Abstract
Cell death has a central role in innate immune responses in both plants and animals. Besides sharing striking convergences and similarities in the overall evolutionary organization of their innate immune systems, both plants and animals can respond to infection and pathogen recognition with programmed cell death. The fact that plant and animal pathogens have evolved strategies to subvert specific cell death modalities emphasizes the essential role of cell death during immune responses. The hypersensitive response (HR) cell death in plants displays morphological features, molecular architectures and mechanisms reminiscent of different inflammatory cell death types in animals (pyroptosis and necroptosis). In this review, we describe the molecular pathways leading to cell death during innate immune responses. Additionally, we present recently discovered caspase and caspase-like networks regulating cell death that have revealed fascinating analogies between cell death control across both kingdoms.
Similar content being viewed by others
Immune Surveillance Systems in Plants and Animals
The interaction between a pathogen and its host is sophisticated and dynamic. Disease develops when the pathogen is able to evade the multiple layers of host defenses. The immune system of an organism has been tailored through evolution by a long history of warfare with its invaders. In contrast to most animals, plants are sessile organisms, they lack a circulatory system and their cells are framed with a rigid cell wall. These evolutionary constraints have resulted in the evolution of a primary cell-autonomous immune system. Despite these fundamental differences between the two kingdoms, plants and animals share striking similarities in their innate immune systems, some of which tell a story of likely convergent evolution.1 Immune systems discriminate self from non-self, and activate tightly regulated pre- and post-invasion defense responses to minimize the damage inflicted by harmful agents. The first line of defense in both plants and animals is provided by pattern recognition receptors (PRRs), which recognize microbe- or danger-associated molecular patterns (MAMPs and DAMPs, respectively), and trigger immune signaling (Figure 1). Plant PRRs are transmembrane receptors.2, 3 The best-studied class of plant PRRs are receptor-like kinases (RLKs), which feature an ectodomain of leucine-rich repeats (LRRs) involved in MAMP perception, and an intracellular kinase domain, involved in signal transduction relay via MAPK cascades, resulting in MAMP-triggered immunity (MTI).4
Innate immune pathways in plants and mammals. In both plants (left) and mammalian (right), cells pathogen detection by membrane and intracellular innate immune receptors leads to signaling cascades that culminate in expression of defense-related genes. Defense mechanisms eventually result in programmed cell death in both kingdoms. This diagram exemplifies hypersensitive response cell death mediated by metacaspase-1 in plants and caspase-1-mediated cell death in animals
Typical animal extracellular PRRs, called Toll-like receptors (TLRs) possess an intracellular Toll-interleukin-1 (IL-1) receptor domain (TIR) that recruits the kinases IRAK or RIP via adaptor proteins, inducing expression of antimicrobial defense molecules.5 These kinases belong to the same functional class of non-RD kinases as plants, and they are linked to innate immune responses in both kingdoms.6 Although plants and animals evolved under very different selective pressures, both evolved similar sensors that converge onto the same generic function: to alert the organism about the presence of non-self.7
By definition, pathogens are microorganisms and viruses that are able to evade or suppress PRR-based defenses. They do so by deploying various effectors, determinants of virulence on susceptible hosts via MTI suppression, into the host cell.4, 8 Successful pathogens are then faced with another hurdle, evolved by hosts to recognize the presence of their effectors and of intracellular MAMPs. In plants, a second intracellular class of innate immune receptors is activated via recognition of pathogen effectors, resulting in effector-triggered immunity (ETI). ETI is mediated by nucleotide-binding domain, LRR (NB-LRR) disease resistance proteins. Their structure is very similar to Nod-like receptors (NLRs) in mammals.9 Both are intracellular proteins that contain a central nucleotide-binding domain involved in activation and multimerization9, 10 and an LRR domain. In addition to structural similarities, NLRs and NB-LRRs have shared function and their stability is regulated by a chaperone complex containing HSP90 and SGT1.11, 12, 13 Both NB-LRRs in plants and NLRs in mammals are classified according to their N-terminal domain architecture. Two main classes of plant NB-LRRs have been described: CC-NB-LRRs contain a predicted coiled-coil N-terminal domain and TIR-NB-LRRs carry N-terminal homology to the intracellular TIR domain of TLRs. In mammals, the NLR family is divided into five subfamilies with different N-terminal effector domains.14 The N-termini of NLRs mediate protein–protein interactions with downstream signaling partners.
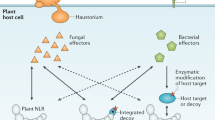
Both NB-LRRs and NLRs act as intracellular immune sentinels. NB-LRRs have evolved to recognize specific pathogen effector proteins, which are delivered into the host cytosol by a broad range of pathogens using various delivery systems. An effective recognition system must be able to sense and respond to a multitude of effectors, since each pathogen delivers its unique repertoire.15 In plants, effector recognition can occur by direct binding of the NB-LRR protein or indirectly, via an intermediate protein. The guard hypothesis16, 17 explains indirect recognition, which occurs after an effector modifies a particular host protein (guardee) that is monitored by a particular NB-LRR (guard). Plants do not appear to express somatic recombination-based diversity generation in their immune system, as do animal cells to generate the familiar T- and B-cell antigen receptors. Therefore, sensing of ‘modified-self’ accounts for a powerful recognition system, that can manage with a limited set of receptors an effective defense response.
In animals, the mechanism by which NLRs sense MAMPs or DAMPs to trigger an appropriate immune response is not fully understood. In vivo direct recognition has not been proven and recent models suggest that NLR activation could occur indirectly as a result of the membrane damage inflicted by pathogens that are either able to reach the cytoplasm, or that accidentally deliver MAMPs via their secretion systems along with effector proteins.18, 19 In this sense, NLRs could be conceived as guard proteins similar to plant NB-LRRs.20
Cell Death at the Center of Immune Responses
Pathogen recognition via NLRs in animals and NB-LRRs in plants leads to inhibition of pathogen growth, which is often, but not always, accompanied in plants by the hypersensitive response (HR), a form of programmed cell death localized at the site of attempted pathogen invasion (Figure 2a). The first observations of HR date back to 1902 in the wheat-Puccinia glumarum pathosystem,21 and the counter-intuitive term ‘hypersensitiveness’ was coined in 1915,22 to describe a pathogen-triggered cell death reaction that correlated with disease resistance in wheat infected with Puccinia graminis. Morphologically, HR is a specific and unique type of cell death. Its hallmarks, some of which are typical for different forms of animal cell death, include cytoplasmic shrinkage, chromatin condensation, mitochondrial swelling, combined with other characteristics that are plant specific, such as vacuolization and chloroplast disruption during the final stages.23
Cell death modalities in response to infection. Diagram representing some of the characteristic features of different types of programmed cell death that can occur in response to infection in plants and mammals. HR cell death in plants (a) and pyroptosis (b) and Necroptosis (c) in mammalian cells. See the text for details
The chloroplast has a central role in defense responses and HR in plants. First, it constitutes a very important source of defense signaling molecules such as reactive oxygen species (ROS), reactive nitrogen oxide intermediates (NOI) and the defense hormones salicylic acid (SA) and jasmonic acid (JA). Second, in many cases, light is required for HR development. Third, several pathogen effectors have chloroplast localization signals,24 and in some cases they have been shown to suppress immunity.25, 26
In plants, the molecular events that lead to HR during ETI are partly overlapping with those associated with MTI, including accumulation of SA, ROS and NOI, activation of MAPK cascades, changes in intracellular calcium levels, transcriptional reprogramming and synthesis of antimicrobial compounds.23 Compared with MTI, ETI is typically an accelerated and amplified response, suggesting that quantitative rather than qualitative differences account for HR induction.4
In animals, naturally occurring cell death was first reported in the 19th century27 and many years later the term apoptosis was defined.28 Caspases, a family of cysteine proteases that cleave their substrates after an aspartic acid residue, emerged as the orchestrators of this cell death process. Remarkably, the first caspase discovered in mammals was the IL-1β-converting enzyme (ICE), later known as caspase-1, which does not participate in apoptosis, but does control inflammation and pyroptotic cell death (see below) downstream of NLR activation.8, 29 During the last decade, an increasing number of cell death morphologies with mixed features have been described in mammals, potentially offering a parallel to plant HR, and programmed cell death classification has become a complex task. A thorough compilation of the morphological/biochemical/functional criteria to define various sorts of cell deaths has recently been published.30
In mammals, several types of cell death have been reported in response to infection. Because these are often associated with NLRs, it may be instructive to view them as possible analogs of pathogen-dependent HR in plants (Figure 2). Pyroptosis is a pro-inflammatory form of cell death initially described as caspase-1-dependent necrosis in macrophages31 (Figure 2b). Pyroptosis has been reported in response to infection with several bacteria32 and viruses.33 Caspase-1 activation occurs within molecular platforms known as inflammasomes.34 The best studied to date are the NLR inflammasomes, which sense mostly MAMPs and DAMPs.14 These supramolecular complexes are assembled via NLR N-terminal domain homotypic interactions. Once activated, the NLRs within the inflammasome bind the N-terminal caspase activation recruitment domain (CARD) of caspase-1 directly or via the adaptor PYD-CARD protein ASC (apoptosis-associated speck-like protein containing a caspase-activating recruitment domain). Once recruited to the inflammasome, caspase-1 is activated by induced proximity and processes the inactive precursors of IL-1β and IL-18 into their mature forms. Caspase-1 also regulates the release of these and other pro-inflammatory cytokines into the extracellular millieu.35 These secreted molecules are instrumental for inflammation, cytoprotection and tissue repair. Interestingly, cytokine maturation is genetically separable from pyroptotic cell death: a recent report has shown that ASC-independent inflammasomes can activate caspase-1 without autoproteolysis, promoting cell death without processing IL-1β/IL-18.36 In contrast to ASC-containing inflammasomes, which form a single large cytoplasmic speckle, no such large structure is generated by ASC-independent inflammasomes.36
The precise mechanism by which caspase-1 leads to cell death has also been investigated by looking at caspase-1 substrates during infection and inflammation. The caspase-1 digestome includes chaperones, cytoskeletal maintenance components and proteins involved in energy metabolism,37 as well as caspase-7,38 which has been shown to be activated downstream of NLRC4 inflammasome during bacterial infection.39 However, it is still not clear why pyroptotic cells die. Permeabilization of the plasma membrane, which presumably participates in protein secretion at early stages of pyroptosis, can be the cause of later cell death due to ruptures caused by cytoplasmic swelling.40 This feature of pyroptosis is shared with another cell death modality: programmed necrosis or necroptosis41 (Figure 2c). This alternative form of programmed cell death is in most cases initiated by stimulation of the extrinsic apoptotic pathway when caspases are absent or inhibited.42 It can also be triggered after PRR activation, by a mechanism not yet characterized.42 Generally, necroptosis is mediated by RIP1–RIP3 kinase complex formation.43, 44 RIP1 is a pleiotropic protein that can mediate both pro-survival (via NF-κB activation) and pro-cell death pathways (apoptosis or necroptosis).45 During apoptosis, active caspase-8 can cleave RIP1 and RIP3 and abolish their kinase activity, preventing them from initiating necroptosis.43 When apoptosis is blocked, necroptosis becomes the predominant form of cell death.42
Increased ROS levels are a hallmark of necroptosis and may be one of the main causes of necroptotic cell death. Enhanced ROS production during necroptosis can be mediated by mitochondria, due to a RIP3-dependent increase in energy metabolism,46 and/or by the NADPH oxidase NOX1, which is recruited to the plasma membrane by RIP1.47 In plants, apoplastic ROS (superoxide) generated by the plasma membrane NADPH oxidases are essential for HR development and activation of systemic immunity,48 drawing a possible mechanistic connection between these two types of cell death. ROS produced in other plant organelles as the chloroplast, mitochondria and peroxisomes also contribute to the HR response and, in fact, compartmentalization might be essential for ROS signaling functions during defense.49
Necroptosis has a pivotal role in inflammation and immunity. Similar to pyroptotic cells, necroptotic cells secrete a broad array of pro-inflammatory molecules that signal through PRRs.50 Necroptosis has been reported to occur in response to infection by certain viruses that block apoptosis in the host cell as a colonization strategy.51 Because of the pro-inflammatory nature of necroptosis, it may constitute not only a backup mechanism for virus clearance when apoptosis is inhibited, but also a way to engage the immune system leading to a systemic response.
Caspase-independent necroptosis and caspase-1-dependent pyroptosis constitute two pro-inflammatory, explosive cell death modalities. In contrast, apoptosis, mediated by apoptotic caspases, is in most cases an immunologically silent process, since cell corpses are cleared by phagocytes. In the context of infection, it might be beneficial to minimize tissue damage during the immune response. Apoptosis can be triggered upon pathogen attack, and several lines of evidence indicate that it is essential for clearance of certain pathogens.52
Pathogen Strategies to Evade Cell Death
The fact that many pathogens have evolved strategies to inhibit different types of cell death further underscores its fundamental role in fighting infections. In mammals, apoptosis can be efficiently blocked by several pathogens via inhibition of apoptotic caspases, prevention of cytochrome c release or activation of pro-survival pathways.53 Necroptosis has been shown to be inhibited by viral inhibitors during infection51 and pyroptosis can be blocked through caspase-1 inhibition by pathogenic bacteria and viruses.32 In some instances, suppression of pyroptosis by a pathogen leads to activation of autophagy,54, 55 highlighting the complex circuitry involved in cell death processes leading to pathogen clearance.
Plant (hemi)biotrophic pathogens feed on living cells, therefore they must evade host detection and death of the invaded plant cells. Thus, they have evolved mechanisms to suppress HR using specific effectors delivered into the cell via diverse secretion systems. Several Pseudomonas syringae pathovar tomato DC3000 effectors are capable of suppressing HR in tobacco and Arabidopsis.56, 57 HR in tobacco can also be suppressed by Xanthomonas campestris pv. vesicatoria effectors.58 Oomycete effectors can also inhibit HR in plants.59, 60, 61 The mechanisms by which HR is suppressed remain unknown, but systematic characterization of the increasing number of effectors identified will help us understand how they interfere with plant defenses, including the control of HR.
In contrast to (hemi)biotrophs, necrotrophic pathogens take their nutrients from dead or dying cells. Necrotrophs have developed mechanisms to induce cell death in their hosts by secreting phytotoxins and cell wall degrading enzymes, resulting in the formation of expanding necrotic lesions in the infected plant tissue.62, 63 While (hemi)biotrophs have evolved strategies to suppress HR, some necrotrophs use the plant HR machinery as a strategy to promote virulence.64 The necrotrophic fungus Cochliobolus victoriae, originally described as the causal agent of Victoria blight in oats,65 secretes the toxin Victorin, required for pathogenicity.66 This fungus hijacks HR via activation of a CC-NB-LRR protein LOV1, which confers sensitivity to victorin and susceptibility to C. victoriae in Arabidopsis.67 In oats, loss of function mutations that eliminate toxin sensitivity and susceptiblility to C. victoriae also eliminate specific recognition and resistance to a biotrophic fungus, Puccinia coronata.68 Thus, as selection favors resistance to the biotrophic fungus, susceptibility to the necrotrophic pathogen is assured. It would be of interest to study the allele frequency of this gene in wild oats and their progenitors.
Regulators of Plant Cell Death
The chain of events leading to cell death in plants after effector recognition via NB-LRR receptors is not fully elucidated. Two separate signaling modules regulate NB-LRR proteins: non-race-specific disease resistance 1 (NDR1) regulates in most cases immune responses mediated by CC-NB-LRR proteins, whereas the enhanced disease susceptibility 1 (EDS1)/phytoalexin deficient 4 (PAD4)/senescence-associated gene 101 (SAG101) complex mediates TIR-NB-LRR signaling.69 These two systems integrate redox signals downstream of NADPH oxidase49 leading to SA accumulation,70, 71 which has a central role in defense responses. ROS and SA act synergistically to drive HR.72
Mutants exhibiting HR-like phenotypes have been long described in many plant species, including corn,73, 74 tomato,75 barley76 and Arabidopsis.77 These mutants, also known as lesion mimic mutants, are classified into initiation and propagation mutants; initiation mutants inappropriately induce PCD and form localized, necrotic spots, whereas propagation mutants cannot stop it, once it has been initiated.78 A forward genetic screen for mutants with HR-like lesions and characteristics of defense responses, including molecular and biochemical markers and enhanced disease resistance, revealed the lesion simulating disease resistance (lsd) class of mutants.79 Two of these genes have been cloned: LSD4, an FtSH protease (PE, Jürg Schmid and JLD, unpublished data; see ref. 80 for details) and the zinc-finger protein LSD1,81 a negative regulator of superoxide-induced cell death.82 LSD1 protects plants from ROS-induced stresses and consequently, lsd1 mutant plants are characterized by runaway cell death (rcd).79, 83 Therefore, lsd1 can be regarded as a sensitized mutant with respect to cell death initiation, and it has been instrumental in identifying other components of the signaling pathway leading to programmed cell death. For example, EDS1 and PAD4 functions are required for lsd1 rcd induced by abiotic stress.83 EDS1, PAD4 and NDR1 are also required for full lsd1 rcd in response to pathogen infection.84 EDS1 and PAD4 regulate a ROS- and SA-dependent signal amplification loop, which in turn is modulated by LSD1.84
The LSD1 protein contains three internally conserved zinc-finger domains of the C2C2 class (consensus: CxxCRxxLMYxxGASxVxCxxC) (Figure 4a).81 This zinc-finger motif is found in plants, algae and protozoa, but not in animals. Only six other Arabidopsis proteins contain one or more LSD1-like zinc-finger domains: the LSD-one-like proteins LOL1 (At1g32540)85 and LOL2 (At4g21610), the metacaspases AtMC1 (At1g02170), AtMC2 (At4g25110) and AtMC3 (At5g64240; although the zinc-finger motif is non-canonical) and LOL6 (At1g79350) (Figure 4a). Yeast-two-hybrid assays demonstrating interaction between the zinc-finger domains of LSD1, LOL1 and AtMC1 (Figure 3) have been validated by genetic approaches: both LOL1 and AtMC1 are required for full lsd1 rcd. Thus, LOL1 and AtMC1 are positive regulators of PCD.85, 86 Surprisingly, AtMC2 functions as a negative regulator of cell death86 (see below). Furthermore, AtbZIP10 (At4g02640) function is required for lsd1 rcd and both R-gene mediated and basal defense responses. Intriguingly, AtbZIP10–LSD1 interaction in planta prevents AtbZIP10 translocation to the nucleus.87 A yeast-two-hybrid screen for LSD1 interactors revealed 10 additional putative LSD1 interaction partners (Mike Richberg, Hironori Kaminaka and JLD, unpublished data; Figure 3). It is conceivable that LSD1 acts as a scaffold protein in the cytoplasm: sequestering positive regulators of cell death (LOL1, AtMC1, AtbZIP10) prevents their function, thereby inhibiting PCD.
The Type I Metacaspase Regulatory Module in HR
Despite the lack of close caspase homologs in plants, several studies using caspase-specific peptide inhibitors suggested the presence of HR-induced caspase-like protease activities in plants.88, 89, 90, 91 The vacuolar processing enzyme VPE in Nicotiana benthamiana and its homolog VPEgamma in Arabidopsis have caspase-1-like activity during HR.89, 91 Additionally, vacuolar fusion to the plasma membrane mediated by a caspase-3-like activity of PBA1, a plant proteasome subunit, was suggested to be a functionally relevant early event in NLR-mediated HR.88
More than a decade ago, two new families of caspase-like proteins, metacaspases and paracaspases, were identified in silico92 (Figure 4b). Similar to caspases, they contain a conserved histidine-cysteine catalytic dyad and homology modeling predicts a caspase-hemoglobinase fold.92, 93 Paracaspases have been found in animals and slime molds, whereas metacaspases are present in plants, fungi, protozoa and cyanobacteria.92 These cysteine proteases differ from caspases in their substrate specificity; caspases cleave their targets after an aspartate residue, while paracaspases are arginine specific94, 95 and metacaspases can cleave both after an arginine or a lysine.96 The human paracaspase, also known as mucosa-associated lymphoid tissue lymphoma translocation protein 1 (MALT1) has an N-terminal extension containing a death domain (DD), which is present in several proteins involved in apoptotic signaling. However, MALT1 seems to act as an anti-apoptotic scaffold protein, bridging several pathways that converge into NF-κB activation during innate and adaptive immune responses.97
Domain structures of (a) Arabidopsis proteins containing LSD1-like zinc-finger domains and (b) caspases and caspase-like proteins. CARD, caspase activation recruitment domain. DD, death domain; IG, immunoglobulin domain; Zn Finger, LSD1-like zinc-finger domain (C2C2 Class); Pro, prolin-rich domain; p20 and p10, caspase (-like putative) catalytic subunits
Eukaryotic metacaspases have been classified into type I if they bear an extension in their N-terminal domain, and type II if there is no such extension and they have a long linker region between the putative catalytic subunits.92 Type II metacaspases are present in algae and land plants, but not in protozoa or fungi. A single metacaspase present in yeast, YCA1, can serve both pro- and anti-cell death functions,98, 99 as well as other functions unrelated to cell death regulation.100, 101 In contrast to a single protein with dual functions, two different Arabidopsis type I metacaspases, AtMC1 and AtMC2, have opposing roles during cell death control (detailed below).86
The N-terminal extension of metacaspases varies among species. Fungi, protozoa and algae generally have a proline-rich domain. Some plant type I metacaspases do not have any recognizable motif in their N-termini, while others feature the conserved, plant-specific LSD1-like zinc-finger domain before the proline-rich domain (Figure 5). These motifs usually participate in protein–protein interactions, and could indicate that oligomerization is important for type I metacaspase activation, analogous to initiator/inflammatory caspases. Recruitment of a limited set of N-terminal extensions through evolution could have driven diversification and functional specialization of this protein family.
Classification of all the metacaspases found in the Viridiplantae phylum into type I (with or without the LSD1-like zinc-finger domain) or type II metacaspases, according to Phytozome (http://www.phytozome.net/)
Several lines of evidence suggest a function for metacaspases in plant HR. Infection was shown to induce the expression of a metacaspase in tomato102 and N. benthamiana103 and several metacaspase genes are pathogen inducible in Arabidopsis.104 Analysis of metacaspase function using knockout or knockdown mutants indicated roles in susceptibility to necrotrophic or hemi-biotrophic pathogens.103, 105 We recently demonstrated that AtMC1 and AtMC2 antagonistically control HR downstream of NB-LRR activation86 (Figure 6a) using Arabidopsis as a model plant to study the immune system in plants.106
Metacaspases/caspases networks in plants and animals. (a) In plants, metacaspase-1 (AtMC1) positively regulates HR cell death mediated by NB-LRR recognition of the invading pathogen at the site of infection. LSD1 negatively regulates cell death propagation in cells surrounding the infection site presumably by binding to AtMC1 and holding it inactive in the cytoplasm. AtMC2 negatively regulates AtMC1 function by an unknown mechanism. (b) In mammals, caspase-12 negatively regulates caspase-1 functions in pathogen clearance and sepsis resistance and (c) cFLIP is a negative regulator of caspase-8 function in the apoptotic extrinsic pathway. Caspase-8 function in lymphocyte proliferation is regulated by both cFLIP and MALT1
AtMC1 is a positive regulator of cell death. It interacts via its N-terminal prodomain with the second and third zinc fingers of LSD1. atmc1 knockout mutants suppress cell death in lsd1 and also bacterial- and oomycete-triggered HR. HR mediated by both CC- and TIR-NB-LRR intracellular immune receptors is severely attenuated in atmc1 plants, indicating convergence of the two pathways into a single cell death output. Interestingly, pathogen growth restriction is not affected by HR suppression, indicating that disease resistance and cell death can be uncoupled. AtMC2 acts genetically as a negative regulator of AtMC1. AtMC2 over-expression mimics atmc1 mutant phenotypes, whereas the lack of AtMC2 results in enhanced HR and accelerates cell death in an lsd1 background. Similar to some animal caspases, the function of both AtMC1 and AtMC2 is negatively regulated by their N-terminal domain. Since AtMC1 interacts with LSD1, prodomain removal could result in release of the putative active form from the LSD1-anti-cell death scaffold into the cytoplasm.
The mechanism by which AtMC2 regulates AtMC1 remains enigmatic. AtMC2 does not interact with LSD1 or AtMC1 in yeast-two-hybrid or in planta co-immunoprecipitation assays. While AtMC1 activity requires caspase-like catalytic residues, AtMC2 function is independent of its putative catalytic cysteines. In mammals, there are several examples of atypical caspases or caspase-like proteins modulating the activity of a caspase independent of their protease activity.107, 108, 109, 110, 111, 112, 113
Caspase-12 has recently emerged as a negative regulator of immune responses in mammals, causing higher susceptibility to colitis, bacterial infection and sepsis.110, 112, 113, 114 Mechanistically, caspase-12 can either inhibit caspase-1, dampening the production of pro-inflammatory cytokines112, 114 or suppress the NF-κB pathway, independent of caspase-1110, 115 (Figure 6b). Cellular FLICE-inhibitory protein (cFLIP) is a proteolytically inactive caspase-8 homolog that acts as a dominant-negative inhibitor of caspase-8 in the apoptotic extrinsic pathway of mammals.108, 111 cFLIP also regulates caspase-8 function in lymphocyte survival and proliferation.107 This non-apoptotic function of caspase-8 can also be mediated by the paracaspase MALT1, independent of its proteolytic activity109 (Figure 6c).
In line with observations using the plant AtMC2, and animal cFLIP and MALT1, the catalytic activity of caspase-12 is not required to exert its regulatory function.110, 112, 113 Caspase-12 inhibition of NLR-mediated innate immunity in mammals 110 recapitulates the role of AtMC2 inhibiting AtMC1-dependent HR, mediated by the analogous NB-LRR proteins in plants.86 The sum of these studies suggests that cell death control mediated by the caspase/metacaspase superfamily is coupled to intracellular innate immune receptor function in both animals and plants.
The HR: Cause or Consequence?
In plants, a fundamental question remains unanswered: why does HR occur? Traditionally, HR was envisioned as the plant mechanism that prevented pathogen growth in incompatible plant–pathogen interactions and therefore causal to disease resistance. This notion was first challenged by Kiraly et al.116 in a study showing that it is not plant cell death that inhibits pathogen proliferation. Since then, several natural examples of plant–pathogen interactions resulting in resistance without cell death have been reported, in particular the potato Rx and barley Rrs1 disease resistance genes.117, 118, 119, 120, 121, 122, 123, 124 Additionally, suppressing caspase-like activities (unrelated to metacaspases) in plants inhibits pathogen-induced cell death without affecting disease resistance.89, 125 As described above, elimination of the metacaspase AtMC1 results in drastically reduced HR after infection with incompatible pathogens, but bacterial growth restriction remains unaffected in this mutant.86 These studies have added new components to the sparsely populated signaling pathways that translate NLR/NB-LRR recognition of pathogens into downstream activation of cell death. The mechanisms by which plants stop pathogen growth require further analysis.
Cell death and restriction of pathogen growth leading to disease resistance are genetically separable in both animals36 and plants,86 at least for the pathogens tested in these studies. In plants, HR cell death may occur simply as a consequence of the escalated signaling at the interface of plant–pathogen interactions, and the consequent rise in toxic intermediates that lead to both host and pathogen cell death. If HR is not adaptive in restricting pathogen growth, it may be adaptive for the generation of long range signals, mediated by ROS and SA, that induce the systemic acquired resistance that primes a plant for secondary infection.126, 127, 128
Abbreviations
- ASC:
-
apoptosis-associated speck-like protein containing a caspase-activating recruiting domain
- AtMC:
-
Arabidopsis metacaspase
- CARD:
-
caspase activation recruitment domain
- CC:
-
coiled-coil domain
- cFLIP:
-
cellular FLICE-inhibitory protein
- DD:
-
death domain
- EDS1:
-
enhanced disease susceptibility 1
- ETI:
-
effector-triggered immunity
- HR:
-
hypersensitive response
- ICE:
-
interleukin-1β-converting enzyme
- IG:
-
immunoglobulin
- IL:
-
interleukin
- JA:
-
jasmonic acid
- LOL:
-
LSD-one-like
- LRR:
-
leucine-rich repeat
- LSD:
-
lesion simulating disease
- MALT1:
-
mucosa-associated lymphoid tissue lymphoma translocation protein 1
- MAMP:
-
microbe-associated molecular pattern
- MAPK:
-
mitogen-activated protein kinase
- MTI:
-
MAMP-triggered immunity
- NB-LRR, nucleotide-binding domain:
-
leucine-rich repeat
- NDR1:
-
non-race specific disease resistance 1
- NLR:
-
nod-like receptor
- NOI:
-
nitrogen oxide intermediates
- PAD4:
-
phytoalexin deficient 4
- PAMP:
-
pathogen-associated molecular pattern
- PCD:
-
programmed cell death
- Pro:
-
proline-rich domain
- PRR:
-
pattern recognition receptor
- rcd:
-
runaway cell death
- RLK:
-
receptor-like kinase
- ROS:
-
reactive oxygen species
- SA:
-
salicylic acid
- SAG101:
-
senescence-associated gene 101
- TIR:
-
Toll-interleukin-1 receptor domain
- TLR:
-
Toll-like receptor
- VPE:
-
vacuolar processing enzyme
References
Ausubel FM . Are innate immune signaling pathways in plants and animals conserved? Nat Immunol 2005; 6: 973–979.
Postel S, Kemmerling B . Plant systems for recognition of pathogen-associated molecular patterns. Semin Cell Dev Biol 2009; 20: 1025–1031.
Zipfel C . Early molecular events in PAMP-triggered immunity. Curr Opin Plant Biol 2009; 12: 414–420.
Jones JD, Dangl JL . The plant immune system. Nature 2006; 444: 323–329.
Kawai T, Akira S . The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nat Immunol 2010; 11: 373–384.
Dardick C, Ronald P . Plant and animal pathogen recognition receptors signal through non-RD kinases. PLoS Pathog 2006; 2: e2.
Ronald PC, Beutler B . Plant and animal sensors of conserved microbial signatures. Science 2010; 330: 1061–1064.
Vance RE, Isberg RR, Portnoy DA . Patterns of pathogenesis: discrimination of pathogenic and nonpathogenic microbes by the innate immune system. Cell Host Microbe 2009; 6: 10–21.
Lukasik E, Takken FL . STANDing strong, resistance proteins instigators of plant defence. Curr Opin Plant Biol 2009; 12: 427–436.
Eitas TK, Dangl JL . NB-LRR proteins: pairs, pieces, perception, partners, and pathways. Curr Opin Plant Biol 2010; 13: 472–477.
da Silva Correia J, Miranda Y, Leonard N, Ulevitch R . SGT1 is essential for Nod1 activation. Proc Natl Acad Sci USA 2007; 104: 6764–6769.
Kadota Y, Shirasu K, Guerois R . NLR sensors meet at the SGT1-HSP90 crossroad. Trends Biochem Sci 2010; 35: 199–207.
Mayor A, Martinon F, De Smedt T, Petrilli V, Tschopp J . A crucial function of SGT1 and HSP90 in inflammasome activity links mammalian and plant innate immune responses. Nat Immunol 2007; 8: 497–503.
Franchi L, Warner N, Viani K, Nunez G . Function of Nod-like receptors in microbial recognition and host defense. Immunol Rev 2009; 227: 106–128.
Collmer A, Schneider DJ, Lindeberg M . Lifestyles of the effector rich: genome-enabled characterization of bacterial plant pathogens. Plant Physiol 2009; 150: 1623–1630.
Dangl JL, Jones JD . Plant pathogens and integrated defence responses to infection. Nature 2001; 411: 826–833.
Van der Biezen EA, Jones JD . Plant disease-resistance proteins and the gene-for-gene concept. Trends Biochem Sci 1998; 23: 454–456.
Miao EA, Warren SE . Innate immune detection of bacterial virulence factors via the NLRC4 inflammasome. J Clin Immunol 2010; 30: 502–506.
Philpott DJ, Girardin SE . Nod-like receptors: sentinels at host membranes. Curr Opin Immunol 2010; 22: 428–434.
Moore CB, Ting JP . Regulation of mitochondrial antiviral signaling pathways. Immunity 2008; 28: 735–739.
Ward HM . On the relations between host and parasite in the bromes and their brown rust, puccinia dispersa (Erikss.). Ann Bot 1902; 16: 233–316.
Stakman EC . Relation between puccinia graminis and plants highly resistant to its attack. J Agric Res 1915; 4: 193–200.
Mur LA, Kenton P, Lloyd AJ, Ougham H, Prats E . The hypersensitive response; the centenary is upon us but how much do we know? J Exp Bot 2008; 59: 501–520.
Guttman DS, Vinatzer BA, Sarkar SF, Ranall MV, Kettler G, Greenberg JT . A functional screen for the type III (Hrp) secretome of the plant pathogen Pseudomonas syringae. Science 2002; 295: 1722–1726.
Fu ZQ, Guo M, Jeong BR, Tian F, Elthon TE, Cerny RL et al. A type III effector ADP-ribosylates RNA-binding proteins and quells plant immunity. Nature 2007; 447: 284–288.
Jelenska J, Yao N, Vinatzer BA, Wright CM, Brodsky JL, Greenberg JT . A J domain virulence effector of pseudomonas syringae remodels host chloroplasts and suppresses defenses. Curr Biol 2007; 17: 499–508.
Clarke PGH, Clarke S . Nineteenth century research on naturally occurring cell death and related phenomena. Anat Embryol 1996; 193: 81–99.
Kerr JF, Wyllie AH, Currie AR . Apoptosis: a basic biological phenomenon with wide-ranging implications in tissue kinetics. Br J Cancer 1972; 26: 239–257.
Martinon F, Tschopp J . Inflammatory caspases and inflammasomes: master switches of inflammation. Cell Death Differ 2007; 14: 10–22.
Kroemer G, Galluzzi L, Vandenabeele P, Abrams J, Alnemri ES, Baehrecke EH et al. Classification of cell death: recommendations of the Nomenclature Committee on Cell Death 2009. Cell Death Differ 2009; 16: 3–11.
Brennan MA, Cookson BT . Salmonella induces macrophage death by caspase-1-dependent necrosis. Mol Microbiol 2000; 38: 31–40.
Bergsbaken T, Fink SL, Cookson BT . Pyroptosis: host cell death and inflammation. Nat Rev Microbiol 2009; 7: 99–109.
Allen IC, Scull MA, Moore CB, Holl EK, McElvania-TeKippe E, Taxman DJ et al. The NLRP3 inflammasome mediates in vivo innate immunity to influenza A virus through recognition of viral RNA. Immunity 2009; 30: 556–565.
Martinon F, Burns K, Tschopp J . The inflammasome: a molecular platform triggering activation of inflammatory caspases and processing of proIL-beta. Mol Cell 2002; 10: 417–426.
Keller M, Ruegg A, Werner S, Beer HD . Active caspase-1 is a regulator of unconventional protein secretion. Cell 2008; 132: 818–831.
Broz P, von Moltke J, Jones JW, Vance RE, Monack DM . Differential requirement for Caspase-1 autoproteolysis in pathogen-induced cell death and cytokine processing. Cell Host Microbe 2010; 8: 471–483.
Shao W, Yeretssian G, Doiron K, Hussain SN, Saleh M . The caspase-1 digestome identifies the glycolysis pathway as a target during infection and septic shock. J Biol Chem 2007; 282: 36321–36329.
Lamkanfi M, Kanneganti TD, Van Damme P, Vanden Berghe T, Vanoverberghe I, Vandekerckhove J et al. Targeted peptidecentric proteomics reveals caspase-7 as a substrate of the caspase-1 inflammasomes. Mol Cell Proteomics 2008; 7: 2350–2363.
Akhter A, Gavrilin MA, Frantz L, Washington S, Ditty C, Limoli D et al. Caspase-7 activation by the Nlrc4/Ipaf inflammasome restricts legionella pneumophila infection. PLoS Pathog 2009; 5: e1000361.
Fink SL, Cookson BT . Caspase-1-dependent pore formation during pyroptosis leads to osmotic lysis of infected host macrophages. Cell Microbiol 2006; 8: 1812–1825.
Degterev A, Huang Z, Boyce M, Li Y, Jagtap P, Mizushima N et al. Chemical inhibitor of nonapoptotic cell death with therapeutic potential for ischemic brain injury. Nat Chem Biol 2005; 1: 112–119.
Vandenabeele P, Galluzzi L, Vanden Berghe T, Kroemer G . Molecular mechanisms of necroptosis: an ordered cellular explosion. Nat Rev Mol Cell Biol 2010; 11: 700–714.
Cho YS, Challa S, Moquin D, Genga R, Ray TD, Guildford M et al. Phosphorylation-driven assembly of the RIP1-RIP3 complex regulates programmed necrosis and virus-induced inflammation. Cell 2009; 137: 1112–1123.
He S, Wang L, Miao L, Wang T, Du F, Zhao L et al. Receptor interacting protein kinase-3 determines cellular necrotic response to TNF-alpha. Cell 2009; 137: 1100–1111.
Declercq W, Vanden Berghe T, Vandenabeele P . RIP kinases at the crossroads of cell death and survival. Cell 2009; 138: 229–232.
Zhang DW, Shao J, Lin J, Zhang N, Lu BJ, Lin SC et al. RIP3, an energy metabolism regulator that switches TNF-induced cell death from apoptosis to necrosis. Science 2009; 325: 332–336.
Kim YS, Morgan MJ, Choksi S, Liu ZG . TNF-induced activation of the Nox1 NADPH oxidase and its role in the induction of necrotic cell death. Mol Cell 2007; 26: 675–687.
Torres MA, Dangl JL . Functions of the respiratory burst oxidase in biotic interactions, abiotic stress and development. Curr Opin Plant Biol 2005; 8: 397–403.
Torres MA . ROS in biotic interactions. Physiol Plant 2010; 138: 414–429.
Zitvogel L, Kepp O, Kroemer G . Decoding cell death signals in inflammation and immunity. Cell 2010; 140: 798–804.
Challa S, Chan FK . Going up in flames: necrotic cell injury and inflammatory diseases. Cell Mol Life Sci 2010; 67: 3241–3253.
Labbe K, Saleh M . Cell death in the host response to infection. Cell Death Differ 2008; 15: 1339–1349.
Faherty CS, Maurelli AT . Staying alive: bacterial inhibition of apoptosis during infection. Trends Microbiol 2008; 16: 173–180.
Hernandez LD, Pypaert M, Flavell RA, Galan JE . A Salmonella protein causes macrophage cell death by inducing autophagy. J Cell Biol 2003; 163: 1123–1131.
Suzuki T, Franchi L, Toma C, Ashida H, Ogawa M, Yoshikawa Y et al. Differential regulation of caspase-1 activation, pyroptosis, and autophagy via Ipaf and ASC in Shigella-infected macrophages. PLoS Pathog 2007; 3: e111.
Guo M, Tian F, Wamboldt Y, Alfano JR . The majority of the type III effector inventory of pseudomonas syringae pv. tomato DC3000 can suppress plant immunity. Mol Plant Microbe Interact 2009; 22: 1069–1080.
Jamir Y, Guo M, Oh HS, Petnicki-Ocwieja T, Chen S, Tang X et al. Identification of pseudomonas syringae type III effectors that can suppress programmed cell death in plants and yeast. Plant J 2004; 37: 554–565.
Fujikawa T, Ishihara H, Leach JE, Tsuyumu S . Suppression of defense response in plants by the avrBs3/pthA gene family of xanthomonas spp. Mol Plant Microbe Interact 2006; 19: 342–349.
Bos JI, Chaparro-Garcia A, Quesada-Ocampo LM, McSpadden Gardener BB, Kamoun S . Distinct amino acids of the phytophthora infestans effector AVR3a condition activation of R3a hypersensitivity and suppression of cell death. Mol Plant Microbe Interact 2009; 22: 269–281.
Bos JI, Kanneganti TD, Young C, Cakir C, Huitema E, Win J et al. The C-terminal half of phytophthora infestans RXLR effector AVR3a is sufficient to trigger R3a-mediated hypersensitivity and suppress INF1-induced cell death in Nicotiana benthamiana. Plant J 2006; 48: 165–176.
Kelley BS, Lee SJ, Damasceno CM, Chakravarthy S, Kim BD, Martin GB et al. A secreted effector protein (SNE1) from phytophthora infestans is a broadly acting suppressor of programmed cell death. Plant J 2010; 62: 357–366.
Alfano JR, Collmer A . Bacterial pathogens in plants: life up against the wall. Plant Cell 1996; 8: 1683–1698.
Walton JD . Host-selective toxins: agents of compatibility. Plant Cell 1996; 8: 1723–1733.
Govrin EM, Levine A . The hypersensitive response facilitates plant infection by the necrotrophic pathogen botrytis cinerea. Curr Biol 2000; 10: 751–757.
Meehan F, Murphy HC . A new helminthosporium blight of oats. Science 1946; 104: 413–414.
Meehan F, Murphy HC . Differential phytotoxicity of metabolic by-products of helminthosporium victoriae. Science 1947; 106: 270–271.
Lorang JM, Sweat TA, Wolpert TJ . Plant disease susceptibility conferred by a ‘resistance’ gene. Proc Natl Acad Sci USA 2007; 104: 14861–14866.
Wolpert TJ, Navarre DA, Moore DL, Macko V . Identification of the 100-kD victorin binding protein from oats. Plant Cell 1994; 6: 1145–1155.
Aarts N, Metz M, Holub E, Staskawicz BJ, Daniels MJ, Parker JE . Different requirements for EDS1 and NDR1 by disease resistance genes define at least two R gene-mediated signaling pathways in arabidopsis. Proc Natl Acad Sci USA 1998; 95: 10306–10311.
Shapiro AD, Zhang C . The role of NDR1 in avirulence gene-directed signaling and control of programmed cell death in arabidopsis. Plant Physiol 2001; 127: 1089–1101.
Wiermer M, Feys BJ, Parker JE . Plant immunity: the EDS1 regulatory node. Curr Opin Plant Biol 2005; 8: 383–389.
Shirasu K, Nakajima H, Rajasekhar VK, Dixon RA, Lamb C . Salicylic acid potentiates an agonist-dependent gain control that amplifies pathogen signals in the activation of defense mechanisms. Plant Cell 1997; 9: 261–270.
Emerson RA . The inheritance of blotch leaf in maize. Cornell Univ Agric Exper 1923; 70: 3–16.
Hoisington DA, Neuffer MG, Walbot V . Disease lesion mimics in maize. I. Effect of genetic background, temperature, developmental age, and wounding on necrotic spot formation with Les1. Dev Biol 1982; 93: 381–388.
Langford AN . Autogenous necrosis in tomatoes immune from cladosporium fulvum cooke. Can J Res 1948; 26: 35–64.
Wolter M, Hollricher K, Salamini F, Schulze-Lefert P . The mlo resistance alleles to powdery mildew infection in barley trigger a developmentally controlled defence mimic phenotype. Mol Gen Genet 1993; 239: 122–128.
Greenberg JT, Ausubel FM . Arabidopsis mutants compromised for the control of cellular damage during pathogenesis and aging. Plant J 1993; 4: 327–341.
Lorrain S, Vailleau F, Balague C, Roby D . Lesion mimic mutants: keys for deciphering cell death and defense pathways in plants? Trends Plant Sci 2003; 8: 263–271.
Dietrich RA, Delaney TP, Uknes SJ, Ward ER, Ryals JA, Dangl JL . Arabidopsis mutants simulating disease resistance response. Cell 1994; 77: 565–577.
lsd4 is a dominant, gain of function mutation characterized by spontaneous lesions and enhanced disease resistance after the onset of lesioning. Two alleles, lsd4-1 and lsd4-2, were used to map LSD4 to a 89 kb interval. Sequencing of the entire region identified At5g53170 as the LSD4 candidate gene. LSD4 is FtSH11, a protease of 806 amino acids with two predicted transmembrane domains, a Walker A and B motif, a SRH domain and a zinc binding motif. lsd4-1 represents a six amino acid deletion (aa 264-269), whereas in lsd4-2 serin277 is mutated to a phenylalanine. FtSH11 is localized to the chloroplast and mitochondrium129, and involved in PSII associated LHCII catabolic processes. Mutations in three other chloroplastic FtSH proteases (At5g42270 (VAR1, FtSH5130)), At2g30950 (VAR2, FtSH2131,132) and At1g50250 (FtSH1133) lead to similar cell death phenotypes.
Dietrich RA, Richberg MH, Schmidt R, Dean C, Dangl JL . A novel zinc finger protein is encoded by the arabidopsis LSD1 gene and functions as a negative regulator of plant cell death. Cell 1997; 88: 685–694.
Jabs T, Dietrich RA, Dangl JL . Initiation of runaway cell death in an arabidopsis mutant by extracellular superoxide. Science 1996; 273: 1853–1856.
Mateo A, Funck D, Muhlenbock P, Kular B, Mullineaux PM, Karpinski S . Controlled levels of salicylic acid are required for optimal photosynthesis and redox homeostasis. J Exp Bot 2006; 57: 1795–1807.
Rusterucci C, Aviv DH, Holt III BF, Dangl JL, Parker JE . The disease resistance signaling components EDS1 and PAD4 are essential regulators of the cell death pathway controlled by LSD1 in arabidopsis. Plant Cell 2001; 13: 2211–2224.
Epple P, Mack AA, Morris VR, Dangl JL . Antagonistic control of oxidative stress-induced cell death in Arabidopsis by two related, plant-specific zinc finger proteins. Proc Natl Acad Sci USA 2003; 100: 6831–6836.
Coll NS, Vercammen D, Smidler A, Clover C, van Breusegem F, Dangl JL et al. Arabidopsis Type I metacaspases control cell death. Science 2010; 330: 1393–1397.
Kaminaka H, Nake C, Epple P, Dittgen J, Schutze K, Chaban C et al. bZIP10-LSD1 antagonism modulates basal defense and cell death in arabidopsis following infection. EMBO J 2006; 25: 4400–4411.
Hatsugai N, Iwasaki S, Tamura K, Kondo M, Fuji K, Ogasawara K et al. A novel membrane fusion-mediated plant immunity against bacterial pathogens. Genes Dev 2009; 23: 2496–2506.
Hatsugai N, Kuroyanagi M, Yamada K, Meshi T, Tsuda S, Kondo M et al. A plant vacuolar protease, VPE, mediates virus-induced hypersensitive cell death. Science 2004; 305: 855–858.
Lam E, del Pozo O . Caspase-like protease involvement in the control of plant cell death. Plant Mol Biol 2000; 44: 417–428.
Rojo E, Martin R, Carter C, Zouhar J, Pan S, Plotnikova J et al. VPEgamma exhibits a caspase-like activity that contributes to defense against pathogens. Curr Biol 2004; 14: 1897–1906.
Uren AG, O'Rourke K, Aravind LA, Pisabarro MT, Seshagiri S, Koonin EV et al. Identification of paracaspases and metacaspases: two ancient families of caspase-like proteins, one of which plays a key role in MALT lymphoma. Mol Cell 2000; 6: 961–967.
Aravind L, Koonin EV . Classification of the caspase-hemoglobinase fold: detection of new families and implications for the origin of the eukaryotic separins. Proteins 2002; 46: 355–367.
Coornaert B, Baens M, Heyninck K, Bekaert T, Haegman M, Staal J et al. T cell antigen receptor stimulation induces MALT1 paracaspase-mediated cleavage of the NF-kappaB inhibitor A20. Nat Immunol 2008; 9: 263–271.
Rebeaud F, Hailfinger S, Posevitz-Fejfar A, Tapernoux M, Moser R, Rueda D et al. The proteolytic activity of the paracaspase MALT1 is key in T cell activation. Nat Immunol 2008; 9: 272–281.
Vercammen D, van de Cotte B, De Jaeger G, Eeckhout D, Casteels P, Vandepoele K et al. Type II metacaspases Atmc4 and Atmc9 of arabidopsis thaliana cleave substrates after arginine and lysine. J Biol Chem 2004; 279: 45329–45336.
Thome M . Multifunctional roles for MALT1 in T-cell activation. Nat Rev Immunol 2008; 8: 495–500.
Madeo F, Carmona-Gutierrez D, Ring J, Buttner S, Eisenberg T, Kroemer G . Caspase-dependent and caspase-independent cell death pathways in yeast. Biochem Biophys Res Commun 2009; 382: 227–231.
Madeo F, Herker E, Maldener C, Wissing S, Lachelt S, Herlan M et al. A caspase-related protease regulates apoptosis in yeast. Mol Cell 2002; 9: 911–917.
Lee RE, Brunette S, Puente LG, Megeney LA . Metacaspase Yca1 is required for clearance of insoluble protein aggregates. Proc Natl Acad Sci USA 2010; 107: 13348–13353.
Lee RE, Puente LG, Kaern M, Megeney LA . A non-death role of the yeast metacaspase: Yca1p alters cell cycle dynamics. PLoS One 2008; 3: e2956.
Hoeberichts FA, ten Have A, Woltering EJ . A tomato metacaspase gene is upregulated during programmed cell death in botrytis cinerea-infected leaves. Planta 2003; 217: 517–522.
Hao L, Goodwin PH, Hsiang T . Expression of a metacaspase gene of Nicotiana benthamiana after inoculation with colletotrichum destructivum or pseudomonas syringae pv. tomato, and the effect of silencing the gene on the host response. Plant Cell Rep 2007; 26: 1879–1888.
Zimmermann P, Hirsch-Hoffmann M, Hennig L, Gruissem W . GENEVESTIGATOR. arabidopsis microarray database and analysis toolbox. Plant Physiol 2004; 136: 2621–2632.
Van Baarlen P, Woltering EJ, Staats M, Van Kan JA . Histochemical and genetic analysis of host and non-host interactions of arabidopsis with three Botrytis species: an important role for cell death control. Mol Plant Pathol 2007; 8: 41–54.
Nishimura MT, Dangl JL . Arabidopsis and the plant immune system. Plant J 2010; 61: 1053–1066.
Budd RC, Yeh WC, Tschopp J . cFLIP regulation of lymphocyte activation and development. Nat Rev Immunol 2006; 6: 196–204.
Irmler M, Thome M, Hahne M, Schneider P, Hofmann K, Steiner V et al. Inhibition of death receptor signals by cellular FLIP. Nature 1997; 388: 190–195.
Kawadler H, Gantz MA, Riley JL, Yang X . The paracaspase MALT1 controls caspase-8 activation during lymphocyte proliferation. Mol Cell 2008; 31: 415–421.
LeBlanc PM, Yeretssian G, Rutherford N, Doiron K, Nadiri A, Zhu L et al. Caspase-12 modulates NOD signaling and regulates antimicrobial peptide production and mucosal immunity. Cell Host Microbe 2008; 3: 146–157.
Rasper DM, Vaillancourt JP, Hadano S, Houtzager VM, Seiden I, Keen SL et al. Cell death attenuation by ‘Usurpin’, a mammalian DED-caspase homologue that precludes caspase-8 recruitment and activation by the CD-95 (Fas, APO-1) receptor complex. Cell Death Differ 1998; 5: 271–288.
Saleh M, Mathison JC, Wolinski MK, Bensinger SJ, Fitzgerald P, Droin N et al. Enhanced bacterial clearance and sepsis resistance in caspase-12-deficient mice. Nature 2006; 440: 1064–1068.
Saleh M, Vaillancourt JP, Graham RK, Huyck M, Srinivasula SM, Alnemri ES et al. Differential modulation of endotoxin responsiveness by human caspase-12 polymorphisms. Nature 2004; 429: 75–79.
Dupaul-Chicoine J, Yeretssian G, Doiron K, Bergstrom KS, McIntire CR, LeBlanc PM et al. Control of intestinal homeostasis, colitis, and colitis-associated colorectal cancer by the inflammatory caspases. Immunity 2010; 32: 367–378.
Labbe K, Miu J, Yeretssian G, Serghides L, Tam M, Finney CA et al. Caspase-12 dampens the immune response to malaria independently of the inflammasome by targeting NF-kappaB signaling. J Immunol 2010; 185: 5495–5502.
Kiraly Z, Barna B, Ersek T . Hypersensitivity as a consequence, not cause, of plant resistance to infection. Nature 1972; 239: 456–458.
Bendahmane A, Kanyuka K, Baulcombe DC . The Rx gene from potato controls separate virus resistance and cell death responses. Plant Cell 1999; 11: 781–791.
Bulgarelli D, Biselli C, Collins CC, Consonni G, Stanca AM, Schulze-Lefert P et al. The CC-NB-LRR-type Rdg2a resistance gene confers immunity to the seed-borne balrey leaf stripe pathogen in the absence of hypersensitive cell death. PLoS One 2010; 5: e12599.
Goulden MG, Baulcombe DC . Functionally homologous host components recognize potato virus X in gomphrena globosa and potato. Plant Cell 1993; 5: 921–930.
Jakobek JL, Lindgren PB . Generalized induction of defense responses in bean is not correlated with the induction of the hypersensitive reaction. Plant Cell 1993; 5: 49–56.
Lehnackers H, Knogge W . Cytological studies on the infection of barley cultivars with known resistance genotypes by rhynchosporium-secalis. Can J Bot 1990; 68: 1953–1961.
Ori N, Eshed Y, Paran I, Presting G, Aviv D, Tanksley S et al. The I2C family from the wilt disease resistance locus I2 belongs to the nucleotide binding, leucine-rich repeat superfamily of plant resistance genes. Plant Cell 1997; 9: 521–532.
Rohe M, Gierlich A, Hermann H, Hahn M, Schmidt B, Rosahl S et al. The race-specific elicitor, NIP1, from the barley pathogen, rhynchosporium secalis, determines avirulence on host plants of the Rrs1 resistance genotype. EMBO J 1995; 14: 4168–4177.
Schiffer R, Gorg R, Jarosch B, Beckhove U, Bahrenberg G, Kogel KH et al. Tissue dependence and differential cordycepin sensitivity of race-specific resistance responses in the barley powdery mildew interaction. Mol Plant Microbe In 1997; 10: 830–839.
del Pozo O, Lam E . Caspases and programmed cell death in the hypersensitive response of plants to pathogens (vol 8, pg 1129, 1998). Curr Biol 1998; 8: R896–R896.
Dorey S, Kopp M, Geoffroy P, Fritig B, Kauffmann S . Hydrogen peroxide from the oxidative burst is neither necessary nor sufficient for hypersensitive cell death induction, phenylalanine ammonia lyase stimulation, salicylic acid accumulation, or scopoletin consumption in cultured tobacco cells treated with elicitin. Plant Physiol 1999; 121: 163–172.
Durrant WE, Dong X . Systemic acquired resistance. Annu Rev Phytopathol 2004; 42: 185–209.
Torres MA, Jones JD, Dangl JL . Pathogen-induced, NADPH oxidase-derived reactive oxygen intermediates suppress spread of cell death in Arabidopsis thaliana. Nat Genet 2005; 37: 1130–1134.
Urantowka A, Knorpp C, Olczak T, Kolodziejczak M, Janska H . Plant mitochondria contain at least two i-AAA-like complexes. Plant Mol Biol 2005; 59: 239–252.
Sakamoto W, Tamura T, Hanba-Tomita Y, Sodmergen, Murata M . The VAR1 locus of Arabidopsis encodes a chloroplastic FtsH and is responsible for leaf variegation in the mutant alleles. Genes Cells 2002; 8: 769–780.
Chen M, Choi Y, Voytas DF, Rodermel S . Mutations in the Arabidopsis VAR2 locus cause leaf variegation due to loss of a chloroplast FtsH protease. Plant J 2000; 4: 303–313.
Bailey S, Thompson E, Nixon PJ, Horton P, Mullineaux CW, Robinson C et al. A critical role for the Var2 FtsH homologue of Arabidopsis thaliana in the photosystem II repair cycle in vivo. J Biol Chem 2002; 3: 2006–2011.
Seo S, Okamoto M, Iwai T, Iwano M, Fukui K, Isogai A et al. Reduced levels of chloroplast FtsH protein in tobacco mosaic virus-infected tobacco leaves accelerate the hypersensitive reaction. Plant Cell 2000; 6: 917–932.
Acknowledgements
We thank Dr Peter Bozhkov (Swedish University of Agricultural Sciences) Dr Vera Bonardi, Dr Marc Nishimura and Dr Tim Eitas (UNC) for critical reading of the manuscript. This work was funded by NIH-GM057171 to JLD.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by P Bozhkov
Rights and permissions
About this article
Cite this article
Coll, N., Epple, P. & Dangl, J. Programmed cell death in the plant immune system. Cell Death Differ 18, 1247–1256 (2011). https://doi.org/10.1038/cdd.2011.37
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2011.37
Keywords
This article is cited by
-
The role and pathway of VQ family in plant growth, immunity, and stress response
Planta (2024)
-
Cathepsin B degrades RbcL during freezing-induced programmed cell death in Arabidopsis
Plant Cell Reports (2024)
-
Fusaric acid-evoked oxidative stress affects plant defence system by inducing biochemical changes at subcellular level
Plant Cell Reports (2024)
-
Disruption of the Novel Small Protein RBR7 Leads to Enhanced Plant Resistance to Blast Disease
Rice (2023)
-
Transcriptome and metabolome analyses revealed the response mechanism of pepper roots to Phytophthora capsici infection
BMC Genomics (2023)