Abstract
Accumulated evidence indicates that the core genetic mechanisms regulating early patterning of the brain rudiment in vertebrates are very similar to those operating during development of the anterior region of invertebrate embryos. However, the mechanisms underlying the morphological differences between the elaborate vertebrate brain and its simpler invertebrate counterpart remain poorly understood. Recently, we hypothesized that the emergence of the most anterior unit of the vertebrate brain, the telencephalon, could be related to the appearance in vertebrates’ ancestors of a unique homeobox gene, Anf/Hesx1(further Anf), which is absent from all invertebrates and regulates the earliest steps of telencephalon development in vertebrates. However, the failure of Anf to be detected in one of the most basal extant vertebrate species, the lamprey, seriously compromises this hypothesis. Here, we report the cloning of Anf in three lamprey species and demonstrate that this gene is indeed expressed in embryos in the same pattern as in other vertebrates and executes the same functions by inhibiting the expression of the anterior general regulator Otx2 in favour of the telencephalic regulator FoxG1. These results are consistent with the hypothesis that the Anf homeobox gene may have been important in the evolution of the telencephalon.
Similar content being viewed by others
Introduction
One of the most important innovations of vertebrates, distinguishing them from other animals, is their complex brain, derived from three main embryonic units: the forebrain, the midbrain and the hindbrain1,2. It has recently been shown that the core genetic mechanisms regulating the early patterning of the brain rudiment in vertebrates are very similar to those operating during the development of the anterior region of invertebrate embryos1,3,4,5,6. However, the mechanisms underlying the obvious differences between the vertebrate brain and its invertebrate homologues are still poorly understood.
In 1992, we identified a previously unknown homeobox gene, named Xanf, based on its expression in the anterior fold of the neural plate of Xenopus laevis embryos, and we then described Xanf orthologues in other species of vertebrates, including humans7,8,9,10,11. Thus, a novel monogenic class of homeoboxes, designated Anf, was described9. Moreover, an Anf orthologue was identified in mice by two other research groups and was designated Hesx1 or Rpx, after its expression in mouse ES cells and in the Rathke pouch12,13,14.
Although Anf genes have been discovered in members of most classes of vertebrates, no orthologues of these genes have been found in invertebrates, even in the closest relatives of vertebrates, the invertebrate chordates. Importantly, this distribution correlates with the presence of the telencephalon in vertebrates, which is a unique region of the forebrain that is apparently absent in all other animals and is derived from the anterior neural fold, where Anf is expressed. Accordingly, gain- and loss-of-function experiments performed in Xenopus and mouse models have confirmed an essential role of Anf homeodomain proteins in the development of the telencephalon10,11,13. Thus, in mouse Anf/Hesx1−/− mutants telencephalic vesicles and eyes are reduced or absent at early somite stages15. Furthermore, using a Xenopus model, we have demonstrated that Anf acts as a transcriptional repressor, and its main function is “cleaning” the prospective rostral forebrain territory of Otx2 homeobox expression, which normally regulates the development of more posterior brain regions11. As a result, genes responsible for telencephalon development, such as FoxG1, can become activated within this territory that has been “cleaned” of Otx2.
On the basis of all of these data, we hypothesized that the appearance of the Anf homeobox at the very beginning of vertebrate evolution could be one of the critical events that provided appropriate conditions for the appearance of the telencephalon11. However, the failure to detect Anf hitherto in the most basal group of extant vertebrates, the cyclostomes, including lampreys and hagfishes, seriously compromised this hypothesis; it began to seem even more questionable after the recent publication of the complete lamprey Petromyzon marinus genome, in which no Anf orthologues were revealed16. Nevertheless, to finally test our hypothesis, we have now made one more attempt to identify this gene in lamprey. As a result, we cloned Anf in this animal and demonstrated that this homeobox gene is indeed expressed in the same pattern as in other vertebrates and executes the same functions (by) inhibiting the expression of Otx2 and promoting the expression of the telencephalic marker gene, FoxG1. These results indicate that Anf likely emerged at the beginning of vertebrate evolution and may have been essential for the evolution of the telencephalon.
Results
Cloning of Anf in lampreys
To clone a possible Anf orthologue in the lamprey, we used a previously elaborated procedure for the mechanical enrichment of putative Anf transcripts9. To this end, we extracted total RNA from the head protrusions of L. camtschaticum and L. fluviatilis, cut at stage 20–2117, and prepared a PCR cDNA library based on this RNA (see Materials and Methods for details). As Anf is only expressed in the anterior neural fold in all gnathostomes studied to date, we employed this method because of its enrichment of Anf transcripts compared with RNA isolated from whole embryos. This approach also helped us to reduce the concentrations of undesirable transcripts of other homeobox genes, which are not expressed within the anterior neural fold.
In addition, given the unique content of the lamprey genome, which is extremely enriched with G and C nucleotides, we used special PCR buffer and Encyclo polymerase (a gift of the Evrogen company), which permits the effective amplification of such sequences.
Subsequent RT-PCR performed based on this RNA sample with a set of Anf-specific degenerate oligos (see Materials and Methods), followed by cloning of the obtained fragments and their sequencing, yielded three clones from L. camtschaticum and two clones from L. fluviatilis containing the same insert, which showed closer homology to Anf sequences in other species than to any of the other known homeoboxes of lampreys. The remaining 5′ and 3′ fragments of this sequence were obtained via the suppression PCR-based Step-Out RACE technique18, which finally allowed us to establish the full coding sequences of the L. camtschaticum and L. fluviatilis Anf cDNAs and to clone them via RT-PCR with corresponding flanking primers. Importantly, with these cDNA sequences in hand, we were able to identify fragments of the genomic sequence of Anf among the L. camtschaticum genomic sequences deposited in GenBank. As a result, we established that similar to all known Anfs of other species, lamprey Anf is disrupted by three introns, two of which are within the homeobox (Fig. 1a, arrows).
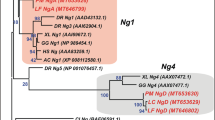
Sequences of lamprey Anfs.
(a) Alignment of Lanfs from L. camtschaticum, L. fluviatilis and P. marinus (GenBank: KX245018, KX245019, KX245020). Homeodomains and the engrailed repressor domain (EnR) are indicated in red and blue frames, respectively; histidine-rich blocks are underlined with a solid black line; conservative motifs flanking the homeodomain are underlined with a dotted line; arrows indicate the positions of introns; the positions of the degenerate primers used for RT-PCR during Lanf cloning are indicated with solid lines. (b) Alignment of the EnR domains and homeodomains of L. camtschaticum Lanf with those of Anfs from different classes of vertebrates. (c) The percentages of similarity (S) and identity (I) between Anfs and some of the homeodomains of other types. The mean difference between the similarity (S) and identity (I) of the Lanf homeodomain with homeodomains of other Anfs (bottom line, highlighted by red) is much greater than the analogous values calculated for Lanf and other types of homeodomains (bottom line, in black). This fact indicates conservation of Lanf function during evolution. (d) Scheme of the neighbour-joining phylogenetic tree of Anf homeodomains and homeodomains of Antp- and Prd-class proteins (full trees constructed via the neighbour-joining and maximum likelihood methods are shown in Fig. S2, Fig. S3 and Fig. S4).
To clone Anf from P. marinus, we used Petromyzon cDNA from the late neurula stage and flanking primers designed based on non-translated regions of the L. camtschaticum Anf mRNA.
The proteins encoded by the identified cDNAs exhibit all of the features characteristic of the Anf class (Fig. 1a). First, their homeodomains show a higher percentage of similarity with the homeodomains of Anfs than with any other type of homeodomain (Fig. 1b). Additionally, the Lanf homeodomain is flanked on both sides by specific short amino acid motifs that are peculiar to Anf-class proteins (Fig. 1a,b). Furthermore, like all other known Anfs, the Lanfs exhibit an engrailed-type repressor domain (EnR) in proximity to the N-terminus (Fig. 1a,b).
In addition, in contrast to the percentage of amino acid identity, the percentage of amino acid similarity between Lanf and homeodomains of the Anf class is much higher than between Lanf and other types of homeodomains, indicating that Lanf belongs to the Anf class. (Fig. 1c). Importantly, the phylogenetic relationships between lamprey Anfs correspond well to the known phylogenetic relationships between these species, with P. marinus being a more distant species from the two closely related species L. camtschaticum and L. fluviatilis (Fig. S2)19.
Finally, a close relationship of Lanf with Anfs from other classes of vertebrates was confirmed by their clustering in phylogenetic trees (Fig. 1d, Fig. S2, Fig. S3, Fig. S4).
In summary, it can be concluded that the identified homeoboxes are indeed lamprey Anfs.
Analysis of the phylogenetic relationships of the Anf homeodomain indicates its possible hybrid origin from Antp and Prd classes
As can be observed from Fig. 1d, analysis of the phylogenetic relationships of Anf homeodomains in different classes of vertebrates, including the most basal species, the lampreys, with other known types of homeodomains revealed an intermediate position of Anf between the homeodomains of two large classes, Antp and Prd. This result led us to analyse the homology of different regions of the Anf homeodomain with the corresponding regions of the Antp and Prd homeodomains.
We found that the most of the Anf homeodomain, from its N-terminus (position 1) to the end of the protein sequence encoded by the 3rd exon (position 46), clustered with the homologous region of the Prd class of homeodomains via the neighbour-joining method (Fig. S1, Fig. S5). Moreover, this finding held true if only an N-fragment of this region (including the 1st alpha-helix) or its central fragment (including the 2nd helix) was used for clustering (not shown). In contrast, the C-terminal region of the Anf homeodomains from position 47 to 60 plus 4 amino acids of the conservative Anf homeodomain-flanking ESQ motif (positions 61–64), encoded by the last (4th) exon, were confidently grouped with Antp-class sequences (Fig. S1, Fig. S6). Thus, at least formally, the Anf homeodomain appears to be a hybrid of Prd- and Antp-class homeodomains.
It is also important to note that among all of the known invertebrate homeodomains, those showing the highest homology to Anf are the homeodomains of the Micro and Pmar proteins of sea urchins (Fig. 1d, Fig. S2, Fig. S3, Fig. S4). However, these proteins lack traits that are characteristic of Anfs, such as specific conservative sequences flanking the homeodomain, the EnR domain near the N-terminus and the location of the homeodomain near the C-terminus. Moreover, the exon-intron structure of their genes is different from that of Anfs. Therefore, one may conclude that Anfs appear to have no orthologues, at least in the genomes of modern invertebrates.
Expression of Lanf in early embryogenesis
The temporal expression pattern of Lanf was investigated via qRT-PCR, and only very low expression of this gene was revealed before gastrulation. At the onset of gastrulation, its expression begins to increase slightly, but before the end of gastrulation returns to its pregastrulation level. Expression increases again, beginning from the late neurula stage (stage 19), reaches a maximum at the pharyngula stage (stage 22) and finally drops down to the background level by the hatching stage. Interestingly, in all other vertebrates, the expression of Anf gradually increases during gastrulation, reaching a maximum by the end of this stage, and then declines to the background level by the onset of the pharyngula stage (i.e., by the stage at which the expression of Lanf reaches its maximum in lamprey)8,9,14. Thus, obvious heterochrony is observed between Lanf expression in lamprey and the expression of its orthologues in other vertebrates.
This heterochrony of Lanf expression is especially evident compared with the expression of another important anterior regulator, the homeobox gene Otx2, which in contrast to Anf, begins its expression at the same stage (i.e., the beginning of gastrulation) in the lamprey and X. laevis (Fig. 2). Interestingly, the telencephalic-specific marker gene FoxG1, whose expression is indirectly regulated by Anf in X. laevis11, begins to increase much later, after the end of neurulation, rather than the late gastrula stage in X. laevis. All of these findings indicate that in the lamprey, the telencephalon specification programme starts much later during embryogenesis than in other vertebrates.
Comparison of the temporal expression pattern of Lanf with those of FoxG1 and Otx2 in lamprey (L. fluviatilis) and frog (X. laevis).
qRT-PCR data obtained for Anfs (Lanf and Xanf1), FoxG1 and Otx2 in whole embryos collected at the stages indicated below were normalized relative to qRT-PCR data obtained for the housekeeping genes EFalfa and ODC. Stage numbers are indicated according to17,34. All data are from three experiments, average values are shown.
The spatial expression of Lanf was studied via whole-mount in situ hybridization. Unfortunately, we were unable to detect expression via this method during gastrulation because of its very low level at this stage. At the same time, Lanf expression was observed beginning from stage 20 within the most anterior part of the brain anlage and in the surface ectoderm covering this brain area. As can be observed in Fig. 3a–c, the expression of Lanf in the neural tissue at this stage was localized just above the expression domain of Shh in the prechordal mesoderm, in the territory corresponding to the forebrain and, thus, to the presumptive telencephalon and rostral diencephalon20. This expression pattern is similar to the patterns of Anf expression in other studied vertebrate embryos9,10,13. As development proceeds, the expression of Lanf progressively ceases in the presumptive di- and telencephalic cells, persisting until stage 23 only in the oral ectoderm and the pituitary placode (Fig. 3d)21.
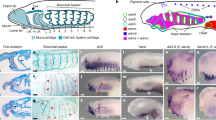
Analysis of the spatial expression pattern of Lanf in sagittal sections of L. fluviatilis embryos.
(a–c) At stage 19, Lanf ((a) - overall view, (b) - enlarged view of anterior region) is expressed in the anterior neurectoderm, in the region corresponding to the stomodeal ectoderm (se) and the forebrain (fb), including the presumptive telencephalon and diencephalon. This region is located just above the region of Shh expression in the notochord (not) and the prechordal plate (pp) (c). An arrow indicates the dorsal limit of Lanf expression. Anterior is to the right; dorsal is at the top. (d) At stage 23, Lanf expression is observed only in the stomodeal ectoderm (se) and the nasohypophysial placode (nhp). (е,f) At the same stage shown for Lanf in (a,b), Otx2 is expressed in a much broader area of the anterior neurectoderm, up to the presumptive mid-hindbrain boundary (MHB), and in the underlying mesoderm. Notably, the most anterior region of the neurectoderm, in which Lanf is expressed, is free of Otx2 expression. (g) At stage 23, Otx2 continues to be expressed in the anterior region, being inhibited in the most rostral part of it. (h) The expression of FoxG1 marks the telencephalon beginning from stage 25 (see Materials and Methods).
As observed in X. laevis embryos, Anf is expressed within a subregion of the broader expression territory of Otx2, whose transcription is suppressed by Anf protein11. Thus, Anf “cleans” the most anterior part of the neuroectoderm of Otx2. As we have demonstrated, this function of Anf is critical for telencephalic development because it permits the telencephalic modulator FoxG1 to be activated within this territory “cleaned” of Otx2.
Based on this assumption, we investigated the early expression patterns of the lamprey orthologues of Otx2 and FoxG1in detail. Similar to Otx2 expression in zebrafish, frog and mouse11,22,23, the expression of Otx2 in lamprey was observed throughout the anterior part of the neural anlage, with a lower level of expression within the most rostral region, from which the telencephalon is derived. Importantly, this region of lower Otx2 expression corresponds to the area of Anf expression, as in other species (Fig. 3e–g).
Unfortunately, we were unable to observe the expression of FoxG1 at the early neurula stage. However, the expression of this gene at later stages indicates that, similar to FoxG1 in other species, lamprey FoxG1 is expressed within the region corresponding to the expression of Lanf (Fig. 3h).
Lanf operates as a transcriptional suppressor
Our data reveal that the expression patterns of Lanf, Otx2 and FoxG1 are similar to those of these genes in other species. This finding indicates that Lanf may play a role similar to that of its orthologues in other species, i.e., permitting the expression of FoxG1 owing to an inhibitory influence upon Otx2 transcription. The presence of an engrailed-type repressor domain in Lanf (Fig. 1a and b) is also in agreement with the possibility that it functions as a transcriptional inhibitor.
To verify whether Lanf inhibits transcription, we tested its influence on the promoter of Xanf1, which is a target of its own protein product and, thus, presents a high probability of being a target of Lanf 24. To this end, we co-injected the Xanf1 promoter-driven luciferase reporter mixed either with Xanf1 or Lanf mRNA into X. laevis embryos at the 4-cell stage and analysed the luciferase signal at the midneurula stage. In both cases, we observed strong inhibition of the reporter compared with the control co-injected with the same reporter with EGFP mRNA (Fig. S7). These experiments confirmed the activity of Lanf as a transcriptional inhibitor.
Lanf inhibits Otx2 expression and promotes the expression of the telencephalon modulator FoxG1
As shown previously, down-regulation of Anf in frog embryos results in anterior expansion of Otx2 domain, accompanied by a reduction of FoxG1 expression, whereas overexpression of Anf elicits opposite effects11,13. To determine whether this result is also true for Lanf, we injected antisense morpholino oligonucleotides targeting Lanf (Lanf MO) or Lanf synthetic mRNA into lamprey embryos. As expected this expanded the Otx2 domain anteriorly, towards the territory, in which its expression was weak in wild-type embryos (Fig. 4a and b). In turn, Lanf mRNA injection frequently reduced Otx2 expression (Fig. 4c and d). In contrast, the opposite effect was observed for FoxG1, which is expressed in cells derived from the Lanf expression domain (Fig. 4e–h). Suppression of Lanf mRNA translation resulted in a reduction of FoxG1 expression (Fig. 4e and f), while injection of Lanf mRNA elicited expansion of the FoxG1 expression area (Fig. 4g and h). The observed expression abnormalities therefore confirmed an inhibitory influence of Lanf upon Otx2 expression and its promotion of the expression of FoxG1.
Lanf is responsible for the suppression of Otx2 in the anterior neural plate and for the induction of FoxG1 expression in this region.
(a–d) In the control (a), the expression of Otx2 is suppressed (arrow) in the anterior neural plate (100%, n = 35), but it is strongly activated (arrow) when Lanf is downregulated with an anti-sense MO targeting Lanf mRNA (70%, n = 50) (b) In contrast, it appears to be more broadly suppressed (arrow) if Lanf mRNA is overexpressed (76%, n = 55) (d) compared with the control EGFP mRNA overexpression (97%, n = 37). Anterior is at the top; dorsal is to the left. (e–h) An effect opposite that caused by Lanf upon Otx2 expression is observed in the case of FoxG1 expression (arrow) (100%, n = 30; 65%, n = 40; 95%, n = 42 and 70%, n = 45, respectively). Anterior is to the right; dorsal is at the top (see Materials and Methods).
Discussion
In this work, we identified an Anf-class homeobox gene in lampreys and confirmed its essential role in lamprey telencephalic development, showing that Anf is indeed present not only in all classes of gnathostomes. but also in cyclostomes (Fig. 5). This finding confirms that the core mechanism responsible for the regionalization of the vertebrate brain arose before the divergence of cyclostomes and gnathostomes25. In turn, the lack of Anf in all invertebrates, including the nearest relatives of vertebrates, tunicates and cephalochordates, which have no anatomical structure homologous to the telencephalon, corroborates our previous hypothesis that the appearance of the Anf homeobox in vertebrate ancestors might be one of the events that provided appropriate conditions for the emergence of the telencephalon.
Interestingly, recent studies have demonstrated that in lancelets, the group of invertebrate chordates whose members exhibit the most similar body morphology to vertebrates, FoxG1 is also expressed at the anterior end of the CNS, i.e., in the brain vesicle, which is thought to be homologous to the vertebrate diencephalon26. At first glance, this result is at odds with the hypothesis that the emergence of Anf in vertebrates was necessary for anterior FoxG1 expression. However, expression of FoxG1 can be observed in this region of the lancelet CNS only on the 3rd day of development, when the brain vesicle already appears to be well formed26. Moreover, FoxG1 is expressed in this region in single scattered cells, whereas no expression of this gene is detected in the brain vesicle within a continuous territory, as observed in the case of FoxG1 expression in the vertebrate telencephalic anlage, including that of the lamprey.
Notably, the expression of FoxG1 and Otx2 in mutually exclusive domains was recently reported in the embryos of Saccoglossus kovalevskii, a member of another invertebrate sister group of vertebrates, the hemichordates27. In embryos of these animals, FoxG1 is expressed in scattered cells of the proboscis, the most anterior part of the Saccoglossus body, whereas Otx2 expression is detected in the more posterior collar region. However, as no Anf was found in Saccoglossus, such mutual exclusion of FoxG1 and Otx2 expression is obviously ensured by other mechanisms.
Additionally, because telencephalic-like structures are not found in the Saccoglossus proboscis6, these results indicate that mutually exclusive expression of FoxG1 and Otx2 at the anterior end of the body is not sufficient by itself to ensure the development of the telencephalon. Importantly, as there is no firm evidence that extant hemichordates are the direct ancestors of vertebrates6, and as they don’t have telencephalon-like structures, this outcome does not contradict our hypothesis that in vertebrates, the mechanism responsible for Otx2 down-regulation in cells of the presumptive telencephalon includes the expression of Anf as one of the necessary conditions.
Similar to other vertebrates, the expression of Anf in lamprey embryos is also observed in the adenohypophysis anlage. As in mouse embryos, the repressive activity of Anf/Hesx1 is required at very early stages for adenohypophysis commitment28. Thus, our detection of Anf expression in the early adenohypophysis anlage is consistent with the previous suggestion that the programme responsible for this organ’s development was already present early in vertebrate evolution, before the divergence of cyclostomes and gnathostomates21,25.
Assuming that Anf is present only in vertebrates, it will be important to understand how this gene might have appeared during the evolution of vertebrates. As we have shown previously, the Anf homeodomain differs from other types of homeodomains in exhibiting an extremely high rate of amino acid substitutions9. Therefore, one possibility regarding the origin of Anf might be rapid evolution from a copy of some other homeobox gene that had undergone duplication in a vertebrate ancestor. Accordingly, early vertebrate evolution is known to have been accompanied by global genome rearrangements, including two rounds of whole-genome duplication29.
Our present analysis shows that the Anf homeodomain appears to be a hybrid of two different homeodomains, belonging to the Prd and Antp classes. Importantly, the N- and C-portions of Anf, which are the most like to the corresponding portions of the Prd and Antp homeodomains, respectively, are encoded by different exons, separated by a 3rd intron in the Anf genes in all species. Moreover, only these portions of the homeobox are separated by an intron in the majority of Prd-class genes and in some Antp-class genes, which is in agreement with the notion of a hybrid origin of the Anf homeodomain. Otherwise, it would be difficult to explain why the alleged connection of different genomic fragments did not result in mistakes in the homeodomain structure owing to a frame shift or the formation of a nonfunctional protein.
In addition, the hybrid hypothesis of the origin of Anf explains some features of its expression pattern and physiological functions. Indeed, if the 5′-region of the genomic sequence of Anf was derived from a Prd-class gene, then it may also have inherited the promoter region of this gene. Additionally, because many genes of the Prd class, such as Gsc, Otx2, Pax6, Pitx, and Rx, are characterized by expression in the anterior region of the embryo, this promoter inherited by Anf from the Prd gene may also govern its anterior expression. For instance, this promoter could come from a copy of some preliminarily duplicated gene of the Rx family, extant members of which show expression closely resembling that of Anf.
On the other hand, as shown by our work, a substantial portion of the Anf recognition helix, including four main residues (at positions 47, 50, 51 and 54) that undergo sequence-specific contacts with DNA, is probably inherited from one of the Antp-class genes. This group of genes is mainly expressed in the trunk region of embryos, and their protein products exert antagonistic effects on genes expressed rostrally. These traits of Antp-class proteins may explain the inhibitory influence of Anf on Otx2 expression.
As we showed previously in X. laevis and have now confirmed in lamprey, Anf inhibits Otx2 expression and thereby “cleans” Otx2 expression from the rostral region of the neural plate, which allows cells in the rostral neural plate to begin to express the telencephalic regulator FoxG1. When Anf is experimentally downregulated, Otx2 expression expands to the presumptive telencephalic territory, which is accompanied by inhibition of FoxG1 expression and posteriorisation of this territory11,13,30.
Accordingly, telencephalic specification appears to represent a peculiar “ground state” that can be reached by removal of Otx2, which could otherwise direct the presumptive telencephalic territory to a more posterior fate. At first glance, such a permissive strategy contradicts the fact that the telencephalon is the evolutionarily youngest brain unit because the “ground state” would intuitively be expected to be the oldest. To imagine how such an “inversion” could arise in evolution, it may be suggested that, as a first step, an inhibitory mechanism suppressing initial specification in a certain group of cells in the anterior region of the neural plate could have emerged. As we discussed above, this might have resulted from a genomic translocation that connected the 5′ region of a rostrally expressed Prd class gene, such as Rx, with the 3′ portion of a gene from the Antp class. Then, the progeny of this mutant ancestor, in which initial cell specification in the rostral region of the neural plate was inhibited, might obtain an advantage in natural selection because anterior cells become free to develop a new structure. Interestingly, the possibility that these ancient Anf-expressing cells could be stem-like cells with an uncertain fate was indirectly confirmed by the expression of Anf in mammalian ES cells, which are known as the cell type with the most uncertain fate12. Accordingly, we may speculate that the supposed translocation event resulting in the generation of Anf-expressing cells in the rostral neural plate could then have been co-opted in subsequent evolution as a necessary regulatory unit in the descendants’ developmental programme. Assuming this scenario to be true, the inhibition of Otx2 by Anf that takes place in the embryos of all extant vertebrates might represent a trace of this ancient inhibitory mechanism that “cleaned out” the territory and was further used by natural selection as a ground state for the “construction” of the telencephalon.
Beginning from Haeckel’s conception of terminal addition, it is thought that evolutionary innovations may be more successfully accepted by natural selection if they appear in later stages of embryogenesis because they may result in less disturbance of the developmental programme in this case31,32. In this respect, the very early expression of Anf within the anterior neurectoderm (beginning from the midgastrula stage) of vertebrates appears to be contradictory to the fact that the telencephalon is the youngest brain unit. However, as shown herein, the expression of Anf in lampreys, the most basal extant vertebrate species, shows heterochrony with its expression in other species, as Anf expression begins within the neurectoderm of the lamprey embryo only at the late neurula stage. Additionally, the much later onset of the expression of the main telencephalic regulator, FoxG1, further confirms the later specification of the telencephalon in lamprey. Importantly, similar significant heterochrony in lamprey for another important gene regulating telencephalon specification, Fgf8, was reported previously33. All of these data are clearly in agreement with the fact that the telencephalon is the youngest brain unit, which could have emerged in vertebrate ancestors at considerably later stages of their embryonic development. This finding in turn suggests that changes occurred during subsequent evolution that pushed telencephalic specification to earlier stages of embryogenesis. Interestingly, this idea is reminiscent of terminal addition, according to which evolutionary innovations are added to the terminal stages of embryogenesis in ancestors and are then pushed into earlier embryonic stages of descendants.
Materials and Methods
Animals
All animal experiments were performed in accordance with guidelines approved by the Shemyakin-Ovchinnikov Institute of Bioorganic Chemistry (Moscow, Russia) Animal Committee and handled in accordance with the 1986 Animals (Scientific Procedures) Act and Helsinki Declaration.
Lampetra fluviatilis, Lethenteron camtschaticum and Petromyzon marinus adult lampreys were collected in the Saint Petersburg, Petropavlovsk-Kamchatski and Archangelsk districts, respectively. Embryos were obtained via artificial fertilization of eggs squeezed from pregnant females. The embryos were staged as described previously17. For in situ hybridization, embryos were fixed in MEMFA (3,7% formaldehyde, 100 mM MOPS, 2 mM EGTA, 1 mM MgSO4), dehydrated in methanol and kept at −20 °C. The Xenopus laevis embryos used for testing Lanf activity were obtained from laboratory frogs through artificial fertilization and staged according to the Nieuwkoop and Faber Normal Tables34.
Cloning of Lanf cDNA
The homeobox-containing fragment of Lanf cDNA was obtained by using the Evrogen Kit for RT-PCR with degenerate oligos (see Supplementary). The positions of these oligos are shown in Fig. 1.
The homeobox sequence showing the highest homology to known Anfs from other vertebrates was chosen to design nested pairs of primers to obtain the 5′ and 3′ ends of Lanf cDNA via the Step-Out RACE method18. The cDNA fragment obtained through this method was cloned into the pGEM-T vector and sequenced. Finally, the cDNA sequence of Lanf was confirmed by obtaining the full-length cDNA with independent pairs of primers (see Supplementary) and sequencing several pGEM-T clones containing this cDNA.
Cloning of other cDNAs for in situ hybridization
Fragments of the cDNAs of FoxG1, Otx2, Shh and Lanf to be used for in situ hybridization were obtained via RT-PCR with the primers shown in Supplementary.
Bioinformatics
Phylogenetic analyses of protein sequences were performed via the neighbor-joining35 and maximum likehood36 methods using the MEGA637 program. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) is shown next to the branches38. The trees were rooted with the UPGMA algorithm or by including the PouII homeobox as the outgroup sequence. Evolutionary distances were computed using the JTT matrix-based method39. Selection of species for analyses in the case of the OTX, Goosecoid, and Anf gene families was performed accordingly40. In other cases, the genes were taken from invertebrates (insect, nematode, sea urchin, sea anemone and ascidia) and vertebrates (lamprey, frog, mouse).
The homeodomain sequences and the names of the species included in the phylogenetic analyses are shown in Supplementary.
RT-PCR and luciferase assays
For qRT-PCR, three groups of L. camtschaticum and X. laevis embryos were collected, obtaining 30 and 5 embryos, respectively, from each of the desired stages. Total RNA was extracted using an RNA isolation kit (MASHEREY-NAGEL) according to the manufacturer’s protocol. The concentration of the extracted RNA was measured with a Qubit® fluorometer (Invitrogen), while RNA integrity was checked visually via gel electrophoresis. The details of qRT PCR preparation and the reaction parameters and primers are shown in Supplementary.
The luciferase assay was performed as described in ref. 41 and Supplementary.
Synthetic mRNA and morpholino
Synthetic Lanf and Xanf1 mRNA was prepared with the mMessage Machine SP6 Kit (Ambion) after linearization of pCS2-based plasmids with NotI.
The following two variants of morpholino antisense oligonucleotides from Gene Tools (Fig. S8a) were injected at a final concentration of 0.4 mM in a volume of 3–4 nl.
1. Lanf MO1 (MO corresponding to positions −20- + 5 of L. fluviatilis mRNA):
5′-GCCATCTCTCGAAAAGTAATTCACG;
2. Lanf MO2 (MO corresponding to positions −46–22 of L. fluviatilis mRNA):
5′-ATTAGTTAATTGATCGGCGGTGGAA.
Importantly, Lanf MO1 and Lanf MO2 induced similar effects when they were injected into the embryos. A mismatched variant of Lanf MO-1 was used as a negative control: misLanf MO1–5′- ACCAAGTCTCGTTAAGAAATTTGCG. The efficiency of the MOs was tested (see Supplementary).
All mRNA and MO were mixed with Fluorescein Lysine Dextran (FLD) (Invitrogen, 40 kD, 5 μg/μl) before injection.
In situ hybridization
Whole-mount in situ hybridization was performed mainly as described previously42 with minor variations (see Supplementary).
Vibratome embryo sections with a thickness of 30 μm, hybridized in whole mounts and mounted in 4% agarose blocks, were prepared with a Microm HM 650 vibratome and photographed with a Leica M205 stereomicroscope.
Additional Information
How to cite this article: Bayramov, A. V. et al. The presence of Anf/Hesx1 homeobox gene in lampreys suggests that it could play an important role in emergence of telencephalon. Sci. Rep. 6, 39849; doi: 10.1038/srep39849 (2016).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Shimeld, S. M. & Holland, P. W. Vertebrate innovations. Proceedings of the National Academy of Sciences of the United States of America 97, 4449–4452 (2000).
Holland, L. Z. Chordate roots of the vertebrate nervous system: expanding the molecular toolkit. Nature reviews 10, 736–746 (2009).
Pani, A. M. et al. Ancient deuterostome origins of vertebrate brain signalling centres. Nature 483, 289–294 (2012).
Tomer, R., Denes, A. S., Tessmar-Raible, K. & Arendt, D. Profiling by image registration reveals common origin of annelid mushroom bodies and vertebrate pallium. Cell 142, 800–809 (2010).
Lowe, C. J., Clarke, D. N., Medeiros, D. M., Rokhsar, D. S. & Gerhart, J. The deuterostome context of chordate origins. Nature 520, 456–465 (2015).
Holland, L. Z. et al. Evolution of bilaterian central nervous systems: a single origin? EvoDevo 4, 27 (2013).
Zaraisky, A. G. et al. A novel homeobox gene expressed in the anterior neural plate of the Xenopus embryo. Developmental biology 152, 373–382 (1992).
Zaraisky, A. G. et al. The homeobox-containing gene XANF-1 may control development of the Spemann organizer. Development (Cambridge, England) 121, 3839–3847 (1995).
Kazanskaya, O. V. et al. Anf: a novel class of vertebrate homeobox genes expressed at the anterior end of the main embryonic axis. Gene 200, 25–34 (1997).
Ermakova, G. V. et al. The homeobox gene, Xanf-1, can control both neural differentiation and patterning in the presumptive anterior neurectoderm of the Xenopus laevis embryo. Development (Cambridge, England) 126, 4513–4523 (1999).
Ermakova, G. V., Solovieva, E. A., Martynova, N. Y. & Zaraisky, A. G. The homeodomain factor Xanf represses expression of genes in the presumptive rostral forebrain that specify more caudal brain regions. Developmental biology 307, 483–497 (2007).
Thomas, P. Q., Johnson, B. V., Rathjen, J. & Rathjen, P. D. Sequence, genomic organization, and expression of the novel homeobox gene Hesx1. The Journal of biological chemistry 270, 3869–3875 (1995).
Dattani, M. T. et al. HESX1: a novel gene implicated in a familial form of septo-optic dysplasia. Acta Paediatr Suppl 88, 49–54 (1999).
Hermesz, E., Mackem, S. & Mahon, K. A. Rpx: a novel anterior-restricted homeobox gene progressively activated in the prechordal plate, anterior neural plate and Rathke’s pouch of the mouse embryo. Development (Cambridge, England) 122, 41–52 (1996).
Andoniadou, C. L. & Martinez-Barbera, J. P. Developmental mechanisms directing early anterior forebrain specification in vertebrates. Cellular and molecular life science 70, 3739–3752 (2013).
Smith, J. J. et al. Sequencing of the sea lamprey (Petromyzon marinus) genome provides insights into vertebrate evolution. Nature genetics 45, 415–421, 421e411-412 (2013).
Tahara, Y. Normal stages of development in the lamprey, Lampetra reissneri (Dybowski). Zoological Science 5, 109–118 (1988).
Matz, M. et al. Amplification of cDNA ends based on template-switching effect and step-out PCR. Nucleic acids research 27, 1558–1560 (1999).
Docker, M. F., Youson, J. H., Beamish, R. J. & Devlin, R. H. Phylogeny of the lamprey genus Lampetra inferred from mitochondrial cytochrome b and ND3 gene sequences. Can. J. Fish. Aquat. 56, 2340–2349. (1999).
Sugahara, F. et al. Involvement of Hedgehog and FGF signalling in the lamprey telencephalon: evolution of regionalization and dorsoventral patterning of the vertebrate forebrain. Development (Cambridge, England) 138, 1217–1226 (2011).
Uchida, K. et al. Development of the adenohypophysis in the lamprey: evolution of epigenetic patterning programs in organogenesis. Journal of experimental zoology 300, 32–47 (2003).
Jaszai, J., Reifers, F., Picker, A., Langenberg, T. & Brand, M. Isthmus-to-midbrain transformation in the absence of midbrain-hindbrain organizer activity. Development (Cambridge, England) 130, 6611–6623 (2003).
Storm, E. E. et al. Dose-dependent functions of Fgf8 in regulating telencephalic patterning centers. Development (Cambridge, England) 133, 1831–1844 (2006).
Eroshkin, F., Kazanskaya, O., Martynova, N. & Zaraisky, A. Characterization of cis-regulatory elements of the homeobox gene Xanf-1. Gene 285, 279–286 (2002).
Sugahara, F. et al. Evidence from cyclostomes for complex regionalization of the ancestral vertebrate brain. Nature 531, 97–100. (2016).
Toresson, H., Martinez-Barbera, J. P., Bardsley, A., Caubit, X. & Krauss, S. Conservation of BF-1 expression in amphioxus and zebrafish suggests evolutionary ancestry of anterior cell types that contribute to the vertebrate telencephalon. Dev Genes Evol. 208, 431–9 (1998).
Lowe, C. J., Wu, M., Salic, A., Evans, L., Lander, E., Stange-Thomann, N., Gruber, C. E., Gerhart, J. & Kirschner, M. Anteroposterior patterning in hemichordates and the origins of the chordate nervous system. Cell 113, 853–65 (2003).
Dasen, J. S. et al. Temporal regulation of a paired-like homeodomain repressor/TLE corepressor complex and a related activator is required for pituitary organogenesis. Genes Dev. 15, 3193–207 (2001).
Holland, P. W., Garcia-Fernandez, J., Williams, N. A. & Sidow, A. Gene duplications and the origins of vertebrate development. Dev Suppl 125–133. (1994).
Fish, M. B. et al. Xenopus mutant reveals necessity of rax for specifying the eye field which otherwise forms tissue with telencephalic and diencephalic character. Dev Biol. 395, 317–330 (2014).
Richardson, M. K. & Keuck, G. Haeckel’s ABC of evolution and development. Biological reviews of the Cambridge Philosophical Society 77, 495–528 (2002).
Raff, R. A. & Raff, E. C. Evolution in the light of embryos: seeking the origins of novelties in ontogeny. In: Forms and Function in Developmental Evolution. Edited by Laubichler, M. D. & Maienshein, J. Cambridge University Press (2009).
Guerin, A. et al. Neurodevelopment genes in lampreys reveal trends for forebrain evolution in craniates. PloS one 4, e5374 (2009).
Nieuwkoop, P. D. & Faber, J. Normal table of Xenopus laevis (Daudin). North-Holland Publ., Co. (1956).
Saitou, N. & Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Molecular Biology and Evolution 4, 406–425 (1987).
Le, S. Q. & Gascuel, O. An Improved General Amino Acid Replacement Matrix. Mol Biol Evol 25, 1307–1320 (2008).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Molecular Biology and Evolution 30, 2725–2729 (2013).
Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 39, 783–791 (1985).
Jones, D. T., Taylor, W. R. & Thornton, J. M. The rapid generation of mutation data matrices from protein sequences. Computer Applications in the Biosciences 8, 275–282 (1992).
Kitamura K., Nishimura, Y., Kubotera, N., Higuchi, Y. & Yamaguchi, M. Transient activation of the micro1 homeobox gene family in the sea urchin (Hemicentrotus pulcherrimus) micromere. Dev Genes Evol. 212, 1–10 (2002).
Bayramov, A. V. et al. Novel functions of Noggin proteins: inhibition of Activin/Nodal and Wnt signaling. Development (Cambridge, England) 138, 5345–5356 (2011).
Sugahara, F., Murakami, Y. & Kuratani, S. Gene expression analysis of lamprey embryo (Springer New York, 2015).
Martynova, N. Y. et al. The LIM-domain protein Zyxin binds the homeodomain factor Xanf1/Hesx1 and modulates its activity in the anterior neural plate of Xenopus laevis embryo. Dev Dyn 237, 736–749 (2008).
Acknowledgements
This work was supported by RFBR grants 15-04-04343 (AVB) and 14-04-10111 (AGZ). Experiments with morpholino were supported by Russian Scientific Foundation (project no. 14-14-00557). We thank Dr. Kovalev M.Yu. and employees of Biological Station “Raduga” (Kamchatka, Russia), Dr. Bazykin G.A., Elena Solovieva, Olga Averianova, Daria Korotkova and Evgeny Orlov for assistance.
Author information
Authors and Affiliations
Contributions
A.V.B. - embryos collection, cloning, injections, PCR, phylogenetic analysis, figures preparation, drawing of the animal outlines on Figure 5; G.V.E. – in situ hybridization, F.M.E. – embryos collection, cloning, injections, A.V.K. - animals, embryos; N.Y.M. - Western blotting; A.G.Z. - embryos, figures preparation, writing of the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Bayramov, A., Ermakova, G., Eroshkin, F. et al. The presence of Anf/Hesx1 homeobox gene in lampreys suggests that it could play an important role in emergence of telencephalon. Sci Rep 6, 39849 (2016). https://doi.org/10.1038/srep39849
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep39849
This article is cited by
-
The evolutionary origin of visual and somatosensory representation in the vertebrate pallium
Nature Ecology & Evolution (2020)
-
Discovery of four Noggin genes in lampreys suggests two rounds of ancient genome duplication
Communications Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.