Abstract
The impact of gut microbiota in eliciting innate and adaptive immune responses beneficial for the host in the context of effective therapies against cancer has been highlighted recently. Chemotherapeutic agents, by compromising, to some extent, the intestinal integrity, increase the gut permeability and selective translocation of Gram-positive bacteria in secondary lymphoid organs. There, anticommensal pathogenic Th17 T-cell responses are primed, facilitating the accumulation of Th1 helper T cells in tumor beds after chemotherapy as well as tumor regression. Importantly, the redox equilibrium of myeloid cells contained in the tumor microenvironment is also influenced by the intestinal microbiota. Hence, the anticancer efficacy of alkylating agents (such as cyclophosphamide) and platinum salts (oxaliplatin, cis-platin) is compromised in germ-free mice or animals treated with antibiotics. These findings represent a paradigm shift in our understanding of the mode of action of many compounds having an impact on the host–microbe mutualism.
Similar content being viewed by others
Facts
-
The anticancer efficacy of cyclophosphamide (CTX) and platinum salts is reduced in germ-free (GF) mice and animals treated with antibiotics.
-
Gut microbiota is involved in the reprogramming of intratumoral myeloid cells.
-
Gram-positive commensal bacteria translocate during chemotherapy and prime pathogenic Th17 (pTh17) cells contributing to the tumoricidal activity of cytotoxic compounds.
-
Intestinal microbiota is important for the bioactivity of immunomodulators (such as a combination of anti-IL-10R and Toll-like receptor 9 (TLR9) agonists).
Open Questions
-
To what extent gut microbiota interferes in the bioactivity of therapeutics?
-
How to noninvasively explore microbial dysbiosis or mucosal barrier dysfunctions at the level of the small intestine (SI) and large intestine?
-
How intestinal and systemic immunity are interconnected to modulate the effects of drugs?
-
How anticommensal immune responses correlate with specific anticancer cellular immunity and patient prognosis?
-
What are the most efficient probiotics capable of eliciting helper T-cell responses against cancer?
Gut Microbiota, Health and Diseases
The distal intestine of humans contains tens of trillions of microbes (of thousands different species representing over 2 kg of material). These microbiota and associated genomes (called ‘microbiome’) have been characterized by metagenomic analyses combining next-generation sequencing of both targeted (16S rRNA hypervariable region) and random (whole-genome shotgun) bacterial DNA sequences.1, 2 The first extensive catalog of the intestinal metagenome outnumbers the size of the human genome by a factor of 150.3 The human gut is colonized by microorganisms belonging to the domains Bacteria, Archaea, Eukarya and their viruses. The three main phyla characterizing the dominant human intestinal microbiota are Firmicutes, Bacteroidetes and Actinobacteria. Metagenomic studies revealed substantial interindividual variations at the genus and species levels, possibly because of genetic and environmental factors.3, 4 Recently, an interesting but still controversial notion has emerged as to the existence of ‘enterotypes’ characterized by dominant genera (Bacteroides, Prevotella and Ruminococcus) and their co-occurring phylogenetic traits that could be associated with long-term dietary habits.5, 6 The Bacteroides enterotype was associated with animal protein and saturated fats, whereas the Prevotella enterotype was predominantly observed with high fiber/plant-based nutrition and high carbohydrates (+low meat and dairy consumption).5, 7, 8 The third enterotype dominated by Ruminococcus often merged with the Bacteroides one.
The microbiome present in the distal gut performs myriad functions protecting the host against pathologies.3 Indeed, the host–microbiota symbiosis has evolved in three directions. First, colonization by commensal microorganisms is key to immune development.9, 10, 11, 12 Second, the commensal community keeps in check invading pathogens and prevents them from expressing virulence.13, 14 Third, the intestinal microbiota appears to digest glycans and regulate fat storage in mice and potentially in humans.15, 16 Exemplifying the host–microbe mutualism, the microbial genome is highly enriched in hundred families of glycoside hydrolases and in more than 20 families of polysaccharide lyases, whereas the human genome is relatively devoid of these carbohydrate-metabolizing enzymes.17 Finally, intestinal bacteria are essential for the postnatal development of the enteric nervous system in the mid-distal SI.18
The growing awareness of the importance of the gut microbiome in health and disease and recognition of the host–microbe mutualism at the immunological and metabolic levels become crucial for a better understanding of immunopathologies such as autoimmune and inflammatory disorders, allergy and obesity. Microbiome differences between controls and cases have been described for a variety of diseases such as inflammatory bowel disease (both Crohn’s disease and ulcerative colitis), obesity, type 2 diabetes, autism and allergies, and involve abnormalities in the relative abundance and representativity of distinct commensal bacteria. A ‘one microbe–one disease’ algorithm has yet only been described for a limited number of pathologies, such as Helicobacter pylori and gastric ulcers.19 However, it remains questionable whether a deviated repertoire of the intestinal microbiota, called ‘dysbiosis’, associated with an expanding list of chronic disorders20 may be seen as a causative agent in disease or is just a by-product of the disease. Transplantation experiments in which microbiota of a disabled mouse is grafted into a GF healthy recipient have highlighted that several disease phenotypes (such as adiposity, metabolic syndrome, colitis eventually causing cancer) can be transmitted by gut microbiota.20, 21, 22 Therefore, gut microbiota becomes progressively considered as a tractable environmental factor highly quantifyable, relatively stable, resilient within an individual and potentially drug targetable (prebiotics, probiotics). Hence, it becomes increasingly important to decipher the genetic potential (metagenomics) as well as the functions (metatranscriptomics) of the gut microbiome and its causal relationship with diseases.
Microbiome and Cancer
Cancer susceptibility and progression results from a complex interplay between gene regulation and the environment. Microbial communities inhabiting our intestine and other portals of entry represent so far unappreciated environmental factors that appear to have a role in carcinogenesis. Pioneering studies performed in GF mice or animals exposed to specific bacteria in specialized facilities (gnotobiotic mice) or in antibiotic-treated rodents revealed an unsuspected role of commensals or pathobionts in tumorigenesis driven or not by inflammation. In the genesis of colon cancer, at least in the 2% cases induced by a pre-existing inflammatory colitis, several studies demonstrated that microbiota can influence inflammation or innate immunity, genomic stability of intestinal epithelial cells (IECs) or the release of metabolites functioning as histone deacetylase (HDAC) inhibitors to regulate epigenetically host gene expression.22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34
Integrating all the current data, Tjalsma et al.35 proposed a bacterial driver–passenger model for microbial involvement in the development of colorectal cancer, implying that bacteria must be incorporated into the genetic paradigm of cancer progression. According to this model, distinct indigenous intestinal bacteria, the ‘driver bacteria’ would create DNA damage and drive genome instability to initiate the first steps of tumorigenesis. Bacterial drivers may progressively disappear in favor of opportunistic bacteria, that is, ‘passenger bacteria’, which then overwhelm the intestinal niche alterations and corrupt the local innate immunity. For instance, the human colonic bacterium, enterotoxigenic B. fragilis, induces colitis and colonic tumors in multiple intestinal neoplasia (Min) mice following signal transducer and activator of transcription 3 (STAT3) induction and IL-23-dependent Th17/γδT17 immune responses.36 Indeed, it appears that the NF-κB–IL-6–STAT3 cascade is a crucial regulator of the proliferation and survival of tumor-initiating IECs.37 Defective expression of several barrier proteins (due to genetic defects initiating the first step of colon cancer) facilitates the adenoma invasion of microbial products, which in turn stimulate tumor-associated myeloid cells. Then, such inflammatory monocytes and precursors become potent producers of IL-23, a protumorigenic cytokine.38
Microbiome and Immune Functions
Gut microbiota is critical for intestinal immune maturation, protecting the host against pathogens and damaging inflammatory reactions. GF mice have smaller Peyer’s patches, fewer plasma cells and impaired immunoglobulin A (IgA) secretion, fewer intraepithelial lymphocytes, as well as compromised release of antimicrobial peptides among other immunologic deficiencies.39 Many of these immune defects are corrected by recolonization with a healthy mouse commensal microbiota. SI immune maturation depends on a coevolved host-specific microbiota. Although gut bacterial numbers and phyla abundance were similar after transplantation of human or rodent fecal material into GF animals, bacterial species among Firmicutes differed and were associated with lower numbers of dendritic cells (DCs) and T cells, mostly proliferating and memory T cells (among which Th17 cells) in the SI lamina propria (LP) and mesenteric lymph nodes (LNs) and weaker protection against pathogens.40 These data underscore that exposure to just any gut commensal microbes or their pathogen-associated molecular patterns (PAMPs) (such as lipopolysaccharide (LPS) or lipoteichoic acid (LTA), etc.) is insufficient to induce intestinal immune maturation.
Commensal microbiota actively shapes intestinal T-cell responses. Among the commensal bacteria, some organisms appear to have a greater impact than others on mucosal immunity. Some studies suggest that there might be a compartmentalization of the microbiota along the gastrointestinal (GI) tract dictating mucosal immune homeostasis with Th17-dominated immune responses in the ileum and Treg in the colon. Commensal bacteria (such as B. fragilis) have a positive effect on Treg numbers. B. fragilis induces IL-10 production by Treg in the colon in a polysaccharide A-dependent manner.41 The Clostridium-forming spore (cluster IV and XIVa) induced robust differentiation of inducible Foxp3+ and IL-10+ Tregs in the mouse colonic LP.42 Segmented filamentous bacteria (SFB), found in numerous vertebrate species but not in humans after 3 years of age, are potent inducers of SI Th17.43, 44 SFB colonization protects mucosae from aggression induced by the enteropathogenic bacterium Citrobacter rodentium through a mechanism involving IL-17 and IL-22.43
Moreover, the intestinal microbiota can also influence systemic immune responses. A recent work highlighted the role of certain metabolites (short-chain fatty acid butyrate and to a lesser extent propionate) produced by commensal bacteria in dictating the extrathymic differentiation of peripheral regulatory T cells.45 Butyrate acts within T cells to enhance acetylation of the Foxp3 locus and protein, as well as DCs to decrease their proinflammatory NF-κB-dependent cytokine secretion profile through an HDAC inhibitory activity. Gut microbiota also controls the systemic Th17 pool. The incidence of an IL-17-dependent autoimmune arthritis developing in genetically susceptible hosts was reduced in GF conditions in which the autoantibodies titers as well as the splenic Th17 cells was also significantly decreased compared with specific pathogen-free (SPF) mice. Reintroduction of SFB into GF mice reconstituted the LP Th17 pool, raised the autoantibody titers and drove the development of arthritis.46 In another example of autoimmune disease, type 1 diabetes (T1D) developing in SPF-free non-obese diabetic (NOD) mice, MyD88 protein was mandatory to drive T1D. The effect was dependent on commensal microbes because GF MyD88-negative NOD mice developed T-cell priming in pancreatic LNs and islet β-cell infiltration by T lymphocytes, whereas colonization of these GF MyD88-negative NOD mice with altered Schaedler flora attenuated T1D. Moreover, MyD88 deficiency changed the composition of the caecum microbiota (reduced Firmicutes to Bacteroidetes ratio, increased Lactobacilli) and transmission of the microbiota of SPF MyD88-negative NOD donors attenuated T1D in GF NOD recipients.47
Altogether, the gut microbiota instructs the local and systemic immune system in a mutualistic/symbiotic manner.
Microbiome and Therapeutics
Mucositis (mucosal barrier injury) is a major oncological problem caused by chemotherapeutic agents used against malignancies. Oral and small (and to a lesser extent large) intestinal mucositis translating into a variety of clinical symptoms (diarrhea, vomiting) can be worsened by neutropenia and antibiotics. As IECs do not regulate intestinal homeostasis in a solely intrinsic manner but require symbiotic coordination with commensal bacteria and local gut leukocytic cells, the role of intestinal microbiota in the development and severity of mucositis induced by chemotherapeutic products has been proposed.48, 49
It is well established that chemotherapeutics elicit their proapoptotic activity against rapidly proliferating cell populations, meaning not only tumor cells but also intestinal stem/progenitor cells.50 However, rapid regeneration occurs between 96 and 168 h after doxorubicin with increased numbers of CD45− stem/progenitor cells, goblet cells, Paneth cells and enteroendocrine cells, crypt fission and crypts.51 The involvement of microbiota in the GI toxicity of irinotecan (CPT-11, 7-ethyl-10-(4-(1-piperidino)-1-piperidino) carbonyloxy-camptothecin) used to treat colorectal and other cancers has been reported.52 In vivo, CPT-11 is converted to the pharmacologically active SN-38, which is responsible for both antitumor activity and dose-limiting toxicity. SN-38 undergoes hepatic glucuronidation and is secreted into the bile as an inactive glucuronide SN-38G.53 Deconjugation of SN-38G in the colon by bacterial β-glucuronidases exposes intestinal epithelia to SN-38, mediating gut toxicity. Moreover, specific bacterial organisms translocate from the intestine of CPT-11-treated animals and cause systemic infection and sepsis. Prophylaxis with antibiotics reduced SN-38 concentration and/or diarrhea both in animal models and patients.54 Glutamine, a key ‘pharmaconutrient’, protects the gut during a variety of stress conditions,55 including cancer chemotherapy.56 Oral glutamine reduced the incidence and severity of late-onset diarrhea following CPT-11 treatment in rats. Dysbiosis induced by CPT-11-based chemotherapy increased the abundance of intestinal Enterobacteriaceae and Clostridium cluster XI. Glutamine mediated several potentially protective responses (such as heat-shock protein induction, reduced β-glucuronidase activity in the caecum), increased in the ratio of reduced to oxidized glutathione and memory CD8+ T cells in mesenteric LNs, eventually mitigating the dysbiosis induced by the tumor status and CPT11 administration.57
Besides the conventional cytotoxic compounds compromising cell cycle to facilitate apoptosis, a new class of therapeutic agents has emerged in the oncological armamentarium, whose mode of action is the effective blockade of T-cell inhibitory receptors (CTLA4, program death-1 (PD-1)). These antibodies also induce diarrhea and eventually colitis that could contribute to systemic inflammatory processes by mitigating intestinal homeostasis. Fagarasan and co-workers58 reported that PD1−/− mice presenting excess numbers of follicular helper T cells (overexpressing TGFβ1 and impaired for IL-21 production) exhibit a reduced affinity maturation of IgA with reduced bacteria-binding capacity, causing a strong bias in the gut microbiota composition (loss of anaerobic bacteria, loss of Bifidobacterium and Bacteroidaceae, increase in Enterobacteriaceae and at the genera level, increase in members of the Erysipelotrichaceae, Prevotellaceae, Alcaligenaceae and TM7 genera incertae sedis). The skewed gut microbial communities and the leaky gut barrier leads to a generalized activation of self-reactive B and T cells and production of autoantibodies.59
Therefore, we surmise that any compound compromising the intestinal barrier integrity and/or the innate mucosal immunity and/or directly the gut microbiota will affect the functional equilibrium of this compartment and cause symptoms, as well as distant immunological perturbations.
The Adjuvant Effects of Bacteria Against Cancer
Contrasting with the above-quoted exemplifications of bacteria driving carcinogenesis, other observations support a beneficial role of distinct bacteria against cancer. Hence, a prolonged combination of metronidazole and ciprofloxacine tripled breast cancer occurrence in proto-oncogene HER-2/neu-driven transgenic (Tg) mice.60 In humans, some epidemiologic studies suggested a dose-dependent association between antibiotic use and risk of breast cancer.61, 62
Since the first report in 1976, accumulated clinical evidence has supported intravesical bacillus Calmette–Guerin (BCG) therapy as one of the standard methods of management of intermediate- and high-risk non-muscle invasive bladder cancer.63 Intravesical immunotherapy with BCG to prevent recurrence of these tumours has been shown to involve the participation of three different TLRs (TLR2, TLR4 and TLR9).64, 65 Despite the fact that BCG is viewed by the immune system as ‘pathogenic’, BCG inoculations performed as classical vaccines failed to prevent relapse of stage II metastatic melanoma in pioneering studies.66
Instead, distinct commensals could have a beneficial role in melanoma. Intratumoral inoculation of 3 mg of heat-killed Propionibacterium acnes in subcutaneous melanoma could promote local and systemic Th1 and Tc1 responses associated with in situ granuloma formation and tumor regression.67 P. acnes is recognized by TLR2 on monocytes, macrophages and DCs, leading to the activation of IL-12 promotor.68 Therefore, it is conceivable that certain commensals be involved in the natural immunosurveillance of malignancies exposed to the portals of entry (such as ulcerated melanoma). Hence, lung tumors presenting tertiary lymphoid organogenesis exhibit a more favorable prognosis, perhaps towing to chronic stimulation with environmental microorganisms.69
Beneficial Effects of Commensals During Cancer Therapy
A role for gut commensals in dictating the response of subcutaneous tumors to cytotoxic or immunomodulatory compounds was unsuspected so far. In recent reports, we and others demonstrated that gut microbiota is indispensable for the immunomodulatory and antitumor effects of certain anticancer therapeutics including CTX and platinum salts.70, 71 We surmise that the scenarii capturing the biology of these effects could be recapitulated as follows.
Alkylating agent CTX mobilizes Gram-positive bacteria from the gut to secondary lymphoid organs
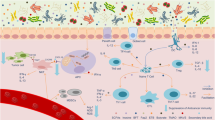
CTX is a DNA-alkylating agent belonging to the family of nitrogen mustards, which entered the clinical practice not only as an anticancer agent but also for the therapy of autoimmune diseases, including systemic lupus erythematosus, rheumatoid arthritis and multiple sclerosis. Its pharmacodynamic profile indicates that CTX mediates immunosuppressive properties at high doses, whereas metronomic CTX regimens exert immunostimulatory effects.72 Several lines of evidence underscore that CTX can also induce a systemic differentiation of Th1 and Th17 cells, as well as γδT17 cells in a dose-dependent manner, in mice and humans, which may interfere in several immunopathologies or malignancies.73 Recently, we and others reported that gut microbiota is indispensable for the immunomodulatory and antitumor effects of certain anticancer therapeutics including CTX and platinum salts (Figure 1).70, 71
Mechanisms involved in the action of gut microbiota during cancer therapies. Certain chemotherapeutics (presumably those affecting the integrity of the gut barrier (1), e.g. CTX and oxaliplatin) facilitate the translocation of distinct bacteria or bacterial products (2) (Gram-positive for CTX), causing a cascade of systemic immune effects translating in the control of tumor outgrowth (3). Viaud et al.70 (2, 3) showed the role of specific Gram-positive commensal bacteria in the elicitation of pTh17 responses (2) associated with Th1 accumulation in tumor beds (3). Iida et al.71 demonstrated the role of ROS and TNF-producing myeloid cells conditioned by the presence of gut commensal bacteria in the efficacy of oxaliplatin and immunomodulators (4, 5), respectively, in triggering early tumor cytotoxicity. These effects account for the reduced efficacy of cytotoxic compounds in animals cotreated with broad-spectrum (or vancomycin or colistin) antibiotics (6)
First, we observed that certain bacterial species of the SI can selectively and efficiently translocate within 24–48 h after exposure to an alkylating agent, CTX, yet administered at a metronomic regimen only reducing B-cell numbers.70, 73 Indeed, cultivation on blood agar plates (in both aerobic and anaerobic conditions) of mesenteric LNs and spleens of CTX-treated mice revealed the specific outgrowth of high numbers of colonies, identified as Lactobacillus johnsonii or Enterococcus hirae using mass spectrometry. Second, at these early time points, the permeability of the intestinal barrier was readily increased, whereas the number of Th17 cells and CD103+ DCs accumulating in the LP significantly decreased, setting the stage for bacterial translocation. Third, within 7 days after CTX administration, the polarity of splenic T cells was geered toward a mixed Th1 and Th17 pattern, whereas a small proportion of CCR6+IL-17+RORγt+ CD4+ Th17 cells became CXCR3+, T-bet+ and IFNγ+, suggesting the acquisition of a ‘pTh17’ phenotype. Interestingly, the elicitation of pTh17 cells was significantly reduced in GF mice or animals treated with broad-spectrum antibiotics (ATB; antibiotics: streptomycin+ampicillin+colistin) or vancomycin (killing Gram-positive bacteria), supporting the notion that gut microbiota was involved in the CTX-mediated splenocyte polarization. Reinforcing this notion, pTh17 cells could not be increased in Myd88−/− mice treated with CTX, whereas somewhat enhanced in Nod2−/− counterparts. Of note, CTX-induced IFNγ-producing αβ+TCR (T-cell receptor) CD8+ T cells and γδTCR+ T cells were not dependent on gut microbiota in the same experimental conditions.
Reprogramming of intratumoral myeloid cells in the absence of gut microbiota
Iida et al.71 compared the gene expression profiling of three transplantable tumor models growing in mice treated with ATBx (versus no ATBx) (Figure 1). The ATBx therapy profoundly downregulated genes related to inflammation, phagocytosis, antigen presentation and adaptive immunity while upregulating those encoding tissue development, cancer and metabolism. Notably, ATBx decreased the recruitment of monocyte-derived major histocompatibility complex (MHC) class II+Ly6Chigh and Ly6G+ cells in tumors and spleens. Based on previous work showing the TNFα- (and CD8+ T-cell-) dependent antitumor efficacy of a combined therapy associating CpG oligodeoxynucleotides and anti-IL-10R antibodies,74 the authors went on to demonstrate that commensal bacteria were required to prime tumor-associated innate myeloid cells for the CpG-induced inflammatory cytokine (IL-1α, IL-1β, IL-12β, CXCL10) production and TNF-mediated necrosis, both indispensable for the tumor regression. In accordance with these data, Tlr4−/− mice failed to fully respond to CpG+anti-IL-10R combinatorial regimen, whereas LPS could partially restore TNF production by intratumoral antigen-presenting cells in ATBx-treated wild-type (WT) mice. Principal component analysis of the microbiota composition versus TNF production in tumors showed a codependence. Among the positive correlates stood out the Alistipes and Ruminococcus genera, whereas Lactobaccili negatively associated with TNF release. Importantly, reconstitution of mice preexposed to ABTx with Alistipes shahii restored the capacity of myeloid cells to secrete TNF, whereas L. fermentum failed to do so.
The authors extended the role of microbiota in modulating the intratumoral myeloid innate cell phenotype in tumors treated with platinum salts. Both oxaliplatin- and cisplatin-mediated tumoricidal activity against MC38 and EL4 tumors were more efficient in SPF mice than in ATBx-treated or GF counterparts. Gene expression analysis revealed that the induction of inflammatory mediators was reduced in the absence of gut microbiota at 18 h after oxaliplatin. ATBx therapy attenuated the oxaliplatin-induced expression of Cybb encoding reactive oxygen species (ROS)-generating nicotinamide adenine dinucleotide phosphate oxidase (NOX2) that mainly resided in intratumoral neutrophils and macrophages. Inhibition of ROS by genetic (Cybb−/−) or pharmacological (N-acetyl cysteine) maneuvers, as well as depletion of Gr1+ cells, interfered in the oxaliplatin-mediated tumoricidal activity. These findings underscore that the reduced efficacy of oxaliplatin in ATBx or GF mice could be attributed, at least partially, to an impaired ROS production by myeloid cells. Altogether, Iida et al.71 unraveled the unsuspected role of commensal bacteria in affecting the type of inflammatory microenvironment required for a TNF- or ROS-dependent therapeutic effect mediated by various immunomodulators or cytotoxics.
Host’s immunization against commensal bacteria in the course of chemotherapy
To address whether ignorance or tolerance toward gut commensals has been broken after CTX, as described following oral infection with Toxoplasma gondii,75 we used two different experimental approaches (Figure 2). First, we performed an adoptive transfer of CBir-specific TCR Tg CD4+ T cells prone to recognize a flagellin epitope of a Clostridium76 into congenic C57BL/6 mice. Then, we monitored their proliferation and polarization as effector cells as well as their memory response directed against the Clostridium peptides. Indeed, CBir-specific TCR Tg lymphocytes accumulated and differentiated into IL-17-producing cells in CTX but not into sham-treated mice. Moreover, following restimulation of splenocytes with CBir peptides, IFNγ secretion was markedly enhanced in CTX-treated recipients but not in control recipients. Second, we addressed whether the Gram-positive translocating bacterial isolates (that could be cultivated and reused in transplantation studies) could mediate pTh17 responses in the spleen after gut recolonization of SPF animals sterilized by a 21-day broad-spectrum ATB regimen (ampicillin, streptomycin, colistin). Indeed, the cocktail of L. johnsonii+E. hirae restored the pTh17 splenic pool generated after CTX, whereas L. plantarum or L. reuteri failed to do so.
Three ways to demonstrate that naive mice can get immunized against their commensal bacteria during CTX therapy. (Upper panel) Reconstitution (by oral gavage using L. johnsonii+E. hirae) of mice presterilized with broad-spectrum antibiotics (ATB) restores the pool of splenic pathogenic Th17 cells (coexpressing the transcription factors T-bet and RORγt). (Middle panel) Restimulation of splenic T cells from mice treated with CTX using syngeneic bone marrow-derived DCs loaded with distinct commensal bacteria (such as L. johnsonii or E. hirae) reactivates memory T cells that produce high levels of interferon-γ (IFNγ). (Lower panel) Adoptive transfer of TCR Tg CD4+ T cells recognizing a flagellin peptide of Clostridium into naive C57BL/6 mice treated or not with CTX. After 1 week, Tg T cells harboring a congenic marker can be analyzed by FACS to determine IL-17 production (in intracellular staining) as well as IFNγ release following restimulation of splenocytes with MHC class II-restricted flagellin peptides
Third, we analyzed memory T-cell responses directed against a variety of Gram-positive bacteria including the translocating bacterial species as well as others (such as E. coli, E. faecalis and LPS). In 50% and 30% cases, CTX could elicit memory Th1 responses against L. johnsonii, and E. hirae or E. faecalis, respectively. Altogether, these data indicate that CTX facilitates the priming of effector pTh17 and memory Th1 cell responses directed against distinct commensals in distant secondary lymphoid organs.
Tumor invasion by Th1 cells is affected by antibiotics, which concomittantly compromised the efficacy of cytotoxic compounds against cancers
We next addressed whether broad-spectrum ATB or vancomycin- or colistin-based antibiotherapy would affect the anticancer efficacy of CTX in various tumor models. In two transplantable tumor models (P815 mastocytoma and MCA205 sarcoma) syngeneic of DBA2 and C57BL/6 mice, respectively, the CTX-mediated control of tumor outgrowth was significantly impaired by either one or all ATB regimen, supporting a beneficial role for intestinal commensals in the tumoricidal activity of CTX. Of note, at this metronomic dosing, CTX induces T-cell-dependent antitumor effects.72 Vancomycin indeed altered the CTX-mediated recruitment of CD3+ TILs into the MCA205 sarcoma and markedly compromised the accumulation of Th1 TILs. We corroborated these results in a spontaneous lung carcinoma model (as initially described by Jacks and co-workers77 and Cortez-Retamozo et al.78, 79). Eight-week-old KP (KrasLSL-G12D/WT; p53Flox/Flox) mice received an adenovirus-expressing Cre recombinase (Ad-cre) by intranasal instillation to initiate lung adenocarcinoma (d0). Mice were either left untreated or received CTX-based chemotherapy (d84, d91 and d98) in the absence or presence of 0.25 mg/ml vancomycin (mixed into drinking water) starting on d77 after Ad-cre and until the end of the experiment to test the inhibitory role of vancomycin-based antibiotherapy on the anticancer efficacy of a successful chemotherapy. In this preclinical model mimicking human tumorigenesis, we validated the concept that the eradication of Gram-positive bacteria by vancomycin compromised the efficacy of CTX-based chemotherapy, correlating with a reduced intratumoral CD8+ T effector/Foxp3+ regulatory T-cell ratio. Thus, Gram-positive bacteria appear to be necessary for the optimal efficacy of the CTX-induced anticancer immune response and tumor mass reduction. Finally, to demonstrate a cause–effect relationship between the lack of elicitation of pTh17 cells by commensals and the loss of chemotherapeutic efficacy observed because of vancomycin cotreatment, we used a transplantable tumor model in which we transferred ex vivo-expanded Th17 derived in various cytokine media to exhibit a regulatory versus pathogenic Th17 phenotype. Indeed, infusion of pTh17 could restore chemosensitivity in vancomycin and CTX cotreated animals, whereas that of regulatory Th17 failed to do so.
Concluding remarks and discussion
These data support the concept that distinct commensals (such as L. johnsonii+E. hirae) niching in the SI of tumor bearers could elicit pTh17 cells in the spleen after translocation into secondary lymphoid organs, such as pTh17 appearing capable of developing into memory Th1 cells eventually accumulating in inflammatory lesions such as growing tumors. These adaptive T-cell responses directed against commensals are occuring consecutively to earlier events modulating the functions of antigen-presenting cells in tumor beds. Several important points are intriguing and remain to be investigated.
First, what molecular or metabolic cues support the cell stress and damage of the intestinal barrier triggered by pharmaceutical compounds (such as CTX, oxaliplatin, TLR9L, anti-IL-10R, etc.) that generate such a ‘helper’ immunity? Wide investigations at several levels (apoptosis, necroptosis, autophagy, activation of the NF-κB pathway, inflammasomes, TLR/NOD-like receptors (NLRs), cytokine receptors (TNFα, IL-17, IL-22, IL-18, IL-22BP, etc.), hematopoietic and/or epithelial-driven signaling pathways (Tables 1 and 2) will be mandatory to nail down the principal components leading to this favorable dysbiosis.
Second, how can microbial structures be recognized by pattern recognition receptors expressed by immune or parenchymal cells and generate innate immune responses, which in turn shape adaptive immunity against microbial and tumoral antigens? These considerations have been largely discussed in the context of organ transplantation by Alegre et al.80 and Chong and Alegre,81 where infections and/or tissue damage directly or indirectly affect alloreactivity and the outcome of transplanted allografts. Hence, three scenarii can be envisioned to explain how T-cell responses elicited by commensals could influence antitumor immunity. First, antigen cross-reactivity or superantigen-driven responses could account for T-cell-dependent tumor regression. Second, cell-intrinsic effects such as direct activation of antigen-presenting cells located in tumor beds by bacterial cell walls or microbial components could reset myeloid cell functions or reprogram Treg bearing TLRs and/or parenchymal cells that respond to TLRs by chemokine or inflammatory cytokine release or apoptosis.82 Third, the differentiation of anticommensal T cells could provide helper cytokines or costimulatory factors for anticancer T cells to be driven. The role of IL-2/IL-15, type 1 or type 2 IFNs or CD40/CD40L interactions could be explored among other candidates.
Third, the precise mode of selection of distinct commensal bacteria/pathobionts by CTX remains obscure. Mucosal integrity (bacterial dysbiosis, loss of Th17 and CD103+CD11b+ DCs, upregulation of lysozyme M) was affected at the level of the SI (more than in the colon) with a positive gradient from the ileum to the duodenum. Many Lactobacilli and E. hirae strains have been described to be highly resistant to acidic pH and bile salts (compared with Lactococci), explaining their relative abundance in the SI downstream of the stomach.83 Physical properties might account for the translocation of distinct bacterial species. Hence, mucosal bacteria that adhere to intestinal mucosal surfaces and epithelial cells84 might be more prone to translocation in case of loss of mucosal permeability. Many Lactobacilli strains and E. hirae belong to this category of ‘mucosal bacteria’, which are more prone to translocation or invasion upon loss of the barrier function of tight junctions85 than ‘luminal’ bacteria. The metabolic properties of distinct bacteria might also explain their selective accumulation or resistance after CTX. Thus, in our study, the identified translocating bacteria were all facultative anaerobes (or microaerophilic ones) and not strict anaerobes, perhaps reflecting the presence of a relative hypoxia (but not total anoxia) during the translocation.
Fourth, the links between bacterial translocation and SI dysbiosis remain unsolved. The kinetics relationship between bacterial translocation and gut dysbiosis indicate that dysbiosis detected in the SI mucosa established late when pTh17 are already primed and might influence other parameters than those described in the first study.
Finally, the functional links between the two pioneering studies70, 71 have to be deciphered. It is conceivable that bacterial products or bacteria could modulate the tone of the tumor microenvironment through metabolic changes, setting the stage for restoration of T-cell functions, anergized in the context of tumor-induced tolerance. pTh17 cells and memory Th1 cells elicited against commensal bacteria might preferentially accumulate in inflammatory tumor microenvironment, already primed by bacterial products or ligands for pathogen recognition receptors. Hence, various commensal or pathobionts may act in concert to modulate systemic and local immunity during cancer regimens.
Future Challenges for the Exploitation of the Host–Bacteria Symbiosis/Dysbiosis During Oncogenesis
Interfering against inflammation has been a source of inspiration for chemoprevention. Apart from aspirin that reduces the incidence of distinct colon cancers and other adenocarcinomas, other strategies based on our expanding knowledge of gut microbiota are being pursued.86 Although a beneficial role for broad-spectrum antibiotics has been shown in many experimental settings to reduce inflammation-induced cancers, this approach would select antibiotic-resistant strains and eliminate species involved in gut homeostasis. Instead of killing indiscriminately all bacteria, restoring an ‘ideal’ microbial composition could theoretically be a more suitable option. Fecal microbiota transplantation was shown to be effective in diarrhea caused by a dysbiosis dominated by Clostridium difficile by suppressing or displacing C. difficile.87, 88
Probiotics and prebiotics represent more common ways to establish/maintain healthy microbiomes. In individuals presenting with lactose intolerance (5–15% frequency in northern Europe), the beneficial effect of live Lactobacilli residing in non-pasteurized yogurt relies on the provision of β-galactosidase activity.89 Genetically modified bacteria may even have stronger effects. A L. acidophilus strain harboring a deletion in the phosphoglycerol transferase gene and unable to synthetize LTA prevented the progression of colonic polyps in ApcDflox mice.90 Elafin-overexpressing L. casei and L. lactis reduced colitis in mice and ex vivo in inflamed epithelial cells from human colitis.91 L. gasseri genetically modified to overexpress superoxide dismutase decreased colitis in IL-10-deficient hosts.92
Prebiotics refer to indigestible food ingredients that selectively promote the colonization of healthy commensals such as the dietary fiber inulin that promotes Bifidobacteria growth. More specifically, cancer-preventive antioxidants include dietary polyphenols (flavonoids, phenolic acids, lignins present in tea, wine, nuts, fruits, and so on, and ellagic acid metabolized by colonic microbiota into urolithins exhibiting antiestrogenic and anti-COX2 activities).93 Another polyphenol called ‘daidzein’, a soy isoflavone metabolized by gut microbiota into equol and only detected in a fraction of individuals (harboring sulfate-reducing bacteria), may protect against breast and prostate cancer, mostly in Asia.94 The fiber has been involved in the prevention of colorectal cancers and butyrate, one of the most abundant short-chain fatty acids resulting from the bacterial fermentation of fibers and selectively transported into the colon epithelium is the most compelling tumor-suppressive molecule. Butyrate has both cell autonomous and cell extrinsic antitumor effects. It decreases proliferation and promotes apoptosis of tumor cells, ameliorates inflammation associated with colitis and favor expansion of peripheral Treg. Most of these effects result from epigenetic regulation, butyrate acting as an endogenous HDAC inhibitor.95, 96
Assuming a causal and linear relationship between translocation of Gram-positive bacteria, pTh17 responses and antitumor immunity leading to tumor control, one might think about exploiting the adjuvanticity of gut commensals to ameliorate the effects of chemotherapeutics. Understanding the molecular cues underlying the immunogenicity of L. johnsonii+E. hirae may unravel novel PAMPs and/or novel patterns of T-cell differentiation that could shape the design of future cancer vaccines. Modifying these bacteria to uncouple their potential pathobiontic from their commensal properties may be mandatory for a future development of such probiotics as ‘adjuvantizers’ of chemotherapeutics.
Future prospects for a better management of cancer patients aim at (i) diagnosing patients dysbiosis (metagenomics, metatranscriptomics, epidemiology on diet, medications and exercise, and so on), (ii) compensating dysbiosis by appropriate ‘immunogenic probiotics’, WT or genetically modified to overexpress specific functions, (iii) prebiotics synergizing with probiotics to set the stage for a healthy intestine that has been compromised by DNA-damaging agents, (iv) monitoring the immune responses raised against the relevant commensals to establish a correlation with long-term benefit and immune fitness (Figure 3).
Future prospects and clinical implementations of this work. To date, patients who are being diagnosed with cancer undergo a pathological biopsy and imaging tomography/magnetic resonance to analyze spreading of the malignant process. In the near future, we will need to investigate their intestinal microbiota and their systemic anticancer and antimicrobial immunity to be able to adapt/personalize the oncological therapy according to their microbial dysbiosis or immune dysfunctions. Specific food intake as well as probiotics composed of immunogenic (and safe) commensal/pathobionts could precede chemotherapeutics to facilitate their tumoricidal activity through bacterial adjuvantization
Abbreviations
- AhR:
-
aryl hydrocarbon receptor
- AOM:
-
azoxymethane
- ASC:
-
apoptosis-associated speck-like protein
- ATB:
-
antibiotics: streptomycin+ampicillin+colistin
- ATBx:
-
antibiotics: imipenem+vancomycin+neomycin
- BCG:
-
bacillus Calmette–Guerin
- BM:
-
bone marrow
- CTX:
-
cyclophosphamide
- DC:
-
dendritic cell
- DSS:
-
dextran sodium sulfate
- GF:
-
germ free
- GI:
-
gastrointestinal
- HDAC:
-
histone deacetylase
- IEC:
-
intestinal epithelial cell
- FADD:
-
Fas-associated protein with death domain
- GC-C:
-
guanylate cyclase C
- hPepT1:
-
human intestinal H-coupled oligonucleotide transporter
- IDO:
-
indoleamine 2,3-dioxygenase
- Ig:
-
immunoglobulin
- LN:
-
lymph node
- LP:
-
lamina propria
- LPS:
-
lipopolysaccharide
- LTA:
-
lipoteichoic acid
- MHC:
-
major histocompatibility complex
- NEMO:
-
NF-κB essential modulator
- NLR:
-
NOD-like receptor
- NLRP:
-
NLR family pyrin domain-containing protein
- PAMP:
-
pathogen-associated molecular pattern
- PD-1:
-
program death-1
- PRR:
-
pathogen recognition receptor
- PSA:
-
polysaccharide A
- pTh17:
-
pathogenic Th17 cells
- ROS:
-
reactive oxygen species
- SFB:
-
segmented filamentous bacteria
- SI:
-
small intestine
- SPF:
-
specific pathogen free
- STAT:
-
signal transducer and activator of transcription
- TAK1:
-
TGF-β-activated kinase 1
- TCR:
-
T-cell receptor
- Tg:
-
transgenic
- TLR:
-
Toll-like receptor
- TNBS:
-
2,4,6-trinitrobenzene sulfonic acid
- VDR:
-
vitamin D receptor
- WASP:
-
Wiskott–Aldrich syndrome protein
- WT:
-
wild type
References
Cancer Genome Atlas Network Comprehensive molecular characterization of human colon and rectal cancer. Nature 2012; 487: 330–337.
Weinstock GM . Genomic approaches to studying the human microbiota. Nature 2012; 489: 250–256.
Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 2010; 464: 59–65.
Costello EK, Lauber CL, Hamady M, Fierer N, Gordon JI, Knight R . Bacterial community variation in human body habitats across space and time. Science 2009; 326: 1694–1697.
Arumugam M, Raes J, Pelletier E, Le Paslier D, Yamada T, Mende DR et al. Enterotypes of the human gut microbiome. Nature 2011; 473: 174–180.
Gophna U . Microbiology. The guts of dietary habits. Science 2011; 334: 45–46.
Wu GD, Chen J, Hoffmann C, Bittinger K, Chen YY, Keilbaugh SA et al. Linking long-term dietary patterns with gut microbial enterotypes. Science 2011; 334: 105–108.
De Filippo C, Cavalieri D, Di Paola M, Ramazzotti M, Poullet JB, Massart S et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc Natl Acad Sci USA 2010; 107: 14691–14696.
Duarte R, Silva AM, Vieira LQ, Afonso LC, Nicoli JR . Influence of normal microbiota on some aspects of the immune response during experimental infection with Trypanosoma cruzi in mice. J Med Microbiol 2004; 53 (Part 8): 741–748.
Neumann E, Oliveira MA, Cabral CM, Moura LN, Nicoli JR, Vieira EC et al. Monoassociation with Lactobacillus acidophilus UFV-H2b20 stimulates the immune defense mechanisms of germfree mice. Braz J Med Biol Res 1998; 31: 1565–1573.
Souza DG, Vieira AT, Soares AC, Pinho V, Nicoli JR, Vieira LQ et al. The essential role of the intestinal microbiota in facilitating acute inflammatory responses. J Immunol 2004; 173: 4137–4146.
Oliveira MR, Tafuri WL, Afonso LC, Oliveira MA, Nicoli JR, Vieira EC et al. Germ-free mice produce high levels of interferon-gamma in response to infection with Leishmania major but fail to heal lesions. Parasitology 2005; 131 (Part 4): 477–488.
van der Waaij D, Berghuis-de Vries JM, Lekkerkerk L-v . Colonization resistance of the digestive tract in conventional and antibiotic-treated mice. J Hyg 1971; 69: 405–411.
Ley RE, Peterson DA, Gordon JI . Ecological and evolutionary forces shaping microbial diversity in the human intestine. Cell 2006; 124: 837–848.
Backhed F, Ley RE, Sonnenburg JL, Peterson DA, Gordon JI . Host–bacterial mutualism in the human intestine. Science 2005; 307: 1915–1920.
Duncan SH, Belenguer A, Holtrop G, Johnstone AM, Flint HJ, Lobley GE . Reduced dietary intake of carbohydrates by obese subjects results in decreased concentrations of butyrate and butyrate-producing bacteria in feces. Appl Environ Microbiol 2007; 73: 1073–1078.
Gill SR, Pop M, Deboy RT, Eckburg PB, Turnbaugh PJ, Samuel BS et al. Metagenomic analysis of the human distal gut microbiome. Science 2006; 312: 1355–1359.
Collins J, Borojevic R, Verdu EF, Huizinga JD, Ratcliffe EM . Intestinal microbiota influence the early postnatal development of the enteric nervous system. Neurogastroenterol Motil 2014; 26: 98–107.
Kinross JM, Darzi AW, Nicholson JK . Gut microbiome–host interactions in health and disease. Genome Med 2011; 3: 14.
Spor A, Koren O, Ley R . Unravelling the effects of the environment and host genotype on the gut microbiome. Nat Rev Microbiol 2011; 9: 279–290.
Couturier-Maillard A, Secher T, Rehman A, Normand S, De Arcangelis A, Haesler R et al. NOD2-mediated dysbiosis predisposes mice to transmissible colitis and colorectal cancer. J Clin Invest 2013; 123: 700–711.
Yoshimoto S, Loo TM, Atarashi K, Kanda H, Sato S, Oyadomari S et al. Obesity-induced gut microbial metabolite promotes liver cancer through senescence secretome. Nature 2013; 499: 97–101.
Mc CW, Mason JM III . Enterococcal endocarditis associated with carcinoma of the sigmoid; report of a case. J Med Assoc State of Alabama 1951; 21: 162–166.
Boleij A, Tjalsma H . The itinerary of Streptococcus gallolyticus infection in patients with colonic malignant disease. Lancet Infect Dis 2013; 13: 719–724.
Huycke MM, Abrams V, Moore DR . Enterococcus faecalis produces extracellular superoxide and hydrogen peroxide that damages colonic epithelial cell DNA. Carcinogenesis 2002; 23: 529–536.
Nougayrede JP, Homburg S, Taieb F, Boury M, Brzuszkiewicz E, Gottschalk G et al. Escherichia coli induces DNA double-strand breaks in eukaryotic cells. Science 2006; 313: 848–851.
Wu S, Rhee KJ, Zhang M, Franco A, Sears CL . Bacteroides fragilis toxin stimulates intestinal epithelial cell shedding and gamma-secretase-dependent E-cadherin cleavage. J Cell Sci 2007; 120 Part 11: 1944–1952.
Goodwin AC, Destefano Shields CE, Wu S, Huso DL, Wu X, Murray-Stewart TR et al. Polyamine catabolism contributes to enterotoxigenic Bacteroides fragilis-induced colon tumorigenesis. Proc Natl Acad Sci USA 2011; 108: 15354–15359.
Mangerich A, Knutson CG, Parry NM, Muthupalani S, Ye W, Prestwich E et al. Infection-induced colitis in mice causes dynamic and tissue-specific changes in stress response and DNA damage leading to colon cancer. Proc Natl Acad Sci USA 2012; 109: E1820–E1829.
Vijay-Kumar M, Aitken JD, Carvalho FA, Cullender TC, Mwangi S, Srinivasan S et al. Metabolic syndrome and altered gut microbiota in mice lacking Toll-like receptor 5. Science 2010; 328: 228–231.
Dapito DH, Mencin A, Gwak GY, Pradere JP, Jang MK, Mederacke I et al. Promotion of hepatocellular carcinoma by the intestinal microbiota and TLR4. Cancer Cell 2012; 21: 504–516.
Kostic AD, Gevers D, Pedamallu CS, Michaud M, Duke F, Earl AM et al. Genomic analysis identifies association of Fusobacterium with colorectal carcinoma. Genome Res 2012; 22: 292–298.
Marchesi JR, Dutilh BE, Hall N, Peters WH, Roelofs R, Boleij A et al. Towards the human colorectal cancer microbiome. PLoS One 2011; 6: e20447.
Sobhani I, Tap J, Roudot-Thoraval F, Roperch JP, Letulle S, Langella P et al. Microbial dysbiosis in colorectal cancer (CRC) patients. PLoS One 2011; 6: e16393.
Tjalsma H, Boleij A, Marchesi JR, Dutilh BE . A bacterial driver–passenger model for colorectal cancer: beyond the usual suspects. Nat Rev Microbiol 2012; 10: 575–582.
Wu S, Rhee KJ, Albesiano E, Rabizadeh S, Wu X, Yen HR et al. A human colonic commensal promotes colon tumorigenesis via activation of T helper type 17 T cell responses. Nat Med 2009; 15: 1016–1022.
Grivennikov S, Karin E, Terzic J, Mucida D, Yu GY, Vallabhapurapu S et al. IL-6 and Stat3 are required for survival of intestinal epithelial cells and development of colitis-associated cancer. Cancer Cell 2009; 15: 103–113.
Grivennikov SI, Wang K, Mucida D, Stewart CA, Schnabl B, Jauch D et al. Adenoma-linked barrier defects and microbial products drive IL-23/IL-17-mediated tumour growth. Nature 2012; 491: 254–258.
Round JL, Mazmanian SK . The gut microbiota shapes intestinal immune responses during health and disease. Nat Rev Immunol 2009; 9: 313–323.
Chung H, Pamp SJ, Hill JA, Surana NK, Edelman SM, Troy EB et al. Gut immune maturation depends on colonization with a host-specific microbiota. Cell 2012; 149: 1578–1593.
Round JL, Mazmanian SK . Inducible Foxp3+ regulatory T-cell development by a commensal bacterium of the intestinal microbiota. Proc Natl Acad Sci USA 2010; 107: 12204–12209.
Atarashi K, Tanoue T, Shima T, Imaoka A, Kuwahara T, Momose Y et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science 2011; 331: 337–341.
Ivanov II, Atarashi K, Manel N, Brodie EL, Shima T, Karaoz U et al. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 2009; 139: 485–498.
Gaboriau-Routhiau V, Rakotobe S, Lecuyer E, Mulder I, Lan A, Bridonneau C et al. The key role of segmented filamentous bacteria in the coordinated maturation of gut helper T cell responses. Immunity 2009; 31: 677–689.
Arpaia N, Campbell C, Fan X, Dikiy S, van der Veeken J, deRoos P et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 2013; 504: 451–455.
Wu HJ, Ivanov II, Darce J, Hattori K, Shima T, Umesaki Y et al. Gut-residing segmented filamentous bacteria drive autoimmune arthritis via T helper 17 cells. Immunity 2010; 32: 815–827.
Wen L, Ley RE, Volchkov PY, Stranges PB, Avanesyan L, Stonebraker AC et al. Innate immunity and intestinal microbiota in the development of type 1 diabetes. Nature 2008; 455: 1109–1113.
van Vliet MJ, Harmsen HJ, de Bont ES, Tissing WJ . The role of intestinal microbiota in the development and severity of chemotherapy-induced mucositis. PLoS Pathogen 2010; 6: e1000879.
Stringer AM, Gibson RJ, Bowen JM, Keefe DM . Chemotherapy-induced modifications to gastrointestinal microflora: evidence and implications of change. Curr Drug Metab 2009; 10: 79–83.
Ijiri K, Potten CS . Further studies on the response of intestinal crypt cells of different hierarchical status to eighteen different cytotoxic agents. Br J Cancer 1987; 55: 113–123.
Dekaney CM, Gulati AS, Garrison AP, Helmrath MA, Henning SJ . Regeneration of intestinal stem/progenitor cells following doxorubicin treatment of mice. Am J Physiol Gastrointest Liver Physiol 2009; 297: G461–G470.
Stringer AM, Gibson RJ, Logan RM, Bowen JM, Yeoh AS, Keefe DM . Faecal microflora and beta-glucuronidase expression are altered in an irinotecan-induced diarrhea model in rats. Cancer Biol Ther 2008; 7: 1919–1925.
Gupta E, Lestingi TM, Mick R, Ramirez J, Vokes EE, Ratain MJ . Metabolic fate of irinotecan in humans: correlation of glucuronidation with diarrhea. Cancer Res 1994; 54: 3723–3725.
Takasuna K, Hagiwara T, Hirohashi M, Kato M, Nomura M, Nagai E et al. Involvement of beta-glucuronidase in intestinal microflora in the intestinal toxicity of the antitumor camptothecin derivative irinotecan hydrochloride (CPT-11) in rats. Cancer Res 1996; 56: 3752–3757.
Manhart N, Vierlinger K, Spittler A, Bergmeister H, Sautner T, Roth E . Oral feeding with glutamine prevents lymphocyte and glutathione depletion of Peyer’s patches in endotoxemic mice. Ann Surg 2001; 234: 92–97.
Decker-Baumann C, Buhl K, Frohmuller S, von Herbay A, Dueck M, Schlag PM . Reduction of chemotherapy-induced side-effects by parenteral glutamine supplementation in patients with metastatic colorectal cancer. Eur J Cancer 1999; 35: 202–207.
Lin XB, Dieleman LA, Ketabi A, Bibova I, Sawyer MB, Xue H et al. Irinotecan (CPT-11) chemotherapy alters intestinal microbiota in tumour bearing rats. PLoS One 2012; 7: e39764.
Maruya M, Kawamoto S, Kato LM, Fagarasan S . Impaired selection of IgA and intestinal dysbiosis associated with PD-1-deficiency. Gut Microbes 2013; 4: 165–171.
Kawamoto S, Tran TH, Maruya M, Suzuki K, Doi Y, Tsutsui Y et al. The inhibitory receptor PD-1 regulates IgA selection and bacterial composition in the gut. Science 2012; 336: 485–489.
Rossini A, Rumio C, Sfondrini L, Tagliabue E, Morelli D, Miceli R et al. Influence of antibiotic treatment on breast carcinoma development in proto–neu transgenic mice. Cancer Res 2006; 66: 6219–6224.
Velicer CM, Heckbert SR, Lampe JW, Potter JD, Robertson CA, Taplin SH . Antibiotic use in relation to the risk of breast cancer. JAMA 2004; 291: 827–835.
Blaser M . Antibiotic overuse: stop the killing of beneficial bacteria. Nature 2011; 476: 393–394.
Kawai K, Miyazaki J, Joraku A, Nishiyama H, Akaza H . Bacillus Calmette–Guerin (BCG) immunotherapy for bladder cancer: current understanding and perspectives on engineered BCG vaccine. Cancer Sci 2013; 104: 22–27.
LaRue H, Ayari C, Bergeron A, Fradet Y . Toll-like receptors in urothelial cells—targets for cancer immunotherapy. Nat Rev Urol 2013; 10: 537–545.
Vacchelli E, Eggermont A, Sautes-Fridman C, Galon J, Zitvogel L, Kroemer G et al. Trial Watch: Toll-like receptor agonists for cancer therapy. Oncoimmunology 2013; 2: e25238.
Lipton A, Harvey HA, Balch CM, Antle CE, Heckard R, Bartolucci AA . Corynebacterium parvum versus bacille Calmette–Guerin adjuvant immunotherapy of stage II malignant melanoma. J Clin Oncol 1991; 9: 1151–1156.
Tsuda K, Yamanaka K, Linan W, Miyahara Y, Akeda T, Nakanishi T et al. Intratumoral injection of Propionibacterium acnes suppresses malignant melanoma by enhancing Th1 immune responses. PLoS One 2011; 6: e29020.
Kim J, Ochoa MT, Krutzik SR, Takeuchi O, Uematsu S, Legaspi AJ et al. Activation of toll-like receptor 2 in acne triggers inflammatory cytokine responses. J Immunol 2002; 169: 1535–1541.
Dieu-Nosjean MC, Antoine M, Danel C, Heudes D, Wislez M, Poulot V et al. Long-term survival for patients with non-small-cell lung cancer with intratumoral lymphoid structures. J Clin Oncol 2008; 26: 4410–4417.
Viaud S, Saccheri F, Mignot G, Yamazaki T, Daillere R, Hannani D et al. The intestinal microbiota modulates the anticancer immune effects of cyclophosphamide. Science 2013; 342: 971–976.
Iida N, Dzutsev A, Stewart CA, Smith L, Bouladoux N, Weingarten RA et al. Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science 2013; 342: 967–970.
Sistigu A, Viaud S, Chaput N, Bracci L, Proietti E, Zitvogel L . Immunomodulatory effects of cyclophosphamide and implementations for vaccine design. Semin Immunopathol 2011; 33: 369–383.
Viaud S, Flament C, Zoubir M, Pautier P, LeCesne A, Ribrag V et al. Cyclophosphamide induces differentiation of Th17 cells in cancer patients. Cancer Res 2011; 71: 661–665.
Vicari AP, Chiodoni C, Vaure C, Ait-Yahia S, Dercamp C, Matsos F et al. Reversal of tumor-induced dendritic cell paralysis by CpG immunostimulatory oligonucleotide and anti-interleukin 10 receptor antibody. J Exp Med 2002; 196: 541–549.
Hand TW, Dos Santos LM, Bouladoux N, Molloy MJ, Pagan AJ, Pepper M et al. Acute gastrointestinal infection induces long-lived microbiota-specific T cell responses. Science 2012; 337: 1553–1556.
Cong Y, Feng T, Fujihashi K, Schoeb TR, Elson CO . A dominant, coordinated T regulatory cell-IgA response to the intestinal microbiota. Proc Natl Acad Sci USA 2009; 106: 19256–19261.
Farago AF, Snyder EL, Jacks T . SnapShot: lung cancer models. Cell 2012; 149: 246–246 e241.
Cortez-Retamozo V, Etzrodt M, Newton A, Rauch PJ, Chudnovskiy A, Berger C et al. Origins of tumor-associated macrophages and neutrophils. Proc Natl Acad Sci USA 2012; 109: 2491–2496.
Cortez-Retamozo V, Etzrodt M, Newton A, Ryan R, Pucci F, Sio SW et al. Angiotensin II drives the production of tumor-promoting macrophages. Immunity 2013; 38: 296–308.
Alegre ML, Bartman C, Chong AS . Microbes and allogeneic transplantation. Transplantation 2014; 97: 5–11.
Chong AS, Alegre ML . The impact of infection and tissue damage in solid-organ transplantation. Nat Rev Immunol 2012; 12: 459–471.
Conforti R, Ma Y, Morel Y, Paturel C, Terme M, Viaud S et al. Opposing effects of toll-like receptor (TLR3) signaling in tumors can be therapeutically uncoupled to optimize the anticancer efficacy of TLR3 ligands. Cancer Res 2010; 70: 490–500.
Faye T, Tamburello A, Vegarud GE, Skeie S . Survival of lactic acid bacteria from fermented milks in an in vitro digestion model exploiting sequential incubation in human gastric and duodenum juice. J Dairy Sci 2012; 95: 558–566.
Mack DR, Ahrne S, Hyde L, Wei S, Hollingsworth MA . Extracellular MUC3 mucin secretion follows adherence of Lactobacillus strains to intestinal epithelial cells in vitro. Gut 2003; 52: 827–833.
Miyauchi E, Morita H, Okuda J, Sashihara T, Shimizu M, Tanabe S . Cell wall fraction of Enterococcus hirae ameliorates TNF-alpha-induced barrier impairment in the human epithelial tight junction. Lett Appl Microbiol 2008; 46: 469–476.
Algra AM, Rothwell PM . Effects of regular aspirin on long-term cancer incidence and metastasis: a systematic comparison of evidence from observational studies versus randomised trials. Lancet Oncol 2012; 13: 518–527.
Lawley TD, Clare S, Walker AW, Stares MD, Connor TR, Raisen C et al. Targeted restoration of the intestinal microbiota with a simple, defined bacteriotherapy resolves relapsing Clostridium difficile disease in mice. PLoS Pathogen 2012; 8: e1002995.
Smits LP, Bouter KE, de Vos WM, Borody TJ, Nieuwdorp M . Therapeutic potential of fecal microbiota transplantation. Gastroenterology 2013; 145: 946–953.
Kolars JC, Levitt MD, Aouji M, Savaiano DA . Yogurt – an autodigesting source of lactose. N Engl J Med 1984; 310: 1–3.
Khazaie K, Zadeh M, Khan MW, Bere P, Gounari F, Dennis K et al. Abating colon cancer polyposis by Lactobacillus acidophilus deficient in lipoteichoic acid. Proc Natl Acad Sci USA 2012; 109: 10462–10467.
Motta JP, Bermudez-Humaran LG, Deraison C, Martin L, Rolland C, Rousset P et al. Food-grade bacteria expressing elafin protect against inflammation and restore colon homeostasis. Sci Transl Med 2012; 4: 158ra144.
Carroll IM, Andrus JM, Bruno-Barcena JM, Klaenhammer TR, Hassan HM, Threadgill DS . Anti-inflammatory properties of Lactobacillus gasseri expressing manganese superoxide dismutase using the interleukin 10-deficient mouse model of colitis. Am J Physiol Gastrointest Liver Physiol 2007; 293: G729–G738.
Davis CD, Milner JA . Gastrointestinal microflora, food components and colon cancer prevention. J Nutr Biochem 2009; 20: 743–752.
Lampe JW . Emerging research on equol and cancer. J Nutr 2010; 140: 1369S–1372S.
Hamer HM, Jonkers D, Venema K, Vanhoutvin S, Troost FJ, Brummer RJ . Review article: the role of butyrate on colonic function. Alim Pharmacol Therap 2008; 27: 104–119.
Smith PM, Howitt MR, Panikov N, Michaud M, Gallini CA, Bohlooly YM et al. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 2013; 341: 569–573.
Nenci A, Becker C, Wullaert A, Gareus R, van Loo G, Danese S et al. Epithelial NEMO links innate immunity to chronic intestinal inflammation. Nature 2007; 446: 557–561.
Steinbrecher KA, Harmel-Laws E, Sitcheran R, Baldwin AS . Loss of epithelial RelA results in deregulated intestinal proliferative/apoptotic homeostasis and susceptibility to inflammation. J Immunol 2008; 180: 2588–2599.
Vereecke L, Sze M, Mc Guire C, Rogiers B, Chu Y, Schmidt-Supprian M et al. Enterocyte-specific A20 deficiency sensitizes to tumor necrosis factor-induced toxicity and experimental colitis. J Exp Med 2010; 207: 1513–1523.
Pickert G, Neufert C, Leppkes M, Zheng Y, Wittkopf N, Warntjen M et al. STAT3 links IL-22 signaling in intestinal epithelial cells to mucosal wound healing. J Exp Med 2009; 206: 1465–1472.
Kajino-Sakamoto R, Inagaki M, Lippert E, Akira S, Robine S, Matsumoto K et al. Enterocyte-derived TAK1 signaling prevents epithelium apoptosis and the development of ileitis and colitis. J Immunol 2008; 181: 1143–1152.
Kim JY, Kajino-Sakamoto R, Omori E, Jobin C, Ninomiya-Tsuji J . Intestinal epithelial-derived TAK1 signaling is essential for cytoprotection against chemical-induced colitis. PLoS One 2009; 4: e4561.
Welz PS, Wullaert A, Vlantis K, Kondylis V, Fernandez-Majada V, Ermolaeva M et al. FADD prevents RIP3-mediated epithelial cell necrosis and chronic intestinal inflammation. Nature 2011; 477: 330–334.
Gunther C, Martini E, Wittkopf N, Amann K, Weigmann B, Neumann H et al. Caspase-8 regulates TNF-alpha-induced epithelial necroptosis and terminal ileitis. Nature 2011; 477: 335–339.
Kaser A, Lee AH, Franke A, Glickman JN, Zeissig S, Tilg H et al. XBP1 links ER stress to intestinal inflammation and confers genetic risk for human inflammatory bowel disease. Cell 2008; 134: 743–756.
Adolph TE, Tomczak MF, Niederreiter L, Ko HJ, Bock J, Martinez-Naves E et al. Paneth cells as a site of origin for intestinal inflammation. Nature 2013; 503: 272–276.
Coulombe G, Leblanc C, Cagnol S, Maloum F, Lemieux E, Perreault N et al. Epithelial tyrosine phosphatase SHP-2 protects against intestinal inflammation in mice. Mol Cell Biol 2013; 33: 2275–2284.
Van der Sluis M, De Koning BA, De Bruijn AC, Velcich A, Meijerink JP, Van Goudoever JB et al. Muc2-deficient mice spontaneously develop colitis, indicating that MUC2 is critical for colonic protection. Gastroenterology 2006; 131: 117–129.
Burger-van Paassen N, van der Sluis M, Bouma J, Korteland-van Male AM, Lu P, Van Seuningen I et al. Colitis development during the suckling-weaning transition in mucin Muc2-deficient mice. Am J Physiol Gastrointest Liver Physiol 2011; 301: G667–G678.
Harmel-Laws E, Mann EA, Cohen MB, Steinbrecher KA . Guanylate cyclase C deficiency causes severe inflammation in a murine model of spontaneous colitis. PLoS One 2013; 8: e79180.
Liu W, Chen Y, Golan MA, Annunziata ML, Du J, Dougherty U et al. Intestinal epithelial vitamin D receptor signaling inhibits experimental colitis. J Clin Invest 2013; 123: 3983–3996.
Chen GY, Liu M, Wang F, Bertin J, Nunez G . A functional role for Nlrp6 in intestinal inflammation and tumorigenesis. J Immunol 2011; 186: 7187–7194.
Elinav E, Strowig T, Kau AL, Henao-Mejia J, Thaiss CA, Booth CJ et al. NLRP6 inflammasome regulates colonic microbial ecology and risk for colitis. Cell 2011; 145: 745–757.
Normand S, Delanoye-Crespin A, Bressenot A, Huot L, Grandjean T, Peyrin-Biroulet L et al. Nod-like receptor pyrin domain-containing protein 6 (NLRP6) controls epithelial self-renewal and colorectal carcinogenesis upon injury. Proc Natl Acad Sci USA 2011; 108: 9601–9606.
Gong J, Xu J, Zhu W, Gao X, Li N, Li J . Epithelial-specific blockade of MyD88-dependent pathway causes spontaneous small intestinal inflammation. Clin Immunol 2010; 136: 245–256.
Frantz AL, Rogier EW, Weber CR, Shen L, Cohen DA, Fenton LA et al. Targeted deletion of MyD88 in intestinal epithelial cells results in compromised antibacterial immunity associated with downregulation of polymeric immunoglobulin receptor, mucin-2, and antibacterial peptides. Mucosal Immunol 2012; 5: 501–512.
Hoshi N, Schenten D, Nish SA, Walther Z, Gagliani N, Flavell RA et al. MyD88 signalling in colonic mononuclear phagocytes drives colitis in IL-10-deficient mice. Nat Commun 2012; 3: 1120.
Turgeon N, Blais M, Gagne JM, Tardif V, Boudreau F, Perreault N et al. HDAC1 and HDAC2 restrain the intestinal inflammatory response by regulating intestinal epithelial cell differentiation. PLoS One 2013; 8: e73785.
Alenghat T, Osborne LC, Saenz SA, Kobuley D, Ziegler CG, Mullican SE et al. Histone deacetylase 3 coordinates commensal-bacteria-dependent intestinal homeostasis. Nature 2013; 504: 153–157.
Dalmasso G, Nguyen HT, Ingersoll SA, Ayyadurai S, Laroui H, Charania MA et al. The PepT1-NOD2 signaling pathway aggravates induced colitis in mice. Gastroenterology 2011; 141: 1334–1345.
Garrett WS, Lord GM, Punit S, Lugo-Villarino G, Mazmanian SK, Ito S et al. Communicable ulcerative colitis induced by T-bet deficiency in the innate immune system. Cell 2007; 131: 33–45.
Garrett WS, Gallini CA, Yatsunenko T, Michaud M, DuBois A, Delaney ML et al. Enterobacteriaceae act in concert with the gut microbiota to induce spontaneous and maternally transmitted colitis. Cell Host Microbe 2010; 8: 292–300.
Kühn R, Lohler J, Rennick D, Rajewsky K, Muller W . Interleukin-10-deficient mice develop chronic enterocolitis. Cell 1993; 75: 263–274.
Rakoff-Nahoum S, Hao L, Medzhitov R . Role of toll-like receptors in spontaneous commensal-dependent colitis. Immunity 2006; 25: 319–329.
Gomes-Santos AC, Moreira TG, Castro-Junior AB, Horta BC, Lemos L, Cruz DN et al. New insights into the immunological changes in IL-10-deficient mice during the course of spontaneous inflammation in the gut mucosa. Clin Dev Immunol 2012; 2012: 560817.
Yoshihara K, Yajima T, Kubo C, Yoshikai Y . Role of interleukin 15 in colitis induced by dextran sulphate sodium in mice. Gut 2006; 55: 334–341.
Zenewicz LA, Yancopoulos GD, Valenzuela DM, Murphy AJ, Stevens S, Flavell RA . Innate and adaptive interleukin-22 protects mice from inflammatory bowel disease. Immunity 2008; 29: 947–957.
Zenewicz LA, Yin X, Wang G, Elinav E, Hao L, Zhao L et al. IL-22 deficiency alters colonic microbiota to be transmissible and colitogenic. J Immunol 2013; 190: 5306–5312.
Takamatsu M, Hirata A, Ohtaki H, Hoshi M, Hatano Y, Tomita H et al. IDO1 plays an immunosuppressive role in 2,4,6-trinitrobenzene sulfate-induced colitis in mice. J Immunol 2013; 191: 3057–3064.
Gurtner GJ, Newberry RD, Schloemann SR, McDonald KG, Stenson WF . Inhibition of indoleamine 2,3-dioxygenase augments trinitrobenzene sulfonic acid colitis in mice. Gastroenterology 2003; 125: 1762–1773.
Nguyen DD, Maillard MH, Cotta-de-Almeida V, Mizoguchi E, Klein C, Fuss I et al. Lymphocyte-dependent and Th2 cytokine-associated colitis in mice deficient in Wiskott–Aldrich syndrome protein. Gastroenterology 2007; 133: 1188–1197.
Nguyen DD, Wurbel MA, Goettel JA, Eston MA, Ahmed OS, Marin R et al. Wiskott–Aldrich syndrome protein deficiency in innate immune cells leads to mucosal immune dysregulation and colitis in mice. Gastroenterology 2012; 143: 719–729 e711-712.
Brenner O, Levanon D, Negreanu V, Golubkov O, Fainaru O, Woolf E et al. Loss of Runx3 function in leukocytes is associated with spontaneously developed colitis and gastric mucosal hyperplasia. Proc Natl Acad Sci USA 2004; 101: 16016–16021.
Furumatsu K, Nishiumi S, Kawano Y, Ooi M, Yoshie T, Shiomi Y et al. A role of the aryl hydrocarbon receptor in attenuation of colitis. Dig Dis Sci 2011; 56: 2532–2544.
Qiao G, Yang L, Li Z, Ying H, Hassen Y, Yin F et al. Program death-1 regulates peripheral T cell tolerance via an anergy-independent mechanism. Clin Immunol 2012; 143: 128–133.
Acknowledgements
We thank our colleagues from the Gustave Roussy animal facility and Institut Pasteur axenic mice facility. This work was supported by Institut National du Cancer (INCa), la Ligue contre le cancer (LIGUE labellisée), SIRIC Socrate, LABEX and PACRI Onco-Immunology, European Research Council starting grant (PGN from SHAPE to VIR no. 202283 to IGB) and ISREC Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by H-U Simon
Rights and permissions
About this article
Cite this article
Viaud, S., Daillère, R., Boneca, I. et al. Gut microbiome and anticancer immune response: really hot Sh*t!. Cell Death Differ 22, 199–214 (2015). https://doi.org/10.1038/cdd.2014.56
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2014.56
This article is cited by
-
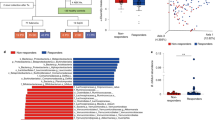
Prospective, longitudinal analysis of the gut microbiome in patients with locally advanced rectal cancer predicts response to neoadjuvant concurrent chemoradiotherapy
Journal of Translational Medicine (2023)
-
Effect of hyperthermia on intestinal microecology, immune function, and progression-free survival in patients with advanced unresectable lung adenocarcinoma
Scientific Reports (2023)
-
Gut Microbiota Metabolites Mediate Bax to Reduce Neuronal Apoptosis via cGAS/STING Axis in Epilepsy
Molecular Neurobiology (2023)
-
The Evolving Landscape of Fecal Microbial Transplantation
Clinical Reviews in Allergy & Immunology (2023)
-
Broad-spectrum antibiotics associated gut microbiome disturbance impairs T cell immunity and promotes lung cancer metastasis: a retrospective study
BMC Cancer (2022)