncol. 27, 3430–3436 (2009).
Claus, E. B., Schildkraut, J. M., Thompson, W. D. & Risch, N. J. The genetic attributable risk of breast and ovarian cancer. Cancer 77, 2318–2324 (1996).
Rennert, G. et al. Clinical outcomes of breast cancer in carriers of BRCA1 and BRCA2 mutations. N. Engl. J. Med. 357, 115–123 (2007).
Byrski, T. et al. Pathologic complete response rates in young women with BRCA1-positive breast cancers after neoadjuvant chemotherapy. J. Clin. Oncol. 28, 375–379 (2010).
Ellis, M. J. et al. Phosphatidyl-inositol-3-kinase alpha catalytic subunit mutation and response to neoadjuvant endocrine therapy for estrogen receptor positive breast cancer. Breast Cancer Res. Treat. 119, 379–390 (2010).
Pérez-Tenorio, G. et al. PIK3CA mutations and PTEN loss correlate with similar prognostic factors and are not mutually exclusive in breast cancer. Clin. Cancer Res. 13, 3577–3584 (2007).
Andre, F. & Delaloge, S. First-generation genomic tests for breast cancer treatment. Lancet Oncol. 11, 6–7 (2010).
van 't Veer, L. J. et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 31, 530–536 (2002).
van de Vijver, M. J. et al. A gene-expression signature as a predictor of survival in breast cancer. N. Engl. J. Med. 347, 1999–2009 (2002).
Buyse, M. et al. Validation and clinical utility of a 70-gene prognostic signature for women with node-negative breast cancer. J. Natl Cancer Inst. 98, 1183–1192 (2006).
Desmedt, C., Ruíz-García, E. & André, F. Gene expression predictors in breast cancer: current status, limitations and perspectives. Eur. J. Cancer 44, 2714–2720 (2008).
Paik, S. et al. Gene expression and benefit of chemotherapy in women with node-negative, estrogen receptor-positive breast cancer. J. Clin. Oncol. 24, 3726–3734 (2006).
Albain, K. S. et al. Prognostic and predictive value of the 21-gene recurrence score assay in postmenopausal women with node-positive, oestrogen-receptor-positive breast cancer on chemotherapy: a retrospective analysis of a randomised trial. Lancet Oncol. 11, 55–65 (2010).
Desmedt, C. et al. Strong time dependence of the 76-gene prognostic signature for node-negative breast cancer patients in the TRANSBIG multicenter independent validation series. Clin. Cancer Res. 13, 3207–3214 (2007).
Leary, A. F. et al. Value and limitations of measuring HER-2 extracellular domain in the serum of breast cancer patients. J. Clin. Oncol. 27, 1694–1705 (2009).
Lipton, A. et al. Elevated serum Her-2/neu level predicts decreased response to hormone therapy in metastatic breast cancer. J. Clin. Oncol. 20, 1467–1472 (2002).
Scaltriti, M. et al. Expression of p95HER2, a truncated form of the HER2 receptor, and response to anti-HER2 therapies in breast cancer. J. Natl Cancer Inst. 99, 628–638 (2007).
Scaltriti, M. et al. Clinical benefit of lapatinib-based therapy in patients with human epidermal growth factor receptor 2-positive breast tumors coexpressing the truncated p95HER2 receptor. Clin. Cancer Res. 16, 2688–2695 (2010).
André, F. et al. Multicenter phase I clinical trial of daily and weekly RAD001 in combination with weekly paclitaxel and trastuzumab in patients with HER2-overexpressing metastatic breast cancer with prior resistance to trastuzumab [abstract]. J. Clin. Oncol. 26 (Suppl.), a1003 (2008).
Nahta, R., Yu, D., Hung, M. C., Hortobagyi, G. N. & Esteva, F. J. Mechanisms of disease: understanding resistance to HER2-targeted therapy in human breast cancer. Nat. Clin. Pract. Oncol. 3, 269–280 (2006).
De Giorgi, U. et al. Circulating tumor cells and [18F]fluorodeoxyglucose positron emission tomography/computed tomography for outcome prediction in metastatic breast cancer. J. Clin. Oncol. 27, 3303–3311 (2009).
Fong, P. C. et al. Inhibition of poly(ADP-ribose) polymerase in tumors from BRCA mutation carriers. N. Engl. J. Med. 361, 123–134 (2009).
Zheng, Z. et al. DNA synthesis and repair genes RRM1 and ERCC1 in lung cancer. N. Engl. J. Med. 356, 800–808 (2007).
Olaussen, K. A. et al. DNA repair by ERCC1 in non-small-cell lung cancer and cisplatin-based adjuvant chemotherapy. N. Engl. J. Med. 355, 983–991 (2006).
Potti, A. et al. A genomic strategy to refine prognosis in early-stage non-small-cell lung cancer. N. Engl. J. Med. 355, 570–580 (2006).
Coate, L. E., John, T., Tsao, M. S. & Shepherd, F. A. Molecular predictive and prognostic markers in non-small-cell lung cancer. Lancet Oncol. 10, 1001–1010 (2009).
Sève, P., Reiman, T. & Dumont, C. The role of betaIII tubulin in predicting chemoresistance in non-small cell lung cancer. Lung Cancer 67, 136–143 (2010).
Chen, H. Y. et al. A five-gene signature and clinical outcome in non-small-cell lung cancer. N. Engl. J. Med. 356, 11–20 (2007).
Zhu, C. Q. et al. Role of KRAS and EGFR as biomarkers of response to erlotinib in National Cancer Institute of Canada Clinical Trials Group Study BR.21. J. Clin. Oncol. 26, 4268–4275 (2008).
Hirsch, F. R. et al. Molecular predictors of outcome with gefitinib in a phase III placebo-controlled study in advanced non-small-cell lung cancer. J. Clin. Oncol. 24, 5034–5042 (2006).
Varella-Garcia, M. et al. EGFR fluorescence in situ hybridisation assay: guidelines for application to non-small-cell lung cancer. J. Clin. Pathol. 62, 970–977 (2009).
Riely, G. J., Marks, J. & Pao, W. KRAS mutations in non-small cell lung cancer. Proc. Am. Thorac. Soc. 6, 201–205 (2009).
Linardou, H. et al. Assessment of somatic k-RAS mutations as a mechanism associated with resistance to EGFR-targeted agents: a systematic review and meta-analysis of studies in advanced non-small-cell lung cancer and metastatic colorectal cancer. Lancet Oncol. 9, 962–972 (2008).
Gomez-Roca, C. et al. Differential expression of biomarkers in primary non-small cell lung cancer and metastatic sites. J. Thorac. Oncol. 4, 1212–1220 (2009).
Cobo, M. et al. Customizing cisplatin based on quantitative excision repair cross-complementing 1 mRNA expression: a phase III trial in non-small-cell lung cancer. J. Clin. Oncol. 25, 2747–2754 (2007).
Rosell, R. et al. BRCA1: a novel prognostic factor in resected non-small-cell lung cancer. PLoS ONE 2, e1129 (2007).
Boukovinas, I. et al. Tumor BRCA1, RRM1 and RRM2 mRNA expression levels and clinical response to first-line gemcitabine plus docetaxel in non-small-cell lung cancer patients. PLoS ONE 3, e3695 (2008).
Soda, M. et al. Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature 448, 561–566 (2007).
Shaw, A. T. et al. Clinical features and outcome of patients with non-small-cell lung cancer who harbor EML4-ALK. J. Clin. Oncol. 27, 4247–4253 (2009).
Sanchez-Cespedes, M. et al. Inactivation of LKB1/STK11 is a common event in adenocarcinomas of the lung. Cancer Res. 62, 3659–3662 (2002).
Koivunen, J. P. et al. Mutations in the LKB1 tumour suppressor are frequently detected in tumours from Caucasian but not Asian lung cancer patients. Br. J. Cancer 99, 245–252 (2008).
Markowitz, S. D. & Bertagnolli, M. M. Molecular origins of cancer: molecular basis of colorectal cancer. N. Engl. J. Med. 361, 2449–2460 (2009).
Walther, A. et al. Genetic prognostic and predictive markers in colorectal cancer. Nat. Rev. Cancer 9, 489–499 (2009).
Wang, Y. et al. Gene expression profiles and molecular markers to predict recurrence of Dukes' B colon cancer. J. Clin. Oncol. 22, 1564–1571 (2004).
Garman, K. S. et al. A genomic approach to colon cancer risk stratification yields biologic insights into therapeutic opportunities. Proc. Natl Acad. Sci. USA 105, 19432–19437 (2008).
Kopetz, S. & Abbruzzese, J. L. Barriers to integrating gene profiling for stage II colon cancer. Clin. Cancer Res. 15, 7451–7452 (2009).
Ribic, C. M. et al. Tumor microsatellite-instability status as a predictor of benefit from fluorouracil-based adjuvant chemotherapy for colon cancer. N. Engl. J. Med. 349, 247–257 (2003).
Loriot, Y., Mordant, P., Deutsch, E., Olaussen, K. A. & Soria, J. C. Are RAS mutations predictive markers of resistance to standard chemotherapy? Nat. Rev. Clin. Oncol. 6, 528–534 (2009).
Eberhard, D. A. et al. Mutations in the epidermal growth factor receptor and in KRAS are predictive and prognostic indicators in patients with non-small-cell lung cancer treated with chemotherapy alone and in combination with erlotinib. J. Clin. Oncol. 23, 5900–5909 (2005).
Roth, A. D. et al. Prognostic role of KRAS and BRAF in stage II and III resected colon cancer: results of the translational study on the PETACC-3, EORTC 40993, SAKK 60–00 trial. J. Clin. Oncol. 28, 466–474 (2010).
Spano, J. P. et al. Epidermal growth factor receptor signaling in colorectal cancer: preclinical data and therapeutic perspectives. Ann. Oncol. 16, 189–194 (2005).
Tol, J., Nagtegaal, I. D. & Punt, C. J. BRAF mutation in metastatic colorectal cancer. N. Engl. J. Med. 361, 98–99 (2009).
Lengauer, C., Kinzler, K. W. & Vogelstein, B. Genetic instability in colorectal cancers. Nature 386, 623–627 (1997).
Ogino, S. et al. CpG island methylator phenotype, microsatellite instability, BRAF mutation and clinical outcome in colon cancer. Gut 58, 90–96 (2009).
Samowitz, W. S. et al. Association of smoking, CpG island methylator phenotype, and V600E BRAF mutations in colon cancer. J. Natl Cancer Inst. 98, 1731–1738 (2006).
Slattery, M. L. et al. Diet and lifestyle factor associations with CpG island methylator phenotype and BRAF mutations in colon cancer. Int. J. Cancer 120, 656–663 (2007).
Iacopetta, B. et al. Functional categories of TP53 mutation in colorectal cancer: results of an International Collaborative Study. Ann. Oncol. 17, 842–847 (2006).
Ogino, S. et al. Prognostic significance and molecular associations of 18q loss of heterozygosity: a cohort study of microsatellite stable colorectal cancers. J. Clin. Oncol. 27, 4591–4598 (2009).
Witte, J. S. Prostate cancer genomics: towards a new understanding. Nat. Rev. Genet. 10, 77–82 (2009).
Kattan, M. W., Wheeler, T. M. & Scardino, P. T. Postoperative nomogram for disease recurrence after radical prostatectomy for prostate cancer. J. Clin. Oncol. 17, 1499–1507 (1999).
Cooperberg, M. R., Broering, J. M. & Carroll, P. R. Risk assessment for prostate cancer metastasis and mortality at the time of diagnosis. J. Natl Cancer Inst. 101, 878–887 (2009).
Stephenson, A. J. et al. Postoperative nomogram predicting the 10-year probability of prostate cancer recurrence after radical prostatectomy. J. Clin. Oncol. 23, 7005–7012 (2005).
Zong, Y. et al. ETS family transcription factors collaborate with alternative signaling pathways to induce carcinoma from adult murine prostate cells. Proc. Natl Acad. Sci. USA 106, 12465–12470 (2009).
Kumar-Sinha, C., Tomlins, S. A. & Chinnaiyan, A. M. Recurrent gene fusions in prostate cancer. Nat. Rev. Cancer 8, 497–511 (2008).
Mosquera, J. M. et al. Prevalence of TMPRSS2-ERG fusion prostate cancer among men undergoing prostate biopsy in the United States. Clin. Cancer Res. 15, 4706–4711 (2009).
Yoshimoto, M. et al. Absence of TMPRSS2:ERG fusions and PTEN losses in prostate cancer is associated with a favorable outcome. Mod. Pathol. 21, 1451–1460 (2008).
Carver, B. S. et al. Aberrant ERG expression cooperates with loss of PTEN to promote cancer progression in the prostate. Nat. Genet. 41, 619–624 (2009).
Squire, J. A. TMPRSS2-ERG and PTEN loss in prostate cancer. Nat. Genet. 41, 509–510 (2009).
Febbo, P. G. Genomic approaches to outcome prediction in prostate cancer. Cancer 115 (Suppl. 13), 3046–3057 (2009).
Rosenbaum, E. et al. Promoter hypermethylation as an independent prognostic factor for relapse in patients with prostate cancer following radical prostatectomy. Clin. Cancer Res. 11, 8321–8325 (2005).
Richiardi, L. et al. Promoter methylation in, APC, RUNX3, and GSTP1 and mortality in prostate cancer patients. J. Clin. Oncol. 27, 3161–3168 (2009).
Taplin, M. E. et al. Androgen receptor mutations in androgen-independent prostate cancer: Cancer and Leukemia Group B Study 9663. J. Clin. Oncol. 21, 2673–2678 (2003).
Kattan, M. W. et al. The addition of interleukin-6 soluble receptor and transforming growth factor beta1 improves a preoperative nomogram for predicting biochemical progression in patients with clinically localized prostate cancer. J. Clin. Oncol. 21, 3573–3579 (2003).
Lopergolo, A. & Zaffaroni, N. Biomolecular markers of outcome prediction in prostate cancer. Cancer 115 (Suppl. 13), 3058–3067 (2009).
Varambally, S. et al. The polycomb group protein EZH2 is involved in progression of prostate cancer. Nature 419, 624–629 (2002).
Rhodes, D. R., Sanda, M. G., Otte, A. P., Chinnaiyan, A. M. & Rubin, M. A. Multiplex biomarker approach for determining risk of prostate-specific antigen-defined recurrence of prostate cancer. J. Natl Cancer Inst. 95, 661–668 (2003).
Khor, L. Y. et al. MDM2 as a predictor of prostate carcinoma outcome: an analysis of Radiation Therapy Oncology Group Protocol 8610. Cancer 104, 962–967 (2005).
Grignon, D. J. et al. p53 status and prognosis of locally advanced prostatic adenocarcinoma: a study based on RTOG 8610. J. Natl Cancer Inst. 89, 158–165 (1997).
Chakravarti, A. et al. Prognostic value of p16 in locally advanced prostate cancer: a study based on Radiation Therapy Oncology Group Protocol 9202. J. Clin. Oncol. 25, 3082–3089 (2007).
Edwards, J. et al. The role of HER1-HER4 and EGFRvIII in hormone-refractory prostate cancer. Clin. Cancer Res. 12, 123–130 (2006).
Lara, P. N. Jr et al. Trastuzumab plus docetaxel in HER-2/neu-positive prostate carcinoma: final results from the California Cancer Consortium Screening and Phase II Trial. Cancer 100, 2125–2131 (2004).
Domingo-Domenech, J. et al. Serum HER2 extracellular domain predicts an aggressive clinical outcome and biological PSA response in hormone-independent prostate cancer patients treated with docetaxel. Ann. Oncol. 19, 269–275 (2008).
Sircar, K. et al. PTEN genomic deletion is associated with p-Akt and AR signalling in poorer outcome, hormone refractory prostate cancer. J. Pathol. 218, 505–513 (2009).
Nakagawa, T. et al. A tissue biomarker panel predicting systemic progression after PSA recurrence post-definitive prostate cancer therapy. PLoS ONE 3, e2318 (2008).
Rajpar, S. et al. Urinary N-telopeptide (uNTx) is an independent prognostic factor for overall survival in patients with bone metastases from castration-resistant prostate cancer. Ann. Oncol. doi:10.1093/annonc/mdq037.
Fizazi, K. et al. Phase II trial of consolidation docetaxel and samarium-153 in patients with bone metastases from castration-resistant prostate cancer. J. Clin. Oncol. 27, 2429–2435 (2009).
de Bono, J. S. et al. Circulating tumor cells predict survival benefit from treatment in metastatic castration-resistant prostate cancer. Clin. Cancer Res. 14, 6302–6309 (2008).
Hruban, R. H. & Adsay, N. V. Molecular classification of neoplasms of the pancreas. Hum. Pathol. 40, 612–623 (2009).
Iacobuzio-Donahue, C. A. et al. DPC4 gene status of the primary carcinoma correlates with patterns of failure in patients with pancreatic cancer. J. Clin. Oncol. 27, 1806–1813 (2009).
Blackford, A. et al. SMAD4 gene mutations are associated with poor prognosis in pancreatic cancer. Clin. Cancer Res. 15, 4674–4679 (2009).
Lubin, M. & Lubin, A. Selective killing of tumors deficient in methylthioadenosine phosphorylase: a novel strategy. PLoS ONE 4, e5735 (2009).
Watanabe, I. et al. Advanced pancreatic ductal cancer: fibrotic focus and beta-catenin expression correlate with outcome. Pancreas 26, 326–333 (2003).
Kurman, R. J. & Shih, I. M. Pathogenesis of ovarian cancer: lessons from morphology and molecular biology and their clinical implications. Int. J. Gynecol. Pathol. 27, 151–160 (2008).
Bast, R. C. Jr, Hennessy, B. & Mills, G. B. The biology of ovarian cancer: new opportunities for translation. Nat. Rev. Cancer 9, 415–428 (2009).
Auner, V. et al. KRAS mutation analysis in ovarian samples using a high sensitivity biochip assay. BMC Cancer 9, 111 (2009).
Singer, G. et al. Mutations in BRAF and KRAS characterize the development of low-grade ovarian serous carcinoma. J. Natl Cancer Inst. 95, 484–486 (2003).
Willner, J. et al. Alternate molecular genetic pathways in ovarian carcinomas of common histological types. Hum. Pathol. 38, 607–613 (2007).
Gamallo, C. et al. Beta-catenin expression pattern in stage I and II ovarian carcinomas: relationship with beta-catenin gene mutations, clinicopathological features, and clinical outcome. Am. J. Pathol. 155, 527–536 (1999).
Madore, J. et al. Characterization of the molecular differences between ovarian endometrioid carcinoma and ovarian serous carcinoma. J. Pathol. 220, 392–400 (2010).
Kolasa, I. K. et al. PIK3CA amplification associates with resistance to chemotherapy in ovarian cancer patients. Cancer Biol. Ther. 8, 21–26 (2009).
Helleman, J. et al. Mismatch repair and treatment resistance in ovarian cancer. BMC Cancer 6, 201 (2006).
de Graeff, P. et al. Factors influencing p53 expression in ovarian cancer as a biomarker of clinical outcome in multicentre studies. Br. J. Cancer 95, 627–633 (2006).
Boyd, J. et al. Clinicopathologic features of BRCA-linked and sporadic ovarian cancer. JAMA 283, 2260–2265 (2000).
Tan, D. S. et al. “BRCAness” syndrome in ovarian cancer: a case-control study describing the clinical features and outcome of patients with epithelial ovarian cancer associated with BRCA1 and BRCA2 mutations. J. Clin. Oncol. 26, 5530–5536 (2008).
Lim, S. L. et al. Promoter hypermethylation of FANCF and outcome in advanced ovarian cancer. Br. J. Cancer 98, 1452–1456 (2008).
Potapova, A., Hoffman, A. M., Godwin, A. K., Al-Saleem, T. & Cairns, P. Promoter hypermethylation of the PALB2 susceptibility gene in inherited and sporadic breast and ovarian cancer. Cancer Res. 68, 998–1002 (2008).
Hanahan, D. & Weinberg, R. A. The hallmarks of cancer. Cell 100, 57–70 (2000).
Hayes, D. F. et al. Tumor marker utility grading system: a framework to evaluate clinical utility of tumor markers. J. Natl Cancer Inst. 88, 1456–1466 (1996).
McShane, L. M. et al. REporting recommendations for tumour MARKer prognostic studies (REMARK). Nat. Clin. Pract. Oncol. 2, 416–422 (2005).
Henry, N. L. & Hayes, D. F. Uses and abuses of tumor markers in the diagnosis, monitoring, and treatment of primary and metastatic breast cancer. Oncologist 11, 541–552 (2006).
Subramanian, J. & Simon, R. What should physicians look for in evaluating prognostic gene-expression signatures? Nat. Rev. Clin. Oncol. 7, 327–334 (2010).
Acknowledgements
The authors wish to thank C. Verjat for preparation of the figures, and C. Massard for useful comments and discussions of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Ferté, C., André, F. & Soria, JC. Molecular circuits of solid tumors: prognostic and predictive tools for bedside use. Nat Rev Clin Oncol 7, 367–380 (2010). https://doi.org/10.1038/nrclinonc.2010.84
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrclinonc.2010.84
This article is cited by
-
Novel immune-related gene signature for risk stratification and prognosis prediction in ovarian cancer
Journal of Ovarian Research (2023)
-
Transformable peptide nanoparticles arrest HER2 signalling and cause cancer cell death in vivo
Nature Nanotechnology (2020)
-
Blank spots on the map: some current questions on nuclear organization and genome architecture
Histochemistry and Cell Biology (2018)
-
UHRF1 overexpression is involved in cell proliferation and biochemical recurrence in prostate cancer after radical prostatectomy
Journal of Experimental & Clinical Cancer Research (2016)
-
INPP4B is an oncogenic regulator in human colon cancer
Oncogene (2016)