Abstract
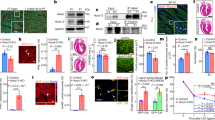
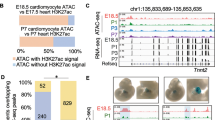
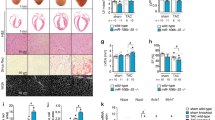
Adult-onset diseases can be associated with in utero events, but mechanisms for this remain unknown1,2. The Polycomb histone methyltransferase Ezh2 stabilizes transcription by depositing repressive marks during development that persist into adulthood3,4,5,6,7,8,9, but its function in postnatal organ homeostasis is unknown. We show that Ezh2 stabilizes cardiac gene expression and prevents cardiac pathology by repressing the homeodomain transcription factor gene Six1, which functions in cardiac progenitor cells but is stably silenced upon cardiac differentiation10. Deletion of Ezh2 in cardiac progenitors caused postnatal myocardial pathology and destabilized cardiac gene expression with activation of Six1-dependent skeletal muscle genes. Six1 induced cardiomyocyte hypertrophy and skeletal muscle gene expression. Furthermore, genetically reducing Six1 levels rescued the pathology of Ezh2-deficient hearts. Thus, Ezh2-mediated repression of Six1 in differentiating cardiac progenitors is essential for stable gene expression and homeostasis in the postnatal heart. Our results suggest that epigenetic dysregulation in embryonic progenitor cells is a predisposing factor for adult disease and dysregulated stress responses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Gluckman, P.D., Hanson, M.A., Buklijas, T., Low, F.M. & Beedle, A.S. Epigenetic mechanisms that underpin metabolic and cardiovascular diseases. Nat. Rev. Endocrinol. 5, 401–408 (2009).
Petronis, A. Epigenetics as a unifying principle in the aetiology of complex traits and diseases. Nature 465, 721–727 (2010).
Surface, L.E., Thornton, S.R. & Boyer, L.A. Polycomb group proteins set the stage for early lineage commitment. Cell Stem Cell 7, 288–298 (2010).
Margueron, R. & Reinberg, D. Chromatin structure and the inheritance of epigenetic information. Nat. Rev. Genet. 11, 285–296 (2010).
Margueron, R. & Reinberg, D. The Polycomb complex PRC2 and its mark in life. Nature 469, 343–349 (2011).
Su, I.H. et al. Ezh2 controls B cell development through histone H3 methylation and Igh rearrangement. Nat. Immunol. 4, 124–131 (2003).
Pereira, J.D. et al. Ezh2, the histone methyltransferase of PRC2, regulates the balance between self-renewal and differentiation in the cerebral cortex. Proc. Natl. Acad. Sci. USA 107, 15957–15962 (2010).
Xu, C.R. et al. Chromatin “prepattern” and histone modifiers in a fate choice for liver and pancreas. Science 332, 963–966 (2011).
Juan, A.H. et al. Polycomb EZH2 controls self-renewal and safeguards the transcriptional identity of skeletal muscle stem cells. Genes Dev. 25, 789–794 (2011).
Guo, C. et al. A Tbx1-Six1/Eya1-Fgf8 genetic pathway controls mammalian cardiovascular and craniofacial morphogenesis. J. Clin. Invest. 121, 1585–1595 (2011).
Heineke, J. & Molkentin, J.D. Regulation of cardiac hypertrophy by intracellular signalling pathways. Nat. Rev. Mol. Cell Biol. 7, 589–600 (2006).
Ahmad, F., Seidman, J.G. & Seidman, C.E. The genetic basis for cardiac remodeling. Annu. Rev. Genomics Hum. Genet. 6, 185–216 (2005).
Bergmann, O. et al. Evidence for cardiomyocyte renewal in humans. Science 324, 98–102 (2009).
Margueron, R. et al. Ezh1 and Ezh2 maintain repressive chromatin through different mechanisms. Mol. Cell 32, 503–518 (2008).
Shen, X. et al. EZH1 mediates methylation on histone H3 lysine 27 and complements EZH2 in maintaining stem cell identity and executing pluripotency. Mol. Cell 32, 491–502 (2008).
Verzi, M.P., McCulley, D.J., De Val, S., Dodou, E. & Black, B.L. The right ventricle, outflow tract, and ventricular septum comprise a restricted expression domain within the secondary/anterior heart field. Dev. Biol. 287, 134–145 (2005).
Weiford, B.C., Subbarao, V.D. & Mulhern, K.M. Noncompaction of the ventricular myocardium. Circulation 109, 2965–2971 (2004).
Swynghedauw, B. Phenotypic plasticity of adult myocardium: molecular mechanisms. J. Exp. Biol. 209, 2320–2327 (2006).
Houweling, A.C., van Borren, M.M., Moorman, A.F. & Christoffels, V.M. Expression and regulation of the atrial natriuretic factor encoding gene Nppa during development and disease. Cardiovasc. Res. 67, 583–593 (2005).
McFadden, D.G. et al. A GATA-dependent right ventricular enhancer controls dHAND transcription in the developing heart. Development 127, 5331–5341 (2000).
Takeuchi, J.K. et al. Chromatin remodeling complex dosage modulates transcription factor function in heart development. Nat. Commun. 2, 187 (2011).
Kumar, J.P. The sine oculis homeobox (SIX) family of transcription factors as regulators of development and disease. Cell. Mol. Life Sci. 66, 565–583 (2009).
Li, X. et al. Eya protein phosphatase activity regulates Six1-Dach-Eya transcriptional effects in mammalian organogenesis. Nature 426, 247–254 (2003).
Schönberger, J. et al. Mutation in the transcriptional coactivator EYA4 causes dilated cardiomyopathy and sensorineural hearing loss. Nat. Genet. 37, 418–422 (2005).
Liu, Y., Chu, A., Chakroun, I., Islam, U. & Blais, A. Cooperation between myogenic regulatory factors and SIX family transcription factors is important for myoblast differentiation. Nucleic Acids Res. 38, 6857–6871 (2010).
Christodoulou, D.C., Gorham, J.M., Herman, D.S. & Seidman, J.G. Construction of normalized RNA-seq libraries for next-generation sequencing using the crab duplex–specific nuclease. Curr. Protoc. Mol. Biol. 94, 4.12.1–4.12.11 (2011).
Acknowledgements
We thank A. Blais for Six1 antiserum, T. Sukonnik for in situ hybridization, J. Wythe for mT/mG embryos, J. Wylie for technical assistance, D. Miguel-Perez for animal husbandry, A. Holloway and A. Williams (Gladstone Bioinformatics Core) for RNA-seq and microarray data analysis, L. Ta (Gladstone Genomics Core) for microarray experiments, J. Zhang (Gladstone Transgenic Core) for oocyte injection, C. Miller (Gladstone Histology Core) for histology, Bruneau lab members for helpful discussions and G. Howard for editorial assistance. This work was supported by a fellowship from the California Institute for Regenerative Medicine (P.D.-O.), US National Institutes of Health grants U01HL098179 (B.G.B.) and U01HL098166 (C.E.S. and J.G.S.), by the Lawrence J. and Florence A. DeGeorge Charitable Trust/American Heart Association Established Investigator Award (B.G.B.) and by William H. Younger, Jr. (B.G.B.). Animal work was supported by a US National Institutes of Health/National Center for Research Resources grant (C06 RR018928) to the J. David Gladstone Institutes.
Author information
Authors and Affiliations
Contributions
P.D.-O. designed the study with B.G.B. and performed most of the experiments. Y.H. performed echocardiography. X.L. provided essential information before publication. X.L. and A.T. provided mouse lines. D.C. generated and analyzed RNA-eq data with C.E.S. and J.G.S. P.D.-O. and B.G.B. wrote the paper with input from all the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1 and 4 and Supplementary Figures 1–14 (PDF 14815 kb)
Supplementary Table 2
RNA-seq data obtained from right ventricular myocardium from Ezh2-deficient hearts (XLSX 1825 kb)
Supplementary Table 3
Signaling pathways in which Six1 target genes missregulated in Ezh2-deficient hearts participate (XLSX 13 kb)
Rights and permissions
About this article
Cite this article
Delgado-Olguín, P., Huang, Y., Li, X. et al. Epigenetic repression of cardiac progenitor gene expression by Ezh2 is required for postnatal cardiac homeostasis. Nat Genet 44, 343–347 (2012). https://doi.org/10.1038/ng.1068
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.1068
This article is cited by
-
Hallmarks of cardiovascular ageing
Nature Reviews Cardiology (2023)
-
KDM8 epigenetically controls cardiac metabolism to prevent initiation of dilated cardiomyopathy
Nature Cardiovascular Research (2023)
-
Inhibition of the cardiac fibroblast-enriched histone methyltransferase Dot1L prevents cardiac fibrosis and cardiac dysfunction
Cell & Bioscience (2022)
-
The function of LncRNA-H19 in cardiac hypertrophy
Cell & Bioscience (2021)
-
Compound heterozygous mutation of the ASXL3 gene causes autosomal recessive congenital heart disease
Human Genetics (2021)