Abstract
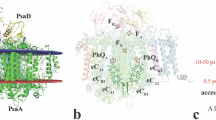
Photosystem II (PSII) is a huge membrane-protein complex consisting of 20 different subunits with a total molecular mass of 350 kDa for a monomer. It catalyses light-driven water oxidation at its catalytic centre, the oxygen-evolving complex (OEC)1,2,3. The structure of PSII has been analysed at 1.9 Å resolution by synchrotron radiation X-rays, which revealed that the OEC is a Mn4CaO5 cluster organized in an asymmetric, ‘distorted-chair’ form4. This structure was further analysed with femtosecond X-ray free electron lasers (XFEL), providing the ‘radiation damage-free’5 structure. The mechanism of O=O bond formation, however, remains obscure owing to the lack of intermediate-state structures. Here we describe the structural changes in PSII induced by two-flash illumination at room temperature at a resolution of 2.35 Å using time-resolved serial femtosecond crystallography with an XFEL provided by the SPring-8 ångström compact free-electron laser. An isomorphous difference Fourier map between the two-flash and dark-adapted states revealed two areas of apparent changes: around the QB/non-haem iron and the Mn4CaO5 cluster. The changes around the QB/non-haem iron region reflected the electron and proton transfers induced by the two-flash illumination. In the region around the OEC, a water molecule located 3.5 Å from the Mn4CaO5 cluster disappeared from the map upon two-flash illumination. This reduced the distance between another water molecule and the oxygen atom O4, suggesting that proton transfer also occurred. Importantly, the two-flash-minus-dark isomorphous difference Fourier map showed an apparent positive peak around O5, a unique μ4-oxo-bridge located in the quasi-centre of Mn1 and Mn4 (refs 4,5). This suggests the insertion of a new oxygen atom (O6) close to O5, providing an O=O distance of 1.5 Å between these two oxygen atoms. This provides a mechanism for the O=O bond formation consistent with that proposed previously6,7.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Shen, J.-R. The structure of photosystem II and the mechanism of water oxidation in photosynthesis. Annu. Rev. Plant Biol. 66, 23–48 (2015)
Yano, J. & Yachandra, V. Mn4Ca cluster in photosynthesis: where and how water is oxidized to dioxygen. Chem. Rev. 114, 4175–4205 (2014)
Najafpour, M. M. et al. Manganese compounds as water-oxidizing catalysts: From the natural water-oxidizing complex to nanosized manganese oxide structures. Chem. Rev. 116, 2886–2936 (2016)
Umena, Y., Kawakami, K., Shen, J.-R. & Kamiya, N. Crystal structure of oxygen-evolving photosystem II at a resolution of 1.9 Å. Nature 473, 55–60 (2011)
Suga, M. et al. Native structure of photosystem II at 1.95 Å resolution viewed by femtosecond X-ray pulses. Nature 517, 99–103 (2015)
Siegbahn, P. E. A structure-consistent mechanism for dioxygen formation in photosystem II. Chemistry 14, 8290–8302 (2008)
Siegbahn, P. E. Structures and energetics for O2 formation in photosystem II. Acc. Chem. Res. 42, 1871–1880 (2009)
Kok, B., Forbush, B. & McGloin, M. Cooperation of charges in photosynthetic O2 evolution-I. A linear four step mechanism. Photochem. Photobiol. 11, 457–475 (1970)
Kern, J. et al. Simultaneous femtosecond X-ray spectroscopy and diffraction of photosystem II at room temperature. Science 340, 491–495 (2013)
Kupitz, C. et al. Serial time-resolved crystallography of photosystem II using a femtosecond X-ray laser. Nature 513, 261–265 (2014)
Kern, J. et al. Taking snapshots of photosynthetic water oxidation using femtosecond X-ray diffraction and spectroscopy. Nat. Commun. 5, 4371 (2014)
Young, I. D. et al. Structure of photosystem II and substrate binding at room temperature. Nature 540, 453–457 (2016)
Sugahara, M. et al. Grease matrix as a versatile carrier of proteins for serial crystallography. Nat. Methods 12, 61–63 (2015)
Shevela, D., Eaton-Rye, J. J., Shen, J. R. & Govindjee . Photosystem II and the unique role of bicarbonate: a historical perspective. Biochim. Biophys. Acta 1817, 1134–1151 (2012)
Rapatskiy, L. et al. Detection of the water-binding sites of the oxygen-evolving complex of Photosystem II using W-band 17O electron-electron double resonance-detected NMR spectroscopy. J. Am. Chem. Soc. 134, 16619–16634 (2012)
Yamanaka, S. et al. Possible mechanisms for the O-O bond formation in oxygen evolution reaction at the CaMn4O5(H2O)(4) cluster of PSII refined to 1.9 Å X-ray resolution. Chem. Phys. Lett. 511, 138–145 (2011)
Saito, K., Rutherford, A. W. & Ishikita, H. Energetics of proton release on the first oxidation step in the water-oxidizing enzyme. Nat. Commun. 6, 8488 (2015)
Takaoka, T., Sakashita, N., Saito, K. & Ishikita, H. pKa of a proton-conducting water chain in photosystem II. J. Phys. Chem. Lett. 7, 1925–1932 (2016)
Isobe, H., Shoji, M., Shen, J. R. & Yamaguchi, K. Chemical equilibrium models for the S3 state of the oxygen-evolving complex of photosystem II. Inorg. Chem. 55, 502–511 (2016)
Siegbahn, P. E. Water oxidation mechanism in photosystem II, including oxidations, proton release pathways, O-O bond formation and O2 release. Biochim. Biophys. Acta 1827, 1003–1019 (2013)
Cox, N. et al. Photosynthesis. Electronic structure of the oxygen-evolving complex in photosystem II prior to O-O bond formation. Science 345, 804–808 (2014)
Pérez-Navarro, M., Neese, F., Lubitz, W., Pantazis, D. A. & Cox, N. Recent developments in biological water oxidation. Curr. Opin. Chem. Biol. 31, 113–119 (2016)
Shoji, M., Isobe, H. & Yamaguchi, K. QM/MM study of the S2 to S3 transition reaction in the oxygen-evolving complex of photosystem II. Chem. Phys. Lett. 636, 172–179 (2015)
Isobe, H. et al. Theoretical illumination of water-inserted structures of the CaMn4O5 cluster in the S2 and S3 states of oxygen-evolving complex of photosystem II: full geometry optimizations by B3LYP hybrid density functional. Dalton Trans. 41, 13727–13740 (2012)
Saito, K., Rutherford, A. W. & Ishikita, H. Mechanism of tyrosine D oxidation in Photosystem II. Proc. Natl Acad. Sci. USA 110, 7690–7695 (2013)
Nakamura, S. & Noguchi, T. Infrared detection of a proton released from tyrosine YD to the bulk upon its photo-oxidation in photosystem II. Biochemistry 54, 5045–5053 (2015)
Shen, J.-R. & Inoue, Y. Binding and functional properties of two new extrinsic components, cytochrome c-550 and a 12-kDa protein, in cyanobacterial photosystem II. Biochemistry 32, 1825–1832 (1993)
Shen, J.-R. & Kamiya, N. Crystallization and the crystal properties of the oxygen-evolving photosystem II from Synechococcus vulcanus. Biochemistry 39, 14739–14744 (2000)
Noguchi, T. & Sugiura, M. Flash-induced FTIR difference spectra of the water oxidizing complex in moderately hydrated photosystem II core films: effect of hydration extent on S-state transitions. Biochemistry 41, 2322–2330 (2002)
Suzuki, H., Sugiura, M. & Noguchi, T. pH dependence of the flash-induced S-state transitions in the oxygen-evolving center of photosystem II from Thermosynechoccocus elongatus as revealed by Fourier transform infrared spectroscopy. Biochemistry 44, 1708–1718 (2005)
Noguchi, T. & Sugiura, M. Flash-induced Fourier transform infrared detection of the structural changes during the S-state cycle of the oxygen-evolving complex in photosystem II. Biochemistry 40, 1497–1502 (2001)
Suzuki, H., Sugiura, M. & Noguchi, T. Monitoring proton release during photosynthetic water oxidation in photosystem II by means of isotope-edited infrared spectroscopy. J. Am. Chem. Soc. 131, 7849–7857 (2009)
Tono, K. et al. Diverse application platform for hard X-ray diffraction in SACLA (DAPHNIS): application to serial protein crystallography using an X-ray free-electron laser. J. Synchrotron Radiat. 22, 532–537 (2015)
Kameshima, T. et al. Development of an X-ray pixel detector with multi-port charge-coupled device for X-ray free-electron laser experiments. Rev. Sci. Instrum. 85, 033110 (2014)
Barty, A. et al. Cheetah: software for high-throughput reduction and analysis of serial femtosecond X-ray diffraction data. J. Appl. Crystallogr. 47, 1118–1131 (2014)
Nakane, T. et al. Data processing pipeline for serial femtosecond crystallography at SACLA. J. Appl. Crystallogr. 49, 1035–1041 (2016)
White, T. A. et al. CrystFEL: a software suite for snapshot serial crystallography. J. Appl. Crystallogr. 45, 335–341 (2012)
Duisenberg, A. J. M. Indexing in single-crystal diffractometry with an obstinate list of reflections. J. Appl. Crystallogr. 25, 92–96 (1992)
Battye, T. G. G., Kontogiannis, L., Johnson, O., Powell, H. R. & Leslie, A. G. W. iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr. D Biol. Crystallogr. 67, 271–281 (2011)
Kabsch, W. Processing of X-ray snapshots from crystals in random orientations. Acta Crystallogr. D Biol. Crystallogr. 70, 2204–2216 (2014)
Karplus, P. A. & Diederichs, K. Assessing and maximizing data quality in macromolecular crystallography. Curr. Opin. Struct. Biol. 34, 60–68 (2015)
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004)
Nango, E. et al. A three-dimensional movie of structural changes in bacteriorhodopsin. Science 354, 1552–1557 (2016)
Acknowledgements
We thank K. Kawakami and N. Kamiya for sharing unpublished information, H. Mino for information on the flash illumination conditions and H. Ago and K. Yamaguchi for discussions. This work was supported by a program for promoting the enhancement of research universities at Okayama University, JSPS KAKENHI Grant Nos JP15H01642, JP16H06162, JP16H06296 (M. Suga), JP16K21181 (F.A.), JP15H05588 (Y.U.), JP15H03841, JP15H01055 (M.K.) and JP24000018 (J.-R.S.), an X-ray Free Electron Laser Priority Strategy Program (J.-R.S., S.Iw.) from MEXT, Japan, an Asahi Glass Foundation (F.A.), a Kato Memorial Bioscience Foundation (F.A.), an Inamori Foundation (M. Suga), the Research Acceleration Program from Japan Science and Technology agency (JST) (S.Iw.), PRESTO from JST (M.K. and F.A.), a grant from Pioneering Project ‘Dynamic Structural Biology’ of RIKEN (M.K.), and the Strategic Priority Research Program of CAS (XDB17030100) (J.-R.S.). The XFEL experiments were performed at beamline 3 of SACLA with the approval of the Japan Synchrotron Radiation Research Institute (JASRI) (proposal nos 2013B1259, 2014A1243, 2014A6927, 2014B1281, 2014B6927, 2014B8048, 2015A1108, 2015A6522, 2015B2108, 2015B6522, 2015B8044, 2016A2542, 2016A6621 and 2016A8033), and we thank the staff at SACLA for their help. We also acknowledge computational support from the SACLA HPC system and the Mini-K supercomputer system.
Author information
Authors and Affiliations
Contributions
J.-R.S. conceived the project; M. Suga, F.A., M.Sugah., M.K., Y.U. and J.-R.S. contributed to the design of the experiment; F.A., Y.N., M.N. and T. Nakano grew the cells and purified the PSII samples; F.A. and Y.N. prepared the PSII crystals; M.Sugah., E.N. and R.T. developed the sample delivery system; F.A., Y.N., M.N., Y.U. and M.Sugah. tested and optimized buffer and crystal suspension conditions for injection; M.Suz., T.Ma., S.In., T.Mo., J.-H.C., H.N., Y.M. and A.Y. operated the injector; K.T., Y.J., T.Ka., T.H., M.Yab., T.I. and S.Iw. developed the diffraction instrumentation; M.K., T.Ki. and T.Nom. designed and optimized the laser excitation scheme and aligned the lasers; M. Suga, F.A., M.Sugah., Y.N., T. Nakane, K.Y., M.N., Y.U., M.Suz., T.Ma., S.In., S.Y., L.-J.Y., T.Mo., J.-H.C., R.T., H.N., Y.M., A.Y. and J.-R.S. participated in collection of the X-ray diffraction data at SACLA; T. Nakane, K.Y., M.Yam., O.N. and S.Iw. developed the data evaluation and/or hit-finding programs; Y.K., F.A. and T.Nog. performed FTIR analysis; M. Suga, T.Y. and T.S. analysed the femtosecond crystallography X-ray diffraction data; M. Suga refined the structure, calculated the electron density maps and made the figures; M. Suga and J.-R.S. wrote the manuscript, and all the authors participated in the discussion of the results and writing of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks R. Debus, J. Murray and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Micro-sized crystals of PSII, its diffraction image and quality of the diffraction data sets.
a, Micro-sized crystals of PSII used for TR-SFX. b, A diffraction image from a micro-sized PSII crystal obtained with a single pulse from SACLA-XFEL. c–f, Quality of the data sets. CC1/2 (c), Rsplit (d), multiplicity (e) and <I/σ(I)> (f) plotted against resolution for the dark data and the 2F data of the pre-flashed samples.
Extended Data Figure 2 Measurement of the S3 state population upon 2F illumination by FTIR.
Top, ATR–FTIR difference spectra upon the first (a) and second (b) flashes of the PSII crystal (red lines) and PSII solution (black lines). Bottom, result of the least-squares fitting analysis of the second flash spectrum of the PSII crystal in the symmetric COO− region (1,470–1,300 cm−1). The spectrum of the PSII crystal (red line) was fit with a linear combination of the first and second flash spectra of the solution sample, F1(ν) and F2(ν), respectively (black lines in the top panel). The resulting fitting spectrum (grey line) and the two components, 0.27F1(ν) (blue line) and 0.64F2(ν) (green line), are also shown.
Extended Data Figure 3 Anomalous signals obtained with the XFEL beam.
a–c, Anomalous difference Fourier maps for OEC atoms (a), a region around the OEC (b), and a newly identified Ca-binding site in CP43 (c), of the non-pre-flashed dark data. The maps are shown in cyan and magenta contoured at 4σ and 12σ, respectively (a), grey contoured at 3σ (b) and cyan contoured at 4σ (c). d, Histogram analysis of the anomalous map of the non-pre-flashed dark data.
Extended Data Figure 4 Electron density maps and structural changes around YD between the non-pre-flashed 2F and dark data sets.
a, Isomorphous difference Fourier map between the non-pre-flashed 2F state and dark state data sets in green (positive) and red (negative) contoured at ±5σ, superimposed with the non-pre-flashed dark structure and 2F structure. The non-pre-flashed dark and 2F structures are coloured yellow and green, respectively. The two different positions of W508 observed in the 1.95 Å resolution XFEL structure (W508I and W508II) are shown as black dots. b, c, 2mFo − DFc map and mFo − DFc map of the non-pre-flashed dark state (b) and 2F state (c). The 2mFo − DFc map is coloured grey, contoured at 1.5σ, and the mFo − DFc map is colored cyan (positive) and brown (negative), contoured at ±4σ.
Extended Data Figure 5 Electron density maps for the O4–water chain.
The mFo − DFc maps calculated for the O4–water chain by omitting the four water molecules W567, W665, W542 and W546 are shown for the non-pre-flashed dark state (a, b) and 2F state (c, d). The maps shown are for monomer A (a, c) and monomer B (b, d) in cyan (positive) and orange (negative) contoured at ±3σ.
Extended Data Figure 6 Distribution of the unit cell parameters and crystal packing of PSII for the non-pre-flashed samples.
a–c, Distributions of unit cell parameters for the dark state images (a) and 2F state images (b) from the non-pre-flashed samples and the dark images when the post-crystallization procedure was not adequate (c) are shown with their average lengths and standard deviations. No large differences were observed between the dark images and the 2F images, whereas a longer c axis was observed in c. d–f, Comparison of the crystal packing between the dark state and the data set with a longer c axis is shown in a view direction perpendicular to the bc plane. Colour codes: PSII and the crystallographic symmetric molecules in the dark state, cyan and blue, respectively; PSII, PsbY subunits in PSII and the crystallographic symmetric molecules in the data set with a long c axis, grey, red and khaki, respectively. Note that PsbY has considerable steric hindrance with the adjacent PSII molecule in the crystal packing of the dark data (e).
Rights and permissions
About this article
Cite this article
Suga, M., Akita, F., Sugahara, M. et al. Light-induced structural changes and the site of O=O bond formation in PSII caught by XFEL. Nature 543, 131–135 (2017). https://doi.org/10.1038/nature21400
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature21400
This article is cited by
-
Dynamic three-dimensional structures of a metal–organic framework captured with femtosecond serial crystallography
Nature Chemistry (2024)
-
Current analysis of cations substitution in the oxygen-evolving complex of photosystem II
Biophysical Reviews (2024)
-
The S1 to S2 and S2 to S3 state transitions in plant photosystem II: relevance to the functional and structural heterogeneity of the water oxidizing complex
Photosynthesis Research (2024)
-
On the simulation and interpretation of substrate-water exchange experiments in photosynthetic water oxidation
Photosynthesis Research (2024)
-
Oxygen-evolving photosystem II structures during S1–S2–S3 transitions
Nature (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.