Abstract
We describe a general approach for refining protein structure models on the basis of cryo-electron microscopy maps with near-atomic resolution. The method integrates Monte Carlo sampling with local density-guided optimization, Rosetta all-atom refinement and real-space B-factor fitting. In tests on experimental maps of three different systems with 4.5-Å resolution or better, the method consistently produced models with atomic-level accuracy largely independently of starting-model quality, and it outperformed the molecular dynamics–based MDFF method. Cross-validated model quality statistics correlated with model accuracy over the three test systems.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Milazzo, A.C. et al. Initial evaluation of a direct detection device detector for single particle cryo-electron microscopy. J. Struct. Biol. 176, 404–408 (2011).
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013).
Cowtan, K. The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr. D Biol. Crystallogr. 62, 1002–1011 (2006).
Langer, G., Cohen, S.X., Lamzin, V.S. & Perrakis, A. Automated macromolecular model building for X-ray crystallography using ARP/wARP version 7. Nat. Protoc. 3, 1171–1179 (2008).
Terwilliger, T.C. et al. Iterative model building, structure refinement and density modification with the PHENIX AutoBuild wizard. Acta Crystallogr. D Biol. Crystallogr. 64, 61–69 (2008).
Tjioe, E., Lasker, K., Webb, B., Wolfson, H.J. & Sali, A. MultiFit: a web server for fitting multiple protein structures into their electron microscopy density map. Nucleic Acids Res. 39, W167–W170 (2011).
Woetzel, N., Lindert, S., Stewart, P.L. & Meiler, J. BCL::EM-Fit: rigid body fitting of atomic structures into density maps using geometric hashing and real space refinement. J. Struct. Biol. 175, 264–276 (2011).
Saha, M. & Morais, M.C. FOLD-EM: automated fold recognition in medium- and low-resolution (4–15 Å) electron density maps. Bioinformatics 28, 3265–3273 (2012).
Lindert, S. et al. EM-fold: de novo atomic-detail protein structure determination from medium-resolution density maps. Structure 20, 464–478 (2012).
Baker, M.L., Baker, M.R., Hryc, C.F., Ju, T. & Chiu, W. Gorgon and pathwalking: macromolecular modeling tools for subnanometer resolution density maps. Biopolymers 97, 655–668 (2012).
Topf, M., Baker, M.L., Marti-Renom, M.A., Chiu, W. & Sali, A. Refinement of protein structures by iterative comparative modeling and cryoEM density fitting. J. Mol. Biol. 357, 1655–1668 (2006).
DiMaio, F., Tyka, M.D., Baker, M.L., Chiu, W. & Baker, D. Refinement of protein structures into low-resolution density maps using Rosetta. J. Mol. Biol. 392, 181–190 (2009).
Trabuco, L.G., Villa, E., Mitra, K., Frank, J. & Schulten, K. Flexible fitting of atomic structures into electron microscopy maps using molecular dynamics. Structure 16, 673–683 (2008).
Song, Y. et al. High-resolution comparative modeling with RosettaCM. Structure 21, 1735–1742 (2013).
Berman, H.M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Egelman, E.H. et al. Structural plasticity of helical nanotubes based on coiled-coil assemblies. Structure 23, 280–289 (2015).
Eswar, N. et al. Comparative protein structure modeling using MODELLER. Curr. Protoc. Bioinformatics 15, 5.6 (2006).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Davis, I.W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
DiMaio, F. et al. Improved low-resolution crystallographic refinement with Phenix and Rosetta. Nat. Methods 10, 1102–1104 (2013).
Wang, R.Y.-R. et al. De novo protein structure determination from near-atomic-resolution cryo-EM maps. Nat. Methods doi:10.1038/nmeth.3287 (23 February 2015).
Fernández, J.J., Luque, D., Castón, J.R. & Carrascosa, J.L. Sharpening high resolution information in single particle electron cryomicroscopy. J. Struct. Biol. 164, 170–175 (2008).
Henderson, R. et al. Outcome of the first electron microscopy validation task force meeting. Structure 20, 205–214 (2012).
Scheres, S.H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Eswar, N. et al. Comparative protein structure modeling using MODELLER. Curr. Protoc. Protein. Sci. 50, 2.9 (2007).
Trabuco, L.G., Villa, E., Schreiner, E., Harrison, C.B. & Schulten, K. Molecular dynamics flexible fitting: a practical guide to combine cryo-electron microscopy and X-ray crystallography. Methods 49, 174–180 (2009).
Peng, L.-M., Ren, G., Dudarev, S.L. & Whelan, M.J. Robust parameterization of elastic and absorptive electron atomic scattering factors. Acta Crystallogr. A 52, 257–276 (1996).
Afonine, P.V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D Biol. Crystallogr. 68, 352–367 (2012).
Radics, J., Königsmaier, L. & Marlovits, T.C. Structure of a pathogenic type 3 secretion system in action. Nat. Struct. Mol. Biol. 21, 82–87 (2014).
Schraidt, O. & Marlovits, T.C. Three-dimensional model of Salmonella's needle complex at subnanometer resolution. Science 331, 1192–1195 (2011).
Mindell, J.A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003).
Yu, Y. et al. Interactions of PAN's C-termini with archaeal 20S proteasome and implications for the eukaryotic proteasome-ATPase interactions. EMBO J. 29, 692–702 (2010).
Roseman, A.M. FindEM—a fast, efficient program for automatic selection of particles from electron micrographs. J. Struct. Biol. 145, 91–99 (2004).
Li, X., Grigorieff, N. & Cheng, Y. GPU-enabled FREALIGN: accelerating single particle 3D reconstruction and refinement in Fourier space on graphics processors. J. Struct. Biol. 172, 407–412 (2010).
Rosenthal, P.B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Acknowledgements
The authors thank K. Laidig and D. Alonso for setting up and maintaining computational resources. This work was supported by the US National Institutes of Health (NIH) grants R01GM092802 (D.B.), R01GM082893 and R01GM098672 (Y.C.), and EB001567 (E.E.).
Author information
Authors and Affiliations
Contributions
F.D. and Y.S. developed the methods and ran experiments; F.D., Y.S. and D.B. wrote the manuscript. X.L. and Y.C. provided the 20S low-resolution data sets and provided feedback on the method. M.J.B. and T.C.M. analyzed the PrgH data set and provided feedback on the method. C.X., V.C. and E.E. collected the fiber data set and provided feedback on the method. All authors helped in editing the final manuscript.
Corresponding author
Ethics declarations
Competing interests
Y.S. is a cofounder of Cyrus Biotechnology, Inc., which will develop and market graphic-interface software for using Rosetta.
Integrated supplementary information
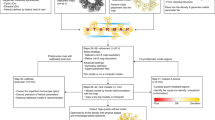
Supplementary Figure 1 Overview of the density-guided rebuilding and refinement protocol.
(a) Flowchart of the protocol. The protocol uses 250 cycles of local backbone rebuilding, followed by five cycles of alternating coordinate and B factor refinement. (b) An inset illustrating the new “density-guided rebuilding” step. A segment with poor fit to density is selected at random. Next, different backbone conformations consistent with the local sequence are first superimposed on the current model, then optimized into the map. Finally, the best fitting backbone conformation is selected, and backbone geometry near the segment is regularized.
Supplementary Figure 2 Model quality as a function of map resolution.
For a given starting model of 20S, we compare refined model accuracy (the fraction of Cα atoms within 1 Å of the reference model on the y-axis) as a function of map resolution for Rosetta-refined models (solid) and MDFF-refined models (dashed). The comparison used (A) 1yar, (B) 1ryp and (C) 1m4y as representatives of easy, medium, and difficult refinement cases, respectively.
Supplementary Figure 3 Identification of model errors using the local density correlation.
For each refined model of 20S, local correlation between each residue and the testing map is calculated. The fraction of residues deviate more than 1Å (black) or 2Å (green) from the reference model (y-axis) is plotted for each local correlation bin (x-axis) at (A) 3.3, (B) 4.1, (C) 4.4, and (D) 6.0Å resolutions.
Supplementary Figure 4 Cross-validation of Rosetta-refined models using FSC.
The FSC (y-axis) is compared to the accuracy of refined models (x-axis). Refinement is carried out with reconstructed maps of (A) 20S proteasome at (magenta) 3.3Å, (cyan) 4.1 Å, (red) 4.4 Å, (blue) 5.0 Å and (green) 6.0 Å, and with maps of (B) prgH needle complex at (red) 4.6 Å, (blue) 5.4 Å and (green) 7.1 Å.
Supplementary Figure 5 An overview of map features necessary for accurate structure determination.
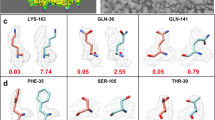
(a) The 20S proteasome dataset at 4.1 Å nicely illustrates the necessary features for structure determination to atomic accuracy. The pitch of helices and individual strands are resolved, and – while sidechains are not uniquely identifiable – density for large sidechains is visible. (b) The same dataset at 6 Å resolution lacks these features: the pitch of helices in not identifiable, and sidechain density for large sidechains is not observed.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Table 1 (PDF 2033 kb)
Rights and permissions
About this article
Cite this article
DiMaio, F., Song, Y., Li, X. et al. Atomic-accuracy models from 4.5-Å cryo-electron microscopy data with density-guided iterative local refinement. Nat Methods 12, 361–365 (2015). https://doi.org/10.1038/nmeth.3286
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3286
This article is cited by
-
Pathway and mechanism of tubulin folding mediated by TRiC/CCT along its ATPase cycle revealed using cryo-EM
Communications Biology (2023)
-
Cryo-EM structure-based selection of computed ligand poses enables design of MTA-synergic PRMT5 inhibitors of better potency
Communications Biology (2022)
-
Unique binding pattern for a lineage of human antibodies with broad reactivity against influenza A virus
Nature Communications (2022)
-
Structural analysis of cross α-helical nanotubes provides insight into the designability of filamentous peptide nanomaterials
Nature Communications (2021)
-
Mapping protein interactions in the active TOM-TIM23 supercomplex
Nature Communications (2021)