Abstract
The complex formed by two members of the S100 calcium-binding protein family, S100A8/A9, exerts apoptosis-inducing activity in various cells of different origins. Here, we present evidence that the underlying molecular mechanisms involve both programmed cell death I (PCD I, apoptosis) and PCD II (autophagy)-like death. Treatment of cells with S100A8/A9 caused the increase of Beclin-1 expression as well as Atg12-Atg5 formation. S100A8/A9-induced cell death was partially inhibited by the specific PI3-kinase class III inhibitor, 3-methyladenine (3-MA), and by the vacuole H+-ATPase inhibitor, bafilomycin-A1 (Baf-A1). S100A8/A9 provoked the translocation of BNIP3, a BH3 only pro-apoptotic Bcl2 family member, to mitochondria. Consistent with this finding, ΔTM-BNIP3 overexpression partially inhibited S100A8/A9-induced cell death, decreased reactive oxygen species (ROS) generation, and partially protected against the decrease in mitochondrial transmembrane potential in S100A8/A9-treated cells. In addition, either ΔTM-BNIP3 overexpression or N-acetyl-L-cysteine co-treatment decreased lysosomal activation in cells treated with S100A8/A9. Our data indicate that S100A8/A9-promoted cell death occurs through the cross-talk of mitochondria and lysosomes via ROS and the process involves BNIP3.
Similar content being viewed by others
Introduction
Type 1 (apoptotic) and type 2 (autophagic) cell death are two common forms of cell demise 1, 2, of which apoptosis appears to be the prevalent form of programmed cell death (PCD) in multicellular organisms. Apoptosis is morphologically characterized by cell shrinkage, chromatin condensation, blebbing, and formation of apoptotic bodies. These processes are influenced by the balance of pro-apoptotic and anti-apoptotic signals, which in turn are regulated by Bcl2-family members 3. Biochemically, the main features of this process are caspase activation and DNA fragmentation 3, 4, 5. Apoptosis can be induced by either death receptors or toxic stimuli such as chemotherapeutic drugs, DNA damage, staurosporine, ultraviolet irradiation, γ-irradiation, growth-factor deprivation, and endoplasmic reticulum stress 4, 6, 7.
Mitochondria play a key role in mediating apoptosis induced by many different stimuli 3, 8. Alterations in mitochondria transmembrane potential (ΔΨm) in response to various triggers lead to the production of reactive oxygen species (ROS) or mitochondrial membrane permeabilization (MMP) 9, 10. MMP can be induced by the interaction of pro-apoptotic Bcl2-family members Bax/Bak with mitochondrial outer membrane. This interaction results in the release of a number of small molecules, including cytochrome c, second mitochondria-derived activator of caspase/direct inhibitor of apoptosis protein-binding protein with low pI (Smac/DIABLO), Omi/HtrA2, apoptosis-inducing factor (AIF), and EndoG that activate caspase-dependent and -independent apoptotic cell death pathways 11, 12.
Autophagy is a regulated process of degradation and recycling of cellular constituents, participating in organelle turnover and in the bioenergetic management of starvation 13. During autophagy, parts of the cytoplasm or entire organelles are sequestered to double-membrane vesicles, referred to as autophagic vacuoles (AV) or autophagosomes, respectively. Autophagosomes ultimately fuse with lysosomes, thereby generating single-membrane autophagolysosomes and degrading their content 14. In addition to its basic role in the turnover of proteins and organelles, autophagy has multiple physiological and pathophysiological functions including roles in cell differentiation, immune defense, and cell death 13. Early activation of autophagy has been described as a frequent form of PCD during embryogenesis, insect metamorphosis, regression of tumors 15, and in human neurodegenerative diseases such as Alzheimer's and Parkinson's diseases 16, 17. On the basis of morphological changes, this autophagic cell death is defined as type II cell death opposed to the non-autophagic apoptotic type I cell death 18.
S100A8 and S100A9 (also known as calgranulins A and B, MRP8 and MRP14, and calprotectin) are members of the S100 multigene sub-family of cytoplasmic EF-hand Ca2+-binding proteins. They are differentially expressed in a wide variety of cell types and are abundant in myeloid cells 19, 20. The S100A8/A9 protein complex is released from activated phagocytes and exhibits antimicrobial activity 21 as well as apoptotic/cytotoxic effect against various tumor cells 9, 19.
The S100A8/A9 complex is located in the cytosol of resting phagocytes and follows two independent translocation pathways when the cells are activated. Therefore, it has been assumed that membrane-associated and soluble S100A8/A9 may have distinct cellular functions. Intracellular S100A8/A9 might be involved in (phagocyte) NADPH oxidase activation 22, whereas the secreted form exerts cell growth-promoting activities at low concentrations 23, and induces cell death at higher concentrations 9. Recently, it has been demonstrated that receptor for advanced glycation end product (RAGE) ligation is involved in the cell growth-promoting activity, while the apoptotic-inducing property relies on a yet unknown receptor 9, 23. Similar to S100A8/A9, it has been shown that S100B, another member of the S100 calcium-binding protein family, displays a bimodal function inasmuch as nanomolar concentrations are anti-apoptotic while 5 μM S100B was pro-apoptotic 24. Moreover, S100B also causes myoblast apoptosis in a RAGE-independent manner 25.
Several reports indicate that the PCD pathways I and II may both be induced by the same stimuli and/or in the same cell types 26, 27. For example, interplay between apoptosis and autophagy has been reported following activation of the death receptor-dependent extrinsic apoptotic pathway by tumor necrosis factor-α and TRAIL 28. Thus, we investigated whether S100A8/A9-induced apoptosis is accompanied by autophagy. Here, we demonstrate that autophagy plays an important role in S100A8/A9-induced cell death and it temporally co-exists with apoptosis. Furthermore, we show that cell death induced by S100A8/A9 involves the atypical pro-apoptotic Bcl2-family member BNIP3, and ROS play an important role in S100A8/A9-triggered cell death and may serve as a messenger between mitochondrial and lysosomal death pathways.
Results
S100A8/A9 induces apoptosis in various cell lines
S100A8/A9 efficiently killed MCF7 (Figure 1A) and SHEP (Figure 1B) cells in a concentration- and time-dependent manner, as determined by the 3-(4,5-dimethyl-2-thiazolyl)-2,5-diphenyl-2H-tetrazolium bromide (MTT) assay. S100A8/A9 was also toxic to L929 (Figure 1D) cells at concentrations ≥ 100 μg/ml after 12 h and ≥ 60 μg/ml after 24 and 36 h. A similar effect was observed in HEK-293 cells (Figure 1C) at concentrations ≥ 60 μg/ml after 12 h and ≥ 40 μg/ml at 24 and 36 h. The apoptosis-specific flow cytometry assay (Nicoletti), which detects apoptosis-typical hypodiploid nuclei 29, confirmed that the S100A8/A9-induced cell death was mostly apoptotic (Figure 1E and 1F).
S100A8/A9 induces cell death with hallmarks of apoptosis in multiple human and rodent cell types from different histological origins. MCF7 (A), SHEP (B), HEK-293 (C), and L929 (D) cells were treated with different concentrations of S100A8/A9 for 12-48 h, and cell viability was assessed by MTT assay. S100A8/A9 killed MCF7 and SHEP cell lines in a concentration and time-dependent manner (P < 0.05). Results are expressed as percentage of corresponding control and represent the mean ± SD of four experiments. (E, F) Flow cytometry analysis of HEK-293 and SHEP cells treated with S100A8/A9. (E) Typical DNA-histogram. M2 (statistical marker) has been placed to mark sub-diploid DNA. The diploid (G1) and tetraploid (G2) DNA are clearly visible in the form of two peaks in the far-right part of the histograms. S100A8/A9 treatment (24 h, 100 μg/ml) showed a typical experiment with apoptosis affecting only a fraction of cells. G1 and G2 peaks are still preserved and sub-diploid peak corresponding to apoptotic cells was also clearly visible to the left from both peaks that represent normal cells. S100A8/A9 induced significant apoptosis at 24 h (P < 0.01). Results are expressed as percentage of apoptotic cells, and represent the mean ± SD of four independent experiments. (G) S100A8/A9 induced caspase activation via the intrinsic pathway. Western blot analysis of cell lysates from SHEP (left panel) and MCF7 (right panel) cells treated with S100A8/A9 (100 μg/ml) for 0 (C) and 24 h. Western blot was performed using specific antibodies toward activated (cleaved fragments) caspase-3, caspase-7, and caspase-9, PARP-1, and BID. Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was used as loading control. (H-K) Luminometric caspase activity assay. White bars represent controls, whereas black bars represent cells treated with 100 μg/ml of S100A8/A9 for indicated periods of time. In an experiment parallel to the one depicted in (G), caspase-8 and caspase-3/caspase-7 activity in MCF7 (H, J) and SHEP (I, K) cells were measured by the Caspase-Glo luminometric assay. The caspase activity is represented as a luminescence activity of each sample. The data represent duplicates of two independent experiments. S100A8/A9 did not induce any significant increase in Caspase-8 activity in MCF7 and SHEP cells (H, I) (P > 0.05), while increased caspase-3/caspase-7 activity was significantly observed at all indicated time points (J, K) (P < 0.05).
Several caspases were investigated to elucidate whether S100A8/A9-triggered cell death involved caspase activation. S100A8/A9 did not induce caspase-8 activation in MCF7 and SHEP cells (Figure 1H and 1I). BID cleavage is a hallmark of either a death receptor to mitochondrial apoptotic signaling or a caspase-9/caspase-3/caspase-6/caspase-8 amplification loop. In our experimental settings, S100A8/A9 did not trigger the cleavage of BID into a 15-kDa tBID fragment in SHEP and MCF7 cells (Figure 1G). The above findings indicate that S100A8/A9-induced apoptosis was independent of the extrinsic receptor-mediated pathways that involve caspase-8.
S100A8/A9 treatment activated caspase-9 and subsequently caspases-3 and caspase-7 resulting in poly(ADP-ribose) polymerase-1 (PARP-1) cleavage in SHEP cells. As expected, we did not observe caspase-3 activation in MCF7 cells, although S100A8/A9 did induce caspase-7 activation and PARP-1 cleavage (Figure 1G). MCF7 cells are known to lack caspase-3; however, they still undergo mitochondria-dependent apoptosis and PARP-1 cleavage via caspase-7 activation 30. Caspase-7, analogous to caspase-3, is activated via the Apaf-1-caspase-9/apoptosome-dependent pathway and it may cleave PARP-1 on apoptosis induction 31, 32, 33. Caspase-3 and caspase-7 activation in the investigated cell lines were also confirmed by a luminometric assay (Figure 1J and 1K).
S100A8/A9 induces autophagy in apoptotic cells
Autophagy involves the sequestration of cytosol or cytoplasmic organelles into double-layered vesicles, called autophagosomes. Autophagosomes subsequently fuse with endosomes and eventually with lysosomes, thereby creating autophago-lysosomes or autolysosomes. In the lumen of these latter structures, lysosomal enzymes operate at low pH and catabolize the autophagic material 34, 35.
To investigate autophagy, SHEP cells were treated with S100A8/A9 (100 μg/ml) for 24 h and studied by transmission electron microscopy (TEM). S100A8/A9-treated SHEP cells, but not untreated cells (Figure 2A), displayed both apoptotic and autophagic ultrastructural characteristics, like pyknotic chromatin, disintegrating nuclear membrane, and cytosolic autophagosomes (Figure 2B). At higher magnification, cytosolic autophagosomes (Figure 2D) and vacuolization (Figure 2C) were clearly visible. The so called Atg genes, involved in the process of autophagosome formation featuring two ubiquitin-like conjugation systems, are well-conserved among eukaryotes. Those are the Atg12-Atg5 and the Atg8/LC3-PE1 (phosphatidylethanolamine) systems 34. Atg12-Atg5 conjugation is a constitutive process since the conjugate Atg12-Atg5 is formed immediately after Atg12 and Atg5 synthesis, independently of starvation or other autophagy-inducing conditions. Free forms of Atg12 and Atg 5 are rarely observed 36, 37, 38. Atg8/LC3 is cleaved by Atg4 (autophagin) to produce the active cytosolic form LC3-I (18 kDa), which is subsequently activated by Atg7, transferred to Atg3, and modified into the active form LC3-II (membrane-bound) that interacts and conjugates with PE 37, 39, 40. Atg6 (and its mammalian ortholog Beclin-1) belong to the class III PI3-kinase complexes, and participate in the regulation of early stages of autophagosome formation 41, 42, 43.
Treatment with S100A8/A9 induces typical hallmarks of autophagy in dying cells. SHEP cells were either left untreated (A) or treated with 100 μg/ml S100A8/A9 (B-D) for 24 h. Cells were then analyzed by Transmission Electron Microscopy (TEM). Magnification: 4.6 × 103 (A, B), 6.4 × 103 (C), and 11.5 ×103 (D). (E) S100A8/A9 induced LC3-β cleavage, increase of Atg12-Atg5 formation, and Beclin-1 expression. Western blot analysis of cell lysates of SHEP and MCF7 cells treated with S100A8/A9 (100 μg/ml) for indicated time intervals using the corresponding specific antibodies. Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was used as loading control. (F) MCF-7 cells were cultured in the absence and presence of S100A8/A9 (100 μg/ml) for 12 h, and the corresponding cell lysates were incubated with specific anti-Bcl2 antibody followed by western blot analysis using specific anti-Beclin-1 antibody. S100A8/A9 treatment induced interaction between Bcl2 and Beclin-1 (right panel), while there was no interaction detectable in the absence of S100A8/A9 (left panel).
We investigated the expression pattern of LC3-I (18 kDa) and LC3-II (16 kDa), Atg12-Atg5 formation, and Beclin-1 expression in MCF7 and SHEP cells after treatment with S100A8/A9 (100 μg/ml) for 24 h using the corresponding specific antibodies as indicated in the Materials and Methods section. As shown in Figure 2E, the levels of LC3-II protein, Atg12-Atg5 formation, and Beclin-1 expression were increased in MCF7 and SHEP cells after exposure to S100A8/A9. These data indicate that S100A8/A9 stimulated the conversion of a significant fraction of LC3-I to LC3-II. To confirm our data, MCF-7 cells were treated with 100 μg/ml S100A8/A9 for 12 h, and Bcl2-Beclin-1 interaction was investigated by co-immunoprecipitation. As shown in Figure 2F (right panel), S100A8/A9 treatment increased Beclin-1 and Bcl2 interaction. In the absence of S100A8/A9 there was no detectable interaction between these two proteins (Figure 2F, left panel).
S100A8/A9-induced cell death is partially reversed by inhibition of PI3-kinase or vacuolar H+-ATPase pump, cathepsin inhibitors, and ATG5 shRNA
Certain forms of apoptosis, e.g. that induced by apoptin, could be efficiently counteracted by the inhibition of PI3-kinase/Akt pathway 44, 45. Similarly, autophagy could be blocked by the inhibition of PI3-kinase and the vacuolar H+-ATPase pump 46. Therefore, we analyzed S100A8/A9-induced cell death in the absence and presence of the class III PI3-kinase inhibitor 3-MA (3-methyladenine) (5 and 10 mM) and the lysosomal hydrogen pump inhibitor bafilomycin-A1 (Baf-A1) (0.05 and 0.1 μM). MTT assays showed that both inhibitors significantly suppressed S100A8/A9-induced cell death in MCF7 (Figure 3A and 3B) and SHEP cells (Figure 3C and 3D) (P < 0.01). In addition, Baf-A1 also inhibited LC3 II formation in SHEP cells treated with S100A8/A9 (Figure 3E). These data confirmed the role of the lysosomal pathway in S100A8/A9-induced autophagy.
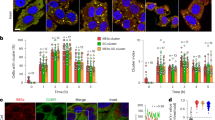
Inhibition of S100A8/A9-induced cell death by PI3-kinase inhibitor 3-MA, and the vacuolar H+-ATPase inhibitor, bafilomycin-A1 (Baf-A1). MCF7 (A, B) and SHEP cells (C, D) were treated for 3 h with 3-methyladenine (3-MA) (A, C) and Baf-A1 (B, D) as indicated, and with S100A8/A9 (100 μg/ml, 24 h). 3-MA and Baf-A1 pre-treatment significantly decreased cytotoxicity of S100A8/A9 in both cell lines (P < 0.01). (E) Baf-A1 inhibited S100A8/A9-induced-LC3-β cleavage. SHEP cells were treated with S100A8/A9 in the absence (right panel) and presence (left panel) of Baf-A1 (0.1 μM) for different time intervals as indicated. Then LC3-β cleavage was investigated by western blot. Baf-A1 inhibited S100A8/A9-induced LC3-β cleavage. (F, G) ATG5 shRNA decreased S100A8/A9-induced cell death. MCF-7 cells infected with either ATG5 scrambled shRNA (left panel) or ATG5 shRNA (right panel) were treated with S100A8/A9 (100 μg/ml) for indicated time intervals. Then western blot analysis of cell lysates was performed with specific anti-ATG5-Atg12 antibody. Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was used as loading control (F). ATG5 scrambled or shRNA infected MCF-7 cells were treated with S100A8/A9 (100 μg/ml) for 24 h, and cell viability was measured using MTT assay (G). ATG5 shRNA infection significantly inhibited S100A8/A9-induced cell death (P < 0.001). Results are expressed as percentage of corresponding control and represent the mean ± SD of three independent experiments. (H-J) S100A8/A9-induced cell death involved lysosomal activation. (H) MCF7 and L929 cells were treated with S100A8/A9 (100 μg/ml, 24 h) and stained with the acidophilic lysosomal probe lysotracker red (LTR). S100A8/A9 caused an increase in the volume and frequency of cytoplasmic granules staining with LTR. (I, J) S100A8/A9-induced cell death was inhibited by cathepsin B and L inhibitors. MCF7 cells were pretreated with CA-074 ME or Z-FF-FMK for 2 h and then cultured in the presence and absence of S100A8/A9 (100 μg/ml) for 24 h. S100A8A8/A9-induced cell death was significantly (P < 0.001) inhibited by these inhibitors. Results are expressed as percentage of corresponding control and represent the mean ± SD of three independent experiments. (K, L) Co-localization of the autophagic marker LC3-β and mitotracker red (MTR) in S100A8/A9-treated cells. MCF7 cells were treated with S100A8/A9 (100 μg/ml, 24 h) followed by staining with MTR (15 min) and then immunostained against LC3-β (FITC conjugated, green) and analyzed by confocal scanning fluorescence microscopy. LC3-β showed higher co-localization with mitochondria in S100A8/A9-treated cells (lower panel) compared to control cells (upper panel). At higher magnification, LC3-β showed a punctated structure in S100A8/A9-treated cells (right panels). (L) LC3-β cleavage was confirmed by immunoblotting of mitochondrial fraction lysates from S100A8/A9-treated cells. MnSOD was used as loading control.
In another approach, Atg5 expression was inhibited in MCF-7 cells by ATG5 shRNA followed by treatment with 100 μg/ml S100A8/A9 for different time intervals as indicated (Figure 3F). Inhibition of Atg5 expression significantly inhibited S100A8/A9-induced cell death in MCF-7 cells (Figure 3G) (P < 0.001).
These data confirmed that autophagic death is involved in S100A8/A9-induced cell death. However, since inhibitors of autophagy did not completely reverse S100A8/A9-induced cell death, we conclude that autophagy is not the exclusive cell death mechanism involved. This is consistent with previous research demonstrating that apoptosis is also involved in S100A8/A9-induced cell death 9, 12, 47, 48.
The participation of lysosomes in autophagic cell death was further confirmed by the analysis of lysosomal activation using lysotracker red (LTR). S100A8/A9 treatment of MCF7 and L929 caused an increase in both volume and frequency of cytoplasmic granules staining with LTR (Figure 3H).
Cathepsins are one of the largest groups of enzymes within the lysosomes. The most abundant are the cysteine cathepsins, which comprise a group of 11 related enzymes in human (B, C, F, H, K, L, O, S, V, W, and X), and the aspartic protease cathepsin D 49, 50. Because cathepsins B, L, and D are abundant among the lysosomal proteases, they are often used as markers of lysosomal activation 51. In order to validate the impact of lysosomal involvement in S100A8/A9-induced cell death, the two cathepsins B and L inhibitors CA-074-ME and Z-FF-FMK were tested in the presence of S100A8/A9. As shown in Figure 3I and 3J, both cathepsin inhibitors significantly inhibited S100A8/A9-induced cell death in MCF-7 cells (P < 0.001).
Next, the co-localization of the mitochondrial marker mitotracker red (MTR) and the autophagosomal marker (LC3-β) were measured in SHEP cells after treatment with S100A8/A9 (100 μg/ml) for 24 h. After exposure to S100A8/A9, levels of LC3-β were significantly increased. At higher magnification, LC3-β fluorescence displayed a punctated pattern that is a characteristic feature of autophagy (Figure 3K, right panels). Three fields were randomly counted, and 7 of 10 cells showed LC3-β punctation in each field. LC3-β also co-localized with mitochondria in these cells, indicating the fusion of mitochondria with AV (Figure 3K). In addition, LC3-β co-localization with mitochondria was confirmed by immunoblotting of mitochondrial fraction lysates from S100A8/A9-treated cells (Figure 3L).
ΔTM-BNIP3 overexpression partially inhibits S100A8/A9-induced cell death, ROS production, and lysosomal activation
BNIP3 is an atypical pro-apoptotic Bcl2-family member that has a single BH3 domain and a C-terminal trans-membrane (TM) domain. Although it belongs to the Bcl2-family, its pro-cell death activity is distinct from those of other family members 52. BNIP3 is too toxic to develop stably transfected cells. Cells transiently transfected with BNIP3 exhibit early plasma membrane permeability, mitochondrial damage, extensive cytoplasmic vacuolation, and mitochondrial autophagy, accompanied by rapid and profound mitochondrial dysfunction characterized by opening of the mitochondrial permeability transition pore and increased ROS production 51, 53. Some levels of BNIP3 expression could be detected in MCF7, SHEP, HEK-293, and L929 cells (data not shown). We have thus investigated whether BNIP3 plays a role in S100A8/A9-induced cell death. ΔTM-BNIP3 (a dominant-negative BNIP3 mutant lacking the TM) overexpression was previously used as a model for functional study of BNIP3. Those studies have shown that ΔTM-BNIP3 antagonizes wild-type (wt) BNIP3-induced effects including mitochondrial dysfunction and cell death 51, 54, 55. The TM domain is required for homodimerization and normal mitochondrial localization, so overexpression of the dominant-negative BNIP3-ΔTM would prevent wt BNIP3 from targeting the mitochondria 54.
The cell death-inducing activity of S100A8/A9 was tested in both L929 and HEK-293 cells overexpressing ΔTM-BNIP3. As shown in Figure 4A, the cells overexpressing ΔTM-BNIP3 were significantly more resistant to S100A8/A9-induced cell death (P < 0.05) compared to the corresponding wt cells. Since ΔTM-BNIP3 overexpression reversed the cell death-inducing activity of S100A8/A9, we investigated ROS production (Figure 4B and 4E) and loss of mitochondrial membrane potential (ΔΨm) using dihydrorhodamine-123 (DHR-123) and 5,5′,6,6′-tetrachloro-1,1′,3,3′-tetraethylbenzimidazolylcarbocyanine iodide (JC-1), respectively (Figure 4C and 4D). S100A8/A9 caused a rapid increase of ROS production and decrease of ΔΨm in wt L929 cells, whereas ROS production and mitochondrial depolarization were significantly reduced in ΔTM-BNIP3-L929 and ΔTM-BNIP3-MCF7 cells.
ΔTM-BNIP3 and antioxidant N-acetyl cysteine protect cells from S100A8/A9-induced cell death. (A) L929 cells overexpressing ΔTM-BNIP3 or Hek-293 overexpressing ΔTM-BNIP3, and the corresponding wt cells were treated with S100A8/A9 (100 μg/ml) for different time intervals as indicated. Cell viability was assessed by MTT assay. Results are expressed as percentage of corresponding control and represented the means ± SD of four repeats. (B, E) ΔTM-BNIP3 overexpression diminished ROS production in S100A8/A9-treated cells. S100A8/A9 (100 μg/ml) triggered marked ROS production in L929 (C) and MCF-7 (F) cells, whereas S100A8/A9 treatment in L929 and MCF-7-ΔTM-BNIP3 cells elicited reduced ROS generation (P < 0.05). ROS was measured using dihydrorhodamine-123. The experiment was repeated four times and the average ROS values are indicated. (C, D) Presence of ΔTM-BNIP3 prevents destabilization of mitochondrial membrane potential (ΔΨm) in L929 (D) and MCF-7 (E) cells on S100A8/A9 treatment. Ψm was determined using the JC-1 fluorescent dye. ΔTM-BNIP3 overexpression significantly protected cells (P < 0.05 for 4 and 8 h and P < 0.01 for 12 h) against a decrease in Ψm when exposed to S100A8/A9 (100 μg/ml). (F) ROS scavenging inhibited S100A8/A9-induced autophagy. MCF-7 cells were pre-treated with NAC (5 mM) and then cultured in the presence and absence of S100A8/A9 (100 μg/ml) for indicated time intervals. Western blot analysis of cell lysates was performed using specific anti-Atg5-Atg12 and anti-LC3-β antibodies. NAC prevented LC3-β cleavage and Atg5-Atg12 formation. Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was used as loading control. (G) ΔTM-BNIP3 overexpression inhibited S100A8/A9-induced autophagy. MCF-7 and MCF-7-ΔTM-BNIP3 overexpressing cells were treated with S100A8/A9 (100 μg/ml) for indicated time intervals followed by western blot analysis using specific anti-Atg5-Atg12 and anti-LC3-β antibodies. ΔTM-BNIP3 overexpressing prevented LC3-β cleavage and Atg5-Atg12 formation (hallmarks of autophagy). GAPDH was used as loading control. (H) Cellular localization of BNIP3 in S100A8/A9-treated SHEP cells. SHEP cells were incubated with S100A8/A9 (100 μg/ml) for 12 h, and subsequently immunostained with anti-BNIP3 and Cy-5-conjugated secondary antibody (magenta), mitotracker red (mitochondria, red), and DAPI (nucleus/DNA, blue). (I) S100A8/A9-treatment induced mitochondrial translocation of BNIP3 in SHEP cells. Cells were treated with 100 μg/ml S100A8/A9 (12 h and 24 h), harvested, and fractionated, and BNIP3 was detected by western blot in mitochondrial and nuclear fraction. HDAC1 (nuclear protein) and MnSOD2 (mitochondrial protein) were used as controls to check the purity of mitochondrial and nuclear fractions, respectively. (J) N-acetyl-L-cysteine (NAC) decreased lysosomal damage in S100A8/A9-treated cells. MCF7 cells were treated with S100A8/A9 (100 μg/ml, 24 h) in the absence (lower panel) or presence of 5 mM NAC (middle panel) followed by staining with the acidophilic lysosomal probe lysotracker red (LTR). (K) ΔTM-BNIP3 overexpression protected the cells against lysosomal damage in S100A8/A9-treated cells. L929 and L929 ΔTM-BNIP3 overexpressing cells were treated with S100A8/A9 (100 μg/ml for 24 h) and stained with the acidophilic lysosomal probe LTR. S100A8/A9 caused an increase in volume and frequency of cytoplasmic granules staining with LTR, which was inhibited in the presence of ΔTM-BNIP3.
We have previously shown that ROS scavenging blocks S100A8/A9-induced apoptosis in HT29/219 and SW742 colon carcinoma cell lines 12, and it has also been reported that ROS could regulate autophagy in various cell models 56, 57. Therefore, we investigated whether ROS scavenging could inhibit S100A8/A9-promoted autophagy.
As shown in Figure 4F, the appearance of autophagy hallmarks (LC3-β cleavage and Atg12-Atg5 formation) was inhibited in MCF7 cells co-treated with N-acetyl cysteine (NAC) (5 mM) and S100A8/A9 (100 μg/ml). Moreover, we also showed that ΔTM-BNIP3 overexpression inhibited autophagy (Figure 4G), which is in accordance with previous studies 58, 59.
BNIP3 needs to integrate into the outer mitochondrial membrane in order to induce cell death 52. This prompted us to examine the subcellular location of BNIP3 after S100A8/A9 treatment by confocal imaging and cell fractionation followed by immunoblotting. When SHEP cells were treated with S100A8/A9, BNIP3 translocated to the mitochondria (Figure 4H). Consistent with this finding, probing of mitochondrial fractions by western blotting indicated the enhanced association of BNIP3 with mitochondria after S100A8/A9-induced apoptosis (Figure 4I). HDAC1 and MnSOD-2 served as nuclear and mitochondrial markers, respectively (quality control of subcellular fractionation).
We next investigated the role of ROS in the cross-talk between mitochondria and lysosomes, and in subsequent lysosomal activation. First, we compared lysosomal activation in MCF7 cells treated with S100A8/A9 alone and in those pretreated with a clinically used antioxidant NAC (5 mM) for 4 h followed by exposure to S100A8/A9. As shown in Figure 4J, lysosomal activation was significantly reduced if the cells were pretreated with NAC, as compared to the corresponding controls. Second, we studied lysosomal activation in wt L929 cells and L929 cells overexpressing ΔTM-BNIP3 after treatment with S100A8/A9. Lysosomal activation was nearly blocked in S100A8/A9-treated L929 cells overexpressing ΔTM-BNIP3 as compared to their corresponding wt controls (Figure 4K). These data provide strong evidence that BNIP3 plays an important role in the S100A8/A9-induced cell death pathway, and that the mitochondria-lysosome cross-talk is mediated by ROS.
Discussion
We have investigated here the molecular mechanisms of S100A8/A9-triggered cell death 9, 12. S100A8/A9 causes a rapid drop in mitochondrial membrane potential, triggers Bak and BNIP3 mitochondrial translocation, induces selective release of Smac/DIABLO and Omi/HtrA2 from mitochondria, and decreases Drp1 expression. Previously published experiments involving both RAGE knockdown and blocking RAGE with specific antibody, excluded the involvement of RAGE in S100A8/A9-induced cell death 9. Thus, the apoptosis-inducing property of S100A8/A9 appeared to be mediated by a yet-to-be identified receptor. We have previously shown that the mitochondrial pathway plays an important role in S100A8/A9-induced cell death 9. Here we show for the first time, that autophagy is also triggered by the S100A8/A9 treatment.
TEM analysis confirmed the presence of ultrastructural characteristics of apoptosis and autophagy in SHEP cells treated with S100A8/A9 (Figure 2A-2D). Autophagosome formation requires two ubiquitin-like conjugation systems, the Atg12 and Atg8 systems, which are tightly associated with the expansion of autophagosomal membrane. Atg12 is conjugated to Atg5 and forms an ∼800 kDa protein complex with Atg16L (referred to as the Atg16L complex). The Atg16L complex contributes to the expansion of autophagosomal membrane by promoting Atg8 lipidation. Microtubule-associated protein 1–light chain 3 (LC3) is the mammalian homolog of the yeast protein Atg8. Upon synthesis, LC3 is processed to its cytosolic form LC3-I that is subsequently conjugated to the lipid phosphatidylethanolamine, generating the LC3-II form. Conjugation to this lipid is required for its association with the autophagosomal membrane. Consistent with these data, we found that the levels of LC3-II and Atg12-Atg5 conjugates were increased after exposure to S100A8/A9 (Figure 2E). This increase could be counteracted by Baf-A1 treatment in our experimental system. While our data on the abundance of LC3-II upon Baf-A1 treatment differs from some previous reports, others also became aware of similar observations (Klionsky, personal communication).
Aside from its role as a waste disposal system, clearing organelles that accumulate from the cytoplasm, autophagy could be involved in clearing proteins and organelles during oxidative stress 60. This may become particularly important in the presence of damaged mitochondria, a circumstance leading to decreased production of ATP and increased accumulation of toxic ROS. LC3-II is an autophagosomal marker in mammals. We could demonstrate that LC3-II co-localized with mitochondria and lysosomes (data not shown) upon treating the cells with S100A8/A9. These data were confirmed by the inhibition of PI3 kinase and vacuolar H+-ATPase pump, which significantly reduced S100A8/A9-induced cell death (Figure 3A-3D) and lysosomal activation (Figure 3E-3L). Thus, S100A8/A9-induced autophagy may play a role in the removal of damaged mitochondria.
Recently published data indicate that apoptosis and autophagy are interconnected, at least under some experimental settings. Pharmacological and genetic inhibition of autophagy delays or partially inhibits cell death in specific conditions 61. Two recent studies provided the first genetic evidence for the involvement of the autophagy pathway in cell death. Gene silencing of atg7 and atg6/beclin 1 blocked cell death in mouse L929 cells treated with the caspase inhibitor zVAD 62, and gene silencing of atg5 and atg6/beclin 1 inhibited cell death of bax−/−bak−/− murine embryonic fibroblasts treated with staurosporine or etoposide 63.
Further evidence for a link between apoptosis and autophagy is given by the recent report that the Bcl2 anti-apoptotic protein inhibits Beclin-1-dependent autophagy 64. Although the role of Bcl2 as autophagy modulator is still in debate, we showed that S100A8/A9 increased Beclin-1 and Bcl2 interaction (Figure 2F), which is in parallel with an increase of autophagy. Thus, the Bcl2/Beclin-1 association may be a rheostat maintaining autophagy at levels that are compatible with cell survival rather than cell death. In a previous report, we have shown that S100A8/A9 at higher concentration induced decrease of Bcl2 expression in treated cells 9, and in the present study, we demonstrated that S100A8/A9 increased Beclin-1 expression and interaction with Bcl2 in the target cells. Therefore, it is likely that the Bcl2/Beclin-1 counterbalance might be a rheostat regulating S100A8/A9-induced autophagy.
BNIP3 is a pro-apoptotic Bcl2-family member. A mutant of BNIP3, ΔTM-BNIP3, lacking the transmembrane domain, blocked BNIP3-induced cell death 51, 65, and as shown here, reversed S100A8/A9-induced cell death, autophagy, and lysosomal activation (Figure 4). S100A8/A9 caused a rapid increase of ROS production and decrease of Ψm in wt L929, whereas ΔTM-BNIP3 overexpression reduced mitochondrial damage. BNIP3 directly interacts with Bcl2 and Bcl-XL, suggesting that BNIP3 could activate Bax and Bak through an indirect mechanism, by sequestering Bcl2 and Bcl-XL 66. Since S100A8/A9, when applied at higher concentration, decreased Bcl2 and Bcl-XL expression in treated cells 9 and this was accompanied by an activation of Bax and Bak, we propose that BNIP3 translocation to mitochondria with simultaneous decrease of Bcl2 and Bcl-XL expression represents a mechanism for Bax and Bak activation in S100A8/A9-treated cells.
Besides its role in removing mitochondria damaged by oxidative stress, autophagy may also play a role in the catabolism of oxidized proteins, in particular in the resolution of large oxidized protein aggregates 67, 68. A role for autophagy in response to ROS is indicated by the accumulation of oxidized proteins during aging 56, 68 and in age-related disorders, such as Alzheimer's disease 69 and diabetes mellitus 70, where autophagic pathways are known to be compromised. Interestingly, BNIP3-ΔTM overexpression also partially inhibited ROS production (Figure 4B and 4E), and vice versa, lysosomal activation was reduced in the presence of the anti-oxidant NAC (Figure 4F and 4J). On the other hand, S100A8/A9 treatment induces ROS production in cells 12, 71, ROS scavenging inhibited S100A8/A9-induced autophagy (Figure 4F), and ΔTM-BNIP3 overexpression delayed S100A8/A9-induced autophagy. Therefore, we conclude that S100A8/A9 induces BNIP3 translocation to mitochondria with subsequent increase in ROS production, which in turn causes autophagy, cell death, and lysosomal activation.
In conclusion, the present study shed new light on S100A8/A9-triggered cell death. We provide the first evidence that S100A8/A9 induces both apoptosis and autophagy. S100A8/A9-induced cell death involves BNIP3 and increase of ROS production by mitochondria, subsequently followed by mitochondrial damage and lysosomal activation. Finally, we suggest ROS as the critical factor that integrates S100-induced mitochondrial and lysosomal death pathways.
Materials and Methods
Materials and reagents
Cell culture media were purchased from Sigma (Oakville, ON, Canada) and Gibco (Canada). Cell culture plastic ware was obtained from Nunc Co. (Canada). Caspase-Glo-8, and caspase-3/caspase-7 assay systems were purchased from Promega, Nepean, ON, Canada. Rabbit anti-human Bcl2, rabbit anti-human cleaved caspase-8, caspase-9, caspase-6, caspase-7, PARP-1, rabbit anti-human Atg12, and rabbit anti-human Beclin-1 were purchased from Cell Signaling (Canada). Mouse anti-human BNIP3, rabbit anti-human LC3-β, 3-MA, Baf-A1, CA-074ME, Z-FF-FMK, puromycin, and MTT were obtained from Sigma. Rabbit anti-human/mouse/rat glyceraldehyde-3-phosphate dehydrogenase (GAPDH) and mouse anti-human/mouse/rabbit histone deactylase 1 (HDAC1) were obtained from Santa Cruz (USA). Mouse anti-human/mouse/rat super-dismutase 2 (MNSOD2) was obtained from R&D systems (Canada). JC-1, MTR, and LTR were obtained from Invitrogen Molecular Probes (Canada). ATG5 shRNA was obtained from Open Biosystem (Manitoba, Canada).
Purification of S100A8 and S100A9
Human neutrophils were prepared from leukocyte-rich blood fractions (“buffy coat”). S100A8/A9 was purified as described earlier 72. Prior to use, the proteins were re-chromatographed by anion exchange using a UnoQ column (BioRad, Munich, Germany). SDS-PAGE revealed a purity of > 95%. Recombinant protein was produced by bacterial overexpression as previously described 73. All experiments were performed first with human neutrophil S100A8/A9, and then results were confirmed using recombinant human S100A8/A9.
Cell culture
MCF-7 (human, estrogen receptor-positive breast cancer), L929 (rodent fibrosarcoma), L929-ΔTM-BNIP3 overexpressing, HEK-293 (human embryonic kidney), HEK-293-ΔTM-BNIP3 overexpressing 54, and SHEP (human neuroblastoma) cells were cultured in RPMI-1640 and DMEM (L929, HEK-293) media supplemented with 10% fetal calf serum, 100 U/ml penicillin, and 100 μg/ml streptomycin. Cells were incubated at 37 °C in a humidified atmosphere of 5% CO2 and 95% air. Cell cultures were maintained under logarithmic growth conditions.
MTT assay
The cytotoxicity of S100A8/A9 (alone or in presence of 3-MA and Baf-A1) toward the indicated above cell lines was determined by MTT assays, as previously described 54, 74. The percentage cell viability was calculated using the equation: (mean OD of treated cells/mean OD of control cells) ×100. For each time point, the treated cells were compared with control cells that had been treated only with medium and PBS (solvent of S100A8/A9).
Measurement of apoptosis by flow cytometry
Apoptosis was measured using the Nicoletti method 29, 54. Briefly, cells grown in 12-well plates were treated with S100A8/A9 (100 μg/ml) for the indicated time intervals. After scraping, the cells were harvested by centrifugation at 800× g for 5 min, washed once with PBS, and resuspended in hypotonic propidium iodide (PI) lysis buffer (1% sodium citrate, 0.1% Triton X-100, 0.5 mg/ml RNase A, 40 μg/ml PI). Cell nuclei were incubated for 30 min at 30 °C and subsequently analyzed by flow cytometry. Nuclei to the left of the G1 peak containing hypodiploid DNA were considered apoptotic.
Luminescence caspase activity assays
Luminometric assays Caspase-Glo-8, caspase-9 and caspase-3/caspase-7 (Promega) were used to measure the proteolytic activity of caspase-3/caspase-7 (DEVD-ase), caspase-8 (IETD-ase), and caspase-9 (LEHD-ase) 51. The assays were performed according to the manufacturer's instructions. Briefly, cells sub-cultured in a 96-well plate (15 000 cells per well) were treated with S100A8/A9 (100 μg/ml) at different time points using freshly prepared caspase reagents containing whole protein cell lysate extract buffer and either z-DEVD-luciferin or z-LETD-Luciferin. In each experiment, negative control cells or cells treated with medium only and blank reagent were included. Plates were gently shaken at 300-500 r.p.m. for 30 sec and incubated for 30 min at room temperature. Then, the solution was transferred to a white-well plate and the luminescence of each sample was measured and compared to the negative controls (Lmax, Molecular Devices).
Short hairpin RNA protocol
The ATG5 shRNA construct was obtained from Open Biosystems (clone V2LHS 195828) distributed by the Manitoba Centre for Proteomics and Systems Biology as a bacterial culture (DH5α). The shRNA constructs include a hairpin of 21 base pair sense and antisense stem and a 6 base pair loop. The hairpin sequence was cloned into the lentiviral vector pGIPZ. Individual colonies were grown in LB broth with Ampicillin (100 μg/ml) and purified by a maxi-prep kit from Qiagen. A VSVG pseudo-typed retrovirus was made with the purified DNA, a packaging vector 8.2Δvpr, and envelope vector VSVG. After 3 days, the supernatant was collected and concentrated by ultracentrifugation (17 000× g, 90 min). The virus was tittered in 293T cells with a MOI of 0.5–10. MCF-7 cells were grown to 70% confluence and then transduced at a MOI of 6. Transfected cells were then selected with 4 μg/ml puromycin for 3 weeks. In tandem, a negative control vector pGIPZ.eGFP was transfected into 293T cells and transduced into the same MCF-7 cell line.
Immunoprecipitation
Cells were washed twice with cold PBS, lysed with ice-cold lysis buffer, incubated for 30 min on ice, and centrifuged for 10 min at 4 °C. Immunoprecipitation was performed using Bcl2 antibody, and the immune complexes were captured with protein A-agarose beads (Amersham Biosciences, Piscataway, NJ, USA). After three washes with cell lysis buffer, bead-bound proteins were subjected to sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and analyzed by western blot analysis according to standard protocols 75. In parallel, interaction of Beclin-1 and Bcl2 was analyzed in cells incubated with a solvent control.
Immunoblotting
Bcl2, Bcl-XL, BNIP3, ATG5-ATG 12, LC3-β, cleaved caspase-3, caspase-9, caspase-7, caspase-6, PARP-1, Bid, and Beclin-1 protein content was determined in lysates of SHEP and MCF7 cells that had been treated with 100 μg/ml S100A8/A9 for different time intervals. Briefly, the harvested cells were washed once with cold PBS and re-suspended for 20 min on ice in a lysis buffer containing 20 mM Tris-HCl (pH 7.5), 0.5% Nonidet P-40, 0.5 mM PMSF, and 0.5% protease inhibitor cocktail (Sigma). The high-speed supernatant (10 000× g) was collected and proteins (30 μg) were separated by SDS-PAGE and transferred onto nitrocellulose membranes. Membranes were blocked in 5% non-fat dry milk in Tris-buffered saline-Tween 1% (TBS; 0.05 M Trizma base, 0.9% NaCl, and 1% Tween-20) and incubated overnight with the primary antibodies at 4 °C. The membranes were incubated at room temperature for 1 h with the relevant secondary antibodies conjugated to HRP and blots were developed by enhanced chemiluminescence detection (Amersham-Pharmacia Biotech).
Measurement of ROS production
L929, L929-ΔTM-BNIP3, MCF-7, and MCF-7-ΔTM-BNIP3 cell lines (1.5 × 104) were treated with S100A8/A9 (100 μg/ml) for different time points. DHR-123 (1 μM) was added to treated cells at 37 °C for 15 min before cells were harvested and washed three times with ice-cold PBS. Cells were left on ice for 15 min to stabilize fluorescence. The fluorescence intensity (FL-1 channel) was then measured by flow cytometry (FACS-Calibur, BD) 51.
Mitochondrial membrane potential assay
The assay was performed using a mitochondria-specific cationic dye (JC-1), which undergoes potential-dependent accumulation in mitochondria. JC-1 exists as a monomer when the membrane potential (Ψm) is lower than 140 mV and emits green light (540 nm) after excitation by blue light (490 nm) 9. At higher membrane potentials, JC-1 monomers are converted to aggregates that emit red light (590 nm) after excitation by green light (540 nm). Normal L929, and 293 cells and ΔTM-BNIP3 overexpressing L929 and 293 cells were seeded in black clear-bottom 96-well plates. Following treatment with 100 μg/ml S100A8/A9 for different time intervals as indicated, cells were loaded with JC-1 by replacing the culture medium with HEPES buffer (40 mM, pH 7.4) containing 4.5 g/l glucose (high glucose medium) or 1.5 g/l glucose (low glucose medium), 0.65% NaCl, and 2.5 μM JC-1 for 30 min at 37 °C, then washed once with HEPES buffer. Fluorescence was measured after a further 90 min (this time period is sufficient for JC-1 to equilibrate between the cytosol and the mitochondrial compartments, as ascertained in preliminary experiments) using a fluorescence plate reader that allows for the sequential measurement of each well at two excitation/emission wavelength pairs, 490/540 and 540/590 nm. Changes in the ratio between the measured red (590 nm) and green (540 nm) fluorescence intensities indicate changes in mitochondrial membrane potential. This ratio was calculated for each well after the fluorescence intensity of wells containing medium and serum without cells was subtracted. The ratio of red to green fluorescence in the same culture exclusively depends on the mitochondrial membrane potential and is independent of factors such as cell number and mitochondrial size, shape, and density.
Cell fractionation
Following induction of apoptosis using S100A8/A9 (100 μg/ml), cytosolic, mitochondrial, and nuclear fractions were generated using a digitonin-based subcellular fractionation technique, essentially as described previously 76. Briefly, 107 cells were harvested by centrifugation at 800× g, washed in PBS (pH 7.2), and re-centrifuged. Cells were digitonin permeabilized for 5 min on ice at a density of 3 × 107 cells per ml in cytosolic extraction buffer (250 mM sucrose, 70 mM KCl, 137 mM NaCl, 4.3 mM Na2HPO4, 1.4 mM KH2PO4 (pH 7.2), 100 μM PMSF, 10 μg/ml leupeptin, 2 μg/ml aprotinin, containing 200 μg/ml digitonin). Plasma membrane permeabilization of cells was confirmed by staining in a 0.2% trypan blue solution. Cells were then centrifuged at 1 000× g for 5 min at 4 °C. The supernatants (cytosolic and mitochondria fractions) were saved and the pellets solubilized in the same volume of nuclear lysis buffer, followed by centrifugation at 12 500× g for 10 min at 4 °C. The mitochondria was separated from the cytosolic fraction by centrifugation at 13 000× g, and the pellets were solubilized in equal volume of mitochondrial lysis buffer (50 mM Tris (pH 7.4), 150 mM NaCl, 2 mM EDTA, 2 mM EGTA, 0.2% Triton X-100, 0.3% NP-40, 100 μM PMSF, 10 μg/ml leupeptin, 2 μg/ml aprotinin). For the detection of proteins, equal amounts of protein were supplemented with 5× SDS-PAGE loading buffer, subjected to 12% SDS-PAGE, and transferred to nitrocellulose membranes.
Immunocytochemistry, confocal imaging and electron microscopy
Cells were grown overnight on coverslips and treated with 100 μg/ml S100A8/A9. After 24 h, cells were washed with PBS, fixed in 4% paraformaldehyde, and permeabilized with 0.1% Triton X-100. To co-localize LC3-β, a primary human anti-rabbit LC3-β antibody (1:200 dilution) and the corresponding FITC-conjugated secondary antibody (Sigma, 1:100 dilutions) were used. Mitochondria and lysosomes were stained with MTR CMXRos (200 nM) and LTR (1:2 500; Molecular Probe), respectively, in culture medium for 15 min prior to fixation with paraformaldehyde (4%). The fluorescent images were analyzed using an Olympus-FV500 multi-laser confocal microscope.
For TEM, cells were fixed in 2.5% glutaraldehyde in PBS (pH 7.4) for 1 h at 4 °C, washed and fixed in 1% osmium tetroxide, before embedding in Epon. TEM was performed with a Philips CM10, at 80 kV, on ultrathin sections (100 nm on 200 mesh grids) stained with uranyl acetate and counterstained with lead citrate.
Statistical analysis
The results were expressed as means ± SD and statistical differences were evaluated by one-way and two-way ANOVA followed by Tukey's post-hoc test, using the software package SPSS 11 and Graph-pad Prism version 4.00. P-values of < 0.05 were considered significant.
Abbreviations
- PCD:
-
(programmed cell death)
- 3-MA:
-
(3-methyladenine)
- Baf-A1:
-
(bafilomycin-A1)
- ROS:
-
(reactive oxygen species)
- MMP:
-
(mitochondrial membrane permeabilization)
- Smac:
-
(second mitochondria-derived activator of caspase)
- DIABLO:
-
(direct inhibitor of apoptosis protein-binding protein with low pI)
- AV:
-
(autophagic vacuoles)
- RAGE:
-
(receptor for advanced glycation end product)
- MTT:
-
(3-(4,5-dimethyl-2-thiazolyl)-2,5-diphenyl-2H-tetrazolium bromide)
- GAPDH:
-
(glyceraldehyde-3-phosphate dehydrogenase)
- MTR:
-
(mitotracker red)
- LTR:
-
(lysotracker red)
- TEM:
-
(transmission electron microscopy)
- PARP-1:
-
(poly(ADP-ribose)
References
Bursch W . The autophagosomal-lysosomal compartment in programmed cell death. Cell Death Differ 2001; 8:569–581.
Leist M, Jaattela M . Four deaths and a funeral: from caspases to alternative mechanisms. Nat Rev Mol Cell Biol 2001; 2:589–598.
Adams JM . Ways of dying: multiple pathways to apoptosis. Genes Dev 2003; 17:2481–2495.
Ghavami S, Hashemi M, Ande SR, et al. Apoptosis and cancer: mutations within caspase genes. J Med Genet 2009; 46:497–510.
Hashemi M, Ghavami S, Eshraghi M, Booy EP, Los M . Cytotoxic effects of intra and extracellular zinc chelation on human breast cancer cells. Eur J Pharmacol 2007; 557:9–19.
Ghavami S, Hashemi M, Kadkhoda K, et al. Apoptosis in liver diseases--detection and therapeutic applications. Med Sci Monit 2005; 11:RA337–RA345.
Maddika S, Ande SR, Panigrahi S, et al. Cell survival, cell death and cell cycle pathways are interconnected: Implications for cancer therapy. Drug Resist Updat 2007; 10:13–29.
Hacker G, Paschen SA . Therapeutic targets in the mitochondrial apoptotic pathway. Expert Opin Ther Targets 2007; 11:515–526.
Ghavami S, Kerkhoff C, Chazin WJ, et al. S100A8/9 induces cell death via a novel, RAGE-independent pathway that involves selective release of Smac/DIABLO and Omi/HtrA2. Biochim Biophys Acta 2008; 1783:297–311.
Gottlieb E, Armour SM, Harris MH, Thompson CB . Mitochondrial membrane potential regulates matrix configuration and cytochrome c release during apoptosis. Cell Death Differ 2003; 10:709–717.
Cheng WC, Leach KM, Hardwick JM . Mitochondrial death pathways in yeast and mammalian cells. Biochim Biophys Acta 2008; 1783:1272–1279.
Ghavami S, Kerkhoff C, Los M, et al. Mechanism of apoptosis induced by S100A8/A9 in colon cancer cell lines: the role of ROS and the effect of metal ions. J Leukoc Biol 2004; 76:169–175.
Klionsky DJ, Emr SD . Autophagy as a regulated pathway of cellular degradation. Science 2000; 290:1717–1721.
Yoshimori T . Autophagy: a regulated bulk degradation process inside cells. Biochem Biophys Res Commun 2004; 313:453–458.
Mizushima N, Levine B, Cuervo AM, Klionsky DJ . Autophagy fights disease through cellular self-digestion. Nature 2008; 451:1069–1075.
Anglade P, Vyas S, Javoy-Agid F, et al. Apoptosis and autophagy in nigral neurons of patients with Parkinson's disease. Histol Histopathol 1997; 12:25–31.
Nixon RA, Cataldo AM, Mathews PM . The endosomal-lysosomal system of neurons in Alzheimer's disease pathogenesis: a review. Neurochem Res 2000; 25:1161–1172.
Zakeri Z, Bursch W, Tenniswood M, Lockshin RA . Cell death: programmed, apoptosis, necrosis, or other? Cell Death Differ 1995; 2:87–96.
Foell D, Wittkowski H, Vogl T, Roth J . S100 proteins expressed in phagocytes: a novel group of damage-associated molecular pattern molecules. J Leukoc Biol 2007; 81:28–37.
Kerkhoff C, Klempt M, Sorg C . Novel insights into structure and function of MRP8 (S100A8) and MRP14 (S100A9). Biochim Biophys Acta 1998; 1448:200–211.
Steinbakk M, Naess-Andresen CF, Lingaas E, et al. Antimicrobial actions of calcium binding leucocyte L1 protein, calprotectin. Lancet 1990; 336:763–765.
Kerkhoff C, Nacken W, Benedyk M, et al. The arachidonic acid-binding protein S100A8/A9 promotes NADPH oxidase activation by interaction with p67phox and Rac-2. FASEB J 2005; 19:467–469.
Ghavami S, Rashedi I, Dattilo BM, et al. S100A8/A9 at low concentration promotes tumor cell growth via RAGE ligation and MAP kinase-dependent pathway. J Leukoc Biol 2008; 83:1484–1492.
Huttunen HJ, Kuja-Panula J, Sorci G, et al. Coregulation of neurite outgrowth and cell survival by amphoterin and S100 proteins through receptor for advanced glycation end products (RAGE) activation. J Biol Chem 2000; 275:40096–40105.
Sorci G, Riuzzi F, Agneletti AL, Marchetti C, Donato R . S100B causes apoptosis in a myoblast cell line in a RAGE-independent manner. J Cell Physiol 2004; 199:274–283.
Boya P, Gonzalez-Polo RA, Casares N, et al. Inhibition of macroautophagy triggers apoptosis. Mol Cell Biol 2005; 25:1025–1040.
Ferraro E, Cecconi F . Autophagic and apoptotic response to stress signals in mammalian cells. Arch Biochem Biophys 2007; 462:210–219.
Rossi D, Gaidano G . Messengers of cell death: apoptotic signaling in health and disease. Haematologica 2003; 88:212–218.
Nicoletti I, Migliorati G, Pagliacci MC, Grignani F, Riccardi C . A rapid and simple method for measuring thymocyte apoptosis by propidium iodide staining and flow cytometry. J Immunol Methods 1991; 139:271–279.
Twiddy D, Cohen GM, Macfarlane M, Cain K . Caspase-7 is directly activated by the approximately 700-kDa apoptosome complex and is released as a stable XIAP-caspase-7 approximately 200-kDa complex. J Biol Chem 2006; 281:3876–3888.
Lippke JA, Gu Y, Sarnecki C, Caron PR, Su MS . Identification and characterization of CPP32/Mch2 homolog 1, a novel cysteine protease similar to CPP32. J Biol Chem 1996; 271:1825–1828.
Philchenkov A, Zavelevich M, Kroczak TJ, Los M . Caspases and cancer: mechanisms of inactivation and new treatment modalities. Exp Oncol 2004; 26:82–97.
Puig B, Tortosa A, Ferrer I . Cleaved caspase-3, caspase-7 and poly (ADP-ribose) polymerase are complementarily but differentially expressed in human medulloblastomas. Neurosci Lett 2001; 306:85–88.
Levine B, Klionsky DJ . Development by self-digestion: molecular mechanisms and biological functions of autophagy. Dev Cell 2004; 6:463–477.
Shintani T, Klionsky DJ . Autophagy in health and disease: a double-edged sword. Science 2004; 306:990–995.
Kuma A, Mizushima N, Ishihara N, Ohsumi Y . Formation of the approximately 350-kDa Apg12-Apg5.Apg16 multimeric complex, mediated by Apg16 oligomerization, is essential for autophagy in yeast. J Biol Chem 2002; 277:18619–18625.
Mizushima N, Yamamoto A, Hatano M, et al. Dissection of autophagosome formation using Apg5-deficient mouse embryonic stem cells. J Cell Biol 2001; 152:657–668.
Nemoto T, Tanida I, Tanida-Miyake E, et al. The mouse APG10 homolog, an E2-like enzyme for Apg12p conjugation, facilitates MAP-LC3 modification. J Biol Chem 2003; 278:39517–39526.
Kabeya Y, Mizushima N, Ueno T, et al. LC3, a mammalian homolog of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J 2000; 19:5720–5728.
Kirisako T, Baba M, Ishihara N, et al. Formation process of autophagosome is traced with Apg8/Aut7p in yeast. J Cell Biol 1999; 147:435–446.
Kihara A, Noda T, Ishihara N, Ohsumi Y . Two distinct Vps34 phosphatidylinositol 3-kinase complexes function in autophagy and carboxypeptidase Y sorting in Saccharomyces cerevisiae. J Cell Biol 2001; 152:519–530.
Liang XH, Jackson S, Seaman M, et al. Induction of autophagy and inhibition of tumorigenesis by Beclin 1. Nature 1999; 402:672–676.
Tassa A, Roux MP, Attaix D, Bechet DM . Class III phosphoinositide 3-kinase—Beclin1 complex mediates the amino acid-dependent regulation of autophagy in C2C12 myotubes. Biochem J 2003; 376 (Part 3):577–586.
Maddika S, Bay GH, Kroczak TJ, et al. Akt is transferred to the nucleus of cells treated with apoptin, and it participates in apoptin-induced cell death. Cell Prolif 2007; 40:835–848.
Maddika S, Wiechec E, Ande SR, et al. Interaction with PI3-kinase contributes to the cytotoxic activity of apoptin. Oncogene 2008; 27:3060–3065.
Kim J, Klionsky DJ . Autophagy, cytoplasm-to-vacuole targeting pathway, and pexophagy in yeast and mammalian cells. Annu Rev Biochem 2000; 69:303–342.
Nakatani Y, Yamazaki M, Chazin WJ, Yui S . Regulation of S100A8/A9 (calprotectin) binding to tumor cells by zinc ion and its implication for apoptosis-inducing activity. Mediators Inflamm 2005; 2005:280–292.
Yui S, Nakatani Y, Hunter MJ, Chazin WJ, Yamazaki M . Implication of extracellular zinc exclusion by recombinant human calprotectin (MRP8 and MRP14) from target cells in its apoptosis-inducing activity. Mediators Inflamm 2002; 11:165–172.
Turk B, Stoka V, Rozman-Pungercar J, et al. Apoptotic pathways: involvement of lysosomal proteases. Biol Chem 2002; 383:1035–1044.
Turk B, Turk D, Turk V . Lysosomal cysteine proteases: more than scavengers. Biochim Biophys Acta 2000; 1477:98–111.
Ghavami S, Asoodeh A, Klonisch T, et al. Brevinin-2R(1) semi-selectively kills cancer cells by a distinct mechanism, which involves the lysosomal-mitochondrial death pathway. J Cell Mol Med 2008; 12:1005–1022.
Ray R, Chen G, Vande Velde C, et al. BNIP3 heterodimerizes with Bcl-2/Bcl-X(L) and induces cell death independent of a Bcl-2 homology 3 (BH3) domain at both mitochondrial and nonmitochondrial sites. J Biol Chem 2000; 275:1439–1448.
Zhang L, Li L, Liu H, et al. HIF-1alpha activation by a redox-sensitive pathway mediates cyanide-induced BNIP3 upregulation and mitochondrial-dependent cell death. Free Radic Biol Med 2007; 43:117–127.
Ghavami S, Eshraghi M, Kadkhoda K, et al. Role of BNIP3 in TNF-induced cell death—TNF upregulates BNIP3 expression. Biochim Biophys Acta 2009; 1793:546–560.
Lamy L, Ticchioni M, Rouquette-Jazdanian AK, et al. CD47 and the 19 kDa interacting protein-3 (BNIP3) in T cell apoptosis. J Biol Chem 2003; 278:23915–23921.
Scherz-Shouval R, Elazar Z . ROS, mitochondria and the regulation of autophagy. Trends Cell Biol 2007; 17:422–427.
Scherz-Shouval R, Shvets E, Elazar Z . Oxidation as a post-translational modification that regulates autophagy. Autophagy 2007; 3:371–373.
Burton TR, Eisenstat DD, Gibson SB . BNIP3 (Bcl-2 19 kDa interacting protein) acts as transcriptional repressor of apoptosis-inducing factor expression preventing cell death in human malignant gliomas. J Neurosci 2009; 29:4189–4199.
Burton TR, Gibson SB . The role of Bcl-2 family member BNIP3 in cell death and disease: NIPping at the heels of cell death. Cell Death Differ 2009; 16:515–523.
Xue L, Fletcher GC, Tolkovsky AM . Mitochondria are selectively eliminated from eukaryotic cells after blockade of caspases during apoptosis. Curr Biol 2001; 11:361–365.
Uchiyama Y . Autophagic cell death and its execution by lysosomal cathepsins. Arch Histol Cytol 2001; 64:233–246.
Yu L, Alva A, Su H, et al. Regulation of an ATG7-beclin 1 program of autophagic cell death by caspase-8. Science 2004; 304:1500–1502.
Shimizu S, Kanaseki T, Mizushima N, et al. Role of Bcl-2 family proteins in a non-apoptotic programmed cell death dependent on autophagy genes. Nat Cell Biol 2004; 6:1221–1228.
Pattingre S, Tassa A, Qu X, et al. Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell 2005; 122:927–939.
Graham RM, Thompson JW, Wei J, Bishopric NH, Webster KA . Regulation of Bnip3 death pathways by calcium, phosphorylation, and hypoxia-reoxygenation. Antioxid Redox Signal 2007; 9:1309–1315.
Kubli DA, Ycaza JE, Gustafsson AB . Bnip3 mediates mitochondrial dysfunction and cell death through Bax and Bak. Biochem J 2007; 405:407–415.
Grune T, Merker K, Sandig G, Davies KJ . Selective degradation of oxidatively modified protein substrates by the proteasome. Biochem Biophys Res Commun 2003; 305:709–718.
Kiffin R, Bandyopadhyay U, Cuervo AM . Oxidative stress and autophagy. Antioxid Redox Signal 2006; 8:152–162.
Cataldo AM, Hamilton DJ, Barnett JL, Paskevich PA, Nixon RA . Properties of the endosomal-lysosomal system in the human central nervous system: disturbances mark most neurons in populations at risk to degenerate in Alzheimer's disease. J Neurosci 1996; 16:186–199.
Sooparb S, Price SR, Shaoguang J, Franch HA . Suppression of chaperone-mediated autophagy in the renal cortex during acute diabetes mellitus. Kidney Int 2004; 65:2135–2144.
Mikami M, Yamazaki M, Yui S . Kinetical analysis of tumor cell death-inducing mechanism by polymorphonuclear leukocyte-derived calprotectin: involvement of protein synthesis and generation of reactive oxygen species in target cells. Microbiol Immunol 1998; 42:211–221.
Kerkhoff C, Klempt M, Kaever V, Sorg C . The two calcium-binding proteins, S100A8 and S100A9, are involved in the metabolism of arachidonic acid in human neutrophils. J Biol Chem 1999; 274:32672–32679.
Hunter MJ, Chazin WJ . High level expression and dimer characterization of the S100 EF-hand proteins, migration inhibitory factor-related proteins 8 and 14. J Biol Chem 1998; 273:12427–12435.
Ghavami S, Barczyk K, Maddika S, et al. Monitoring of programmed cell death in vivo and in vitro–new and old methods of cancer therapy assessment. In: Los M, Gibson SB, eds. Apoptotic Pathways as Target for Novel Therapies in Cancer and Other Diseases. New York: Springer Science+Business Media Inc., 2005:323–341.
Maddika S, Panigrahi S, Wiechec E, et al. Unscheduled Akt-triggered activation of cyclin-dependent kinase 2 as a key effector mechanism of apoptin's anticancer toxicity. Mol Cell Biol 2009; 29:1235–1248.
Maddika S, Ande SR, Wiechec E, et al. Akt-mediated phosphorylation of CDK2 regulates its dual role in cell cycle progression and apoptosis. J Cell Sci 2008; 121 (Part 7):979–988.
Acknowledgements
This work was supported by grants from the MHRC, MICH, NTPAA, CLA/CIHR/GSK, and Parker B Francis (to SG), “Interdisziplinäres Zentrum für Klinische Forschung (IZKF)” of the University of Muenster (project Ker3/086/04; to CK), “Deutsche Forschungsgemeinschaft (DFG)” (projects KE 820/6-1 and KE 820/2-4; both to CK), the US National Institutes of Health (RO1 GM62112; to WJC), CCMF (ME). ML acknowledges the support from Deutsche Forschungsgemeinschaft (SFB 773, GRK 1302) and the Deutsche Krebshilfe.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Ghavami, S., Eshragi, M., Ande, S. et al. S100A8/A9 induces autophagy and apoptosis via ROS-mediated cross-talk between mitochondria and lysosomes that involves BNIP3. Cell Res 20, 314–331 (2010). https://doi.org/10.1038/cr.2009.129
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2009.129
Keywords
This article is cited by
-
S100A9 plays a key role in Clostridium perfringens beta2 toxin-induced inflammatory damage in porcine IPEC-J2 intestinal epithelial cells
BMC Genomics (2023)
-
S100a9 inhibits Atg9a transcription and participates in suppression of autophagy in cardiomyocytes induced by β1-adrenoceptor autoantibodies
Cellular & Molecular Biology Letters (2023)
-
Serum proteomics unveil characteristic protein diagnostic biomarkers and signaling pathways in patients with esophageal squamous cell carcinoma
Clinical Proteomics (2022)
-
The S100A10–AnxA2 complex is associated with the exocytosis of hepatitis B virus in intrauterine infection
Laboratory Investigation (2022)
-
S100A8 inhibition in leukemic lymphoblasts induces sensitivity to chemotherapy and inhibition of disease relapse
Medical Oncology (2022)