Abstract
Seed size is a major determinant of seed yield but few is known about the genetics controlling of seed size in plants. Phytohormones cytokinin and brassinosteroid were known to be involved in the regulation of herbaceous plant seed development. Here we identified a homolog of Auxin Response Factor 19 (JcARF19) from a woody plant Jatropha curcas and genetically demonstrated its functions in controlling seed size and seed yield. Through Virus Induced Gene Silencing (VIGS), we found that JcARF19 was a positive upstream modulator in auxin signaling and may control plant organ size in J. curcas. Importantly, transgenic overexpression of JcARF19 significantly increased seed size and seed yield in plants Arabidopsis thaliana and J. curcas, indicating the importance of auxin pathway in seed yield controlling in dicot plants. Transcripts analysis indicated that ectopic expression of JcARF19 in J. curcas upregulated auxin responsive genes encoding essential regulators in cell differentiation and cytoskeletal dynamics of seed development. Our data suggested the potential of improving seed traits by precisely engineering auxin signaling in woody perennial plants.
Similar content being viewed by others
Introduction
Plant seeds and seed products comprise more than 70% of calories and 60% of all proteins consumed by the human population. Global seed-derived food security is now facing serious problems caused by multiple factors, such as continuous population increase, reduced arable land, global climate change, and the demands for the production of biofuels1. The United States Department of Energy estimates that global demand for energy will increase by approximately 35% between 2005 and 2030. Use of sustainable biofuels as energy is predicted to be a major contributor to the increase and a further threat to global food security. The use of agricultural plants such as corn and soybean as feedstock for large-scale biofuel production would also conflict with food production, causing food supply shortages, increased food prices and ethical conflicts2.One potential solution to the food crisis is to produce biofuels from plant species capable of growing in marginal lands. Such species should be easy to grow and propagate with high oil yield, and their cultivation should be environmentally beneficial or neutral. Life cycle assessment (LCA) is a quantitative approach to estimating the environmental sustainability of biofuels3. However, the widespread commercialization of biodiesel is limited by low seed yields, which drive up costs.
Seed yield is related to seed size, which is predominantly determined by genetic factors. Genetic studies with the model species Arabidopsis thaliana and rice (Oryza sativa) have identified a number of genes affecting seed size1,4. Current evidences suggest that seed size is mainly controlled by epigenetics and by genetic pathways, including proteasomal degradation, phytohormones, G protein signaling and small RNA regulation1,4,5,6,7,8,9. Although manipulation of these processes could maximize seed yield, how they are regulated and integrated is poorly understood. Furthermore, as a consequence of the complex organization of seeds, only a few studies report yield improvements by direct engineering of genes for either seed size or final seed yield10. Therefore, it is necessary to identify more genes from model or non-model plants to understand the genetic network that controls seed size11.
Plant hormones play pivotal roles in the developmental processes of diverse traits related to yield12. For example, the phytohormones cytokinin and brassinosteroid are involved in the regulation of herbaceous plant seed development1. Auxin regulates cell division and is further involved in almost every process of plant growth and development, including seed size. Auxin signaling initiates from hormone perception by an F-box protein receptor Transport Inhibitor Response 1 (TIR1) followed by degradation of negative regulators AUX/IAA proteins13. Auxin Response Factors (ARFs) are then released. In the current auxin signaling model, Aux/IAAs dimerize with and repress ARF transcription factors in the absence of auxin. In the presence of auxin, Aux/IAAs interact with TIR1 resulting in repressor degradation, freeing ARFs for auxin-responsive gene transcription14. The interaction between ARFs and IAAs is a key aspect of auxin regulation and occurs through the highly conserved COOH-terminal (CT) Phox and Bem1p (PB1) domain present in both types of proteins15,16,17,18.Genetically, auxin induces the formation of Arabidopsis pluripotent cells via a root development pathway by ARF7, ARF19 and other downstream transcription factors such as Lateral Organ Boundaries-domain (LBD) LBD16, LBD17, LBD18 and LBD2919. They bind to auxin-responsive elements (AuxREs) in promoter regions to enhance or repress auxin-regulated genes and auxin-inducible genes. In Arabidopsis, ARF2 negatively regulates auxin signaling and seed size via downregulation of cell division in the integument region of endosperm. However, arf2 plants exhibit pleiotropic effects, e.g. reduced seed set, which make these plants undesirable for agriculture20,21,22. Rice THOUSAND-GRAIN WEIGHT 6 (TGW6) encodes an IAA-glucosehydrolase and plays an essential role in the regulation of auxin homeostasis during endosperm development. Downregulation of this negative regulator in the auxin pathway increases grain weight and yield in rice23. Activation of the BIG GRAIN1 (BG1) gene, which encodes a regulator in auxin transporting in rice, leads to significantly increased seed size and plant biomass, seed weight and yield24. The YUCCA1 (ZmYUC1) gene in maize (Zea mays) encodes a seed-specific flavin monooxygenase that is involved in tryptophan-dependent IAA biosynthesis; its mutant has 40% less dry mass than wild-type seeds25. However, which ARFs and how the ARFs and AUX/IAA proteins control seed development remains largely unknown12.
Compared with conventional herbaceous vegetative oil, seed oil from perennial woody plants has notable nutritional advantages as well as potential raw materials for biodiesel production. Woody plants are usually perennials with strong secondary xylem that can efficiently transport water and nutrients from root to leaf. Jatropha curcas (Jatropha) is one promising new energy crop with high seed oil content, tolerance to drought and an ability to thrive in poor soil26. Furthermore, the reduction of green house gas emission for generating 1 gigajoule energy can be at least 40–107% with respect to fossil diesel27,28,29. However, Jatropha has some undesirable traits, such as a high male-flower ratio, high sensitivity to viral diseases, high polyunsaturated fatty acid level and low seed yields30,31,32. To improve the inadequacy, biotechnological progress had been made in the seed oil profile, flowering traits and virus resistance of Jatropha31,33,34,35,36, however, seed size and seed yield of Jatropha cultivars would need to improve to meet seed production demands34. It is known that auxin plays an important role in the growth of Jatropha. Pretreated cuttings Jatropha with indole-3-butyric acid (IBA) and 1-naphthalene acetic acid (NAA) increased rooting, sprouting, and seeds yield. Auxin pretreated cuttings produced much longer and healthier plants that flowered and produced fruits and seeds a year earlier than the control37. However the detailed molecular mechanism how extrogenous application of auxin can increase seed yield is still unknown.
Seed size in J. curcas is greater than in the congener J. integerrima (Fig. 1a). Using the inter-species crossing population J. curcas × J. integerrima, we conducted a whole-genome scan for quantitative trait loci (QTLs) and expression QTLs that affect seed oil traits in Jatropha11,38,39. We screened key genes in auxin pathway including ARFs, IAAs and downstream effectors to identify candidate genes controlling seed size. Importantly, we detected a major QTL qSL11-a controlling seed length with a high likelihood of odd score (LOD score) of 16.69 and percentage of variance explained (PVE) 29.6% on Linkage group (LG) 11, were harboring the JcARF19 gene11.
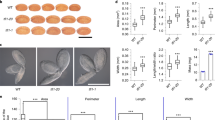
(a) Seed cross-sections of J. curcas (left) and J. integerrima (right). En: Endosperm; Em: Embryo. Size bar: 10 mm. (b) Expression of JcARF19 in different organs. Root, stem and leaf samples were harvested at six weeks after fertilization and fruit shell and endosperm samples were collected at five weeks after fertilization. Each organ had three biological replicates and each biological replicate with three technical replicates. Values (n = 3) were shown as mean ± SD. The relative expression level of JcARF19 in Jatropha was normalized with the expression level of 18S RNA in Jatropha. (c) Phenotypes of Jatropha plants leaves at 27 days post-infiltration (dpi) with various synthetic TRV (sTRV) constructs: Empty vector (EV), sTRV: JcARF19 (JcARF19), sTRV: JcCH42 (JcCH42). Bar: 10 mm. (d) The width of leaves from Jatropha plants infiltrated with various synthetic TRV constructs at 27 dpi. Values (n = 5) were shown as mean ± SD and statistic analysis with Student T-test. **Indicates P < 0.01. *Indicates P < 0.05. (e) Relative expression levels of JcLBD18 and JcLBD29 in systemic leaves of plants infiltrated with sTRV empty vector (EV) and sTRV: JcARF19 (JcARF19). Samples were collected after treatment of IAA (10 nM) and each treatment had three biological replicates. For qRT-PCR, each biological replicate was replicated three times. Numbers represent mean relative values from three independent experiments with standard deviation. The relative expression level of JcIAA9 in sTRV- silenced plants was normalized as 1.
In this study, we functionally characterized a key genetic factor ARF19 which regulates seed size and seed yield in Jatropha. We found that JcARF19 functions as a positive upstream modulator. Its ectopic expression leads to increase in cell size and cell number in Jatropha. Transgenic expression of JcARF19 can also increase seed size and seed weight both in Arabidopsis and Jatropha.
Results
Down regulation of JcARF19 affected auxin signalling transduction in Jatropha
According to the whole-genome scan results, we hypothesized that JcARF19 might participate in controlling seed size. To test the hypothesis, firstly, we investigated potential tissue-specific roles of JcARF19 in Jatropha by profiling its expression patterns in roots, stems, leaves, fruits and seeds using quantitative real-time PCR. JcARF19 showed highly similar expression profiles to ubiquitous expression in all organs and highest in endosperm (Fig. 1b), providing evidence that JcARF19 might be related to seed traits in Jatropha. To test the function of JcARF19 in auxin signalling transduction and therefore plant development in Jatropha, we used synthetic tobacco rattle virus (sTRV) based virus-induced gene silencing (VIGS) method, which we developed previously and allowed us to rapidly identify gene function in various plants40,41. At 27 days after inoculation of agrobacterium containing VIGS vectors of JcARF19 and positive control Jatropha Chlorata 42 (JcCH42) or empty vector control (EV), we observed distinct smaller newly expanding leaves in JcARF19-silenced plants than EV-treated plants (Fig. 1c) and the difference between them was remarkably as shown in Fig. 1d34. Smaller leaf size was also observed in the positive control treatment containing the silenced marker gene, JcCH42, which encodes a subunit of Magnesium (Mg) chelatase, involved in photosynthesis (Fig. 1c)42. Recent research showed that auxin treatment enhances ARF19 binding to its target gene promoters, which correlates with the enhancement of transcriptional activity of the ARF19 in Arabidoposis thaliana (Arabidopsis)43.We next tested the gene expression of downstream transcription factors esp. Lateral Organ Boundaries-domain (LBDs). Upon auxin treatment, two genes JcLBD18 and JcLBD29 which encode putative LBDs of Jatropha were up-regulated differentially (Fig. 1e), especially JcLBD18 which had 7-fold higher expression in EV-treated plants. But in JcARF19-silenced plants, there was no obvious induction of JcLBD18 or JcLBD29 upon auxin treatment, indicating that JcARF19 was essential for proper auxin-mediated signalling transduction process in leaf cells of Jatropha.
Transgenic overexpression of JcARF19 increased seed size and dry seed weight in Arabidopsis
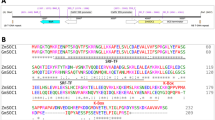
To further investigate the function of JcARF19, at first, we got partial cDNA sequence from a database of sequenced cDNA library prepared from Jatropha seeds11. By integrating known genomic sequence of JcARF19 and cloned sequenced information from 5′RACE and 3′RACE, we finally got the full-length coding sequence of JcARF19 (Genbank accession NO. KX988008, detailed sequence information could be found in Supplementary file). JcARF19 protein encoded 1133 amino acids and consists of major functional domains, an amine-terminus B3 DNA binding domain and a carboxyl-terminus (CT) Phox and Bem1p (PB1) domain. Amino acid sequence alignment showed that ARF19s from various plants had high sequence similarity in the three conserved domains (Fig. S1), suggesting functional conservation of ARF19 family proteins. To further investigated the function of JcARF19, we generated thirty transgenic lines for CaMV35S:JcARF19 in Arabidoposis44. An obvious increase in plant size and seed size was observed in the 35S:JcARF19 plants compared to wild-type (WT) Col-0 plants (Fig. 2a–d). Seed length and seed dry weight of 35S:JcARF19 lines were increased remarkably compared with those of WT Col-0 plants as well (Fig. 2e,f). We further conducted oil traits analysis to check the effects of JcARF19 on oil yield or oil composition. There was no significant change in either oil content per dry seed weight or oil composition in transgenic line of 35S:JcARF19 compared with those of WT Col-0 plants (Fig. S2). Since seed size and weight were significantly increased in transgenic line of 35S:JcARF19, the lipid content per seed was obviously increased. These results indicated that overexpression of JcARF19 increased seed size, seed dry weight and oil yield, but had no effect on oil composition of seeds in Arabidopsis.
Fourteen-day-old WT Columbia ecotype (Col-0) Arabidopsis (a) and 35S:JcARF19 overexpression plants (b). T3 generation plants were used for observations. Bar: 1 cm. SEM observation seeds of WT Col-0 (c) and 35S:JcARF19 overexpression plants (d). T3 generation plant seeds were used for observations. Bar: 100 μm. (e) Seed length of WT Col-0 and 35S: JcARF19 overexpression Arabidopsis. Values are mean ± SD (n = 10) and statistic analysis with Student T-test. **Indicates P < 0.01. (f) Dry seed weight of 100 seeds of WT Col-0 and 35S: JcARF19 overexpression Arabidopsis. Values are mean ± SD (n = 10) and statistic analysis with Student T-test. **Indicates P < 0.01.
Increased seed size and seed numbers by overexpression of JcARF19 in Jatropha
Jatropha was also transformed with 35S:JcARF19 vector31. The ectopic expression of JcARF19 affected calluses formation during transformation procedure (Fig. S3), producing bigger calluses than that of empty vector (EV). After transferring to soil, the ectopic expression of JcARF19 also affected flowering time (Fig. 3a) and other characteristics (Fig. 3c–f). Under normal growth conditions in a greenhouse, WT Jatropha Jc-MD required around 8 months to produce the first inflorescence whereas 10 primary independent JcARF19 overexpression lines formed their first inflorescences after only 5 months, with a 3 months reduction (Figs 3a and 4a–c). Quantitative reverse transcriptase PCR (qRT-PCR) analysis verified the presence of the transgene and ectopic expression of JcARF19 in transgenic Jatropha plants (Fig. 3b). JcARF19 overexpression lines also had greater branching compared with WT Jc-MD control plants (Figs 3c and 4a). A four-fold increase in seed set from JcARF19 ectopic expression plants was collected within one year of transplanting. Furthermore, a 17.2% increase of single seed weight and 17.1% increase in fruit size were found in JcARF19 ectopic expression plants (Figs 3d–f and 4d–f). The longer cell length (18.4%) and higher cell number (16.5%) in JcARF19 ectopic expression plants explained the increased seed weights and lengths (Figs 3g,h and 4g,h). These results indicated that overexpression of JcARF19 increased length, number and weight of seeds and also oil yield by increasing seed cell number and length in Jatropha.
(a) Flowering time in 10 independent T0 transgenic Jatropha plants overexpressing JcARF19 and wild-type plant Jc-MD. Values are mean ± SD (n = 10), and Student T-test statistic was used for analysis. **Indicates P < 0.01, the same for the below traits analysis. (b) Relative JcARF19 expression level in leaves of T0 generation 35S:JcARF19 transgenic Jatropha lines (JcARF19OE#1, #10 and #13) and wild-type plant Jc-MD. Values are mean ± SD (n = 3). (c) Comparison of branch number of WT plant Jc-MD and ten T0 JcARF19 overexpression lines. Values are mean ± SEM (n = 10). (d) Comparison of seed number per tree per year of WT plant Jc-MD and ten T0 JcARF19OE lines. Values are mean ± SEM (n = 10). (e) Comparison of single seed weight of WT plant Jc-MD and three T0 JcARF19 overexpression (JcARF19OE #1, #10 and #13) line. Values are mean ± SEM (n = 50). (f) Comparison of seed length of WT plant Jc-MD and three JcARF19OE #1, #10 and #13 lines. Values are mean ± SEM (n = 50). (g) Comparison of cell length of WT plant Jc-MD and JcARF19OE #1 line. Values are mean ± SEM (n = 10). (h) Comparison of seed cell number of WT plant Jc-MD and JcARF19OE #1 line. Values are mean ± SEM (n = 10).
(a) Comparison of plant architecture of JcARF19 overexpression transgenic T0 plant (#1, right) with WT Jatropha plant (Jc-MD, left). (b) No inflorescence was found in WT Jatropha plant Jc-MD in the same stage as JcARF19-overexpressing transgenic T0 plant (#1). (c) JcARF19-overexpressing transgenic T0 plant (#1) with early inflorescence. Bar: 10 cm for (a–c). (d,e) Comparison of fruit size of WT Jatropha (Jc-MD, left) and transgenic T1 Jatropha plant overexpressing JcARF19 (right). Bar: 1 cm. (f) Comparison of seed size of WT Jatropha (Jc-MD, left) and transgenic T1 Jatropha plant overexpressing JcARF19 (right). Dashed line indicates the position of the cross-section. Bar: 1 cm. Comparison of cell size of WT Jatropha Jc-MD (g) and transgenic T1 Jatropha plant overexpressing JcARF19 (h). Bar: 50 μm.
We further germinated T1 JcARF19-overexpression Jatropha seeds and found that the germination percentage of T1 JcARF19-overexpression Jatropha seeds was higher than these of the wild type control (Fig. S4).
Expression of Auxin responsible genes upon the ectopic expression of JcARF19 in developing seeds
In VIGS assay, we have demonstrated that JcARF19 is essential for auxin signaling transduction on downstream transcription factors JcLBD18 and JcLBD29 in Jatropha leaves (Fig. 1e). To further understand its role in auxin pathway during seed developing, we analyzed the expression of JcLBD18 and JcLBD29 in JcARF19 overexpression seeds and found that both of two LBD genes were higher expressed either at early stage or middle stage compared with those of WT control. We further checked the expression of other downstream genes in auxin pathway such as cell cycle and cell number controlling. We found that the expression level of Expansin1 (JcEXP1) and AUXIN-REGULATED GENE INVOLVED IN ORGAN SIZE (JcARGOS)45, together with cell cycle regulators JcCDKA1, JcCYCD2 and JcCYCD5, are induced in JcARF19-ectopic expression Jatropha seeds endosperm, at either early or middle stage of seed development (Fig. 5a), providing molecular explanations for the increased cell number in JcARF19 overexpression Jatropha seeds. We found that the expression of several genes encoding important regulators in cell differentiation and cytoskeletal dynamics have been enhanced including ARGOS46, small GTPases auxin-Rho of Plants (ROP), ROP-interactive protein RIC and receptor-like auxin-(Transmembrane Kinase) TMK47,48, either in early or middle stages of seed development of J. curcas (Fig. 5b), which also explain the cell size expansion in JcARF19 ectopic expression Jatropha seeds.
Early stage: 3WAF, Middle stage: 6WAF. Values are mean ± SD (n = 3). (a) Relative expression folds of cell cycle and cell number related genes (JcARF19, JcIAA9, JcLBD18, JcLBD29, JcEXP1, JcARGOS, JcCDKA1, JcCYCD2 and JcCYCD5) in JcARF19 #1 overexpression plants normalized with the wild type. (b) Relative expression folds of cell differentiation and cytoskeletal dynamics related genes (JcRIC1, JcRIC4, JcROP2, JcROP6, JcTMK1, JcTMK2 and JcTMK3) in JcARF19 #1 overexpression plants normalized with the wild type.
Characterization of interaction between JcARF19 and JcIAA9
Each ARF transcription factor forms dimer with corresponding Aux/IAAs. Stimulated by eQTL clue of the genetic interaction of JcARF19 and JcIAA9 we identified previously, we hypothesized that ARF19 may function via direct protein-protein interaction with IAA9. The secondary structure of JcARF19 and JcIAA9 were predicted to have 4 α-helix and 5 β-sheet folding module (Fig. 6a). We found that JcARF19 interacts with JcIAA9 physically in vitro pull-down assays (Fig. 6b). We used glutathione S-transferase (GST) fused JcIAA9 as bait and JcARF19 as prey. We found that the GST-fused JcIAA9 COOH-terminal protein (JcIAA9-CT) could interact strongly with 6*Histidine-tagged ARF19 COOH-terminal protein (JcARF19-CT), in contrast to much weaker interaction found on the protein pair of J. integerrima (JiARF19 and JcIAA9). This difference was surprising because the ARF19-PB1 and IAA9-PB1 polypeptides of the two species differ by only one amino acid. The S → G mutation which located in interaction interface β5 of ARF19 proteins affects ARF19-IAA9 binding ability. We also provided evidence to show the physical interaction between JcARF19 and JcIAA9 in vivo and the vital role of key amino acid in heterodimer formation by Bimolecular Fluorescence Complementation (BiFC) assays (Fig. 6c). However, although we identified a putative protein-protein interaction pair of IAA9-ARF19 here (Fig. 6), it is still unclear what the significance of this interaction is so far and how IAA9 affects the function of ARF19 in auxin signaling pathway.
(a) Predicated secondary structure of JcIAA9, JcARF19 and other related proteins. (b) GST pull-down assay. Recombinant bait protein GST-JcIAA9 COOH-terminal (residues 229–368 shows JcIAA9-CT containing protein-protein interaction domain III and domain IV of Aux/IAA protein family) strongly binds to prey protein JcARF19 COOH-terminal (residues 1005–1276), but weakly binds to prey protein JiARF19-CT (residues 1008–1279). (c) BiFC assay. Very strong fluorescence signal was only found in the combination of C-terminal of JcIAA9 and JcARF19 (nYFP-JcIAA9+JcARF19-cYFP), but not in the combination of C-terminal of JcIAA9 and JiARF19 (nYFP-JcIAA9+JiARF19-cYFP). Bar: 50 μm.
Discussion
In herbaceous plants, species with small seeds sometimes have larger seed set than larger-seed species, assuming a limited total amount of energy. This energetic trade-off has been observed in genetic mutants such as APETALA249,50,51 and CURLY LEAF2852. In this study we showed that manipulation of the auxin pathway in J.curcas not only increased seed size but also enhanced total seed yield. Previous knowledge on the molecular mechanisms of seed size was mainly limited to model herbaceous plants, particularly Arabidopsis and rice53. Our work suggests that manipulation of auxin is an alternative approach to increase seed size in woody plants.
Several lines of evidence support the involvement of JcARF19 in seed size determination. First, JcARF19 was mapped in the major quantitative trait locus (QTL) region and was significantly associated with seed size11. Second, by using expression QTL (eQTL) analysis to link variants with functional candidate genes, we provided evidences that seed traits were affected by the genetic interaction of JcARF19 and JcIAA911. Third, the C-terminal of JcARF19, which is essential for protein-protein interaction among ARF proteins, was identical between ARF19 homologs of bigger seed size in Jatropha. Point mutations on the single-nucleotide polymorphisms (SNPs) of JcARF19 affected their direct physical interactions (Fig. 6). Fourth, overexpression of JcARF19 increased seed size in both Arabidopsis and Jatropha by increasing both cell numbers and cell length. Fifth, ectopic expression of JcARF19 upregulated auxin responsive genes encoding important regulators involving in cell differentiation and cytoskeletal dynamics.
Woody plants such as Jatropha had longer life cycle than herbaceous ones. It took as long as two years for Jatropha to get the first flower blooming under our current lab condition in Beijing North of China, on contrast of 4–5 months in tropical countries such as Singapore. We tried to plant Jatropha in South of China and so far they do not flower yet. For this reason we used a few independent T0 Jatropha plants and we also performed genetic analyses on laboratory model plant Arabidopsis. The increased seeds yield of T3 JcARF19 ectopic Arabidopsis was also consist with the results of overexpression in Jatropha, confirming our claims of improved seed traits with JcARF19 ectopic expression strategy. Furthermore, we found that the germination percentage of T1 JcARF19-overexpression Jatropha seeds was obviously higher than that of the wild type (Fig. S4). Early research had reported that the auxin has close relationship with the seeds germination rate54. It indicated that the traits we observed at T0 generation can be inherited into the next generation and our improved seed traits by JcARF19 ectopic expression are reliable as well55. Nevertheless, the observed impact on plant architecture, seed size and yield by manipulation of ARF19 need not only more observations on T1 and T2 generations plants under greenhouse condition, more researches under field condition are also necessary for the feasibility of a big scale commercialization of this strategy. Auxin is a multifunctional hormone that regulates pattern formation in plants56. The location and timing of auxin accumulation and signal transduction play critical roles in various aspects of plant development57,58. In the future, to avoid growth abnormalities in auxin signaling pathway transgenic plants, it is advisable to use a weaker or an organ and developmental specific promoter rather than a stronger promoter like CaMV 35S promoter because the amount of the hormone produced by the transgene and the response should be confined to the target tissue at an appropriate level as did in cotton fiber cell and other success reports59.
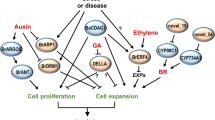
We present a common sharing genetic framework for the control of cell division, differentiation and size for various plant organs, e.g. seed and root. Given the early working model of ARF19 in auxin signaling transduction53,55,60, the seed size controlling results in this study can be best integrated as the working model presented in Fig. S5. In this seed size controlling model, auxin activates the transcription of JcARF19 via RETINOBLASTOMA-RELATED (RBR) protein and cytokinin-dependent transcription factor ARABIDOPSIS RESPONSE REGULATOR12 (ARR12)60. JcARF19 is involved in promoting cell differentiation and thus cell number increasing in early stage of seed development by regulating the transcription of LBD18 and LBD2955. ARGOS gene family is auxin-induced and involved in the regulation of cell number for the duration of organ growth45. Ectopic expression of ARGOS prolongs the expression of AINTEGUMENTA (ANT) and cell cycle regulator CycD3; as well as the neoplastic activity of leaf cells45. Overexpression of ARGOS genes modifies plant sensitivity to Ethylene, leading to improved drought tolerance in both Arabidopsis and maize46. The auxin-(Transmembrane Kinase) TMK sensing and auxin-Rho of Plants (ROP) signaling networks have been demonstrated to control auxin signaling pathway47,48. JcARF19 might be also involved in enlarged cell size by TMK Auxin-Sensing and ROP GTPase signaling complex in middle stage of seed development. Considering that orthologs of JcARF19 exist in many other plant species, including castor bean, alfalfa (Medicago sativa), soybean and apple, the manipulation of ARF19 may provide a broad application to increase plant biomass and seed productivity in many other species.This study provides evidence that an auxin signaling integrator ARF19 plays vital roles in determining seed size. ARF19 is conserved in higher plants and involved in auxin pathway signal transduction55. Nevertheless, it is necessary to test the ARF19 ectopic expression Jatropha lines under field conditions to get conclusive statements of its commercial viability. Meanwhile besides of plant genome, plant rhizospheric or leaf-residing microbiomes via plant endogenous auxin signalling pathway have been successfully to improve crop yield dramatically including Jatropha61,62. ARF19 transcription factors mediated auxin pathway is essential for growth and yield promoting effect by beneficial microbes63. Manipulation of the auxin signaling pathway can result in larger seed sizes and improved seed yield in J. curcas, this ARF19-ectopic expression plant may become a more attractive commercial plant.
Materials and Methods
Plant materials and growth condition
Three species of plants were used in this study. For Jatropha, seeds were obtained from the Jatropha curcas (Jc-MD) elite plants which were pre-selected by Drs. Yan Hong and Chenxin Yi64. The seeds were germinated on ½ Murashige and Skoog salt medium at 25 °C under a 16 h light/8 h dark photoperiod with a light intensity of 100 μmol·m−2 s−1. When two or three true leaves were grown64, the seedlings were transplanted into pots filled with soil and grown at 25 °C in greenhouse under natural light condition. Plant management, including pesticide spraying, watering and artificial fertilization, was carried out according to normal practice31. For Arabidopsis, seeds of ecotype Col-0 were vernalized on ½ Murashige and Skoog salt medium at 4 °C in darkness for about three days and transferred into a growth chamber at 22 °C under16 h dark/8 h light photoperiod. The seedling with four true leaves were transplanted into soil and grown in the same condition. Arabidopsis transformation was performed according to the floral-dipping method previously described65. For Nicotiana benthamiana, seeds were sown in soil to germinate. The seedlings were transplanted in separated pots and grown in a greenhouse at 25 °C with 14 h dark/10 h light photoperiod.
RNA extraction and analysis
RNA was isolated and analyzed according to previously described methods23. 100 mg samples from different organs were harvested and extracted with plant RNA purification reagent (Invitrogen, Carlsbad, CA, USA). Nanodrop (Thermo Scientific, Wilmington, DE, USA) was used to quantify RNA concentration. M-MLV reverse transcriptase (Promega, Madison, USA) was used for reverse transcription. For quantitative PCR analysis, THUNDERBIRD SYBR qPCR Mix (TOYOBO) was used and run in Bio-Rad CFX96 qPCR machine. Each treatment was repeated with three biological replicates, and with three technical replicates for each biological sample. The Jatropha UBQ transcript was served as an internal control for RNA samples. The primers for target genes are listed in Table S1. Standard deviation was calculated based on the three biological replicates.
Virus induced gene silencing
We used the sTRV method described by Ye et al.40,41, using psTRV1 and psTRV2. PCR-based cloning was used to clone partial cDNAs of JcARF19 to psTRV2 to generate psTRV2 derivatives. Another psTRV2 clone with insertion of Jatropha Chlorata 42 (JcCH42) was served as a positive control34. psTRV1, psTRV2 and psTRV2 derivatives were electroporated into Agrobacterium strain AGL1. Vacuum agroinfiltration was used to inoculate those Agrobacterium into Jatropha seedling with two or three true leaves. At least 5 Jatropha seedlings were agroinfiltrated with psTRV1 and psTRV2-JcARF19, psTRV2-JcCH42 or psTRV2 vector only accordingly. After infiltration, plants were grown in a growth chamber at 25 °C with a 16 h light ⁄8 h dark photoperiod40. Phenotypes of Jatropha plants at 27 days post-infiltration (dpi) with various sTRV constructs were recorded and leaves in same leaf position were picked and leaf width were measured. Values (n = 5) were shown as mean ± SD and statistic analysis with Student T-test. **Indicates P < 0.01, *indicates P < 0.05.
IAA sensitivity test
Detached silenced systemic Jatropha leaves were kept in liquid MS medium containing 10 nM IAA following treated with vacuum infiltration. Treated leaves were incubated at 25 °C in an incubator for 6 hrs. After the IAA treatment, the leaves were subjected to RNA isolation and following quantitative real-time PCR analysis.
Transgenic plant plasmid construction
JcARF19 gene was identified from a database of sequenced cDNA library prepared from Jatropha seeds (detailed sequence information could be found in Supplementary file). The full-length cDNA fragment of JcARF19 was PCR-amplified with primers (Table S1). The PCR fragment was inserted in the sense orientation into suitable sites of pCABMIA1300-3HA vector34.
Scanning electronic microscopy (SEM) and light microscopy
For observation of Arabidopsis seeds with the scanning electron microscope (SEM), collected seeds from WT Col-0 and 35S:JcARF19 overexpression plants respectively were fixed with a tape inside a sample chamber, following freezing in liquid N2. Images were collected using a SEM (JSM-6360LV, JEOL, USA) with an acceleration voltage of 20 kV. For observation of Jatropha seed endosperm with light microscopy, endosperm discs from WT Jc-MD and 35S:JcARF19 overexpression plants respectively were excised from mature Jatropha endosperm (7 WAF, weeks after fertilization) and fixed overnight in 2.5% glutaraldehyde in 0.1 M phosphate buffer, pH 7.2 as described previously40. Endosperm discs were rinsed three times in 0.1 M phosphate buffer for 15 min each, and were then post-fixed in 1% (w/v) aqueous OsO4 for 1 h. Tissues were dehydrated in an ethanol series and embedded in Spurr’s resin. Semi-thin sections with thickness of 500 nm were stained in 0.1% toluidine blue and photographed with a Zeiss Axioplan 2 microscope (Carl Zeiss, Germany). Cell size and cell number per disc were analyzed with ImageJ and calculated, followed with statistic analysis with Student T-test. **Indicates P < 0.01. Values are mean ± SEM (n = 10).
Arabidopsis seed size and weight measurement
Mature seeds were harvested from WT Col-0 and 35S:JcARF19 overexpression plants grown under the same conditions. 100 seeds from ten independent transgenic lines and WT Col-0 were weighted and recorded with three technical replicates. Values (n = 10) are given as mean ± SD. A DM5000B microscope (Leica) and ImageJ analysis software were used to measure seed sizes. Values (n = 10) are given as mean ± SD. Statistic analysis with Student T-test. **Indicates P < 0.01.
Fatty acid analysis
Total lipid was extracted and transmethylated from 100 dry Arabidopsis seeds as described previously66. The resulting FAMEs were separated and detected by GCMS-QP2010 Ultra (Shimadzu, Kyoto, Japan). The GC analysis was performed under conditions described before31. The data were presented based on three biological replicates and each biological replicate had three technical replicates. Values (n = 3) are given as means ± standard deviation.
Explant material preparation and Jatropha transformation method
Cotyledons were harvested from WT plants Jc-MD sterilized seedlings that were 7–9 days old and were cut into small pieces (5 mm × 5 mm) used as explants. After co-cultivation, shoot regeneration, shoot elongation and rooting, we got the JcARF19 overexpression line. Detailed protocol can be found in Qu et al.31.
JcARF19-overexpressing Jatropha agronomic traits measurement and statistical analysis
Wild-type Jc-MD and JcARF19 transgenic overexpression Jatropha plants were grown in the same condition. Flowering time was scored by the number of days from transplantation to soil to the day of first inflorescence emergence. 10 independent T0 JcARF19 overexpression plants and WT plant Jc-MD should be calculated and values are mean ± SD (n = 10). Branch number of each plants after 1 year of transplanting was recorded for either Jc-MD (n = 10) or T0 JcARF19 overexpression plants (n = 10). Values are mean ± SEM (n = 10). Single seed weight and seed length for each seeds were measured for three of WT plants Jc-MD and T1 JcARF19 overexpression seeds of three lines, JcARF19OE #1, #10 and #13. Values are mean ± SEM (n = 50). Germination percentage were measured for five of WT plants Jc-MD and T1 JcARF19 overexpression seeds of JcARF19 OE #10 and #13. Values are mean ± SEM (n = 5). Student T-test was used for statistical analyses for all agronomic traits. **Indicates P < 0.01, *indicates P < 0.05.
In vitro GST pull-down assay
The C-terminal sequence of JcIAA9 and JcARF19 were amplified by PCR using Phusion High-Fidelity DNA Polymerase (Thermo-Fisher, Finnzymes, Espoo, Finnland) and subcloned into pGEX6P-1 or pET28-SUMO vectors to generate GST fusion or 6*His fusion constructs. Point mutations were performed to generate vector of JiARF19 fusion with 6*His tag by QuikChange Site-Directed Mutagenesis Kit (Stratagene, Agilent, Wilmington, DE, USA). In vitro pull-down assays were performed with 2 μg of GST fusion proteins and 2 μg of His-tagged proteins. GST fusion proteins were incubated in a binding buffer (50 mM Tris-HCl at pH 7.5, 100 mM NaCl, 0.25% Triton X-100, 35 mM β-mercaptoethanol) with 25 μL of glutathione sepharose 4B (GE Healthcare, Uppsala, Sweden) for 3 h at 4 °C and GST beads were washed six times with binding buffer. His-tagged JcARF19-CT and JiARF19-CT proteins were added into GST beads and the mixture was incubated overnight at 4 °C. After washing again with binding buffer six times, pulled-down proteins were separated on 12% SDS–polyacrylamide gel and detected by Western blotting using anti-His or anti-GST antibody as previously described67,68,69.
Bimolecular fluorescence complementation (BiFC)
BiFC was carried using previously described vectors and methods67,68. The C-terminal sequence of JcARF19 and JcIAA9 were cloned in corresponding restrict enzyme sites of BiFC vectors. Point mutations were performed to generate JiARF19 by QuikChange Site-Directed Mutagenesis Kit (Stratagene, Agilent, Wilmington, DE, USA). The resulting cassettes including fusion proteins and constitutive promoters were cloned into pGreen binary vector HY105 and transformed into Agrobacterium. For BiFC experiments, 3-week-old Nicotiana benthamiana leaves were co-infiltrated with Agrobacterium as previously described. Two days after incubation, fluorescence and DAPI staining were analyzed by confocal microscopy68,70. The confocal laser scanning microscope technique we used was referred to the Leica SP8 microscope instruction.
Additional Information
How to cite this article: Sun, Y. et al. Manipulation of Auxin Response Factor 19 affects seed size in the woody perennial Jatropha curcas. Sci. Rep. 7, 40844; doi: 10.1038/srep40844 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Zuo, J. & Li, J. Molecular genetic dissection of quantitative trait loci regulating rice grain size. Annual review of genetics 48, 99–118 (2014).
Marriott, P. E., Gómez, L. D. & McQueen‐Mason, S. J. Unlocking the potential of lignocellulosic biomass through plant science. New Phytologist 209, 1366–1381 (2016).
Kazamia, E. & Smith, A. G. Assessing the environmental sustainability of biofuels. Trends in plant science 19, 615–618 (2014).
Bartrina, I., Otto, E., Strnad, M., Werner, T. & Schmülling, T. Cytokinin regulates the activity of reproductive meristems, flower organ size, ovule formation, and thus seed yield in Arabidopsis thaliana. The Plant cell 23, 69–80 (2011).
Li, J., Nie, X., Tan, J. L. H. & Berger, F. Integration of epigenetic and genetic controls of seed size by cytokinin in Arabidopsis. Proceedings of the National Academy of Sciences 110, 15479–15484 (2013).
Wang, A. et al. The VQ motif protein IKU1 regulates endosperm growth and seed size in Arabidopsis. The Plant Journal 63, 670–679 (2010).
Li, J. & Berger, F. Endosperm: food for humankind and fodder for scientific discoveries. New Phytologist 195, 290–305 (2012).
Lu, J., Zhang, C., Baulcombe, D. C. & Chen, Z. J. Maternal siRNAs as regulators of parental genome imbalance and gene expression in endosperm of Arabidopsis seeds. Proceedings of the National Academy of Sciences 109, 5529–5534 (2012).
Garcia, D. & Berger, F. Arabidopsis haiku mutants reveal new controls of seed size by endosperm. Plant Physiology 131, 1661–1670 (2003).
Ruan, Y.-L., Patrick, J. W., Bouzayen, M., Osorio, S. & Fernie, A. R. Molecular regulation of seed and fruit set. Trends in plant science 17, 656–665 (2012).
Ye, J. et al. Identification of candidate genes JcARF19 and JcIAA9 associated with seed size traits in Jatropha. Functional & integrative genomics 14, 757–766 (2014).
Durbak, A., Yao, H. & McSteen, P. Hormone signaling in plant development. Current opinion in plant biology 15, 92–96 (2012).
Tan, X. et al. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446, 640–645 (2007).
Wang, R. & Estelle, M. Diversity and specificity: auxin perception and signaling through the TIR1/AFB pathway. Current opinion in plant biology 21, 51–58 (2014).
Guilfoyle, T. J. & Hagen, G. Getting a grasp on domain III/IV responsible for Auxin Response Factor–IAA protein interactions. Plant Science 190, 82–88 (2012).
Wang, R. & Estelle, M. Diversity and specificity: auxin perception and signaling through the TIR1/AFB pathway. Current opinion in plant biology 21C, 51–58 (2014).
Korasick, D. A. et al. Molecular basis for AUXIN RESPONSE FACTOR protein interaction and the control of auxin response repression. Proceedings of the National Academy of Sciences of the United States of America 111, 5427–5432 (2014).
Guilfoyle, T. J. & Hagen, G. Auxin Response Factors. Journal of Plant Growth Regulation 10, 453–460 (2001).
Kang, N. Y., Lee, H. W. & Kim, J. The AP2/EREBP gene PUCHI co-acts with LBD16/ASL18 and LBD18/ASL20 downstream of ARF7 and ARF19 to regulate lateral root development in Arabidopsis. Plant and cell physiology 54, 1326–1334 (2013).
Wang, L. et al. Auxin Response Factor2 (ARF2) and its regulated homeodomain gene HB33 mediate abscisic acid response in Arabidopsis. PLoS Genet 7, e1002172 (2011).
Schruff, M. C. et al. The AUXIN RESPONSE FACTOR 2 gene of Arabidopsis links auxin signalling, cell division, and the size of seeds and other organs. Development 133, 251–261 (2006).
Okushima, Y., Mitina, I., Quach, H. L. & Theologis, A. AUXIN RESPONSE FACTOR 2 (ARF2): a pleiotropic developmental regulator. The Plant Journal 43, 29–46 (2005).
Ishimaru, K. et al. Loss of function of the IAA-glucose hydrolase gene TGW6 enhances rice grain weight and increases yield. Nature genetics 45, 707–711 (2013).
Liu, L. et al. Activation of Big Grain1 significantly improves grain size by regulating auxin transport in rice. Proceedings of the National Academy of Sciences 112, 11102–11107 (2015).
Chourey, P. S., Li, Q.-B. & Kumar, D. Sugar–hormone cross-talk in seed development: two redundant pathways of IAA biosynthesis are regulated differentially in the invertase-deficient miniature1 (mn1) seed mutant in maize. Molecular plant 3, 1026–1036 (2010).
Maghuly, F. & Laimer, M. Jatropha curcas, a biofuel crop: Functional genomics for understanding metabolic pathways and genetic improvement. Biotechnology journal 8, 1172–1182 (2013).
Kumar, S., Singh, J., Nanoti, S. & Garg, M. A comprehensive life cycle assessment (LCA) of Jatropha biodiesel production in India. Bioresource Technology 110, 723–729 (2012).
Liang, S., Xu, M. & Zhang, T. Life cycle assessment of biodiesel production in China. Bioresource Technology 129, 72–77 (2013).
Ajayebi, A., Gnansounou, E. & Raman, J. K. Comparative life cycle assessment of biodiesel from algae and jatropha: A case study of India. Bioresource technology 150, 429–437 (2013).
Gao, S., Qu, J., Chua, N.-H. & Ye, J. A new strain of Indian cassava mosaic virus causes a mosaic disease in the biodiesel crop Jatropha curcas. Archives of virology 155, 607–612 (2010).
Qu, J. et al. Development of marker-free transgenic Jatropha plants with increased levels of seed oleic acid. Biotechnology for biofuels 5, 1 (2012).
Wang, G. et al. DNA-A of a highly pathogenic Indian cassava mosaic virus isolated from Jatropha curcas causes symptoms in Nicotiana benthamiana. Virus genes 48, 402–405 (2014).
Ye, J. et al. Engineering geminivirus resistance in Jatropha curcus. Biotechnology for biofuels 7, 1 (2014).
Ye, J. et al. The Jatropha FT ortholog is a systemic signal regulating growth and flowering time. Biotechnology for biofuels 7, 1 (2014).
Misra, A., Khan, K., Niranjan, A., Nath, P. & Sane, V. A. Over-expression of JcDGAT1 from Jatropha curcas increases seed oil levels and alters oil quality in transgenic Arabidopsis thaliana. Phytochemistry 96, 37–45 (2013).
Maghuly, F. & Laimer, M. Jatropha curcas, a biofuel crop: Functional genomics for understanding metabolic pathways and genetic improvement. Biotechnology journal 8, 1172–1182 (2013).
Kochhar, S., Singh, S. P. & Kochhar, V. K. Effect of auxins and associated biochemical changes during clonal propagation of the biofuel plant— Jatropha curcas. Biomass & Bioenergy 32, 1136–1143 (2008).
Wang, C. M. et al. Isolation and Identification of miRNAs in Jatropha curcas. Int J Biol Sci 8, 418–429 (2012).
Sun, F. et al. An approach for jatropha improvement using pleiotropic QTLs regulating plant growth and seed yield. Biotechnology for biofuels 5, 1 (2012).
Ye, J., Qu, J., Bui, H. T. N. & Chua, N. H. Rapid analysis of Jatropha curcas gene functions by virus-induced gene silencing. Plant biotechnology journal 7, 964–976 (2009).
Qu, J. et al. Dissecting functions of KATANIN and WRINKLED1 in cotton fiber development by virus-induced gene silencing. Plant physiology 160, 738–748 (2012).
Koncz, C. et al. Isolation of a gene encoding a novel chloroplast protein by T-DNA tagging in Arabidopsis thaliana. The EMBO journal 9, 1337 (1990).
Cho, H. et al. A secreted peptide acts on BIN2-mediated phosphorylation of ARFs to potentiate auxin response during lateral root development. Nat Cell Biol 16, 66–76, doi: 10.1038/ncb2893 (2014).
Zhang, X., Henriques, R., Lin, S.-S., Niu, Q.-W. & Chua, N.-H. Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nature protocols 1, 641–646 (2006).
Hu, Y., Xie, Q. & Chua, N.-H. The Arabidopsis auxin-inducible gene ARGOS controls lateral organ size. The Plant Cell 15, 1951–1961 (2003).
Shi, J., Drummond, B., Wang, H., Archibald, R. L. & Habben, J. E. Maize and Arabidopsis ARGOS proteins interact with ethylene receptor signaling complex, supporting a regulatory role for ARGOS in ethylene signal transduction. Plant physiology, pp. 00347.02016 (2016).
Lin, D., Ren, H. & Fu, Y. ROP GTPase‐mediated auxin signaling regulates pavement cell interdigitation in Arabidopsis thaliana. Journal of integrative plant biology 57, 31–39 (2015).
Korasick, D. A., Jez, J. M. & Strader, L. C. Refining the nuclear auxin response pathway through structural biology. Current opinion in plant biology 27, 22–28 (2015).
Würschum, T., Groß-Hardt, R. & Laux, T. APETALA2 regulates the stem cell niche in the Arabidopsis shoot meristem. The Plant cell 18, 295–307 (2006).
Ohto, M.-a., Fischer, R. L., Goldberg, R. B., Nakamura, K. & Harada, J. J. Control of seed mass by APETALA2. Proceedings of the National Academy of Sciences of the United States of America 102, 3123–3128 (2005).
Jofuku, K. D., Omidyar, P. K., Gee, Z. & Okamuro, J. K. Control of seed mass and seed yield by the floral homeotic gene APETALA2. Proceedings of the National Academy of Sciences of the United States of America 102, 3117–3122 (2005).
Liu, J. et al. CURLY LEAF Regulates Gene Sets Coordinating Seed Size and Lipid Biosynthesis. Plant physiology 171, 424–436 (2016).
Ioio, R. D. et al. A genetic framework for the control of cell division and differentiation in the root meristem. Science 322, 1380–1384 (2008).
de Brito, C. D. et al. Osmoconditioning prevents the onset of microtubular cytoskeleton and activation of cell cycle and is detrimental for germination of Jatropha curcas L. seeds. Plant Biology 18, 1053–1057, doi: 10.1111/plb.12482 (2016).
Okushima, Y., Fukaki, H., Onoda, M., Theologis, A. & Tasaka, M. ARF7 and ARF19 regulate lateral root formation via direct activation of LBD/ASL genes in Arabidopsis. The Plant cell 19, 118–130 (2007).
Xu, T. et al. Cell surface- and rho GTPase-based auxin signaling controls cellular interdigitation in Arabidopsis. Cell 143, 99–110 (2010).
Dubrovsky, J. G. et al. Auxin acts as a local morphogenetic trigger to specify lateral root founder cells. Proceedings of the National Academy of Sciences of the United States of America 105, 8790–8794 (2008).
Vanneste, S. & Friml, J. Auxin: a trigger for change in plant development. Cell 136, 1005–1016 (2009).
Zhang, M. et al. Spatiotemporal manipulation of auxin biosynthesis in cotton ovule epidermal cells enhances fiber yield and quality. Nature biotechnology 29, 453–458 (2011).
Schaller, G. E., Bishopp, A. & Kieber, J. J. The yin-yang of hormones: cytokinin and auxin interactions in plant development. The Plant cell 27, 44–63 (2015).
Madhaiyan, M., Alex, T. H. H., Te Ngoh, S., Prithiviraj, B. & Ji, L. Leaf-residing Methylobacterium species fix nitrogen and promote biomass and seed production in Jatropha curcas. Biotechnology for biofuels 8, 1 (2015).
Madhaiyan, M. et al. Improvement of plant growth and seed yield in Jatropha curcas by a novel nitrogen-fixing root associated Enterobacter species. Biotechnology for biofuels 6, 1 (2013).
Garnica-Vergara, A. et al. The volatile 6-pentyl-2H-pyran-2-one from Trichoderma atroviride regulates Arabidopsis thaliana root morphogenesis via auxin signaling and ETHYLENE INSENSITIVE 2 functioning. New Phytologist 209, 1496–1512 (2016).
Yi, C. Z. S., Liu, X. et al. Does epigenetic polymorphism contribute to phenotypic variances in Jatropha curcas L.? BMC Plant Biology 10, 1 (2010).
Zhang, X., Henriques, R., Lin, S.-S., Niu, Q.-W. & Chua, N.-H. Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nature protocols 1, 641–646 (2006).
Kim, M. J. et al. Gene silencing of Sugar-dependent 1 (JcSDP1), encoding a patatin-domain triacylglycerol lipase, enhances seed oil accumulation in Jatropha curcas. Biotechnology for biofuels 7, 1 (2014).
Ye, J. et al. Geminivirus Activates ASYMMETRIC LEAVES 2 to Accelerate Cytoplasmic DCP2-Mediated mRNA Turnover and Weakens RNA Silencing in Arabidopsis. PLoS Pathog 11, e1005196 (2015).
Li, R. et al. Virulence factors of geminivirus interact with MYC2 to subvert plant resistance and promote vector performance. The Plant cell 26, 4991–5008 (2014).
Ye, J., Qu, J., Zhang, J.-F., Geng, Y.-F. & Fang, R.-X. A critical domain of the Cucumber mosaic virus 2b protein for RNA silencing suppressor activity. FEBS letters 583, 101–106 (2009).
Liu, C., Xi, W., Shen, L., Tan, C. & Yu, H. Regulation of floral patterning by flowering time genes. Developmental cell 16, 711–722 (2009).
Acknowledgements
We would like to thank Professor Nam-Hai Chua, Rockefeller University, USA and Drs Yan Hong, Chengxin Yi (JOil) for their kind help during the whole research process and Mr Khar Meng Ng and Mr. Xuezhi Ouyang (Temasek Life Sciences Laboratory, Singapore) for providing materials and valuable technical assistance on this work. This work was supported by the State Key Laboratory of Plant Genomics, Institute of Microbiology, Chinese Academy of Sciences, Temasek Life Sciences Laboratory, JOil and the Singapore Millennium Foundation.
Author information
Authors and Affiliations
Contributions
J.Y. and C.M.W. conceived of the study, and design the experiments and draft the manuscript. C.M.W. carried out the gene expression analysis and analysis of the study. Z.F.X. participated in data interpretation and revised the manuscript. Y.W.S. helped to design the experiments, analyzed the data, drafted the manuscript and performed molecular cloning and analysis, genotyping. N.W. participated in the design of the study, performed molecular analysis and statistical analysis and drafted the manuscript. X.Y.J. performed chemical analysis, field traits collection and analysis. X.H.W. and Z.J.L. participated in the molecular analysis. H.Z.M. did the Jatropha transformation. G.H.Y. provided research materials. C.X.Z. provided important reagents. F.J.W. provided important reagents and help to revise the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Sun, Y., Wang, C., Wang, N. et al. Manipulation of Auxin Response Factor 19 affects seed size in the woody perennial Jatropha curcas. Sci Rep 7, 40844 (2017). https://doi.org/10.1038/srep40844
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep40844
This article is cited by
-
JcSEUSS1 negatively regulates reproductive organ development in perennial woody Jatropha curcas
Planta (2023)
-
Options for the generation of seedless cherry, the ultimate snacking product
Planta (2022)
-
Improving seed size, seed weight and seedling emergence in Camelina sativa by overexpressing the Atsob3-6 gene variant
Transgenic Research (2020)
-
geneHummus: an R package to define gene families and their expression in legumes and beyond
BMC Genomics (2019)
-
Impact of farming awareness and delay on the dynamics of mosaic disease in Jatropha curcas plantations
Computational and Applied Mathematics (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.