Abstract
MicroRNAs (miRNAs) are a class of naturally occurring small non-coding RNAs that target protein-coding mRNAs at the post-transcriptional level. Our previous studies suggest that mir-21 functions as an oncogene and has a role in tumorigenesis, in part through regulation of the tumor suppressor gene tropomyosin 1 (TPM1). Given that TPM1 has been implicated in cell migration, in this study we further investigated the role of mir-21 in cell invasion and tumor metastasis. We found that suppression of mir-21 in metastatic breast cancer MDA-MB-231 cells significantly reduced invasion and lung metastasis. Consistent with this, ectopic expression of TPM1 remarkably reduced cell invasion. Furthermore, we identified two additional direct mir-21 targets, programmed cell death 4 (PDCD4) and maspin, both of which have been implicated in invasion and metastasis. Like TPM1, PDCD4 and maspin also reduced invasiveness of MDA-MB-231 cells. Finally, the expression of PDCD4 and maspin inversely correlated with mir-21 expression in human breast tumor specimens, indicating the potential regulation of PDCD4 and maspin by mir-21 in these tumors. Taken together, the results suggest that, as an oncogenic miRNA, mir-21 has a role not only in tumor growth but also in invasion and tumor metastasis by targeting multiple tumor/metastasis suppressor genes. Therefore, suppression of mir-21 may provide a novel approach for the treatment of advanced cancers.
Similar content being viewed by others
Introduction
MicroRNAs (miRNAs) are a class of naturally occurring small non-coding RNAs that target protein-coding mRNAs by repressing translation or causing mRNA degradation 1, 2. Mature miRNAs comprise about 22 nucleotides, derived from long transcripts pri-miRNAs and pre-miRNAs. Although miRNAs operate in a similar fashion as short interfering RNAs (siRNAs) 3, 4, they typically target a cluster of genes instead of one specific gene. It is predicted that an average miRNA can have more than 100 targets 5. Emerging evidence demonstrates an important role of miRNAs in regulating diverse cellular processes including differentiation, proliferation and apoptosis 6, 7.
Deregulation of miRNAs affects normal cell growth and development, leading to a variety of disorders including human malignancies 6, 8, 9, 10, 11. Of particular interest, the overall miRNA expression profile of normal tissues is distinct from that of tumor tissues 12. Specific sets of miRNAs may serve as biomarkers to predict clinical outcomes 13. miRNAs can function as tumor suppressors or oncogenes, depending on whether they specifically target oncogenes or tumor suppressor genes 14, 15, 16. In this regard, tumor suppressive miRNAs are usually underexpressed in tumors. For instance, let-7, mir-15 and mir-16 are downregulated or deleted in lung cancer and leukemia 16, 17, 18. By contrast, oncogenic miRNAs, such as mir-21 and mir-155, are overexpressed in tumors or tumor cell lines 12, 19, 20.
Despite substantial progress in identifying the miRNA signatures for particular types of cancer by profiling miRNA expression in the past years, how miRNAs affect cancer development and progression is still not well understood. We have previously shown that mir-21 promotes growth of the breast cancer cell line MCF-7 both in vitro and in vivo 21, which may be due to the ability of mir-21 to suppress the expression of the tumor suppressor PTEN 15, 22. Consistent with this, we subsequently discovered that mir-21 also directly represses the expression of the tumor suppressor gene tropomyosin 1 (TPM1) 23.
As tumor suppressor genes, PTEN and TPM1 are also implicated in cell migration and invasion 24, 25, suggesting that mir-21 may also have a role in invasion and metastasis. However, little is known as to how mir-21 affects these processes. The present study provides the first evidence that mir-21 regulates invasion and metastasis, at least in part, by targeting metastasis-related tumor suppressor genes such as TPM1, programmed cell death 4 (PDCD4) and maspin. Furthermore, examination of human breast tumor specimens revealed an inverse correlation of mir-21 with PDCD4 and maspin. These data raise the possibility that mir-21 targeting therapy may be more effective than blocking a single oncogenic pathway.
Results
Mir-21 affects cell invasion and metastasis
We have previously shown that breast cancer MCF-7 cells transfected with antisense mir-21 oligonucleotide (anti-mir-21) are inhibited in growth both in vitro and in vivo, which may be due in part to the negative regulation of TPM1 by mir-21 23. TPM1 might also have a role in cell invasion 24. In addition, clinical studies of pancreatic cancer specimens suggest that mir-21 correlates with both tumor growth and metastasis 26. Therefore, we investigated whether and how mir-21 affects invasion and metastasis. We chose MDA-MB-231 because it is a metastatic breast cancer cell line, but also because it expresses a high level of mir-21, similar to MCF-7 cells (Figure 1A). Thus, we used the same anti-mir-21 approach to suppress mir-21 expression and found that the level of mature mir-21 was reduced by approximately 80% in the anti-mir-21-transfected cells (Figure 1B). Interestingly, anti-mir-21 had no effect on the growth of MDA-MB-231 in vitro (Figure 2A). Furthermore, anti-mir-21 had no significant effect on the orthotopic growth of MDA-MB-231 cells (Figure 2B), although anti-mir-21-treated tumors were slightly smaller than the control tumors. These results are very different from those with MCF-7 cells 21. One possibility could be that the manifestation of the biological effects of mir-21 is further defined by the specific genetic and epigenetic background of each cell line. MCF-7 cells are hormone-sensitive, but not metastatic unless implanted with matrigel. MDA-MB-231 cells, on the other hand, are highly tumorigenic and metastatic. While the differential effects of mir-21 on the growth of MCF-7 and MDA-MB-231 need to be further investigated, MDA-MB-231 cells offer a good model to study the effect of mir-21 on tumor invasion and metastasis.
MDA-MB-231 cells express a high level of mature mir-21, which can be suppressed by anti-mir-21. (A) Detection of mature mir-21 by TaqMan real-time PCR using U6 RNA for normalization. Both MCF-7 and MDA-MB-231 cells express higher levels of mir-21 compared with 293T cells, which express low levels of mir-21. (B) Suppression of mir-21 by anti-mir-21 in MDA-MB-231 cells. The cells were first transfected with anti-mir-21 or a negative control. After 2 days, they were harvested and total RNA was extracted using Trizol reagent. Mature mir-21 was detected by TaqMan real-time PCR using U6 RNA for normalization. Values in (A) and (B) are the means from three independent experiments ±SE. **P < 0.01.
Effect of anti-mir-21 on MDA-MB-231 cell growth in vitro and in vivo. (A) MDA-MB-231 cells were transfected with anti-mir-21 or a negative control and then plated out in 96-well plates. Relative cell growth was measured by MTT assay at the indicated times. No difference in cell growth between the negative control and anti-mir-21 was detected. The values are the means from three separate experiments. (B) MDA-MB-231 cells were transfected with anti-mir-21 or a negative control and then injected into mice in the mammary fat pads (1.5 million cells per injection). Tumors were harvested 28 days after injection of the cells. No significant difference in tumor weight between the negative control (Neg) and anti-mir-21 (Anti) was detected. The values are the means from 5 animals ±SE.
An in vitro invasion assay indicated that the invasiveness of MDA-MB-231 cells transfected with anti-mir-21 was specifically reduced by approximately 60% (Figure 3A and 3B). Since mir-21 is overexpressed in many types of solid tumors, including prostate cancer 12, 20, 22, 26, 27, 28, 29, 30, mir-21-mediated cell invasion and metastasis may also occur in other types of tumors. Thus, we tested the effect of anti-mir-21 on metastatic prostate cancer PC-3MM cells 25, as determined by the same invasion assay. We found that anti-mir-21 also inhibited the invasiveness of PC-3MM (not shown).
Suppression of cell invasion and tumor metastasis by anti-mir-21. (A, B) Cell invasion assay. MDA-MB-231 cells were first transfected with anti-mir-21 (Anti) or a negative control (Neg) and then subject to matrigel chamber assays, as described in Materials and Methods. After 20 h, invasion cells were counted after staining with crystal violet. (A) Average invasive cell number from three independent experiments ±SE. (B) Representative fields of invasive cells on the membrane. (C, D) In vivo metastasis assay. MDA-MB-231 cells were first transfected with anti-mir-21 (Anti) or a negative control (Neg) and then injected into animals through tail veins, as described in Materials and Methods. Animals were killed 3 weeks after injection. (C) Average tumor nodules in the lung (n = 5). (D) Representative tumor nodules. ** P < 0.01.
To further confirm the mir-21 effects, mir-21-transfected and negative control-transfected MDA-MB-231 cells were tail-vein injected into young female nude mice. Mouse lungs were harvested to evaluate tumor metastasis 3 weeks after the injection. As shown in Figure 3C and 3D, anti-mir-21 significantly reduced the number of lung metastases. For instance, an average of 22 nodules were detected per lung in mice bearing negative control-transfected cells, whereas mice bearing anti-mir-21-transfected tumor cells produced an average of 2 tumor nodules per lung (Figure 3D). These data demonstrate an important role of mir-21 in tumor metastasis.
Mir-21 directly targets PDCD4 and maspin in addition to TPM1
Although mir-21 has been previously shown to suppress the expression of PTEN 15, 22 and TPM1 23, it is not clear whether the negative regulation of the target genes, in particular TPM1, contributes to the suppression of invasion and metastasis by anti-mir-21. Furthermore, given that miRNAs usually target multiple genes at the post-transcriptional level, it is highly likely that mir-21 exerts its effects by negatively regulating multiple genes that are involved in invasion and metastasis. Therefore, we searched for additional mir-21 targets using a genetic screening/selection system, and identified PDCD4 and maspin as candidate mir-21 target genes (Wu et al., unpublished data). Of interest, PDCD4 has been suggested to be a putative target for mir-21 and implicated in the metastasis of pancreatic tumors 26. Maspin is a well-known tumor suppressor that can suppress tumor growth, invasion and metastasis 31. Therefore, we examined the effect of TPM1, PDCD4 and maspin on the invasiveness of MDA-MB-231 cells. We first transfected MDA-MB-231 cells with Myc-tagged TPM1, PDCD4 or Flag-tagged maspin and confirmed their expression by western blotting with anti-Myc or anti-Flag antibodies (Figure 4A). As shown in Figure 4B, overexpression of TPM1 suppressed cell invasion by up to 75% compared with the vector control, suggesting that TPM1 is involved in cell invasion. Similarly, both PDCD4 and maspin also significantly suppressed cell invasion in MDA-MB-231 cells (Figure 4B). These results suggest that mir-21 probably affects cell invasion and metastasis through regulating multiple target genes, such as TPM1, PDCD4 and maspin.
Effect of TPM1, PDCD4 and maspin on cell invasion. (A) Detection of TPM1-Myc, PDCD4-Myc and maspin-Flag by western blotting. Anti-Myc antibody was used to detect TPM1 and PDCD4, and anti-Flag antibody was used to detect maspin. (B) Cell invasion assays. MDA-MB-231 cells were transfected with plasmid constructs as indicated and then subjected to matrigel chamber assays. The same transfected cells in (A) were aliquoted out for the invasion assay. The values are the means from three independent experiments ±SE. ** P < 0.01.
To further confirm that PDCD4 and maspin are direct mir-21 targets, we tested whether mir-21 can suppress the 3′-UTR (untranslated region)-mediated gene expression, because, like TPM1, the 3′-UTR of each of PDCD4 and maspin carries a putative mir-21 binding site (Figure 5A). Thus, we cloned the PDCD4 3′-UTR and the maspin 3′-UTR into the downstream of the luciferase gene (pGL3 control vector), respectively. We tested the effect of mir-21 on luciferase activity for each construct in 293T cells that express a low level of mir-21. As shown in Figure 5B, mir-21 suppressed the luciferase activity of the Luc-PDCD4 3′-UTR by more than 50% compared with the vector control, whereas deletion of the mir-21 binding site (underlined in Figure 5A) blocked this suppression (Figure 5B). Similarly, mir-21 also suppressed the luciferase activity of the Luc-Maspin 3′-UTR by more than 40% compared with the vector control (Figure 5B), and deletion of the mir-21 binding site (underlined) reversed this suppression. To determine whether anti-mir-21 affects the expression of the endogenous target genes, we transfected MDA-MB-231 cells with negative control or anti-mir-21. Western blotting indicated that the suppression of mir-21 by anti-mir-21 in MDA-MB-231 cells causes upregulation of endogenous PDCD4 and maspin (Figure 5C). In addition, we detected upregulation of endogenous TPM1 as has been shown previously 23, as well as of the endogenous PTEN by anti-mir-21 at the protein level (not shown). These results suggest that both PDCD4 and maspin are direct mir-21 targets.
PDCD4 and maspin are direct mir-21 targets. (A) The PDCD4 UTR sequence reveals a putative mir-21 binding site (http://microrna.sanger.ac.uk/cgi-bin/targets/v4/detail_view.pl?transcript_id=ENST00000280154). The underlined sequence denotes a deletion in Luc-PDCD4-UTR-d. Also shown is the maspin UTR sequence, which reveals a putative mir-21 binding site (http://microrna.sanger.ac.uk/cgi-bin/targets/v4/detail_view.pl?transcript_id=ENST00000382771); the underlined portion was deleted in Luc-Maspin-UTR-d. (B) Effect of mir-21 on the luciferase activity of Luc-PDCD4-UTR and Luc-PDCD4-UTR-d, and the luciferase activity of Luc-Maspin-UTR and Luc-Maspin-UTR-d. The assay was done in 293T cells as described in Materials and Methods. V, vector (pCMV); 21, pCMV-mir-21. The values are the means from three independent experiments ±SE. **P < 0.01. (C) Suppression of mir-21 upregulates the endogenous PDCD4 and maspin in MDA-MB-231 cells, respectively. N, negative control; A, anti-mir-21.
PDCD4 protein level negatively correlates with mir-21 expression in tumor specimens
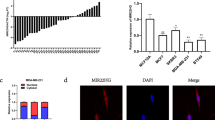
To determine the clinical significance of mir-21 target genes, we examined PDCD4 protein levels in 8 pairs of matched breast tumor specimens by western blotting. As expected, lower levels of PDCD4 were detected in tumors in all cases (Figure 6A). To determine whether there is any correlation between PDCD4 and mir-21, we also measured mir-21 expression in these cases using TaqMan real-time PCR and found that all tumors had higher levels of mir-21 expression. Statistical analysis using the Pearson's method confirmed the inverse correlation between PDCD4 protein and mir-21 in these tissue samples, with a correlation coefficient of −0.824 (Figure 6C). Similar to PDCD4, we also detected downregulation of maspin in tumors in 3 of 4 cases (Figure 6B).
Expression of PDCD4, maspin and mir-21 in matched breast tumor specimens. (A) Expression of PDCD4 protein in eight pairs of matched breast normal tissue and breast tumor tissues, as detected by western blotting. The specimen number is shown above each pair comprising normal (N) and tumor (T). (B) Expression of maspin in the matched tumor tissues. (C) A negative correlation between PDCD4 and mir-21 in matched tumor specimens.
Therefore, although it is possible that other cancer-related cellular factors are also altered in breast tumors, and more samples may be required to make a firm conclusion, upregulation of mir-21 might at least in part be attributive to the downregulation of metastasis-related tumor suppressors, such as PDCD4 and maspin.
Discussion
Metastasis is a major cause of cancer-related death. Hence, identifying the role of mir-21 in invasion and metastasis has direct clinical implications. Although it is now known that miRNAs might have a key role in cancer development, the underlying mechanism is largely unknown. We show here that the overexpression of TPM1 suppresses the invasiveness of MDA-MB-231, and identify PDCD4 and maspin as direct targets for mir-21 through interacting with their 3′-UTR. More importantly, our present study suggests that, as an oncogenic miRNA, mir-21 affects not only tumor growth but also invasion and metastasis. The biological effects of mir-21 are probably due to the simultaneous repression of multiple tumor suppressor genes, including TPM1, PDCD4, maspin and PTEN.
PTEN is well known for its role in tumor growth, invasion and metastasis, while information regarding the role of TPM1, maspin and PDCD4 in these aspects is only recently emerging. PTEN is capable of restricting growth and survival signals by limiting the activity of the phosphoinositide 3-kinase (PI3K) pathway. A decrease in functional PTEN causes constitutive activation of downstream components of the PI3K pathway including Akt, leading to tumor progression and metastasis 32. Thus, PTEN downregulation by mir-21 may contribute to transformation and increased tumor cell survival 32. TPM1 is an actin-binding protein and is capable of stabilizing microfilaments 33. Moreover, TPMs regulate both microfilament organization and anchorage-independent growth, highlighting the importance of TPMs in cell transformation 34. Since overexpression of TPM1 suppresses cell invasion, our study supports the notion that TPM1 can have a role in cell invasion.
A previous study has suggested that PDCD4 is a potential target for mir-21 26. In support of this, our study further confirms that PDCD4 is a direct target for mir-21 through the mir-21 binding site at the 3′-UTR. As a tumor suppressor protein, PDCD4 interacts with eukaryotic initiation factor 4A (EIF4A) and inhibits protein synthesis 35, 36. PDCD4 also suppresses the transactivation of AP-1-responsive promoters by c-Jun, and phosphorylation of PDCD4 by Akt has been shown to decrease the ability of PDCD4 to interfere with the transactivation of the AP-1-responsive promoter by c-Jun 37. The ability of PDCD4 to suppress cell invasion may involve the downregulation of MAP4K1 transcription 38. In addition, PDCD4 represses the transcription of the mitosis-promoting factor cyclin-dependent kinase (CDK)1/cdc2 through upregulating p21 39. However, until now little has been known about the post-transcriptional regulation of PDCD4 expression. Our study suggests that PDCD4 is subject to miRNA regulation.
Maspin has tumor-suppressive activity and its expression is substantially downregulated in mammary carcinomas, possibly through DNA methylation or loss of transcription factors 31. Since miRNAs only target mRNAs, they are not expected to exert any effect on target gene expression when the target genes are already silenced by DNA modification and/or transcriptional repression. Interestingly, anti-mir-21 can increase the expression of maspin in MDA-MB-231 cells. This evidence raises the possibility that complete silencing of maspin also requires miRNAs.
Interestingly, both PDCD4 and maspin have been implicated in the regulation of urokinase receptor (uPAR) 40, 41, in addition to their association with apoptosis 42, 43. uPAR is a cell invasion and metastasis-promoting factor and has been shown to be a potential prognosis marker 44. In this regard, maspin has been shown to interact with cell-surface-associated uPA and stimulates the internalization of the uPA/uPAR complex 41. On the other hand, PDCD4 is able to downregulate uPAR via its promoter region 40. Therefore, the consequence of these actions by either PDCD4 or maspin is the reduction of uPAR level such that cell invasion or metastasis is inhibited. Our study shows that transient ectopic expression of PDCD4 or maspin leads to significant inhibition of cell invasion (Figure 4B). Therefore, downregulation of PDCD4 and maspin by mir-21 may promote tumor invasion and metastasis. Interestingly, maspin is known to suppress tumor invasion and metastasis without affecting cell growth. It remains to be determined whether the mir-21 effect on maspin is a major determinant of the observed MDA-MB-231 phenotype.
Since MCF-7 and MDA-MB-231 cells respond so differently to anti-mir-21, it would be interesting to investigate the underlying mechanisms. In addition to their difference in metastatic potential and hormone sensitivity, MCF-7 and MDA-MB-231 may differ in other specific genetic and epigenetic backgrounds. Thus, gene profiling of these cells, after transfection with anti-mir-21, may provide some indication of the possible mechanisms. In this regard, mir-21 is similar to the tumor suppressor Drg-1, but in an opposite way because Drg-1 has been shown to suppress primary tumor growth in bladder cancer cells 45, functioning as a tumor suppressor. However, in prostate and colon cancer cells Drg-1 is able to suppress distant metastasis, but has no effect on primary tumor growth 46, 47, and functions as a metastasis suppressor. It remains to be determined whether mir-21 can target such similar tumor suppressor genes.
In summary, the identification of PDCD4 and maspin as direct mir-21 targets provides an exciting new insight into the molecular mechanism underlying the oncogenic effects of mir-21. Our data suggest that mir-21 may promote tumor invasion and metastasis by simultaneously downregulating multiple metastasis-related tumor suppressor genes operating at distinct steps of tumor progression. It remains an intriguing possibility that nature designed miRNAs to coordinate the expression of functionally regulated cluster of genes. Given that an miRNA can target over 100 genes 5, we expect that there are additional mir-21 targets yet to be identified in different cellular systems, which could also have a role in invasion and metastasis. Therefore, our biological evidence suggests that targeting mir-21 may provide a better strategy to block tumor metastasis.
Materials and Methods
Cell culture, reagents and human tissues
Human breast cancer cell line MDA-MB-231 (American Type Cell Collection (ATCC), Manassas, VA) and human prostate carcinoma PC-3-derived subline PC-3MM (from Dr Watabe, SIU) were grown in RPMI-1640 (Cambrex, Walkersville, MD), supplemented with 10% fetal bovine serum (Sigma-Aldrich, St Louis, MO), 2 mM glutamine, 100 U of penicillin/ml and 100 μg of streptomycin/ml (Cambrex). The cultivation of immortalized kidney epithelial cells 293T (ATCC) was as described 5. All cells were incubated at 37 °C in a humidified chamber supplemented with 5% CO2.
Control negative oligonucleotide and anti-mir-21 oligonucleotide were purchased from Ambion (Huston, TX). Anti-maspin antibody was purchased from BD Biosciences (San Jose, CA). Anti-PDCD4 antibody was from Rockland (Gilbertsville, PA). Anti-Myc antibody was from Applied Biological Materials (Vancouver, Canada). Anti-Flag antibody was from Sigma (St Louis, MO). All PCR primers were purchased from Sigma-Genosys (Woodland, TX). Breast tumor specimens and their matching normal breast specimens were obtained from the Cooperative Human Tissue Network (CHTN) (Midwestern Division, Columbus, OH).
Construction of expression vectors
Expression vectors of maspin 48, TPM1 and mir-21 23 have been described previously. To construct a vector to express Myc-tagged PDCD4, a DNA fragment covering the PDCD4 coding region was first amplified from MCF10A cDNA by PCR (forward primer 5′-GAA TTC TGG ATG TAG AAA ATG AGC AGA and reverse primer 5′-GCG GCC GCT CAG TAG CTC TCT GGT TTA AG) and was then cloned into pCR8 (Invitrogen). After verification by DNA sequencing, the full-length PDCD4-coding fragment was finally cloned into pCMV-Myc (Clontech) at the EcoR1 and Not1 sites.
Transfection
Cells were seeded in six-well plates at 30% confluence and grown overnight before transfection. Transfection of MDA-MB-231 and PC-3MM cells with anti-mir-21 or negative control oligonucleotide was performed using the RNAifectin reagent (Applied Biological Materials) according to the manufacturer's protocol. Transfection of MDA-MB-231 cells with the expression vectors for Flag-maspin 48, Myc-TPM1 23 and Myc-PDCD-4, and with mir-21, was performed using the DNAfectin Kit (Applied Biological Materials). Transfection of 293T cells was performed using the calcium phosphate method as described previously 49.
Detection of mature mir-21 by TaqMan real-time PCR
PCR-based detection of mature mir-21 was performed by the TaqMan miRNA assays (ABI, Forest City, CA) as described previously 50, 51. The real-time PCR results, recorded as threshold cycle numbers (Ct), were normalized against an internal control (U6 RNA), and then expressed as fold changes 50.
Luciferase reporter assay
The 3′-untranslated mRNA sequences of maspin and PCDC-4 containing the mir-21 binding site were amplified by PCR using the following primers:
PDCD4-UTR-5.1 (forward) 5′-ATA TAA GAA CTC TTG CAG TCT TAG A
PDCD4-UTR-3.1 (reverse) 5′-GAA TAA GTC TCA TGA CTA ATT TTT T
Maspin-UTR-5.1 (forward), 5′-GTG GCA TAG CCC ATG TTA AGT CC
Maspin-UTR-3.1 (reverse), 5′-AAA TTC AAT ATT GAA TCT CAG AAC
After amplification, PCR products were first cloned into pCR8 and then individually subcloned downstream to the luciferase coding sequence in the luciferase reporter pGL3-control vector (Promega, Madison, WI). To make mir-21 binding site deletion mutants, we used the two-step PCR approach where the seed sequences were deleted in the primers used for PCR reactions. All constructs were verified by DNA sequencing. 293T cells were seeded in six-well plates and transfected with luciferase reporters using the calcium phosphate method as described 48. After transfection, the cells were split into 12-well plates in duplicates, harvested 24 h later, and analyzed for luciferease activity using the assay kit from Promega. β-Galactosidase activity was used for normalizing the transfection efficiency and protein input.
Western blotting
SDS-PAGE and western blotting were performed according to standard procedures. Western blotting of β-actin on the same membrane was used as a loading control. Signals were detected by secondary antibodies labeled with either Alex Fluor 680 (Invitrogen) or IRDye 800 (Rockland Immunochemicals, Gilbertsville, PA) and signal intensity was determined by Odyssey Infrared Imaging System (Li-Cor Biosciences, Lincoln, NE) as described previously 21.
Cell proliferation assay
After transfection with negative control or anti-mir21, MDA-MB-231 cells were seeded in 96-well plates at 2 500 cells/well. MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl-tetrazoliumbromide) assay was used to detect viable proliferating cells at various time points 21.
In vitro invasion assay
The invasive potential of cells was measured in 12-well matrigel-coated invasion chambers (BD Biosciences, Bedford, MA). The lower chambers were filled with 0.75 ml of RPMI-1640 medium containing 10% fetal bovine serum (FBS). A cell suspension of 2.5 × 104 in 0.5 ml RPMI-1640 medium was added into each well of the upper chamber. After the cells were incubated for 20 h at 37 °C in a humidified incubator with 5% CO2, the non-invading cells that remained on the upper surface of the membrane were removed by scraping. The invasive cells attached to the lower surface of the membrane insert were fixed in 10% formalin at room temperature for 30 min and stained with 0.05% crystal violet. The number of invasive cells on the lower surface of the membrane was then counted under a microscope.
Tumor metastasis in vivo
Female athymic nude (nu/nu) mice (4-5 weeks old) were purchased from Harlan Sprague Dawley (Indianapolis, IN) and were maintained in the SIU School of Medicine's accredited animal facility. All animal studies were conducted in accordance with NIH animal use guidelines and a protocol approved by the SIU Animal Care Committee. Exponentially growing MDA-MB-231 cells were first transfected with negative control or anti-mir-21. One day after transfection, the cells were split and grown in a fresh medium for one more day before harvest for inoculation. A total of 0.8 million cells were injected into each mouse through tail veins. At 3 weeks after injection, the animals were killed, and lungs were harvested and fixed in Bouin's solution before tumor nodules were counted. Alternatively, 1.5 million of the transfected MDA-MB-231 cells were injected into the mammary fat pad of nude mice to determine the primary tumor growth.
Statistical analysis
The statistical significance of the in vitro studies was analyzed using Student's t test. Differences with P values of less than 0.01 are considered significant. The linear correlation coefficient (Pearson's r) was calculated to estimate the correlation between mir-21 values and PDCD4 levels in the matched breast tumor specimens.
Abbreviations
- miRNA:
-
(microRNA)
- siRNA:
-
(short interfering RNA)
- PDCD4:
-
(programmed cell death 4)
- TPM1:
-
(tropomyosin 1)
References
Pillai RS . MicroRNA function: multiple mechanisms for a tiny RNA? RNA 2005; 11:1753–1761.
Zamore PD, Haley B . Ribo-gnome: the big world of small RNAs. Science 2005; 309:1519–1524.
Bartel DP . MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004; 116:281–297.
Fitzgerald K . RNAi versus small molecules: different mechanisms and specificities can lead to different outcomes. Curr Opin Drug Discov Dev 2005; 8:557–566.
Brennecke J, Stark A, Russell RB, Cohen SM . Principles of microRNA-target recognition. PLoS Biol 2005; 3:e85.
Croce CM, Calin GA . miRNAs, cancer, and stem cell division. Cell 2005; 122:6–7.
Chen CZ, Li L, Lodish HF, Bartel DP . MicroRNAs modulate hematopoietic lineage differentiation. Science 2004; 303:83–86.
Hwang HW, Mendell JT . MicroRNAs in cell proliferation, cell death, and tumorigenesis. Br J Cancer 2006; 94:776–780.
Hammond SM . MicroRNAs as oncogenes. Curr Opin Genet Dev 2006; 16:4–9.
Esquela-Kerscher A, Slack FJ . Oncomirs – microRNAs with a role in cancer. Nat Rev Cancer 2006; 6:259–269.
Gregory RI, Shiekhattar R . MicroRNA biogenesis and cancer. Cancer Res 2005; 65:3509–3512.
Iorio MV, Ferracin M, Liu CG, et al. MicroRNA gene expression deregulation in human breast cancer. Cancer Res 2005; 65:7065–7070.
Yanaihara N, Caplen N, Bowman E, et al. Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell 2006; 9:189–198.
Cimmino A, Calin GA, Fabbri M, et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci USA 2005; 102:13944–13949.
O'Donnell KA, Wentzel EA, Zeller KI, Dang CV, Mendell JT . c-Myc-regulated microRNAs modulate E2F1 expression. Nature 2005; 435:839–843.
Johnson SM, Grosshans H, Shingara J, et al. RAS is regulated by the let-7 microRNA family. Cell 2005; 120:635–647.
Takamizawa J, Konishi H, Yanagisawa K, et al. Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res 2004; 64:3753–3756.
Calin GA, Dumitru CD, Shimizu M, et al. Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci USA 2002; 99:15524–15529.
Eis PS, Tam W, Sun L, et al. Accumulation of miR-155 and BIC RNA in human B cell lymphomas. Proc Natl Acad Sci USA 2005; 102:3627–3632.
Chan JA, Krichevsky AM, Kosik KS . MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res 2005; 65:6029–6033.
Si ML, Zhu S, Wu H, Lu Z, Wu F, Mo YY . miR-21-mediated tumor growth. Oncogene 2007; 26:2799–2803.
Meng F, Henson R, Lang M, et al. Involvement of human micro-RNA in growth and response to chemotherapy in human cholangiocarcinoma cell lines. Gastroenterology 2006; 130:2113–2129.
Zhu S, Si ML, Wu H, Mo YY . MicroRNA-21 targets the tumor suppressor gene Tropomyosin 1 (TPM1). J Biol Chem 2007; 282:14328–14336.
Varga AE, Stourman NV, Zheng Q, et al. Silencing of the Tropomyosin-1 gene by DNA methylation alters tumor suppressor function of TGF-beta. Oncogene 2005; 24:5043–5052.
Bandyopadhyay S, Pai SK, Hirota S, et al. PTEN up-regulates the tumor metastasis suppressor gene Drg-1 in prostate and breast cancer. Cancer Res 2004; 64:7655–7660.
Roldo C, Missiaglia E, Hagan JP, et al. MicroRNA expression abnormalities in pancreatic endocrine and acinar tumors are associated with distinctive pathologic features and clinical behavior. J Clin Oncol 2006; 24:4677–4684.
Tran N, McLean T, Zhang X, et al. MicroRNA expression profiles in head and neck cancer cell lines. Biochem Biophys Res Commun 2007.
Lee EJ, Gusev Y, Jiang J, et al. Expression profiling identifies microRNA signature in pancreatic cancer. Int J Cancer 2007; 120:1046–1054.
Wang T, Zhang X, Obijuru L, et al. A micro-RNA signature associated with race, tumor size, and target gene activity in human uterine leiomyomas. Genes Chromosomes Cancer 2007; 46:336–347.
Volinia S, Calin GA, Liu CG, et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci USA 2006; 103:2257–2261.
Lockett J, Yin S, Li X, Meng Y, Sheng S . Tumor suppressive maspin and epithelial homeostasis. J Cell Biochem 2006; 97:651–660.
Sansal I, Sellers WR . The biology and clinical relevance of the PTEN tumor suppressor pathway. J Clin Oncol 2004; 22:2954–2963.
Perry SV . Vertebrate tropomyosin: distribution, properties and function. J Muscle Res Cell Motil 2001; 22:5–49.
Boyd J, Risinger JI, Wiseman RW, Merrick BA, Selkirk JK, Barrett JC . Regulation of microfilament organization and anchorage-independent growth by tropomyosin 1. Proc Natl Acad Sci USA 1995; 92:11534–11538.
Yang HS, Jansen AP, Komar AA, et al. The transformation suppressor Pdcd4 is a novel eukaryotic translation initiation factor 4A binding protein that inhibits translation. Mol Cell Biol 2003; 23:26–37.
Dorrello NV, Peschiaroli A, Guardavaccaro D, Colburn NH, Sherman NE, Pagano M . S6K1- and betaTRCP-mediated degradation of PDCD4 promotes protein translation and cell growth. Science 2006; 314:467–471.
Palamarchuk A, Efanov A, Maximov V, Aqeilan RI, Croce CM, Pekarsky Y . Akt phosphorylates and regulates Pdcd4 tumor suppressor protein. Cancer Res 2005; 65:11282–11286.
Yang HS, Matthews CP, Clair T, et al. Tumorigenesis suppressor Pdcd4 down-regulates mitogen-activated protein kinase kinase kinase kinase 1 expression to suppress colon carcinoma cell invasion. Mol Cell Biol 2006; 26:1297–1306.
Goke R, Barth P, Schmidt A, Samans B, Lankat-Buttgereit B . Programmed cell death protein 4 suppresses CDK1/cdc2 via induction of p21(Waf1/Cip1). Am J Physiol Cell Physiol 2004; 287:C1541–C1546.
Leupold JH, Yang HS, Colburn NH, Asangani I, Post S, Allgayer H . Tumor suppressor Pdcd4 inhibits invasion/intravasation and regulates urokinase receptor (u-PAR) gene expression via Sp-transcription factors. Oncogene 2007; 26:4550–4562.
Yin S, Lockett J, Meng Y, et al. Maspin retards cell detachment via a novel interaction with the urokinase-type plasminogen activator/urokinase-type plasminogen activator receptor system. Cancer Res 2006; 66:4173–4181.
Jurisicova A, Latham KE, Casper RF, Casper RF, Varmuza SL . Expression and regulation of genes associated with cell death during murine preimplantation embryo development. Mol Reprod Dev 1998; 51:243–253.
Li X, Chen D, Yin S, et al. Maspin augments proteasome inhibitor-induced apoptosis in prostate cancer cells. J Cell Physiol 2007; 212:298–306.
Stephens RW, Nielsen HJ, Christensen IJ, et al. Plasma urokinase receptor levels in patients with colorectal cancer: relationship to prognosis. J Natl Cancer Inst 1999; 91:869–874.
Kurdistani SK, Arizti P, Reimer CL, Sugrue MM, Aaronson SA, Lee SW . Inhibition of tumor cell growth by RTP/rit42 and its responsiveness to p53 and DNA damage. Cancer Res 1998; 58:4439–4444.
Bandyopadhyay S, Pai SK, Gross SC, et al. The Drg-1 gene suppresses tumor metastasis in prostate cancer. Cancer Res 2003; 63:1731–1736.
Guan RJ, Ford HL, Fu Y, Li Y, Shaw LM, Pardee AB . Drg-1 as a differentiation-related, putative metastatic suppressor gene in human colon cancer. Cancer Res 2000; 60:749–755.
Yin S, Li X, Meng Y, et al. Tumor-suppressive maspin regulates cell response to oxidative stress by direct interaction with glutathione S-transferase. J Biol Chem 2005; 280:34985–34996.
Mo YY, Beck WT . Association of human DNA topoisomerase IIalpha with mitotic chromosomes in mammalian cells is independent of its catalytic activity. Exp Cell Res 1999; 252:50–62.
Chen C, Ridzon DA, Broomer AJ, et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res 2005; 33:e179.
Lao K, Xu NL, Yeung V, Chen C, Livak KJ, Straus NA . Multiplexing RT-PCR for the detection of multiple miRNA species in small samples. Biochem Biophys Res Commun 2006; 343:85–89.
Acknowledgements
This study was supported by grants to Y Mo from the National Cancer Institute (CA102630) and from Department of defense (BC045418 and BC052294), and to S Sheng from the National Cancer Institute (CA84176).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhu, S., Wu, H., Wu, F. et al. MicroRNA-21 targets tumor suppressor genes in invasion and metastasis. Cell Res 18, 350–359 (2008). https://doi.org/10.1038/cr.2008.24
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2008.24
Keywords
This article is cited by
-
Linking cell mechanical memory and cancer metastasis
Nature Reviews Cancer (2024)
-
Nucleic acid drug vectors for diagnosis and treatment of brain diseases
Signal Transduction and Targeted Therapy (2023)
-
Pterostilbene–isothiocyanate reduces miR-21 level by impeding Dicer-mediated processing of pre-miR-21 in 5-fluorouracil and tamoxifen-resistant human breast cancer cell lines
3 Biotech (2023)
-
The Significant Role of microRNAs in Gliomas Angiogenesis: A Particular Focus on Molecular Mechanisms and Opportunities for Clinical Application
Cellular and Molecular Neurobiology (2023)
-
Understanding crosstalk of organ tropism, tumor microenvironment and noncoding RNAs in breast cancer metastasis
Molecular Biology Reports (2023)