Abstract
HSP90 inhibition represents a promising route to cancer therapy, taking advantage of cancer cell-inherent proteotoxic stress. The HSP90-inhibitor ganetespib showed benefit in advanced clinical trials. This raises the need to identify the molecular determinants of treatment response. We tested the efficacy of ganetespib on a series of colorectal cancer (CRC)-derived cell lines and correlated their sensitivities with comprehensive gene expression analysis. Notably, the drug concentration required for 50% growth inhibition (IC50) varied up to 70-fold (from 36 to 2500 nM) between different cell lines. Correlating cell line-specific IC50s with the corresponding gene expression patterns revealed a strong association between ganetespib resistance (IC50>500 nM) and high expression of the UDP glucuronosyltransferase 1A (UGT1A) gene cluster. Moreover, CRC tumor samples showed a comparable distribution of UGT1A expression levels. The members of the UGT1A gene family are known as drug-conjugating liver enzymes involved in drug excretion, but their function in tumor cells is hardly understood. Chemically unrelated HSP90 inhibitors, for example, 17-N-allylamino-17-demethoxygeldanamycin (17-AAG), did not show correlation of drug sensitivities with UGT1A levels, whereas the ganetespib-related compound NVP-AUY922 did. When the most ganetespib-resistant cell line, HT29, was treated with ganetespib, the levels of HSP90 clients were unaffected. However, HT29 cells became sensitized to the drug, and HSP90 client proteins were destabilized by ganetespib upon siRNA-mediated UGT1A knockdown. Conversely, the most ganetespib-sensitive cell lines HCT116 and SW480 became more tolerant toward ganetespib upon UGT1A overexpression. Mechanistically, ganetespib was rapidly glucuronidated and excreted in resistant but not in sensitive CRC lines. We conclude that CRC cell-expressed UGT1A inactivates ganetespib and other resorcinolic Hsp90 inhibitors by glucuronidation, which renders the drugs unable to inhibit Hsp90 and thereby abrogates their biological activity. UGT1A levels in tumor tissues may be a suitable predictive biomarker to stratify CRC patients for ganetespib treatment.
Similar content being viewed by others
Main
Constant proteotoxic stress is a frequent occurrence in cancer cells and is derived from an adverse external microenvironment (hypoxia, acidosis) and internally from conformationally aberrant oncoproteins, high reactive oxygen species (ROS) levels, genomic instability, and stoichiometric imbalances in multi-protein machines. This stress condition raises the need for massive heat-shock chaperone support, especially from the heat-shock protein 90 (HSP90) system, to prevent protein aggregation and illicit interactions and promote tumor cell survival. Cancer-associated factors, such as mutant p53,1, 2 ErbB2,3 AKT,4 and macrophage migration inhibitory factor (MIF),5, 6 among others, represent HSP90 clients and require HSP90 for their stabilization in tumors. Hence, the multi-component HSP90 chaperone is highly upregulated and activated specifically in cancer cells as an adaptive response to malignancy.7
HSP90 inhibitors have emerged as a highly promising class of anti-cancer compounds because of their ability to interfere with broadly active molecular networks, rather than a narrowly defined signaling pathway8, 9 and they enhance proteotoxic stress.10 Geldanamycin-based compounds represented the mainstay of HSP90 inhibition for the last 20 years.8 Clinically, however, these compounds proved to be of limited value due to their inherent liver and ocular toxicity coupled with only modest potency in vivo.11 Major advances came with recently developed second-generation synthetic inhibitors such as the resorcinol containing compounds ganetespib (STA-9090)12 and NVP-AUY92213 that are considerably more potent and less toxic. These compounds are currently being tested in phase II/III trials for their efficacy against various cancer types. Anaplastic lymphoma kinase (ALK)-driven NSCLC cancers showed particular clinical responsiveness to ganetespib.14, 15
Colorectal carcinoma (CRC) represents one of the most frequent malignancies worldwide, with a correspondingly high death toll. Moreover, the identification of predictive markers for patient stratification has proven to be difficult.16 HSP90 inhibition might be an attractive strategy for therapeutic improvement. However, this requires studies on how HSP90 inhibitors act on tumor cells in this cancer type.
One approach to identify genes that determine cancer drug response was provided by highly parallel analysis of many cancer cell lines, specifically comparing their response with a large variety of small compounds with their gene expression patterns.17, 18 This strategy led to the identification of genes with expression patterns correlating with drug sensitivity. Notably, for many anticancer drugs this approach did not yield strong candidates, but in the case of the HSP90 inhibitor 17-N-allylamino-17-demethoxygeldanamycin (17-AAG) a strong correlation was seen between drug sensitivity and the expression of functional NAD(P)H dehydrogenase quinone 1 (NQO1), an enzyme that can metabolize and activate 17-AAG.19, 20 Hence, correlating drug sensitivity and gene expression patterns in cell lines can identify mechanisms that determine drug response.
Drugs are subjected to metabolic turnover, and a major route of excretion from the body consists in conjugation with a hydrophilic sugar moiety within the liver parenchyma, followed by secretion into the bile. A major group of enzymes that carry out such conjugations are the UDP glucuronosyltransferases (UGTs).21, 22, 23 These enzymes are the products of gene clusters that cover various substrate specificities. UGT substrates include bilirubin, amines, and phenol structures.24 The existence of such mechanisms for drug conjugation in the liver raises the question if and under what circumstances they can be found directly in tumor cells, and presumably cause drug resistance when highly expressed.
Here, we show that human CRC-derived cell lines fall into ganetespib-sensitive and -resistant groups. While the majority of CRC lines were sensitive, two lines were highly resistant. Importantly, resistant cancer cells show a high expression of the UGT1A gene, and high levels of UGT1A were shown to be critical for ganetespib turnover, drug inactivation, and cell resistance. Thus, UDP glucuronosyl conjugation detoxifies ganetespib not only in the liver but also in a subset of CRC cells, representing a potential predictive biomarker for ganetespib response in CRC and possibly other tumor types.
Results
Expression levels of UGT1A vary in CRC-derived cell lines, correlating with resistance to ganetespib
We tested the sensitivity of a panel of 11 CRC-derived cell lines toward ganetespib by analyzing their proliferation over a period of 4 days through quantitative light microscopy (Supplementary Figure 1). Drug concentrations that inhibited the growth rate by 50% were determined and found to vary strongly between cell lines, ranging from 36 to 2500 nM (Figure 1a). Cell lines with an IC50 of above 500 nM (SW1463 and HT29) were considered as resistant to ganetespib. Although different definitions of resistance versus sensitivity are employed throughout the literature, we will be using the above-described definition by proliferation-based IC50 throughout this manuscript. Next, we compared the pattern of ganetespib resistance with the respective whole-genome gene expression profiles that we had previously established for these cell lines.25 A number of genes were found to correlate in their expression with ganetespib resistance (Supplementary Table 1). Among them, the UGT1A gene stood out due to its known broad-range drug-metabolizing activity.26 We therefore independently assessed the expression levels of UGT1A by quantitative RT-PCR and confirmed that it closely correlated with ganetespib resistance (Figure 1a). Of note, HT29 cells and SW1463 cells showed high levels of UGT1A and were highly resistant to (i.e., proliferated despite high concentrations of) ganetespib. Corresponding elevated UGT1A protein levels were detected by immunoblot analysis in these two resistant lines but not in the sensitive cell lines (Figure 1b).
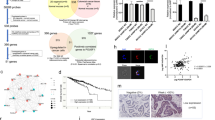
Ganetespib resistance and correlating expression levels of UGT1A in CRC-derived cell lines and primary tumors. (a) The growth inhibitory concentration of ganetespib was determined for 11 cell lines derived from CRC (blue columns), as detailed in Supplementary Figure S1. The corresponding expression levels of pan-UGT1A, initially found to correlate with ganetespib resistance through microarray analysis (Supplementary Table S1), were re-determined by reverse transcription and quantitative PCR (red columns). Correlation between ganetespib resistance and UGT1A expression levels was highly significant (R=0.976). (b) UGT1A protein levels were determined by immunoblot analysis for the same cell lines as in (a), to show that ganetespib-resistant cell lines have high levels of UGT1A protein. GAPDH detection serves as a loading control. (c) To analyze the distribution of UGT1A mRNA expression levels in primary colorectal tumors, microarray hybridization data sets from 217 rectal carcinomas were obtained.28, 29 The mean hybridization intensities from three probes corresponding to UGT1A (log2 scale) were normalized according to their deviation from the overall mean intensity of all tumors. The distribution of UGT1A levels was determined in the same way for 11 CRC cell lines.25 Overlay of both distributions reveals that tumors and cell lines display similar variations of UGT1A expression
UGT1A is highly expressed in normal colonic tissue,27 yet from our CRC cell line data the frequency of UGT1A expression was low. To better understand this, we analyzed UGT1A expression levels from a microarray database of >200 CRC patient tumors.28, 29, 30 From the results, a small but distinct subset of CRC tumors show elevated UGT1A expression that is comparable to the ganetespib-resistant cell lines (Figure 1c), suggesting that transformation may select against UGT1A expression in most but not in all CRCs.
Sensitivity to the HSP90 inhibitor 17-AAG does not correlate with UGT1A expression
We next sought to determine whether the UGT1A expression levels also correlate with the resistance of CRC cells toward a structurally different class of HSP90 inhibitor. However, as shown in Figure 2a, this was not the case. Although the HSP90 inhibitor 17-AAG (17-N-allylamino-17-demethoxygeldanamycin) also showed varying activity against the different cell lines, its sensitivity distribution did not correlate with the expression levels of UGT1A and only CaoCo2 cells did show pronounced resistance to 17-AAG. For geldanamycin derivatives such as 17-AAG31 and 17-DMAG (17-Dimethylaminoethylamino-17-demethoxygeldanamycin),32 it is known that the activation of these prodrugs is facilitated by NAD(P)H:quinone oxidoreductase 1 (NQO1), an oxidoreductase that reduces their quinone moieties. The 17-AAG-resistant cell line CaCo2 was previously reported to possess a polymorphism of the NQO1 gene (the *2 polymorphism), leading to rapid proteasomal degradation of the NQO1 enzyme and hence strongly reduced NQO1 activity,33 providing a mechanistic basis for their resistance to 17-AAG. Such pronounced resistance patterns were not observed for HSP90 inhibitors of different chemical classes such as PU-H71, an inhibitor based on a purine scaffold, and SNX-2112, an inhibitor that represents a class of its own (Figure 2b). In contrast, however, NVP-AUY922, a compound that is related to ganetespib, was again far less efficient when used on SW1463 and HT29 cells (Figure 2b, right), as already seen for ganetespib (Figure 1a). Cell viability assays largely reflected the effects observed with cell proliferation (Figure 2c). In sum, we conclude that UGT1A mRNA levels correlate with the resistance to ganetespib and its relative NVP-AUY922, but not to chemically different HSP90 inhibitors. Also the comparison of ganetespib, NVP-AUY922, 17-AAG, and 17-DMAG revealed that ganetespib- or NVP-AUY992-resistant cell lines could be highly sensitive to 17-AAG or its more hydrophilic relative 17-DMAG (e.g., HT29) or vice versa (e.g., SW480) (Figure 2c; see Supplementary Figure S2 for full concentration-response curves). Taken together, HSP90 inhibitors of different chemical classes show distinct activities against a panel of CRC cell lines, arguing that the resistance mechanisms are not directly related to the intrinsic properties of HSP90 and its clients, but rather result from an upstream phenomenon such as active drug stability.
Lack of correlation between UGT1A expression and 17-AAG sensitivity. (a) IC50 inhibitory concentrations of 17-AAG do not correlate to UGT1A levels and therefore not to ganetespib resistivity. The same cell lines described in Figure 1a were also assayed for 17-AAG sensitivity using the same cell proliferation assay, as described in Supplementary Figure S1. (b) IC50 determination with the Celigo system was performed as described in Supplementary Figure S1, for PU-H71, SNX-2112, NVP-AUY922, and ganetespib. The ganetespib-resistant cell lines SW1463 and HT29 are also resistant to NVP-AUY922, but do not exhibit resistance to drugs from different inhibitor classes. (c) IC50s vary strongly between colon cancer cell lines, but do not correlate between the resorcinol containing drugs ganetespib and NVP-AUY922 or the benzoquinone ansamycins 17-AAG and 17-DMAG. Full concentration-response curves are shown in Supplementary Figure 2
UGT1A knockdown sensitizes previously resistant CRC cells to ganetespib
To determine whether elevated UGT1A expression was directly responsible for ganetespib resistance, we reduced UGT1A mRNA and protein levels in the ganetespib-resistant cell line HT29 with two distinct siRNAs (s75 and s76), as validated by quantitative RT-PCR and immunoblot analysis (Figure 3a) and evaluated cell number and viability. From the results, decreased UGT1A expression enhanced the sensitivity toward ganetespib treatment, as revealed by reduced cell proliferation (Figure 3b) and reduced cell viability (Figure 3c). These data indicate that elevated UGT1A levels causally contribute to ganetespib resistance.
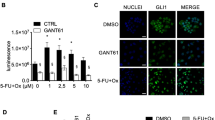
Sensitization toward ganetespib by UGT1A knockdown. (a) Reduction of UGT1A mRNA and protein levels by siRNA-mediated knockdown, validated by quantitative RT-PCR (left) and immunoblot analysis (right). (b) Diminished proliferation of HT29 cells in response to ganetespib upon UGT1A knockdown. Twenty-four hours after siRNA transfection, ganetespib was added to the cells at the indicated concentrations; ganetespib-containing medium was refreshed every 24 h. Cell confluence was determined by quantitative light microscopy (Celigo) 72 h after first addition of the drug. For (a and b) *P<0.05, **P<0.01, ***P<0.005 (Student’s T-test). (c) Diminished viability by ganetespib upon UGT1A knockdown. Upon treatment as in (b), cell viability was determined by assessing the ATP concentration in cell lysates using a luciferase assay
Several isoforms of UGT1A correlate with ganetespib resistance, and overexpressed UGT1A10 renders previously sensitive cells resistant
The UGT1A gene can be expressed in at least nine different but related isoforms, allowing the glucuronyl conjugation of a broad palette of substrates.26 To test which isoforms might be responsible for ganetespib resistance, we first performed selective RT-PCR assays to amplify groups of closely related isoform. This cluster approach had to be taken since some isoforms differ from each other only in a few nucleotide positions, making it virtually impossible to distinguish them by PCR primers. To account for this fact, primers were chosen to amplify groups of UGT1A isoforms based on primer sequences corresponding to UGT1A isoform 1, isoforms 3–5, isoform 6, and isoforms 7–10 (note that isoform 2 does not exist; for sequence alignments and primer sequences, cf. Supplementary Figure S4 and Supplementary Table S2). We found that the expression levels of isoform cluster 7–10 as well as cluster 3–5 correlated with ganetespib resistance (Figure 4a). It is important to keep in mind, however, that ganetespib resistance could simultaneously be conferred by several individual isoforms within a cluster, given the high similarity between them. As additional proof of principle for the causal role of UGT1A in ganetespib resistance, we chose isoform 10 as an example. We transfected the sensitive cell lines HCT116 and SW480 (both express low endogenous UGT1A levels, cf. Figure 1a) to overexpress UGT1A10. Indeed, this led to increased cell proliferation (Figure 4b) and increased cell viability (Figure 4c) in the presence of ganetespib. In conclusion, UGT1A expression is not only necessary for maintaining ganetespib resistance, but it is also sufficient to convert a sensitive cell line into a resistant one.
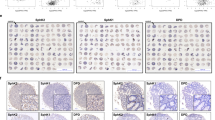
Ganetespib resistance by UGT1A overexpression. (a) To identify UGT1A isoforms that correlate with ganetespib resistance, we quantified the UGT1A gene clusters by selective RT-PCR analysis, and compared with a primer pair that amplifies all isoforms. This analysis revealed that the cluster composed of isoforms 7, 8, 9, and 10 is expressed at a particularly high level in the ganetespib-resistant cell line HT29. Thus, isoform 10 was chosen for overexpression in Figures 4b and c. (b and c) Overexpression of UGT1A10 confers ganetespib resistance. SW480 and HCT116 cells, both sensitive towards ganetespib, were transfected to overexpress UGT1A10. Cells were subsequently treated with ganetespib, (b) their confluence was determined 72 h later by Celigo, and (c) their cell viability was determined by ATP quantification via luciferase as in Figure 3c. *P<0.05, **P<0.01, ***P<0.005 (t-test)
UGT1A-expressing CRC cells fail to destabilize HSP90 client proteins in response to ganetespib, but not to 17-AAG
HSP90 stabilizes a number of cancer-driving proteins, and HSP90 inhibitors are known to decrease the abundance of such HSP90 clients by enhancing their proteasomal degradation.9, 12 We sought to determine whether UGT1A-mediated resistance to ganetespib is also mediated by maintaining the stability of HSP90 client oncoproteins. Indeed, immunoblot analysis revealed that upon treatment with ganetespib, resistant cells, which expressed high levels of UGT1A (HT29, SW1463, see Figure 1b) failed to decrease the levels of the representative HSP90 clients Wee1 and AKT. In contrast, sensitive cells, which expressed undetectable levels of UGT1A (SW480, HCT116, see Figure 1b) did destabilize their HSP90 clients (Figure 5a). In contrast, 17-AAG destabilized HSP90 clients in HT29, SW1463 and HCT116 cells but not in SW480 cells (Figure 5b), exactly reflecting the overall 17-AAG resistance pattern (Figure 2). Moreover, in direct support of a causal mechanism, UGT1A knockdown led to destabilization of HSP90 client proteins in ganetespib-resistant HT29 cells (Figure 5c). Conversely, UGT1A overexpression in originally ganetespib-sensitive HCT116 and SW480 cells stabilized HSP90 clients in the presence of ganetespib (Figure 5d). Taken together, cellular resistance to HSP90 inhibitors is reflected by the failure of such inhibitors to destabilize HSP90 client proteins.
UGT1A-expressing CRC cells fail to decrease HSP90 client protein levels in response to ganetespib, but not to 17-AAG. (a) Ganetespib-sensitive (SW480, HCT116) and -resistant (SW1463, HT29) cells were each treated with ganetespib, followed by immunoblot analysis of the HSP90 clients AKT and Wee1. Ganetespib addition reduced the levels of HSP90 clients only in sensitive cells, indicating that resistant cells preserve HSP90 function despite ganetespib. (b) Cells were treated and analyzed as in (a), but 17-AAG was used instead of ganetespib. Immunoblot analysis revealed that all cells except the 17-AAG-resistant SW480 cells (see Figure 2c) reduced the levels of HSP90 clients. (c) Knocking down UGT1A protein levels by siRNA in the originally ganetespib-resistant HT29 cells destabilizes HSP90 clients upon treatment with ganetespib in a dose-dependent manner. HT29 cells were first transfected with siRNA for 24 h, followed by incubation with ganetespib for 48 h at the indicated concentrations and immunoblot analysis of HSP90 clients. To avoid complete ganetespib turnover, ganetespib-containing media were renewed every 8 h during the incubation period. (d) Overexpression of UGT1A10 in ganetespib-sensitive cells stabilizes HSP90 clients despite the presence of the drug. HCT116 cells (top) or SW480 cells (bottom) were transfected with the overexpression vector or β-Galactosidase (βGal) vector as a control, and subjected to the indicated concentration of ganetespib for 48 h before harvesting. (a–d) GAPDH as a loading control
Ganetespib is a substrate for UDP-glucuronosyl conjugation by tumor cells
The resistance mediated by UGT1A strongly suggests that UGT1A conjugates ganetespib, leading to its inactivation and excretion from the tumor cells. To test this, we measured intracellular and secreted levels of ganetespib and its glucuronidated metabolites kinetically over 8 h in ganetespib-sensitive and -resistant CRC cell lines (Figures 6a and b). Ganetespib was highly stable in the sensitive cell lines but rapidly metabolized and excreted in the resistant cell lines, evident by a decrease in ganetespib levels and a rise in ganetespib glucuronides. Interestingly, the kinetics for ganetespib metabolism were slower in SW1463 compared with HT29, with ∼190 nM ganetespib present in SW1463 at 8 h compared with just 13 nM in HT29. This correlated with the level of UGT1A expression in these two lines (Figure 1b) and thus the greater ganetespib sensitivity of SW1463 cells compared with HT29 cells (IC50 of 100 nM versus 2000 nM, respectively, based on cell viability; cf. Figure 2c). Therefore, both the presence and level of UGT1A expression may be important indicators of ganetespib activity in CRC. In addition, a similar bioanalysis was performed on cell lysates and supernatant of HCT116 and HT29 cells treated with NVP-AUY922, another resorcinol-based Hsp90 inhibitor. An analogous accumulation of NVP-AUY922-glucuronide in the supernatant medium of HT29 cells was detected, whereas the HCT116 cells were unable to glucuronidate the drug (Figures 6c and d).
UDP glucuronosyl conjugation of ganetespib. (a and b) HCT116, SW480, SW1463, and HT29 cells were treated with 1 μM ganetespib. Cell lysates and culture media were collected at 5, 15, 30, 60, and 480 min. The total concentration of ganetespib and its glucuronidated metabolites in the lysates (a) and the supernatant media (b) were determined by LC-MS/MS. (c and d) HCT116 and HT29 cells were treated with 1 μM NVP-AUY922. Cell lysates and culture media were collected at 5, 15, 30, 60, and 480 min. The total concentration of NVP-AUY922 and its glucuronidated metabolites in the lysates and the supernatant media were determined by LC-MS/MS. The levels of glucuronidated NVP-AUY922 are given as NVP-AUY922 glucuronide peak area ratios
Discussion
Our results show that the level of UGT1A expression in colorectal cancer (CRC) cells correlates with their tolerance to high levels of ganetespib. The intracellular inactivation of the resorcinol containing Hsp90 inhibitors ganetespib and NVP-AUY922 by UGT1A-mediated glucuronidation and cellular excretion renders the drugs unable to effectively interact with HSP90. This in turn abrogates the biological activity of these drugs (i.e., makes them unable to degrade HSP90 clients), and prevents their cancer cell growth suppressing function.
Drug metabolism has long been known to be a primary determinant of pharmaceutical efficacy. Traditionally, however, the liver is regarded as the major source of drug metabolizing and deactivating enzymes. Genetic polymorphisms lead to individual variations of drug turnover rates in the liver, as exemplified by Cytochrome P45034, 35 and also by UGT1A.21, 23 Our findings contrast with this general view by identifying a highly variable drug metabolism directly in the tumor cells of a major cancer species, that is, CRC. This raises the specter that not only the patient’s overall genetic make-up, but also tumor cell-associated metabolism can strongly influence the efficacy of an anti-cancer drug. We show that this is true for a very promising class of drug candidates, the HSP90 inhibitors.
The clear correlation between UGT1A expression and resistance to ganetespib and NVP-AUY922 suggests that UGT1A detection in tumor biopsy specimens might allow the development of a specific biomarker that predicts the response to resorcinolic Hsp90 inhibitors. Our results argue that as soon as the clinical efficacy of ganetespib will be tested in a large cohort of CRC patients, this opportunity should be seized to evaluate UGT1A levels as a resistance marker, applying rigorous assessment according to the REMARK criteria.36
Aside from liver, it is the gastro-intestinal mucosa including normal colon cells that stand out as main tissues that synthesize UGT1A enzymes,37 perhaps as a first line to metabolize substrates before they even reach the liver. We speculate that some CRCs have maintained a subset of their gene expression patterns, thus causing enhanced UGT1A expression. In agreement, it has been reported that UGT1A levels drop in colon cancer as compared with normal colonic mucosa, albeit not in all cancer samples.38 If true, this would argue that UGT1A expression represents more of a problem when treating colon or liver cancers with ganetespib, as compared with other tumors that do not originate from the gastrointestinal tract. Accordingly, gene expression data from the human protein atlas (www.proteinatlas.org) reveal that UGT1A products are found in the gastrointestinal tract (mostly liver, gall bladder, small intestine, and colon), with some isoforms also detectable in the urinary tract (kidney and urinary bladder; cf. Giuliani et al.38). Similar patterns are observed in the corresponding cell lines derived from cancers of these two origins (protein atlas and Liu et al.39). Taken together, our data argue that UGT1A expression should be measured in pre-treatment biopsies of these two cancer species before treating them with ganetespib.
The UGT1A-deficient CRC cell lines used in the presented study are highly susceptible to ganetespib treatment in vitro. However, it has been previously shown in a xenograft model using a ganetespib-sensitive cell line (HCT116) that ganetespib as a single agent has only relatively modest effects on cancer growth rate (decrease of ∼50%). Rather, the drug has to be used in combination with conventional chemotherapeutics to exert a full effect in xenograft models and perhaps also in human cancer.40 Still, in-cell glucuronidation of the drug may predict susceptibility to at least one part of such a regimen, that is, the efficacy of HSP90 inhibition.
The comparison of drug sensitivity with gene expression patterns in a series of cell lines has previously led to the identification of some resistance-mediating genes.17, 18 However, this had been carried out on hundreds of different cell lines. In the novel example presented here, a relatively small number of cell lines already turned out to suffice for yielding a candidate gene that then proved to be responsible for drug resistance. We expect that, especially in the case of drug metabolizing genes, this approach may continue to yield clinically relevant resistance mechanisms that represent promising candidates for much-needed predictive biomarkers to properly stratify individual patients before therapy.
Materials and Methods
Statistical analysis of gene expression patterns and their correlation with drug sensitivities
mRNA expression levels were analyzed using log2 transformation and quantile normalization.41 Except for control spots, all 43 376 features were used without any a priori filtering. To determine significant differences of expression levels between the pooled groups of ganetespib-sensitive and ganetespib-resistant cell lines, a moderated Student’s t test was computed on a gene-by-gene basis. We applied an empiric Bayes estimator42 to compute the linear models for thousands of genes in parallel and assess their significance.
To not exceed a false-discovery rate (FDR) of 5%, the P-values were adjusted for multiple testing using the Benjamini-Hochberg method.43 All analyses were performed using the free statistical software R (version 2.15.2; available from www.r-project.org). Linear models were computed using the limma package.44
Cell culture, transfections, and treatments
Eleven cell lines derived from human colorectal cancers (CRCs) were cultured in RPMI (Invitrogen, Darmstadt, Germany) supplemented with 10% FCS (ThermoScientific, Waltham, MA, USA), 2 mM L-glutamine, and Penicillin-Streptomycin (Invitrogen). Cell-line cross-contamination was excluded using short tandem repeat profiling, as described.45 Specific knockdown of target genes was performed using pre-designed Silencer Select siRNAs (s231075 and s231076, designated s75 and s76) or control siRNAs (scrambled, all from Ambion, Hamburg, Germany) with a final concentration of 5 nM. Overexpression was performed with pCMV-SPORT6 vector constructs. For both upregulation and downregulation, Lipofectamine 2000 (Invitrogen) was applied in a forward transfection protocol. Ganetespib (Synta Pharma, Lexington, MA, USA) and 17-AAG (Calbiochem, Darmstadt, Germany) were dissolved in DMSO (50 and 5 mM stocks, respectively) and diluted to the indicated concentrations with culture medium.
Gene expression analysis by RT-PCR
Total RNA from cells was isolated using Trizol reagent (Invitrogen) according to manufacturer’s guidelines. Equal amounts of RNA were reverse-transcribed using M-MuLV Reverse Transcriptase (NEB) and real-time PCR analysis was performed using qPCR Master-Mix (75 mM Tris-HCl, pH 8.8, 20 mM (NH4)2SO4, 3 mM MgCl2, 0.3 M Trehalose (all from Roth, Karlsruhe, Germany), 0.01% Tween-20, 0.25% Triton X-100 (both from Applichem, Darmstadt, Germany), SYBR Green 1:80 000 (Fermentas, Schwerte, Germany), 0.2 mM dNTPs, 20 U/ml Taq-polymerase (both from Primetech, Minsk, Belarus) and 0.3 mM primers. Primers for total UGT1A: 5′-ATCTGCTTGGTCACCCGATG-3′ and 5′-TCCATGCGCTTTGCATTGTC-3′; for UGT1A1: 5′-GCCATTCCAAAGGGAGGATGTG-3′ and 5′-TGGGAACAGCCAGACAAAAGC-3′; for cluster UGT1A3-5: 5′-CATAATGAGGCCCTGATCAGGC-3′ and 5′-AATCGACAGGTACTTAGCCAG-3′; for UGT1A6: 3′-GCTGGTGGTCCCTCAGGAC-5′ and 5′-CAGCTCTTCTTGGTCATACGGC-3′; for cluster UGT1A7-10: 5′-CACAGTGCCCTGCTCCTC-3′ and 5′-GTTTGGAGAATTTCAGAGGCTATTTC-3′; for normalizer HPRT1: 5′-ATGCTGAGGATTTGGAAAGG-3′, and 5′-TCATCACATCTCGAGCAAGAC-3′. Primers (Metabion, Mansried, Germany) were used in a two-step protocol (2 min at 95 °C pre-heating; 40 cycles at 95 °C for 15 s followed by 58 °C for 1 min).
Immunoblot analysis and antibodies
Whole-cell lysates were made with RIPA buffer (1% Triton X-100, 1% Desoxycholat, 0.1% SDS, 150 mM NaCl, 10 mM EDTA, 20 mM Tris-HCl, pH 7.5, and complete protease inhibitor mix) and homogenized by sonication. After protein determination by BCA protein assay (Pierce, Bonn, Germany), equal amounts of total protein were separated by SDS gel electrophoresis, transferred onto nitrocellulose membrane (Millipore, Darmstadt, Germany), blocked and probed with the following antibodies (all in 5% milk in Tris-buffered saline solution containing 0.1% Tween-20): Wee1 (cs4936), AKT (cs9272; both from Cell Signaling, Frankfurt, Germany), GAPDH (ab8245; Abcam, Cambridge, UK), UGT1A (sc-271268), phos-ERK (sc-7383), ERK (sc-94; Santa Cruz, Heidelberg, Germany). Antibodies were detected with peroxidase-coupled secondary antibodies (Jackson, Newmarket, UK).
Proliferation Assay
For cell proliferation analysis, cells were seeded at 105 cells/well in 12-well plates 24 h before treatment. Cells were then treated with inhibitors with continuous change of inhibitor-containing medium and confluence measurement every 24 h. Over the course of the 4-day treatment, cell confluence was measured by bright-field microscopy using a Celigo Adherent Cell Cytometer (Brooks, Chelmsford, MA, USA). Confluence was calculated with the Celigo software program.
Viability Assay
Twenty-four hours after transfection, 5000 cells (HCT116, SW480) to 10 000 cells (HT29) were seeded in triplicate in 96-well plates 1 day before treatment. Cells were treated with inhibitors and drugs for 48 h and subjected to the CellTiterGlo Luminescent Cell Viability assay (Promega, Madison, WI, USA). This luciferase assay that determines the remaining ATP concentration thus reflects cell viability.
Bioanalysis
Colon cancer cells were treated with 1 μM ganetespib or NVP-AUY922 for 5, 15, 30, 60, or 480 min. At each time point, media was collected and cell lysates were generated for subsequent bioanalysis of secreted and intracellular concentrations of ganetespib and its glucuronides, or NVP-AUY922 and its glucuronides. Equal protein concentrations from cell lysates or equal volumes of media were used for the bioanalysis.
Samples were extracted by protein precipitation with methanol containing the internal standards (150 ng/ml [13C, 3H3]-ganetespib, 100 ng/ml [13C, 3H3]-ganetespib glucuronide #1 and [13C, 3H3]-ganetespib glucuronide #2), and analyzed by LC-MS/MS using an Agilent 1100 HPLC (Agilent Technologies, Santa Clara, CA, USA) interfaced to an API 4000 tandem mass spectrometer (Applied Biosystems, Foster City, CA, USA). The separation of NVP-AUY922 and its glucuronide were performed on a Kinetex 2.6 μm C18 (30 × 2.1 mm) column (Phenomenex, Torrance, CA, USA) with a run time of 3.5 min per sample using the mobile phase consisted of 0.1% formic acid (A) and acetonitrile including 0.1% formic acid (B). The conditions for elution were as follows: 20 95% B (0–1.7 min), 95% B (1.7–2.0 min), 95–20% B (2.0–2.1 min), and 20% B (2.1–3.5 min). The flow rate was 0.5 ml/min. Detection was accomplished in the positive electrospray ionization mode by selected reaction monitoring of the mass transitions m/z 365.3 323.3 for ganetespib, m/z 541.3 365.3 for ganetespib glucuronides, m/z 466.3 308.1 for NVP-AUY922, and m/z 642.3 466.3 for NVP-AUY922 glucuronide. Quantitation was done by extrapolation from a standard curve ranging from 2.5 to 5000 nM for NVP-AUY922 and from 1.0 to 5000 nM for ganetespib with 1/x2 weighting. Due to the lack of the authentic reference material, concentrations of NVP-AUY922 glucuronide were reported as analyte/internal standard peak area ratios.
Abbreviations
- 17-AAG:
-
17-N-allylamino-17-demethoxygeldanamycin
- 17-DMAG:
-
17-dimethylaminoethylamino-17-demethoxygeldanamycin
- CRC:
-
colorectal cancer
- FDR:
-
false-discovery rate
- HPLC:
-
high-performance liquid chromatography
- HSP90:
-
heat-shock protein 90
- MIF:
-
migration inhibitory factor
- NQO1:
-
NAD(P)H dehydrogenase quinone 1
- ROS:
-
reactive oxygen species
- RT-PCR:
-
quantitative real-time polymerase chain reaction
- UDP:
-
uridine diphosphate
- UGT1A:
-
UDP glucuronosyltransferase 1A
References
Li D, Marchenko ND, Moll UM . SAHA shows preferential cytotoxicity in mutant p53 cancer cells by destabilizing mutant p53 through inhibition of the HDAC6-Hsp90 chaperone axis. Cell Death Differ 2011; 18: 1904–1913.
Li D, Marchenko ND, Schulz R, Fischer V, Velasco-Hernandez T, Talos F et al. Functional inactivation of endogenous MDM2 and CHIP by HSP90 causes aberrant stabilization of mutant p53 in human cancer cells. Mol Cancer Res 2011; 9: 577–588.
Xu W, Mimnaugh E, Rosser MF, Nicchitta C, Marcu M, Yarden Y et al. Sensitivity of mature Erbb2 to geldanamycin is conferred by its kinase domain and is mediated by the chaperone protein Hsp90. J Biol Chem 2001; 276: 3702–3708.
Sato S, Fujita N, Tsuruo T . Modulation of Akt kinase activity by binding to Hsp90. Proc Natl Acad Sci USA 2000; 97: 10832–10837.
Schulz R, Streller F, Scheel AH, Ruschoff J, Reinert MC, Dobbelstein M et al. HER2/ErbB2 activates HSF1 and thereby controls HSP90 clients including MIF in HER2-overexpressing breast cancer. Cell Death Dis 2014; 5: e980.
Schulz R, Dobbelstein M, Moll UM . HSP90 inhibitor antagonizing MIF: the specifics of pleiotropic cancer drug candidates. Oncoimmunology 2012; 1: 1425–1426.
Kamal A, Thao L, Sensintaffar J, Zhang L, Boehm MF, Fritz LC et al. A high-affinity conformation of Hsp90 confers tumour selectivity on Hsp90 inhibitors. Nature 2003; 425: 407–410.
Whitesell L, Lindquist SL . HSP90 and the chaperoning of cancer. Nat Rev Cancer 2005; 5: 761–772.
Neckers L . Hsp90 inhibitors as novel cancer chemotherapeutic agents. Trends Mol Med 2002; 8 (4 Suppl): S55–S61.
Dobbelstein M, Moll U . Targeting tumour-supportive cellular machineries in anticancer drug development. Nat Rev Drug Discov 2014; 13: 179–196.
Supko JG, Hickman RL, Grever MR, Malspeis L . Preclinical pharmacologic evaluation of geldanamycin as an antitumor agent. Cancer Chemother Pharmacol 1995; 36: 305–315.
Proia DA, Bates RC . Ganetespib and HSP90: translating preclinical hypotheses into clinical promise. Cancer Res 2014; 74: 1294–1300.
Eccles SA, Massey A, Raynaud FI, Sharp SY, Box G, Valenti M et al. NVP-AUY922: a novel heat shock protein 90 inhibitor active against xenograft tumor growth, angiogenesis, and metastasis. Cancer Res 2008; 68: 2850–2860.
Sang J, Acquaviva J, Friedland JC, Smith DL, Sequeira M, Zhang C et al. Targeted inhibition of the molecular chaperone Hsp90 overcomes ALK inhibitor resistance in non-small cell lung cancer. Cancer Discov 2013; 3: 430–443.
Socinski MA, Goldman J, El-Hariry I, Koczywas M, Vukovic V, Horn L et al. A multicenter phase II study of ganetespib monotherapy in patients with genotypically defined advanced non-small cell lung cancer. Clin Cancer Res 2013; 19: 3068–3077.
Van Schaeybroeck S, Allen WL, Turkington RC, Johnston PG . Implementing prognostic and predictive biomarkers in CRC clinical trials. Nat Rev Clin Oncol 2011; 8: 222–232.
Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 2012; 483: 603–607.
Garnett MJ, Edelman EJ, Heidorn SJ, Greenman CD, Dastur A, Lau KW et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 2012; 483: 570–575.
Siegel D, Yan C, Ross D . NAD(P)H:quinone oxidoreductase 1 (NQO1) in the sensitivity and resistance to antitumor quinones. Biochem Pharmacol 2012; 83: 1033–1040.
Ross D . Quinone reductases multitasking in the metabolic world. Drug Metab Rev 2004; 36: 639–654.
Guillemette C . Pharmacogenomics of human UDP-glucuronosyltransferase enzymes. Pharmacogenomics J 2003; 3: 136–158.
Wu B, Kulkarni K, Basu S, Zhang S, Hu M . First-pass metabolism via UDP-glucuronosyltransferase: a barrier to oral bioavailability of phenolics. J Pharmaceutical Sci 2011; 100: 3655–3681.
Maruo Y, Iwai M, Mori A, Sato H, Takeuchi Y . Polymorphism of UDP-glucuronosyltransferase and drug metabolism. Curr Drug Metab 2005; 6: 91–99.
King CD, Rios GR, Green MD, Tephly TR . UDP-glucuronosyltransferases. Curr Drug Metab 2000; 1: 143–161.
Spitzner M, Emons G, Kramer F, Gaedcke J, Rave-Frank M, Scharf JG et al. A gene expression signature for chemoradiosensitivity of colorectal cancer cells. Int J Radiat Oncol Biol Phys 2010; 78: 1184–1192.
Rowland A, Miners JO, Mackenzie PI . The UDP-glucuronosyltransferases: their role in drug metabolism and detoxification. Int J Biochem Cell Biol 2013; 45: 1121–1132.
Strassburg CP, Manns MP, Tukey RH . Expression of the UDP-glucuronosyltransferase 1A locus in human colon. Identification and characterization of the novel extrahepatic UGT1A8. J Biol Chem 1998; 273: 8719–8726.
Grade M, Hormann P, Becker S, Hummon AB, Wangsa D, Varma S et al. Gene expression profiling reveals a massive, aneuploidy-dependent transcriptional deregulation and distinct differences between lymph node-negative and lymph node-positive colon carcinomas. Cancer Res 2007; 67: 41–56.
Ghadimi BM, Grade M, Difilippantonio MJ, Varma S, Simon R, Montagna C et al. Effectiveness of gene expression profiling for response prediction of rectal adenocarcinomas to preoperative chemoradiotherapy. J Clin Oncol 2005; 23: 1826–1838.
Gaedcke J, Grade M, Jung K, Camps J, Jo P, Emons G et al. Mutated KRAS results in overexpression of DUSP4, a MAP-kinase phosphatase, and SMYD3, a histone methyltransferase, in rectal carcinomas. Genes Chromosomes Cancer 2010; 49: 1024–1034.
Gaspar N, Sharp SY, Pacey S, Jones C, Walton M, Vassal G et al. Acquired resistance to 17-allylamino-17-demethoxygeldanamycin (17-AAG, tanespimycin) in glioblastoma cells. Cancer Res 2009; 69: 1966–1975.
Reigan P, Siegel D, Guo W, Ross D . A mechanistic and structural analysis of the inhibition of the 90-kDa heat shock protein by the benzoquinone and hydroquinone ansamycins. Mol Pharmacol 2011; 79: 823–832.
Siegel D, Anwar A, Winski SL, Kepa JK, Zolman KL, Ross D . Rapid polyubiquitination and proteasomal degradation of a mutant form of NAD(P)H:quinone oxidoreductase 1. Mol Pharmacology 2001; 59: 263–268.
Ingelman-Sundberg M . Pharmacogenetics of cytochrome P450 and its applications in drug therapy: the past, present and future. Trends Pharmacol Sci 2004; 25 (4): 193–200.
Hu L, Zhuo W, He YJ, Zhou HH, Fan L . Pharmacogenetics of P450 oxidoreductase: implications in drug metabolism and therapy. Pharmacogenet Genomics 2012; 22: 812–819.
Dancey JE, Dobbin KK, Groshen S, Jessup JM, Hruszkewycz AH, Koehler M et al. Guidelines for the development and incorporation of biomarker studies in early clinical trials of novel agents. Clin Cancer Res 2010; 16: 1745–1755.
Strassburg CP, Nguyen N, Manns MP, Tukey RH . UDP-glucuronosyltransferase activity in human liver and colon. Gastroenterology 1999; 116: 149–160.
Giuliani L, Ciotti M, Stoppacciaro A, Pasquini A, Silvestri I, De Matteis A et al. UDP-glucuronosyltransferases 1A expression in human urinary bladder and colon cancer by immunohistochemistry. Oncol Rep 2005; 13: 185–191.
Liu M, Wang Q, Liu F, Cheng X, Wu X, Wang H et al. UDP-glucuronosyltransferase 1A compromises intracellular accumulation and anti-cancer effect of tanshinone IIA in human colon cancer cells. PLoS One 2013; 8: e79172.
He S, Smith DL, Sequeira M, Sang J, Bates RC, Proia DA . The HSP90 inhibitor ganetespib has chemosensitizer and radiosensitizer activity in colorectal cancer. Invest New Drugs 2014.
Bolstad BM, Irizarry RA, Astrand M, Speed TP . A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 2003; 19: 185–193.
Smyth GK . Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 2004; 3 : Article3.
Hochberg Y, Benjamini Y . More powerful procedures for multiple significance testing. Stat Med 1990; 9: 811–818.
Wettenhall JM, Smyth GK . limmaGUI: a graphical user interface for linear modeling of microarray data. Bioinformatics 2004; 20: 3705–3706.
Han JY, Lim HS, Shin ES, Yoo YK, Park YH, Lee JE et al. Comprehensive analysis of UGT1A polymorphisms predictive for pharmacokinetics and treatment outcome in patients with non-small-cell lung cancer treated with irinotecan and cisplatin. J Clin Oncol 2006; 24: 2237–2244.
Acknowledgements
This work was supported by the Deutsche Forschungsgemeinschaft (DFG), the Klinische Forschergruppe 179 (KFO 179) ‘Biological Basis of Individual Tumor Response in Patients with Rectal Cancer’, the Deutsche Krebshilfe, the German José Carreras Leukemia Foundation, and the Wilhelm Sander foundation. HL was supported by the Göttingen Graduate School for Neurosciences, Biophysics and Molecular Biosciences.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
DAP, SH, and LSO are employed by Synta Pharmaceuticals.
Additional information
Edited by G Melillo
Supplementary Information accompanies this paper on Cell Death and Disease website
Rights and permissions
Cell Death and Disease is an open-access journal published by Nature Publishing Group. This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Landmann, H., Proia, D., He, S. et al. UDP glucuronosyltransferase 1A expression levels determine the response of colorectal cancer cells to the heat shock protein 90 inhibitor ganetespib. Cell Death Dis 5, e1411 (2014). https://doi.org/10.1038/cddis.2014.378
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2014.378
This article is cited by
-
Unravelling the clinicopathological and functional significance of replication protein A (RPA) heterotrimeric complex in breast cancers
npj Breast Cancer (2023)
-
Safety and efficacy of HSP90 inhibitor ganetespib for neoadjuvant treatment of stage II/III breast cancer
npj Breast Cancer (2022)
-
The HSP-RTK-Akt axis mediates acquired resistance to Ganetespib in HER2-positive breast cancer
Cell Death & Disease (2021)
-
Untangling the clinicopathological significance of MRE11-RAD50-NBS1 complex in sporadic breast cancers
npj Breast Cancer (2021)
-
Emerging roles for UDP-glucuronosyltransferases in drug resistance and cancer progression
British Journal of Cancer (2020)