Abstract
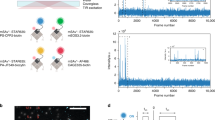
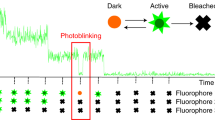
We describe a localization microscopy analysis method that is able to extract results in live cells using standard fluorescent proteins and xenon arc lamp illumination. Our Bayesian analysis of the blinking and bleaching (3B analysis) method models the entire dataset simultaneously as being generated by a number of fluorophores that may or may not be emitting light at any given time. The resulting technique allows many overlapping fluorophores in each frame and unifies the analysis of the localization from blinking and bleaching events. By modeling the entire dataset, we were able to use each reappearance of a fluorophore to improve the localization accuracy. The high performance of this technique allowed us to reveal the nanoscale dynamics of podosome formation and dissociation throughout an entire cell with a resolution of 50 nm on a 4-s timescale.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Heintzmann, R. & Ficz, G. Breaking the resolution limit in light microscopy. Methods Cell Biol. 81, 561–580 (2007).
Hell, S.W. Microscopy and its focal switch. Nat. Methods 6, 24–32 (2009).
Klar, T.A., Jakobs, S., Dyba, M., Egner, A. & Hell, S.W. Fluorescence microscopy with diffraction resolution barrier broken by stimulated emission. Proc. Natl. Acad. Sci. USA 97, 8206–8210 (2000).
Heintzmann, R., Jovin, T.M. & Cremer, C. Saturated patterned excitation microscopy—a concept for optical resolution improvement. J. Opt. Soc. Am. A 19, 1599–1609 (2002).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Rust, M.J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–796 (2006).
Wombacher, R. et al. Live-cell super-resolution imaging with trimethoprim conjugates. Nat. Methods 7, 717–719 (2010).
Shroff, H., Galbraith, C.G., Galbraith, J.A. & Betzig, E. Live-cell photoactivated localization microscopy of nanoscale adhesion dynamics. Nat. Methods 5, 417–423 (2008).
Westphal, V. et al. Video-rate far-field optical nanoscopy dissects synaptic vesicle movement. Science 320, 246–249 (2008).
Jones, S.A., Shim, S.-H., He, J. & Zhuang, X. Fast, three-dimensional super-resolution imaging of live cells. Nat. Methods 8, 499–505 (2011).
Kner, P., Chhun, B.B., Griffis, E.R., Winoto, L. & Gustafsson, M.G.L. Super-resolution video microscopy of live cells by structured illumination. Nat. Methods 6, 339–342 (2009).
Heilemann, M., Dedecker, P., Hofkens, J. & Sauer, M. Photoswitches: key molecules for subdiffraction-resolution fluorescence imaging and molecular quantification. Laser Photonics Rev. 3, 180–202 (2009).
Zhuang, X. Nano-imaging with STORM. Nat. Photonics 3, 365–367 (2009).
Heilemann, M., van de Linde, S., Mukherjee, A. & Sauer, M. Super-resolution imaging with small organic fluorophores. Angew. Chem. Int. Ed. 48, 6903–6908 (2009).
Lippincott-Schwartz, J. & Manley, S. Putting super-resolution fluorescence microscopy to work. Nat. Methods 6, 21–23 (2009).
Dertinger, T., Colyera, R., Iyera, G., Weissa, S. & Enderleind, J. Fast, background-free, 3D super-resolution optical fluctuation imaging (SOFI). Proc. Natl. Acad. Sci. USA 106, 22287–22292 (2009).
Dertinger, T., Heilemann, M., Vogel, R., Sauer, M. & Weiss, S. Superresolution optical fluctuation imaging with organic dyes. Angew. Chem. 122, 9631–9633 (2010).
Holden, S., Uphoff, S. & Kapanidis, A. DaoSTORM: an algorithm for high-density super-resolution microscopy. Nat. Methods 8, 279–280 (2011).
Huang, F., Schwartz, S.L., Byars, J.M. & Lidke, K.A. Simultaneous multiple-emitter fitting for single molecule super-resolution imaging. Biomedical Optics Express 2, 1377–1393 (2011).
Linder, S. & Aepfelbacher, M. Podosomes: adhesion hot-spots of invasive cells. Trends Cell Biol. 13, 376–385 (2003).
Linder, S. & Kopp, P. Podosomes at a glance. J. Cell Sci. 118, 2079–2082 (2005).
Monypenny, J. et al. Role of WASP in cell polarity and podosome dynamics of myeloid cells. Eur. J. Cell Biol. 90, 198–204 (2011).
Ghahramani, Z. & Jordan, M.I. Factorial hidden Markov models. Mach. Learn. 29, 245–273 (1997).
Rabiner, L.R. A tutorial on hidden Markov models and selected applications in speech recognition. Proc. IEEE 77, 257–286 (1989).
Wiesner, C., Faix, J., Himmel, M., Bentzien, F. & Linder, S. KIF5B and KIF3A/KIF3B kinesins drive MT1-MMP surface exposure, CD44 shedding, and extracellular matrix degradation in primary macrophages. Blood 116, 1559–1569 (2010).
Burnette, D.T. et al. A role for actin arcs in the leading-edge advance of migrating cells. Nat. Cell Biol. 13, 371–382 (2011).
Xie, X.S. Optical studies of single molecules at room temperature. Annu. Rev. Phys. Chem. 49, 441–480 (1998).
MacKay, D.J.C. Information Theory, Inference, and Learning Algorithms (Cambridge Univ. Press, 2003).
Acknowledgements
We acknowledge helpful discussions with A. Fraser, F. Viola, P. Fox-Roberts, O. Mandula and J. Sleep. We thank M. Kielhorn for assistance in aligning the optical system and K. Glocer for critical reading of the manuscript. We thank M. Parsons (King's College London) for providing the template plasmid. We acknowledge support from the EU Seventh Framework Programme (FP7 Project GA 215597 (S.C. and R.H.), EU FP7 Project ITN 237946 T3Net (G.E.J.)), the Wellcome Trust (S.C., J.M. and G.E.J.), the Medical Research Council (UK) (G.E.J.) and the Royal Society (S.C.).
Author information
Authors and Affiliations
Contributions
S.C., J.M., T.J.–T., D.T.B., J.L.-S., G.E.J. and R.H. conceived of and designed the experiments. S.C. and E.R. conceived of and designed the analysis. J.M. prepared the podosome samples, and T.J.–T. and D.T.B. prepared the samples for correlative measurements. S.C. and J.M. performed live-cell experiments, S.C. carried out fixed-cell experiments on podosomes, and T.J.–T. and D.T.B. carried out the correlative measurements. E.R. and S.C. carried out the data analysis and wrote the manuscript, and all authors revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Note (PDF 425 kb)
Supplementary Video 1
Raw data video of vinculin in fixed podosome samples. Samples were labeled with Alexa 488 and mounted in PBS with 100 mM mercaptoethanol added as a reducing agent to induce blinking. Sample was illuminated using a laser at 488nm with a nominal power of 1 kW/cm2. A series of 300 images were collected, taken at 50 frames per second. Scalebar is 500 nm. (MOV 766 kb)
Supplementary Video 2
Raw data video of live THP-1 cells stably expressing an mCherry tagged, truncated talin construct. Sample was illuminated with a Xenon arc lamp in the wavelength range 615-687 nm. Video shows the first 500 of 8,000 images taken at frame rates of 50 fps. Scalebar is 2 μm. (MOV 2552 kb)
Supplementary Video 3
Widefield (left) and 3B (right) video of a podosome being dissociated by unwinding. A truncated talin construct is labeled. This video corresponds to Figure 3a in the main text. Each widefield and 3B image is generated from 200 frames (4 seconds) of raw data, and are spaced 50 frames (1 second) apart. Widefield images are created by averaging. Cells were maintained at 37C during imaging. Scalebar is 1 μm. (MOV 406 kb)
Supplementary Video 4
Widefield (left) and 3B (right) video of a podosome being dissociate by being drawn into its center. A truncated talin construct is labeled. This video corresponds to Figure 3b in the main text. Each widefield and 3B image is generated from 200 frames (4 s) of raw data, and are spaced 50 frames (1 s) apart. Widefield images are created by averaging. Cells were maintained at 37C during imaging. Scalebar is 1 μm. (MOV 577 kb)
Supplementary Video 5
Widefield (left) and 3B (right) video of podosomes being constructed. A truncated talin construct is labeled. This video corresponds to Figure 3c in the main text. Each widefield and 3B image is generated from 200 frames (4 seconds) of raw data, and are spaced 50 frames (1 second) apart. Widefield images are created by averaging. Cells were maintained at 37C during imaging. Scalebar is 1 μm. (MOV 975 kb)
Supplementary Video 6
Widefield (left) and 3B (right) video of a steady state podosome in which a truncated talin construct is labeled. This video corresponds to Figure 3d in the main text. Each widefield and 3B image is generated from 200 frames (4 seconds) of raw data, and are spaced 50 frames (1 second) apart. Widefield images are created by averaging. Cells were maintained at 37C during imaging. Scalebar is 500 nm. (MOV 343 kb)
Supplementary Video 7
Widefield (left) and 3B (right) video reveals podosomes in a motile cell to be highly dynamic. A truncated talin construct is labeled in these cells. This video includes, as a small part, the areas shown in Figure 4a–c in the main text. Each widefield and 3B image is generated from 200 frames (4 seconds) of raw data, and are spaced 100 frames (2 s) apart. Widefield images are created by averaging. Cells were maintained at 37C during imaging. Scalebar is 2 μm. (MOV 5126 kb)
Supplementary Software
3B analysis software. Contains source code, test data and instructions for use. (ZIP 1681 kb)
Rights and permissions
About this article
Cite this article
Cox, S., Rosten, E., Monypenny, J. et al. Bayesian localization microscopy reveals nanoscale podosome dynamics. Nat Methods 9, 195–200 (2012). https://doi.org/10.1038/nmeth.1812
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1812
This article is cited by
-
Spectroscopic single-molecule localization microscopy: applications and prospective
Nano Convergence (2023)
-
High-fidelity 3D live-cell nanoscopy through data-driven enhanced super-resolution radial fluctuation
Nature Methods (2023)
-
Single-frame deep-learning super-resolution microscopy for intracellular dynamics imaging
Nature Communications (2023)
-
Extending resolution within a single imaging frame
Nature Communications (2022)
-
Simultaneous spatiotemporal super-resolution and multi-parametric fluorescence microscopy
Nature Communications (2021)