Abstract
Effective population size (Ne) is a measure of genetic drift and is thus a central parameter in evolution, conservation genetics and invasion biology. Interestingly, in native marine species, Ne is typically several orders of magnitude lower than the census size. This pattern has often been explained by high fecundity, variation in reproductive success and pronounced early mortality, resulting in genetic drift across generations. Data documenting genetic drift and/or Ne in marine invasive species are, however, still scarce. We examined the importance of genetic drift in the invasive species Crepidula fornicata by genotyping 681 juveniles sampled during each annual recruitment peak over nine consecutive years in the Bay of Morlaix (Brittany, France). Observed variations in genetic diversity were partially explained by variation in recruitment intensity. In addition, substantial temporal genetic differentiation was documented (that is, genetic drift), and was attributed to nonrandom variance in the reproductive success of different breeding groups across years in the study species. Using a set of single-sample and temporal estimators for Ne, we estimated Ne to be three or four orders of magnitude smaller than the census size (Nc). On one hand, this reduction in Ne relative to Nc appeared congruent with, although slight higher than, values commonly observed in native marine species. Particular life-history traits of this invasive species may play an important role in buffering genetic drift. On the other hand, Ne still remained far below Nc, hence, possibly reducing the efficiency of selection effects.
Similar content being viewed by others
Introduction
Genetic drift influences genetic diversity, the efficiency of selection and the potential for adaptation to environmental change (Frankham, 1995; Berthier et al., 2002; Hare et al., 2011). Its intensity depends on the effective size of a population (Ne), the size of an ideal population in which genetic drift occurs at the same rate as in the study population (Wright, 1931). Although Ne has been frequently investigated in the context of conservation and fisheries management (for example, Chapman et al., 2002; Hauser et al., 2002; Hoarau et al., 2005), studies of temporal genetic changes and effective size have seldom been investigated in nonnative marine species, despite their importance for examining the relationship between founder events, invasiveness and evolutionary processes acting on marine invasive species following their introduction (Blanchet, 2012; Rius et al., 2015).
Estimates of Ne are typically lower than the population census size (Nc) (Frankham, 1995; Hauser et al., 2002; Palstra and Ruzzante, 2008). Many factors are likely to lower the Ne-to-Nc ratio, such as variance in reproductive success, uneven sex ratio and variation in population size over successive generations (Hedrick, 2000; Turner et al., 2006). Low Ne/Nc ratios are especially common in marine species, where Ne can be several orders of magnitude smaller than Nc (for example, Ne/Nc=2.10−5 in Pleuronectes platessa (Hoarau et al., 2005); Ne/Nc=10−5 in Crassostreas gigas (Hedgecock et al., 1992); see Hauser and Carvalho, 2008 for a review). One explanation for the low Ne/Nc ratio found in marine species with a free-swimming larval stage is high variation in individual reproductive success, particularly because of high interfamily variance in the mortality of larvae dispersed in a fluctuating environment (see, for example, Siegel et al., 2008). Therefore, a large proportion of reproductive individuals eventually fail to reproduce, potentially leading to a familial lottery or ‘sweepstakes’ of reproductive success (Hedgecock, 1994). Given that adults who effectively reproduce are only a fraction of the total population, reduction of the effective population size relative to the census size is one of the key predictions of the ‘sweepstakes reproductive success’ (SRS) hypothesis (Hedgecock and Pudovkin, 2011). The SRS hypothesis leads to other testable predictions in term of genetic diversity, genetic structure and relatedness (reviewed in Hedgecock and Pudovkin, 2011). For instance, genetic diversity of offspring from a single reproduction event will be reduced relative to the adult population, because only a subset of the total population succeeds in reproducing (see, for example, Li and Hedgecock, 1998). In addition, viewed as a lottery with different sets of individuals successfully reproducing at each reproductive event, allele frequencies will shift from one reproductive event to the next (see, for example, Lee and Boulding, 2007). Furthermore, related individuals will be observed in the pool of dispersing larvae as the offspring from the few ‘lucky winner’ adults (that is, juveniles; see for example, Christie et al., 2010), although the vast majority are likely unrelated (see, for example, Buston et al., 2009). The SRS hypothesis has been intensively studied, both empirically and theoretically, in marine species and used to explain the chaotic genetic patchiness observed in these species (Hedgecock, 1994). Studies that have tested the predictions of the SRS hypothesis have led to conflicting results that can be not only attributed to the methods used and the predictions tested, but also to variation in the intensity of SRS across time (that is, ‘the right place, but the wrong time’ as stated by Hedgecock and Pudovkin, 2011). Using temporal replicates is one way of resolving potential discrepancies when testing SRS predictions.
Life-history traits and mating systems are important components of SRS and, more generally, of the low Ne/Nc ratio. In addition to the particular properties of marine species outlined above, the effect of unequal reproductive success among families may be further enhanced in species forming small perennial breeding groups with restricted mating opportunities, a situation found in the invasive marine mollusk Crepidula fornicata. Native to the eastern coasts of North America, C. fornicata was introduced in Europe in the late-nineteenth century where it now reaches high densities in several bays (Blanchard, 2009). C. fornicata is long-lived (6–8 years; Blanchard, 1995) and displays a benthopelagic life cycle, including a free-swimming larval stage (2 to 7 weeks; Rigal, 2009) and a sessile adult stage. With high fecundity (5000–20 000 larvae released per female at each reproduction event; Richard et al., 2006) and high mortality during early life stages (Pechenik and Levine, 2007), this species has a typical type III survivorship curve (that is, early life stages experience the greatest mortality rates, whereas older life stages experience high survival probabilities). Hedrick (2005) demonstrated that Ne is small when there are few successful breeders (Nb) in a population during one reproductive event. In C. fornicata, several biological characteristics may reduce Nb. This mollusk shows extreme gregarious behavior, with individuals settling on the top of each other to form long-lived ‘stacks’ (that is, individuals are attached on top of each other for their entire life). It is also a protandrous species in which males change into females according to various parameters including age and stack sex ratio (Coe, 1938; Hoagland, 1978). The sexes are not randomly distributed within a stack: (older) females are at the bottom of the stack. In addition, fertilization is internal and females store the sperm transferred by several males, using mainly sperm from one male over a period lasting several months (Le Cam et al., 2009). Because of these characteristics, reproductive individuals have access to a limited number of mates (Dupont et al., 2006; Broquet et al., 2015). Although females release larvae between three and five times per year (Richard et al., 2006), each set of released larvae has a small number of fathers, and this remains stable over the reproductive season (Broquet et al., 2015). Temporal polyandry is therefore unlikely to compensate for reproductive bias at each mating (Pearse and Anderson, 2009). Along with high variance in the mortality of early life stages, this species is particularly prone to nonrandom variance in reproductive success. The effective population size of C. fornicata can be further reduced by a strongly male-biased sex ratio at both the stack and population levels (Nunney, 1993; Caballero, 1994). The reproductive features listed above (iteroparity, high fecundity, reproductive bias due to protandry, gregariousness and sperm storage) are expected to decrease Ne relative to Nc. Nonetheless, other features may buffer these effects because the reproductive pool is composed of adults that are long-lived and belong to overlapping generations.
The aim of this study was to compare genetic diversity and genetic structure across successive cohorts of settlers (that is, new recruits) in the invasive marine species C. fornicata. Although reproduction occurs continuously over 7–9 months, there is only one major recruitment event observed each year in late summer/early autumn. This peak of recruits allows clear identification of a single generation, that is, the youngest generation composed of immature individuals that are <1 year old (Le Cam, 2009). However, older generations cannot be identified with accuracy because of a poor correlation between size and age (Le Cam, 2009). To test the hypothesis of substantial genetic drift across successive cohorts and a moderate effective number of breeders in the wild, we examined temporal variation in genetic diversity of juvenile pools of C. fornicata that were sampled during the main annual settlement event for 9 consecutive years (2002–2010). Genetic data were also used to estimate contemporary effective population size (that is, short-term Ne estimates among samples collected over 10 years). Our experiment tested the following three hypotheses: (1) an increase in recruitment success reduces the loss of genetic diversity at the population level; (2) temporal changes of genetic diversity occur across successive episodes of juvenile settlement; and (3) effective population size is substantially lower than census size, in light of predictions of the SRS hypothesis.
Materials and methods
Sampling design
Reproduction in C. fornicata is known to occur during spring and summer (Henry et al., 2010) and, following larval dispersal and metamorphosis, recruits (that is, juveniles—both terms will be used interchangeably thereafter) settle en masse during a 2–4-month period in late summer/early fall on stacks in response to chemical cues released by adults (Pechenik and Heyman, 1987; McGee and Targett, 1989). For this study, C. fornicata was sampled in the Bay of Morlaix (Brittany, France; 48°40′18′′N, 3°53′17′′W), a site where the benthic phase of the species has been surveyed monthly since 2004 as part as a long-term monitoring program. Each month, 30 stacks (minimum 300 individuals) are sampled along a 50-m transect by scuba diving. Males, females and juveniles found on these stacks are counted to determine the proportion of juveniles each month and the onset of the settlement period. Individuals are preserved in 96% ethanol for subsequent genetic analyses. The proportion of juveniles to adults in the population varies across years (Table 1; Le Cam, 2009; S Le Cam et al., unpublished data) with two extreme values: 1.6% in 2009 and 21.0% in 2006. For this study, for each year except 2009, an ‘annual recruits’ sample was selected: it included all juveniles collected in three successive months during the major recruitment episode. Because of low recruitment in 2009, we selected all the juveniles collected before the start of the next reproductive period (March 2010). Additional samples were available for 2002 and 2003, that is, before the start of the monthly monitoring, with samples collected three times only (autumn, winter and spring). We used the autumn sample that is the most similar to the ‘annual recruits’ sample described above.
DNA extraction and microsatellite genotyping
Genomic DNA extraction of juveniles was performed using a Nucleospin Multi-96 Tissue Kit (MACHEREY-NAGEL, GmbH & Co. KG, Düren, Germany) following the manufacturer’s protocol. Each individual was genotyped at 15 microsatellite loci; 12 loci were expressed sequence tag–simple sequence repeats (Riquet et al., 2011) and 3 markers were isolated from anonymous nuclear libraries: CfH7 (Dupont et al., 2006), Cf8 (Proestou, 2006) and CfCA4 (Dupont et al., 2003). Loci were amplified by PCR following Riquet et al. (2011), but with 2 μl of stock DNA diluted 1:20. Amplification products were separated by electrophoresis on an ABI 3130xl DNA sequencer (Applied Biosystems, Waltham, MA, USA). Scoring of alleles was performed using GeneMapper v. 4.0 (Applied Biosystems) and manually verified.
Analyses of genetic diversity
Allele frequencies and allelic richness (Ar, that is, the expected number of alleles corrected for sampling size based on a rarefaction procedure) were estimated for each sample using FSTAT 2.9.3.2 (Goudet, 1995). We used GENEPOP 4.1 (Rousset, 2008) to estimate expected heterozygosity (He), fixation index (FIS) and to test departure from Hardy–Weinberg equilibrium (HWE; 10 000 dememorization steps, 500 batches and 5000 iterations per batch). Juvenile samples that departed from mean Ar and He values computed over all samples were tested using a bootstrap procedure (10 000) implemented in the statistical program R (R Development Core Team, 2011). The correlation between single-locus Ar or He estimates and the percentage of recruits observed during monitoring was assessed using Spearman’s correlation analyses using R. Correlations were independently assessed and P-values were then combined using Fisher’s combined test statistic (Fisher, 1932). Genotypic data from adults were available for the same 15 loci; the adult samples have been collected four times (in 2002, 2006, 2007 and 2010; F Riquet et al., unpublished data). Indices of genetic diversity were compared between juveniles and adults for each of these 4 years, using the permutation procedure implemented in FSTAT (5000 permutations). Correspondence analysis implemented in GENETIX (Belkhir et al., 1996–2004) was performed to graphically visualize the genetic distance between each sample of juveniles over the 9 study years. As recommended by Jombart et al. (2009), alleles with frequencies 5% were excluded. Genetic structure between samples was assessed using FST estimates (Weir and Cockerham, 1984) calculated using GENEPOP 4.1 (Rousset, 2008). Exact tests for population differentiation (10 000 dememorization steps, 500 batches and 5000 iterations per batch) were carried out to test for differences in allele distributions among samples and between pairwise samples. To adjust P-values for multiple tests, q-values were computed using the QVALUE package in the R software (Storey, 2002). Isolation By Time was tested using the Mantel test (Mantel, 1967) implemented in GENEPOP on the web (http://genepop.curtin.edu.au/) correlating temporal distance (years or months between recruitment) with FST/1−FST.
Effective size estimation
Estimating Ne is challenging and the use of several independent methods is recommended to improve accuracy (Waples, 2005; Fraser et al., 2007; Waples and Do, 2010; Hare et al., 2011). Two categories of methods were used to estimate effective population size: temporal methods and single-sample methods.
First, we used methods in which Ne is estimated by measuring temporal changes in allele frequencies between two or more samples separated by one or more generations (that is, ‘temporal methods’; Waples, 1989). Four temporal methods were used: (1) moment-based F-statistics (Waples, 1989) (method 1); (2) a pseudo-likelihood-based approach (Wang, 2001) (method 2); (3) a maximum-likelihood coalescent-based method (Beaumont, 2003) (method 3); and (4) a method identical to (2) but in which migration rate is jointly estimated (Wang and Whitlock, 2003) (method 4), relaxing the assumption that the population is completely closed to immigration. Sampling plan II was used (Waples, 2005) because individuals were sampled before reproduction and not replaced. Sampling periods spanned 9 years (2002–2010) and we assumed a generation time of 2 years. The effective size was estimated between the most temporally spaced samples (that is, recruits of 2002 and 2010), hence, separated by four generations, a timespan short enough to ignore the effects of mutation (Waples, 1989; Beaumont, 2003) and reduce bias due to overlapping generations (Waples and Yokota, 2007). The moment-based temporal estimates (method 1) and their 95% confidence intervals (Waples, 1989) were computed using NeEstimator v2 (Ovenden et al., 2007; Do et al., 2014) using a modified formula for the temporal variance F (Jorde and Ryman, 2007). The pseudo-likelihood methods (methods 2 and 4) were implemented in MLNe 2.3 (Wang and Whitlock, 2003). We set maximum Ne to 1 500 000, a value that is higher than Nc estimated in the Bay of Morlaix. Based on a spatial survey of the site, density of C. fornicata ranges from 20 to 200 ind m−2 (Dupont, 2004), spread over 6000 m2 (L Lévêque and T Broquet, personal communication), resulting in ca. 120 000 to 1 200 000 individuals in the study site. Migration was set to zero, a condition under which MLNe estimates the likelihood estimator proposed by Wang (2001) (method 2). For method 4, samples used as the ‘source population’ were adults collected at four sites within the Bay of Morlaix and two sites from the closest adjacent bay (Saint-Brieuc) (F Riquet et al., unpublished data). Finally, we used a coalescent-based maximum likelihood method implemented in TMVP (method 3; Beaumont, 2003), and used Markov chain Monte Carlo simulations (25 000 replicates) to generate a posterior distribution of Ne assuming a Bayesian prior on a maximum Ne of 1 500 000. A combined point estimator based on the harmonic mean of these four temporal methods was calculated according to strategy 1 described in Appendix A of Waples and Do (2010).
Second, we also used methods based on the analysis of a single sample to estimate the effective number of breeders (Nb) that produced the sample (Waples, 2005): (1) a method measuring the nonrandom association of alleles (method 5) at different loci within a sample (Hill, 1981) using a bias correction (Waples, 2006; Waples and Do, 2008) and implemented in NeEstimator v2 (Do et al., 2014); (2) a method based on kinship reconstruction (method 6) as proposed by Wang (2009) and available in the COLONY software (Wang, 2004; Jones and Wang, 2010); and (3) a method based on molecular coancestry (method 7) (Nomura, 2008) implemented in NeEstimator v2 (Do et al., 2014). For method 5, a criterion (Pcrit) was used for excluding rare alleles and balancing the precision–bias tradeoff due to rare alleles. Given the size of each sample and following recommendations by Waples and Do (2010), we excluded alleles with frequencies below Pcrit=0.05 for 2009; Pcrit=0.02 for 2002 to 2005, 2008 and 2010; and Pcrit=0.01 for 2006 and 2007. Similar to temporal estimates, point estimates from the three single-sample methods were combined into a joint-point estimate (unweighted harmonic mean) per sample that was used to evaluate variations in Ne over the sample period (Waples and Do, 2010).
Finally, to obtain an overall estimate of effective size, a joint-point estimate based on all single-sample and temporal estimates was computed (Waples and Do, 2010). The single-sample estimators provide an estimate of the effective number of breeders in the previous year. To avoid combining inferences over different timeframes, we computed single-sample and temporal estimates over comparable timeframes and excluded the 2009 sample, in which only 17 individuals were genotyped. This selection resulted in three comparable timeframes: 2002/2006, 2003/2007 and 2002/2008, for which estimates were combined based on (1) temporal methods applied to the recruits of 2002 and 2006 and single-sample methods applied to the samples from 2003 to 2007; (2) temporal methods applied to the recruits of 2003 and 2007 and single-sample methods applied to the samples from 2004 to 2008; and (3) temporal methods applied to the recruits of 2002 and 2008 and single-sample methods applied to the samples from 2003 to 2009. Computations were performed following recommendations and equations provided in Waples and Do (2010) and are shown in Supplementary Appendix S1.
For examining the influence of generation time on our conclusions, we also computed joint-point estimates for each of three timeframes (2002/2006, 2003/2007 and 2002/2008) but increasing generation time, that is, 4 years (using 2002/2006 and 2003/2007 data sets) and 6 years for the 2002/2008 data set.
Results
Genetic diversity and recruitment success
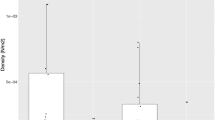
A total of 681 juveniles were genotyped at 15 highly polymorphic microsatellites (2 to 68 alleles/locus). Estimates of genetic diversity for each sample are summarized in Table 1. After correcting for variation in sample size across years, allelic richness ranged over the 2002–2010 data set from 6.73 in 2009 (J09) to 7.25 in 2006 (J06). He ranged from 0.648 (J03) to 0.680 (J05 and J06; Table 1). Two samples departed from the mean Ar (J04 and J09; Figure 1a), whereas three samples departed from the mean He (J03, J05 and J06; Figure 1b). The most genetically diverse pool of recruits was observed in 2006. Significant correlations were observed between the percentage of yearly recruits and both Ar (Fisher’s combined test statistic, P=0.015) and He (Fisher’s combined test statistic, P=0.007). No reduced genetic diversity was observed in juveniles compared with adults sampled the same year (that is, the putative parental generations; Ar: P= 0.91; He: P=0.15); the indices obtained on adults are given in Supplementary Appendix S2. Departures from HWE were found in each sample (Table 1) and out of the nine study samples, three (J03, J07, J08) exhibited an estimate of FIS higher than the mean value (FIS-mean=0.130). Each locus showed heterozygosity deficiencies in at least one sample, and, although few HWE tests within a sample were statistically significant, combined heterozygosity deficiencies were highly significant across the 15 loci.
Temporal genetic variation across juvenile pools over consecutive generations (2002–2010)
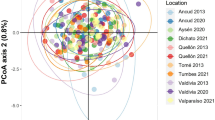
Significant temporal genetic differentiation was observed over the study period (FST-all samples=0.001, P<0.001). Juveniles of 2006 and 2007 are clearly distinguished on the first axis of the correspondence analysis (Figure 2), a pattern also observed with FST estimates (Table 2) that revealed significant genetic differentiation between J07 and all the other samples as well as between J06 and all samples other than J08 and J09. Although the 2003 sample is separated along the second axis of the correspondence analysis, this heterogeneity was not supported by pairwise FST estimates: J03 was only genetically distinct from the J06 and J07 samples. As expected from the above results, no isolation by time was detected across the 9-year sampling (P-Mantel=0.22).
First factorial axes of a correspondence analysis based on 15 microsatellite markers used to analyze recruits sampled from 2002 to 2010. The barycenter of each population is shown. J06 and J07, samples that showed significant genetic differentiation, are distinguishable from all other samples along the first axis. Black arrows perpendicular to the first axis outline these dissimilar samples. J03 stood out along the second axis that is outlined by black arrow perpendicular to the second axis. Samples are labeled according to Table 1.
Convergence of effective size estimators toward moderate values
Temporal changes in allele frequencies were used to estimate effective population size per generation (2002/2010 in Table 3). Over all methods, point estimates of Ne ranged from 136.4 (method 4) to 3 576.0 (method 2) with overlapping confidence intervals of methods 1 and 2, and 1, 3 and 4. The harmonic mean of the four temporal methods yielded Ne=205.7 when taking into account a generation time of 2 years, decreasing to 134.3 and to 82.3 years with a 4-year and 8-year generation time, respectively.
Single-sample methods yielded estimate of Nb ranging from 27.2 (J03, method 7) to infinite estimates (several year samples for methods 5 and 7; Table 4). Confidence intervals of these methods overlapped for only 3 out of the 9 years (J02, J03 and J09). Combined point estimates based on single-sample methods were variable across years, and an unweighted harmonic mean of estimates across all 9 years yielded an estimated Ne of 200.0.
Considering the three comparable timeframes (see ‘Effective size estimation’ in Materials and methods), point estimates yielded a Ne of 174.1 and 205.1 for 2002–2006, 135.0 and 343.5 for 2003–2007 and 310.3 and 211.2 for 2002–2008, across temporal and single-sample methods respectively (details in Supplementary Appendix S1).
Combining the two categories of methods by taking into account variance and weight estimates produced Ne point estimates of 199.8, 157.9 and 214.3 (2002–2006, 2003–2007 and 2002–2008, respectively; Supplementary Appendix S1). Across temporal methods, increasing the generation time reduced the point estimates from 1.80 (2003–2007) to 2.04 (2002–2006; Table 3), resulting in joint-point estimates of Ne across temporal and single-sample methods of 127.1 (2002–2006), 81.6 (2003–2007) and 207.7 (2002–2008), these values being 1.03 to 1.93 lower than with a 2-year generation time (Supplementary Appendix S1).
To summarize, whatever the generation time considered and the method used, with the exception of methods 2 and 7, the majority of Ne estimates, as well as the combined methods, converged to a few hundred individuals.
Discussion
Genetic analysis of the C. fornicata recruits that settled annually over 9 years in the Bay of Morlaix (Brittany, France) revealed departure from HWE, variation in genetic diversity and shifts in allele frequencies over time. The large departure from HWE observed cannot be explained only by the presence of null alleles (Riquet et al., 2011), and can also be attributed to a temporal Wahlund effect (Johnson and Black, 1982, 1984a, b). A temporal Wahlund effect can arise from the juxtaposition of several groups with different allele frequencies, that is, offspring from different families. The temporal genetic changes observed in this bay likely reflect variance in adult reproductive success over time rather than fluctuation in the geographic origin of larvae because the Bay of Morlaix has been shown to receive limited immigration from adjacent bays (Dupont et al., 2007a; see details below).
C. fornicata produces a high number of offspring that undergo high mortality during early life, a pattern typical of many marine species and thought to reflect variability in ocean conditions (Siegel et al., 2008; Hedgecock and Pudovkin, 2011). Many individuals from the parental generation ultimately fail to reproduce because only a small random subset of larvae survives dispersal, metamorphosis and settlement, that is, the so-called SRS hypothesis (Hedgecock, 1994). The effective number of breeders is thus reduced relative to the number of reproductive adults and shift in allele frequencies can be observed from one reproductive event to the next (Johnson and Black, 1991). Temporal genetic variation across years however varied in intensity, suggesting that the extent of this reproductive lottery in C. fornicata may vary among years, and result in temporally variable genetic drift. Similar levels of genetic diversity and low genetic differentiation were observed among the four adult samples of the study population (Supplementary Appendix S2). This similarity suggested a stable reproductive pool (adult samples contributed up to 6–7 annual recruitment events) over time that may buffer genetic variation among settlers. In addition, although predicted by the SRS hypothesis, no reduced genetic diversity was observed in new recruits compared with the adult population, as found in other marine species (see, for example, Ruzzante et al. 1996; but see Hedgecock et al., 2007), a result likely because of the difficulty of detecting such changes with highly polymorphic markers.
Interestingly, recruitment success was variable across years (χ2 test: P<0.001), and 3 years showed particularly successful recruitment with a higher percentage of juveniles (17.35%, 21.01% and 15.71% in 2003, 2006 and 2007, respectively; Table 1) than the mean value (11.6%) observed across the full study period. In contrast, very few recruits were produced in 2009 (⩽2% of the total population; Table 1). Using the proportion of recruits as a proxy for settlement success, we observed a significant correlation between settlement success and indices of genetic diversity (Ar and He), suggesting that higher levels of recruitment yield a more genetically diverse population, a result in favor of the SRS hypothesis. This correlation was heavily influenced by the 2006 and 2009 samples, the most extreme recruitments: upon removal of these samples, the correlation was no longer significant (r=0.47, P=0.78). Therefore, extreme recruitment events highly influence juvenile genetic diversity, but slight interannual fluctuations do not. Based on the joint combined estimates, Ne estimates fluctuated slightly according to the timeframe used: it varied from 135 (2003–2007 timeframe) to 310 (2002–2008 timeframe). These variations may be due to the influence of recruitment success. For instance, the 2002–2008 timeframe, which showed the highest Ne value, included the 3 years with high recruitment success (2003, 2006, 2007). Conversely, the 2002–2010 timeframe included the year with the lowest recruitment (2009) and displayed the lowest Ne estimates. These extreme events seemed also to influence Ne estimations based on single-sample methods, with larger Ne estimates in years of high recruitment success (for example, 2006 and 2007) and smaller Ne estimates in years of low recruitment success (for example, 2009). The overall correlation between Ne estimates and recruitment success is however not significant. Altogether, genetic drift is likely driven by extreme events.
Temporal genetic changes in populations closed to migration are largely driven by genetic drift, an outcome of reductions in effective population size relative to census population size. Migration is usually neglected in most methods estimating Ne, although ignoring it can lead to underestimates or overestimates (Wang and Whitlock, 2003; Fraser et al., 2007; Gilbert and Whitlock, 2015). Migration can induce rapid shifts in allele frequency if immigration involves a genetically different population, and such rapid changes can be wrongly attributed to severe genetic drift, hence, underestimating Ne. Conversely, when source populations are similar to recipient populations, Ne can be overestimated (Gilbert and Whitlock, 2015). The Bay of Morlaix is unlikely to be strongly influenced by migration from neighboring bays. Larval distribution studies and hydrodynamic modeling has shown that this bay mostly exports larvae and that the larval supply entering the bay is small (Rigal et al., 2010). In addition, genetic studies have shown significant genetic differentiation between the Bay of Morlaix and its western and eastern neighboring bays (FST=0.0015; P-value on exact test <0.001; F Riquet et al., unpublished data; Dupont et al., 2007a) that may indicate limited gene flow (and/or large effective population size). Estimates from methods 2 and 4, which are identical except for the presence of migration were, however, inconsistent. The highest Ne values were observed when ignoring gene flow (that is, overestimating Ne), a result congruent with many other studies (references in Fraser et al., 2007; Gilbert and Whitlock, 2015). Thus, although the study population is thought to experience limited immigration from neighboring bays, we cannot exclude some biases due to immigration. However, using seven methods differing in mathematical properties and assumptions (review in Luikart et al., 2010; Gilbert and Whitlock, 2015), and combined estimates following Waples and Do (2010), the overall majority of estimates was of the same order and yielded a mean value of ca. 194.4 (detailed comparisons of Ne estimates are provided in Supplementary Appendix S3). Congruent with the study of Gilbert and Whitlock (2015) based on simulated data using seven methods and various demographic scenarios, two methods (methods 3 and 4), plus method 6 not assessed in Gilbert and Whitlock (2015), appeared to be the most reliable given their robustness to changes in timeframe and their confidence intervals.
Considering the census size estimated in the field for the sampled population and the mean Ne estimate, the Ne/Nc ratio is 1.6 × 10−3 when using the lower bound of the estimated Nc (Nc-min=120 000) and 1.6 × 10−4 when using the upper bound of the estimated Nc (Nc-max=1 200 000). The effective size of the population was thus estimated to be at most three or four orders of magnitude less than the number of potential breeders. This ratio is only slightly modified if we consider a longer generation time, a parameter difficult to estimate in the study species. In laboratory studies, maturity takes place within 4 months for males (Warne, 1956, cited in Fretter and Graham, 1981) and 300 days for females (Nelson et al., 1983), suggesting a 2-year generation time. When assuming longer generation times (4 or 6 years), Ne estimates were 1.03 to 1.93 times lower than those estimated with a 2-year generation time, thus suggesting higher genetic drift from one generation to the next, but Ne/Nc is of the same order because of large Nc values. Theoretically, the Ne/Nc ratio is expected to be ∼0.5 for a wide range of demographic and reproductive scenarios (Nunney, 1993, 1996). By analyzing Ne/Nc ratios of different species characterized by different life-history traits, Palstra and Ruzzante (2008) showed that life-history traits such as high fecundity and high mortality in the early stage (that is, traits generating high variance in reproductive success) may result in a 86% reduction in effective size relative to the census size. However, the Ne/Nc ratio estimated in C. fornicata is <0.14, whatever the Ne/Nc value considered, as reported in the majority of studies in the marine realm that have documented even lower Ne/Nc ratios (see, for example, Hedgecock, 1994; Hauser et al., 2002; Hoarau et al., 2005). However, our ratio is much higher than those detected in other marine species, especially in fishes and oysters (Hauser and Carvalho, 2008; Hare et al., 2011 and references herein), for which Ne is several orders of magnitude smaller than Nc. This suggests that the Ne/Nc ratio in C. fornicata is moderate compared with other marine species.
Census population size estimates used in this study may have been underestimated because the spatial distribution of the species is highly heterogeneous, and density values, measured by scuba diving along transects (Dupont, 2004), may have missed local dense patches. There are thus uncertainties in the Nc estimates used here. However, interestingly, the population seems stable over time: although Nc has not been carefully monitored, monthly scuba diving over 13 years, plus two surveys carried out in the entire Bay of Morlaix in 2006 and 2010, have not reported significant changes in C. fornicata population size and density (that is, no major boom or bust). Despite the large number of larvae released per reproductive event, most larvae are exported out of the bay, and only a fraction recruits into the study population, contributing to the demographic stability in the study site (Rigal et al., 2010). A large effective size (relative to other marine species) may also explain the large Ne/Nc ratio in the study species relative to other marine species. Differences in life-history traits may have important impacts on Ne (Gaggiotti and Vetter, 1999) and particular life-history traits of the study species may have positively affected Ne. C. fornicata displays stacking behavior, and at the stack level, that is, within a breeding group (Dupont et al., 2006; Proestou et al., 2008; Le Cam et al., 2009), individuals from several generations co-occur and mating between different generations is de rigueur (Dupont et al., 2007b; Le Cam et al., 2009; Broquet et al., 2015). These particular age–sex structures in which generations overlap and mate should reduce temporal genetic variation (Ryman, 1997; Gaggiotti and Vetter, 1999). The stability of the reproductive pool over successive years makes successive annual cohorts of recruits likely to be partly issued from genetically similar reproductive pools. These particular life traits may be sufficient to buffer the effects of reproductive variation and sweepstake events, and thus genetic drift at the population level (Dupont et al., 2007b).
Despite moderate genetic drift relative to other marine species, we estimated a small effective population size compared with the census size. Responses to selection have been hypothesized to be important processes in explaining invasion success of nonindigenous species, especially in primary introduced species with large genetic diversity on which selection could act (Lee, 2002; Roman and Darling, 2007; Barrett and Schluter, 2008; Lawson-Handley et al., 2011; Rius et al., 2015). Introduced populations that have experienced founder events and lost genetic variation following introduction may also lose their ability to adapt to new conditions. Based on a genome-scan approach, no selection effects have been shown in C. fornicata following its introduction (Riquet et al., 2013). This pattern may be because of a limited effective population size across several generations. Note, however, that we estimated contemporary Ne. Further studies investigating Ne over longer timescales and at a species-wide level should be pursued. In particular, sampling distinct adult generations (that is, non-overlapping; for instance every 10 years) would be needed, although hardly feasible on such a long timescale. To our knowledge, no other study has examined Ne estimates in this introduced species, and rigorous Ne comparisons of native and introduced populations are also needed to ascertain this putative link between (the lack of) founder events, (large) effective size and invasion success.
Conclusion
A reduced effective population size relative to the census size was observed in C. fornicata. In addition, temporal genetic variations between cohorts of settlers were here documented, both of which are signatures left by a reproductive lottery. This footprint of genetic drift and SRS seems, however, weak in C. fornicata when considering values reported in other marine species (that is, Ne/Nc <<0.01). These differences may reflect particular life-history traits of C. fornicata that minimize the effect of reproductive variation among adults relative to other marine species. Estimating effective size is challenging in marine species with larval dispersal and large population sizes because of the risks of sampling bias and long-distance immigration from distant populations. Investigating SRS should thus not only include estimating Ne/Nc ratios, but also investigate all of its other predictions such as higher relatedness within larval cohorts (see, for example, Planes et al., 2002) that require temporal sampling at different stages of the life cycle, particularly between parents and their offspring before dispersal.
Data archiving
Microsatellite data available from the Dryad Digital Repository: http://dx.doi.org/10.5061/dryad.6g6mj.
References
Barrett RDH, Schluter D . (2008). Adaptation from standing genetic variation. Trends Ecol Evol 23: 38–44.
Beaumont MA . (2003). Estimation of population growth or decline in genetically monitored populations. Genetics 164: 1139–1160.
Belkhir K, Borsa P, Chikhi L, Raufaste N, Bonhomme F . (1996-2004) GENETIX 4.05, logiciel sous Windows TM pour la génétique des populations. Laboratoire Génome, Populations, Interactions, CNRS UMR 5171, Université de Montpellier II: Montpellier, France.
Berthier P, Beaumont MA, Cornuet JM, Luikart G . (2002). Likelihood-based estimation of the effective population size using temporal changes in allele frequencies: a genealogical approach. Genetics 160: 741–751.
Blanchard M . (1995). Origine et état de la population de Crepidula fornicata (Gastropoda Prosobranchia) sur le littoral français. Haliotis 24: 75–86.
Blanchard M . (2009). Recent expansion of the slipper limpet population (Crepidula fornicata in the Bay of Mont-Saint-Michel (Western Channel, France). Aquat Living Resour 22: 11–19.
Blanchet S . (2012). The use of molecular tools in invasion biology: an emphasis on freshwater ecosystems. Fish Manag Ecol 19: 120–132.
Broquet T, Barranger A, Billard E, Bestin A, Berger R, Honnaert G, Viard F . (2015). The size advantage model of sex allocation in the protandrous sex-changer Crepidula fornicata: role of the mating system, sperm storage, and male mobility. Am Nat 186: 404–420.
Buston PM, Fauvelot C, Wong MYL, Planes S . (2009). Genetic relatedness in groups of the humbug damselfish Dascyllus aruanus: small, similar-sized individuals may be close kin. Mol Ecol 18: 4707–4715.
Caballero A . (1994). Developments in the prediction of effective population-size. Heredity 73: 657–679.
Chapman RW, Ball AO, Mash LR . (2002). Spatial homogeneity and temporal heterogeneity of red drum (Sciaenops ocellatus microsatellites: effective population sizes and management implications. Mar Biotechnol 4: 589–603.
Coe WR . (1938). Conditions influencing change of sex in mollusks of the genus Crepidula. J Exp Zool 77: 401–424.
Christie MR, Johnson DW, Stallings CD, Hixon MA . (2010). Self-recruitment and sweepstakes reproduction amid extensive gene flow in a coral-reef fish. Mol Ecol 19: 1042–1057.
Do C, Waples RC, Peel D, Macbeth GM, Tillett BJ, Ovenden JR . (2014). NeEstimator v2: re-implementation of software for the estimation of contemporary effective population size (Ne) from genetic data. Mol Ecol Resour 14: 209–214.
Dupont L . (2004). Invasion des côtes françaises par le mollusque exotique Crepidula fornicata: contribution de la dispersion larvaire et du système de reproduction au succès de la colonisation. MSc Thesis, University of Pierre and Marie Curie.
Dupont L, Jollivet D, Viard F . (2003). High genetic diversity and ephemeral drift effects in a successful introduced mollusc (Crepidula fornicata: Gastropoda). Mar Ecol Prog Ser 253: 183–195.
Dupont L, Richard J, Paulet YM, Thouzeau G, Viard F . (2006). Gregariousness and protandry promote reproductive insurance in the invasive gastropod Crepidula fornicata: evidence from assignment of larval paternity. Mol Ecol 15: 3009–3021.
Dupont L, Ellien C, Viard F . (2007a). Limits to gene flow in the slipper limpet Crepidula fornicata as revealed by microsatellite data and a larval dispersal model. Mar Ecol Prog Ser 349: 125–138.
Dupont L, Bernas D, Viard F . (2007b). Sex and genetic structure across age groups in populations of the European marine invasive mollusc, Crepidula fornicata L. (Gastropoda). Biol J Linn Soc Lond 90: 365–374.
Fisher RA . (1932) Statistical Methods for Research Workers, 4th edn. Oliver and Boyd: Edinburgh.
Frankham R . (1995). Effective population-size adult-population size ratios in wildlife - a review. Genet Res 66: 95–107.
Fraser DJ, Hansen MM, Ostergaard S, Tessier N., Legault M, Bernatchez L . (2007). Comparative estimation of effective population sizes and temporal gene flow in two contrasting population systems. Mol Ecol 16: 3866–3889.
Fretter V, Graham A . (1981). The prosobranch molluscs of Britain and Denmark. Molluscs of Britain and Denmark. part 6. J Mollusc Stud Sup 9: 309–313.
Gaggiotti OE, Vetter RD . (1999). Effect of life history strategy, environmental variability, and overexploitation on the genetic diversity of pelagic fish populations. Can J Fish Aquat Sci 56: 1376–1388.
Gilbert KJ, Whitlock MC . (2015). Evaluating methods for estimating local effective population size with and without migration. Evolution 69: 2154–2166.
Goudet J . (1995). FSTAT (Version 1.2): a computer program to calculate F-statistics. J Hered 86: 485–486.
Hare MP, Nunney L, Schwartz MK, Ruzzante DE, Burford M, Waples RS et al. (2011). Understanding and estimating effective population size for practical application in marine species management. Conserv Biol 25: 438–449.
Hauser L, Adcock GJ, Smith PJ, Ramirez JHB, Carvalho GR . (2002). Loss of microsatellite diversity and low effective population size in an overexploited population of New Zealand snapper (Pagrus auratus. Proc Natl Acad Sci USA 99: 11742–11747.
Hauser L, Carvalho GR . (2008). Paradigm shifts in marine fisheries genetics: ugly hypotheses slain by beautiful facts. Fish Fish 9: 333–362.
Hedgecock D . (1994) Does variance in reproductive success limit effective population sizes of marine organisms? In: Beaumont A (ed). Genetic and Evolution of Aquatic Organisms. Chapman and Hall: London, pp 122–134.
Hedgecock D, Chow V, Waples RS . (1992). Effective population numbers of shellfish broodstocks estimated from temporal variance in allelic frequencies. Aquaculture 108: 215–232.
Hedgecock D, Launey S, Pudovkin AI, Naciri Y, Lapegue S, Bonhomme F . (2007). Small effective number of parents (Nb) inferred for a naturally spawned cohort of juvenile European flat oysters Ostrea edulis. Mar Biol 150: 1173–1182.
Hedgecock D, Pudovkin AI . (2011). Sweepstakes reproductive success in highly fecund marine fish and shellfish: a review and commentary. Bull Mar Sci 87: 971–1002.
Hedrick P . (2000) Genetics of Populations, 2nd edn. Jones and Bartlett Publishers, Inc.: Boston.
Hedrick P . (2005). Large variance in reproductive success and the Ne/N ratio. Evolution 59: 1596–1599.
Henry JJ, Collin R, Perry KJ . (2010). The slipper snail, Crepidula: an emerging lophotrochozoan model system. Biol Bull 218: 211–229.
Hill WG . (1981). Estimation of effective population-size from data on linkage disequilibrium. Genet Res 38: 209–216.
Hoagland KE . (1978). Protandry and evolution of environmentally-mediated sex change - study of mollusca. Malacologia 17: 365–391.
Hoarau G, Boon E, Jongma DN, Ferber S, Palsson J, Van der Veer HW et al. (2005). Low effective population size and evidence for inbreeding in an overexploited flatfish, plaice (Pleuronectes platessa L.). Proc Biol Sci 272: 497–503.
Jombart T, Pontier D, Dufour AB . (2009). Genetic markers in the playground of multivariate analysis. Heredity 102: 330–341.
Jones OR, Wang J . (2010). COLONY: a program for parentage and sibship inference from multilocus genotype data. Mol Ecol Res 10: 551–555.
Johnson MS, Black R . (1982). Chaotic genetic patchiness in an intertidal limpet, Siphonaria sp. Mar Biol 70: 157–164.
Johnson MS, Black R . (1984a). Pattern beneath the chaos: the effect of recruitment on genetic patchiness in an intertidal limpet. Evolution 38: 1371–1383.
Johnson MS, Black R . (1984b). The Wahlund effect and the geographical scale of variation in the intertidal limpet Siphonaria sp. Mar Biol 79: 295–302.
Johnson MS, Black R . (1991). Genetic subdivision of the intertidal snail Bembicium vittatum (Gastropoda: Littorinidae) varies with habitat in the Houtman Abrolhos Islands, Western Australia. Heredity 67: 205–213.
Jorde PE, Ryman N . (2007). Unbiased estimator for genetic drift and effective population size. Genetics 177: 927–935.
Lawson-Handley LJ, Estoup A, Evans DM, Thomas CE, Lombaert E, Facon B et al. (2011). Ecological genetics of invasive alien species. Biocontrol 56: 409–428.
Le Cam S . (2009). Grégarité, changement de sexe et polyandrie: modalités de la reproduction chez une espèce invasive Crepidula fornicata. MSc Thesis, University of Pierre and Marie Curie.
Le Cam S, Pechenik JA, Cagnon M, Viard F . (2009). Fast versus slow larval growth in an invasive marine mollusc: does paternity matter? J Hered 100: 455–464.
Lee CE . (2002). Evolutionary genetics of invasive species. Trends Ecol Evol 17: 386–391.
Lee HJ, Boulding EG . (2007). Mitochondrial DNA variation in space and time in the northeastern Pacific gastropod, Littorina keenae. Mol Ecol 16: 3084–3103.
Li G, Hedgecock D . (1998). Genetic heterogeneity, detected by PCR-SSCP, among samples of larval Pacific oysters (Crassostrea gigas supports the hypothesis of large variance in reproductive success. Can J Fish Aquat Sci 55: 1025–1033.
Luikart G, Ryman N, Tallmon DA, Schwartz MK, Allendorf FW . (2010). Estimation of census and effective population sizes: the increasing usefulness of DNA-based approaches. Conserv Genet 11: 355–373.
Mantel N . (1967). Detection of disease clustering and a generalized regression approach. Cancer Res 27: 209.
McGee BL, Targett NM . (1989). Larval habitat selection in Crepidula (L) and its effect onadult distribution patterns. J Exp Mar Biol Ecol 131: 195–214.
Nelson DA, Calabrese A, Greig RA, Yevich PP, Chang S . (1983). Long term silver effects on the marine gastropod Crepidula fornicata. Mar Ecol Prog Ser 12: 155–165.
Nomura T . (2008). Estimation of effective number of breeders from molecular coancestry of single cohort sample. Evol Appl 1: 462–474.
Nunney L . (1993). The influence of mating system and overlapping generations on effective population-size. Evolution 47: 1329–1341.
Nunney L . (1996). The influence of variation in female fecundity on effective population size. Biol J Linn Soc 59: 411–425.
Ovenden JR, Peel D, Street R, Courtney AJ, Hoyle SD, Peel SL et al. (2007). The genetic effective and adult census size of an Australian population of tiger prawns (Penaeus esculentus. Mol Ecol 16: 127–138.
Palstra FP, Ruzzante DE . (2008). Genetic estimates of contemporary effective population size: what can they tell us about the importance of genetic stochasticity for wild population persistence? Mol Ecol 17: 3428–3447.
Pearse DE, Anderson EC . (2009). Multiple paternity increases effective population size. Mol Ecol 18: 3124–3127.
Pechenik JA, Heyman WD . (1987). Using KCL to determine size at competence for larvae of the marine gastropod Crepidula fornicata (L). J Exp Mar Biol Ecol 112: 27–38.
Pechenik JA, Levine SH . (2007). Estimates of planktonic larval mortality using the marine gastropods Crepidula fornicata and C. plana. Mar Ecol Prog Ser 344: 107–118.
Planes S, Lecaillon G, Lenfant P, Meekan M . (2002). Genetic and demographic variation in new recruits of Naso unicornis. J Fish Biol 61: 1033–1049.
Proestou DA . (2006). Isolation and characterization of microsatellite markers in the Atlantic slipper shell Crepidula fornicata for use in paternity analysis. Mol Ecol Notes 6: 437–439.
Proestou DA, Goldsmith MR, Twombly S . (2008). Patterns of male reproductive success in Crepidula fornicata provide new insight for sex allocation and optimal sex change. Biol Bull 214: 192–200.
R Development Core Team. (2011). R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria. Available at: http://www.R-project.org/.
Richard J, Huet M, Thouzeau G, Paulet YM . (2006). Reproduction of the invasive slipper limpet, Crepidula fornicata, in the Bay of Brest, France. Mar Biol 149: 789–801.
Rigal F . (2009). Dynamique spatio-temporelle du nuage larvaire du gastéropode introduit Crepidula fornicata au sein d'une baie mégatidale, la baie de Morlaix (France). MSc Thesis, University of Pierre and Marie Curie.
Rigal F, Viard F, Ayata SD, Comtet T . (2010). Does larval supply explain the low proliferation of the invasive gastropod Crepidula fornicata in a tidal estuary? Biol Inv 12: 3171–3186.
Riquet F, Ballenghien M, Tanguy A, Viard F . (2011). In silico mining and characterization of 12 EST-SSRs for the invasive slipper limpet Crepidula fornicata. Mar Genom 4: 291–295.
Riquet F, Daguin-Thiébaut C, Ballenghien M, Bierne N, Viard F . (2013). Contrasting patterns of genome-wide polymorphism in the native and invasive range of the marine mollusk Crepidula fornicata. Mol Ecol 22: 1003–1018.
Rius M, Turon X, Bernard G, Volckaert F, Viard F . (2015). Marine invasion genetics: from spatial and temporal patterns to evolutionary outcomes. Biol Inv 17: 869–885.
Roman J, Darling JA . (2007). Paradox lost: genetic diversity and the success of aquatic invasions. Trends Ecol Evol 22: 454–464.
Rousset F . (2008). GENEPOP'007: a complete re-implementation of the GENEPOP software for Windows and Linux. Mol Ecol Res 8: 103–106.
Ruzzante DE, Taggart CT, Cook D . (1996). Spatial and temporal variation in the genetic composition of a larval cod (Gadus morhua aggregation: cohort contribution and genetic stability. Can J Fish Aquat Sci 53: 2695–2705.
Ryman N . (1997). Minimizing adverse effects of fish culture: understanding the genetics of populations with overlapping generations. J Mar Sci 54: 1149–1159.
Siegel DA, Mitarai S, Costello CJ, Gaines SD, Kendall BE et al. (2008). The stochastic nature of larval connectivity among nearshore marine populations. Proc Natl Acad Sci USA 105: 8974–8979.
Storey J . (2002). A direct approach to false directory rates. J R Soc Series B 64: 479–498.
Turner TF, Osborne MJ, Moyer GR, Benavides MA, Alo D . (2006). Life history and environmental variation interact to determine effective population to census size ratio. J R Soc Series B 273: 3065–3073.
Wang JL . (2001). A pseudo-likelihood method for estimating effective population size from temporally spaced samples. Genet Res 78: 243–257.
Wang JL . (2004). Sibship reconstruction from genetic data with typing errors. Genetics 166: 1963–1979.
Wang J . (2009). A new method for estimating effective population sizes from a single sample of multilocus genotypes. Mol Ecol 18: 2148–2164.
Wang JL, Whitlock MC . (2003). Estimating effective population size and migration rates from genetic samples over space and time. Genetics 163: 429–446.
Waples RS . (1989). A generalized-approach for estimating effective population-size from temporal changes in allele frequency. Genetics 121: 379–391.
Waples RS . (2005). Genetic estimates of contemporary effective population size: to what time periods do the estimates apply? Mol Ecol 14: 3335–3352.
Waples RS . (2006). A bias correction for estimates of effective population size based on linkage disequilibrium at unlinked gene loci. Conserv Genetics 7: 167–184.
Waples RS, Do C . (2008). LDNE: a program for estimating effective population size from data on linkage disequilibrium. Mol Ecol Res 8: 753–756.
Waples RS, Do C . (2010). Linkage disequilibrium estimates of contemporary Ne using highly variable genetic markers: a largely untapped resource for applied conservation and evolution. Evol Appl 3: 244–262.
Waples RS, Yokota M . (2007). Temporal estimates of effective population size in species with overlapping generations. Genetics 175: 219–233.
Weir B, Cockerham C . (1984). Estimating F-statistics for the analysis of population structure. Evolution 38: 1358–1370.
Wright S . (1931). Evolution in Mendelian populations. Genetics 16: 0097–0159.
Acknowledgements
We are very grateful to the divers of ‘Marine Operations Department’ at the Biological Station of Roscoff who carried out the sampling. We thank the Biogenouest Genomics core facility for its technical support. We are also grateful to Frieda Benun Sutton, Jimiane Ashe and Tony Wilson for their help in improving the manuscript. We also thank three anonymous referees for their useful comments on an earlier version of this manuscript. This work was funded by the French National Research Agency (Project Hi-Flo, ANR-08-BLAN-0334), the AXA Research Fund (project ‘Marine Aliens and Climate Change’) and the Interreg project Marinexus. FR acknowledges a PhD grant from the Centre National pour la Recherche Scientifique and the Conseil Régional de Bretagne (SEA-FLO ARED project).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Heredity website
Supplementary information
Rights and permissions
About this article
Cite this article
Riquet, F., Le Cam, S., Fonteneau, E. et al. Moderate genetic drift is driven by extreme recruitment events in the invasive mollusk Crepidula fornicata. Heredity 117, 42–50 (2016). https://doi.org/10.1038/hdy.2016.24
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2016.24
This article is cited by
-
Comparative population genetics of habitat-forming octocorals in two marine protected areas: eco-evolutionary and management implications
Conservation Genetics (2023)
-
Origin and genetic diversity of the invasive mussel Semimytilus algosus in South Africa, relative to source populations in Chile and Namibia
Biological Invasions (2020)
-
Effective population size and heterozygosity-fitness correlations in a population of the Mediterranean lagoon ecotype of long-snouted seahorse Hippocampus guttulatus
Conservation Genetics (2019)