Abstract
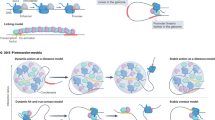
How a complex animal can arise from a fertilized egg is one of the oldest and most fascinating questions of biology, the answer to which is encoded in the genome. Body shape and organ development, and their integration into a functional organism all depend on the precise expression of genes in space and time. The orchestration of transcription relies mostly on surrounding control sequences such as enhancers, millions of which form complex regulatory landscapes in the non-coding genome. Recent research shows that high-order chromosome structures make an important contribution to enhancer functionality by triggering their physical interactions with target genes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Banerji, J., Rusconi, S. & Schaffner, W. Expression of a β-globin gene is enhanced by remote SV40 DNA sequences. Cell 27, 299–308 (1981).
Bulger, M. & Groudine, M. Functional and mechanistic diversity of distal transcription enhancers. Cell 144, 327–339 (2011).
Spitz, F. & Furlong, E. E. Transcription factors: from enhancer binding to developmental control. Nature Rev. Genet. 13, 613–626 (2012).
Dunham, I. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Stamatoyannopoulos, J. A. What does our genome encode? Genome Res. 22, 1602–1611 (2012).
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Kleinjan, D. A. & van Heyningen, V. Long-range control of gene expression: emerging mechanisms and disruption in disease. Am. J. Hum. Genet. 76, 8–32 (2005).
Weischenfeldt, J., Symmons, O., Spitz, F. & Korbel, J. O. Phenotypic impact of genomic structural variation: insights from and for human disease. Nature Rev. Genet. 14, 125–138 (2013).
Montavon, T. & Duboule, D. Landscapes and archipelagos: spatial organization of gene regulation in vertebrates. Trends Cell Biol. 22, 347–354 (2012).
Lagha, M., Bothma, J. P. & Levine, M. Mechanisms of transcriptional precision in animal development. Trends Genet. 28, 409–416 (2012).
Maeda, R. K. & Karch, F. Gene expression in time and space: additive vs hierarchical organization of cis-regulatory regions. Curr. Opin. Genet. Dev. 21, 187–193 (2011).
Levine, M. Transcriptional enhancers in animal development and evolution. Curr. Biol. 20, R754–R763 (2010).
Duboule, D. & Wilkins, A. S. The evolution of 'bricolage'. Trends Genet. 14, 54–59 (1998).
Kirschner, M. & Gerhart, J. Evolvability. Proc. Natl Acad. Sci. USA 95, 8420–8427 (1998).
Ohno, S. Evolution by Gene Duplication. (Springer, 1970).
Ruf, S. et al. Large-scale analysis of the regulatory architecture of the mouse genome with a transposon-associated sensor. Nature Genet. 43, 379–386 (2011). In this paper, the authors use a transposable reporter gene system in mice to probe for enhancer activity in vivo and show widely varying reporter expression patterns at hundreds of genomic integration sites.
O'Kane, C. J. & Gehring, W. J. Detection in situ of genomic regulatory elements in Drosophila. Proc. Natl Acad. Sci. USA 84, 9123–9127 (1987).
Pennacchio, L. A. et al. In vivo enhancer analysis of human conserved non-coding sequences. Nature 444, 499–502 (2006).
Odom, D. T. et al. Tissue-specific transcriptional regulation has diverged significantly between human and mouse. Nature Genet. 39, 730–732 (2007).
Wilson, M. D. et al. Species-specific transcription in mice carrying human chromosome 21. Science 322, 434–438 (2008).
Zinzen, R. P., Girardot, C., Gagneur, J., Braun, M. & Furlong, E. E. Combinatorial binding predicts spatio-temporal cis-regulatory activity. Nature 462, 65–70 (2009).
Lee, D., Karchin, R. & Beer, M. A. Discriminative prediction of mammalian enhancers from DNA sequence. Genome Res. 21, 2167–2180 (2011).
Gorkin, D. U. et al. Integration of ChIP-seq and machine learning reveals enhancers and a predictive regulatory sequence vocabulary in melanocytes. Genome Res. 22, 2290–2301 (2012).
Crawford, G. E. et al. DNase-chip: a high-resolution method to identify DNase I hypersensitive sites using tiled microarrays. Nature Methods 3, 503–509 (2006).
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 1 (2012).
Hardison, R. C. & Taylor, J. Genomic approaches towards finding cis-regulatory modules in animals. Nature Rev. Genet. 13, 469–483 (2012).
Arnold, C. D. et al. Genome-wide quantitative enhancer activity maps identified by STARR-seq. Science 339, 1074–1077 (2013). In this paper, the authors report a genome-wide screening method that quantifies the self-transcription of random DNA segments to identify sequence modules with enhancer activity.
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
de Wit, E. & de Laat, W. A decade of 3C technologies: insights into nuclear organization. Genes Dev. 26, 11–24 (2012).
Carter, D., Chakalova, L., Osborne, C. S., Dai, Y. F. & Fraser, P. Long-range chromatin regulatory interactions in vivo. Nature Genet. 32, 623–626 (2002).
Tolhuis, B., Palstra, R. J., Splinter, E., Grosveld, F. & de Laat, W. Looping and interaction between hypersensitive sites in the active β-globin locus. Mol. Cell 10, 1453–1465 (2002).
Palstra, R. J. et al. Maintenance of long-range DNA interactions after inhibition of ongoing RNA polymerase II transcription. PloS ONE 3, e1661 (2008).
Deng, W. et al. Controlling long-range genomic interactions at a native locus by targeted tethering of a looping factor. Cell 149, 1233–1244 (2012). In this study, it is shown that the artificial physical tethering of a distant enhancer to a gene promoter leads to highly activated transcription, illustrating that enhancer contacts causally underlie transcriptional output.
Noordermeer, D. et al. Variegated gene expression caused by cell-specific long-range DNA interactions. Nature Cell Biol. 13, 944–951 (2011). This article shows that inter-chromosomal gene activation by an ectopic enhancer leads to variegated gene expression in subsets of mouse cells.
Andrey, G. et al. A switch between topological domains underlies HoxD genes collinearity in mouse limbs. Science 340, 1234167 (2013). In this paper, the authors show that different regulatory landscapes are recruited during development by the same genes, which switch contacts from one to the other.
Palstra, R. J. et al. The β-globin nuclear compartment in development and erythroid differentiation. Nature Genet. 35, 190–194 (2003).
Vernimmen, D., De Gobbi, M., Sloane-Stanley, J. A., Wood, W. G. & Higgs, D. R. Long-range chromosomal interactions regulate the timing of the transition between poised and active gene expression. EMBO J. 26, 2041–2051 (2007).
Drissen, R. et al. The active spatial organization of the β-globin locus requires the transcription factor EKLF. Genes Dev. 18, 2485–2490 (2004).
Vakoc, C. R. et al. Proximity among distant regulatory elements at the β-globin locus requires GATA-1 and FOG-1. Mol. Cell 17, 453–462 (2004.12.028 (2005).
van de Werken, H. J. et al. Robust 4C-seq data analysis to screen for regulatory DNA interactions. Nature Methods 9, 969–972 (2012).
Riddle, R. D., Johnson, R. L., Laufer, E. & Tabin, C. Sonic hedgehog mediates the polarizing activity of the ZPA. Cell 75, 1401–1416 (1993).
Lettice, L. A. et al. A long-range Shh enhancer regulates expression in the developing limb and fin and is associated with preaxial polydactyly. Hum. Mol. Genet. 12, 1725–1735 (2003).
Sagai, T., Hosoya, M., Mizushina, Y., Tamura, M. & Shiroishi, T. Elimination of a long-range cis-regulatory module causes complete loss of limb-specific Shh expression and truncation of the mouse limb. Development 132, 797–803 (2005).
Amano, T. et al. Chromosomal dynamics at the Shh locus: limb bud-specific differential regulation of competence and active transcription. Dev. Cell 16, 47–57 (2009).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Lettice, L. A. et al. Opposing functions of the ETS factor family define Shh spatial expression in limb buds and underlie polydactyly. Dev. Cell 22, 459–467 (2012).
Eijkelenboom, A. et al. Genome-wide analysis of FOXO3 mediated transcription regulation through RNA polymerase II profiling. Mol. Syst. Biol. 9, 638 (2013).
Melo, C. A. et al. eRNAs are required for p53-dependent enhancer activity and gene transcription. Mol. Cell 49, 524–535 (2012).
Nora, E. P., Dekker, J. & Heard, E. Segmental folding of chromosomes: a basis for structural and regulatory chromosomal neighborhoods? BioEssays 5, 201300040 (2013).
Sanyal, A., Lajoie, B. R., Jain, G. & Dekker, J. The long-range interaction landscape of gene promoters. Nature 489, 109–113 (2012). This report is the first systematic analysis, based on 5C, of promoter-centred DNA contacts, revealing complex and dynamic contact networks, which change during development.
Li, G. et al. Extensive promoter-centered chromatin interactions provide a topological basis for transcription regulation. Cell 148, 84–98 (2012).
Splinter, E. et al. CTCF mediates long-range chromatin looping and local histone modification in the β-globin locus. Genes Dev. 20, 2349–2354 (2006).
Montavon, T. et al. A regulatory archipelago controls Hox genes transcription in digits. Cell 147, 1132–1145 (2011). In this paper, the authors report the existence of a regulatory archipelago that controls transcription in developing digits and contains several islands with various quantitative and qualitative contributions.
Bender, M. A. et al. Flanking HS-62.5 and 3-HS1, and regions upstream of the LCR, are not required for β-globin transcription. Blood 108, 1395–1401 (2006).
Noordermeer, D. et al. Transcription and chromatin organization of a housekeeping gene cluster containing an integrated β-globin locus control region. PLoS Genet. 4, e1000016 (2008).
Marinić, M., Aktas, T., Ruf, S. & Spitz, F. An integrated holo-enhancer unit defines tissue and gene specificity of the Fgf8 regulatory landscape. Dev. Cell 24, 530–542 (2013).
Cajiao, I., Zhang, A., Yoo, E. J., Cooke, N. E. & Liebhaber, S. A. Bystander gene activation by a locus control region. EMBO J. 23, 3854–3863 (2004).
Spitz, F., Gonzalez, F. & Duboule, D. A global control region defines a chromosomal regulatory landscape containing the HoxD cluster. Cell 113, 405–417 (2003).
Zuniga, A. et al. Mouse limb deformity mutations disrupt a global control region within the large regulatory landscape required for Gremlin expression. Genes Dev. 18, 1553–1564 (2004).
Lower, K. M. et al. Adventitious changes in long-range gene expression caused by polymorphic structural variation and promoter competition. Proc. Natl Acad. Sci. USA 106, 21771–21776 (2009).
Tschopp, P., Fraudeau, N., Bena, F. & Duboule, D. Reshuffling genomic landscapes to study the regulatory evolution of Hox gene clusters. Proc. Natl Acad. Sci. USA 108, 10632–10637 (2011).
De Gobbi, M. et al. A regulatory SNP causes a human genetic disease by creating a new transcriptional promoter. Science 312, 1215–1217 (2006).
Bender, M. A. et al. Targeted deletion of 5-HS1 and 5-HS4 of the β-globin locus control region reveals additive activity of the DNaseI hypersensitive sites. Blood 98, 2022–2027 (2001).
Wijgerde, M., Grosveld, F. & Fraser, P. Transcription complex stability and chromatin dynamics in vivo. Nature 377, 209–213 (1995).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Sexton, T. et al. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 148, 458–472 (2012). Refs 45, 65 and 66 report that mammalian and fly chromosomes are structurally subdivided into topological domains with flanking boundaries that hamper DNA contacts across.
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Phillips-Cremins, J. E. et al. Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell 153, 1281–1295 (2013).
Williams, A., Spilianakis, C. G. & Flavell, R. A. Interchromosomal association and gene regulation in trans. Trends Genet. 26, 188–197 (2010).
Fuss, S. H., Omura, M. & Mombaerts, P. Local and cis effects of the H element on expression of odorant receptor genes in mouse. Cell 130, 373–384 (2007).
Gaetz, J. et al. Evidence for a critical role of gene occlusion in cell fate restriction. Cell Res. 22, 848–858 (2012).
Miller, J. C. et al. A TALE nuclease architecture for efficient genome editing. Nature Biotechnol. 29, 143–148 (2011).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Kleinjan, D. J. & Coutinho, P. Cis-ruption mechanisms: disruption of cis-regulatory control as a cause of human genetic disease. Brief. Funct. Genomic Proteomic 8, 317–332 (2009).
Ovcharenko, I. et al. Evolution and functional classification of vertebrate gene deserts. Genome Res. 15, 137–145 (2005).
Ahituv, N. et al. Deletion of ultraconserved elements yields viable mice. PLoS Biol. 5, e234 (2007).
Acknowledgements
The authors thank Y. Oz for preparing the figures, and G. Geeven, M. Janssen, G. Andrey, L. Beccari, T. Montavon and M. Leleu for their help. The authors' laboratories are supported by funds from the Dutch Scientific Organization (NWO) (714012002 and 724012003 (VICI)), EU grant 2010-259743 (MODHEP), a KWF Dutch Cancer Foundation grant (2009-4459) and a NanoNextNL grant (to W.dL) and the Ecole Polytechnique Fédérale de Lausanne, the University of Geneva, the Swiss National Research Foundation, the ERC grant SystemsHox.ch and the FP7 program IDEAL (to D.D.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reprints and permissions information is available at www.nature.com/reprint.
Rights and permissions
About this article
Cite this article
de Laat, W., Duboule, D. Topology of mammalian developmental enhancers and their regulatory landscapes. Nature 502, 499–506 (2013). https://doi.org/10.1038/nature12753
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12753
This article is cited by
-
Enhancer–promoter interactions become more instructive in the transition from cell-fate specification to tissue differentiation
Nature Genetics (2024)
-
The spatial organization of transcriptional control
Nature Reviews Genetics (2023)
-
Identification of non-coding silencer elements and their regulation of gene expression
Nature Reviews Molecular Cell Biology (2023)
-
A Prox1 enhancer represses haematopoiesis in the lymphatic vasculature
Nature (2023)
-
Enhancer–promoter interactions can bypass CTCF-mediated boundaries and contribute to phenotypic robustness
Nature Genetics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.