Abstract
To identify what long non-coding RNAs (lncRNAs) are involved in non-small cell lung cancer (NSCLC), we analyzed microarray data on gene expression and methylation. Gene expression chip and HumanMethylation450BeadChip were used to interrogate genome-wide expression and methylation in tumor samples. Differential expression and methylation were analyzed through comparing tumors with adjacent non-tumor tissues. LncRNAs expressed differentially and correlated with coding genes and DNA methylation were validated in additional tumor samples using RT-qPCR and pyrosequencing. In vitro experiments were performed to evaluate lncRNA’s effects on tumor cells. We identified 8,500 lncRNAs expressed differentially between tumor and non-tumor tissues, of which 1,504 were correlated with mRNA expression. Two of the lncRNAs, LOC146880 and ENST00000439577, were positively correlated with expression of two cancer-related genes, KPNA2 and RCC2, respectively. High expression of LOC146880 and ENST00000439577 were also associated with poor survival. Analysis of lncRNA expression in relation to DNA methylation showed that LOC146880 expression was down-regulated by DNA methylation in its promoter. Lowering the expression of LOC146880 or ENST00000439577 in tumor cells could inhibit cell proliferation, invasion and migration. Analysis of microarray data on gene expression and methylation allows us to identify two lncRNAs, LOC146880 and ENST00000439577, which may promote the progression of NSCLC.
Similar content being viewed by others
Introduction
Lung cancer is one of the most common malignant tumors worldwild. According to the GLOBOCAN 2012, around 1.8 million people are diagnosed with lung cancer every year, consisting of 13% of all newly diagnosed cancer patients. Lung cancer is the number one cause of cancer deaths, about 1.6 million deaths a year, accounting for 19% of all cancer deaths1. Non-small cell lung cancer (NSCLC) is the most common form of lung cancer, which accounts for approximately 85% of all lung cancer cases. Because of the high mortality, NSCLC is one of the mostly studied malignancies. Despite extensive research, our understanding of the disease remains limited. One of the challenges is tumor heterogeneity due not only to individual’s variations in disease characteristics, but also cellular differences in response to therapeutic regimens. Rapid advancement in high-throughput technologies has provided new tools to study the disease in a more comprehensive fashion. Many researchers have begun to integrate omics data from different high-throughput platforms to investigate the disease mechanisms and to search for robust biomarkers2,3,4,5,6.
Epigenetic regulation of coding and non-coding genes has been shown to play important roles in genome functions. Research has revealed significant interactions betweeen and alterations in DNA methylation and gene expression in many types of cancer, including the breast7, prostate8, and lung9. Besides DNA methylation, non-coding RNA transcripts are also essential components of epigenetic regulation. Recently, long non-coding RNAs (lncRNAs) have drawn much attention in cancer research as this class of transcripts has complex and diverse biologic actions on cell activities and functions10. LncRNAs regulate gene expression through various mechanisms, including complementary binding to DNA, as well as to other coding and non-coding RNA transcripts11. Although there are significant interactions between DNA methylation and expression of protein-coding and non-coding RNAs, little is known about their alterations in lung cancer. An integrated genome-wide analysis of both DNA methylation and expression of lncRNAs and mRNAs may help to understand the complex regulatory network that underlies the tumorigenic mechanisms of the lung.
We report here our study of integrated genome-wide analyses of DNA methylation, lncRNA and gene expression in search for new regulatory and functional connections among DNA methylation, lncRNA and mRNA expression in lung cancer. Through exploring novel networks and mechanisms, we anticipate to identify new biomarkers and targets for lung cancer diagnosis, prognosis and treatment. Our study also offers clues for integration of genome-wide data from different platforms which have cross-talk in biological signaling and function, and advocates the use of this approach to further investigation of tumorigenesis and search for reliable tumor markers for clinical management of cancer patients.
Results
Identification of lncRNA-mRNA pairs
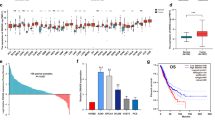
The workflow of our microarray analysis, data integration, gene identification and bioinformatic evaluation is shown in Supplementary Fig. S1. From the expression microarray, we found 10,206 transcripts differentially expressed between tumor and adjacent non-tumor tissues (P < 0.05 after FDR adjustment). Of these transcripts, 8,500 were lncRNAs and 1,685 were mRNAs. PCA plots (Fig. 1a) show complete separations of tumor and non-tumor samples in terms of lncRNA expression profiles. From the profiles, we selected top 3,690 lncRNAs to show the differences in lncRNA expression between tumor and non-tumor tissues, and the expression patterns were clearly different between tumor and non-tumor samples (Fig. 1b).
Bioinformatics analysis of lncRNA.
(a) PCA of all lncRNA expression results from the microarray analysis. (b) Heatmap results based on the top 3,690 lncRNAs significantly associated with tumor samples from the microarray analysis of gene expression. (c) IPA results based on the 1,345 cis-pairs of lncRNAs and mRNAs which were differentially expressed in NSCLC. (d) Distributions of the methylation loci that were significantly correlated with lncRNA expression and their associated lncRNAs on chromosome 17. X axis is the correlation coefficient; Y axis is the p value; Z axis is the location of CpG sites in reference to TSS.
We next investigated the correlation of differentially expressed lncRNAs with differentially expressed mRNAs in NSCLC. The analyses yielded 1,504 pairs of lncRNAs and mRNAs with R2 > 0.5 (P < 0.05). Of these pairs, 1,345 were in cis- and 159 were in trans-correlation. IPA was performed on the cis-pairs. The top ten significant pathways were shown in Fig. 1c, and nearly half of the pathways were directly related to cancer, including cell death and survival, cellular development, and cellular growth and proliferation, suggesting the critical involvement of lncRNAs in tumorigenesis. From the list of correlated lncRNAs and mRNAs which also had large fold-change in expression and known biological annotation, we selected two pairs of lncRNAs and mRNAs (LOC146880/KPNA2 and ENST00000439577/RCC2) for further validation. The coding genes in these pairs are known to have significant biological implications in NSCLC. The chromosomal location of the lncRNAs and their correlated genes are shown in Supplementary Fig. S2.
Identification of lncRNA-methylation pairs
In the genome-wide methylation analysis, we identified 113,644 CpG with differential methylation (DM) between tumor and adjacent non-tumor tissues (P < 0.05 after FDR adjustment). We performed correlation analysis for the differentially expressed lncRNAs with their DM loci corresponding to the same chromosome. Among these DM loci, one methylation site (cg12562461) was located in the promoter of LOC146880 on chromosome 17. Distribution of the methylation sites and their associated lncRNAs on chromosome 17 revealed that more methylation sites around TSS were correlated with lncRNA expression (Fig. 1d). Results suggested that cg12562461 methylation was significantly lower in tumor than in non-tumor tissues, and the expression of LOC146880 was inversely correlated with methylation.
Validation of lncRNA/mRNA relations and analysis of their associations with survival
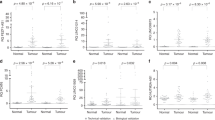
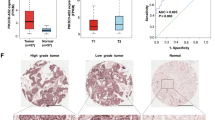
Using qPCR, we validated our microarray findings of two pairs of correlated lncRNAs and mRNAs in total 402 tumors and 105 adjacent non-tumor tissues. The qPCR results shown in Fig. 2 were similar to those of microarray results, showing that LOC146880 and KPNA2 expression was higher in tumor than in adjacent non-tumor tissues (Fig. 2a). We also observed a significant correlation between LOC146880 and KPNA2 expression (r = 0.306, P = 0.011) (Fig. 2b). We further analyzed the associations of LOC146880 and ENST00000439577 expression with patient’s clinical and pathological characteristics, and found that LOC146880 expression was associated with histological type and tumor size, while ENST00000439577 expression was associated with metastasis status and disease stage (Table 1). Kaplan-Meier survival analysis revealed that high expression of LOC146880 was associated with poor overall survival both in the training (P < 0.001) and validation sets (P = 0.044) (Fig. 2c). In addition, we analyzed the methylation of cg12562461 located in TSS200 of LOC146880 in 126 NSCLC tumors and 30 adjacent non-tumor tissues, and found low cg12562461 methylation in tumor than in adjacent non-tumor tissues (Fig. 2d). Our analysis also showed low LOC146880 expression in high methylation and high expression in low methylation tissue samples (P < 0.01) (Fig. 2e). Similarly, ENST00000439577 and RCC2 expression were higher in tumor than in adjacent non-tumor tissues (Fig. 2f), and ENST00000439577 and RCC2 expression were positively correlated (r = 0.288, P = 0.011) (Fig. 2g). Furthermore, high expression of ENST00000439577 was associated with poor overall survival both in the training (P = 0.028) and validation sets (P = 0.006) (Fig. 2h).
Validation of LOC146880 and ENST00000439577 expression in tumor and adjacent non-tumor tissues of NSCLC.
(a) Higher expression of LOC146880 and KPNA2 in tumor than in adjacent non-tumor tissues. (b) A positive correlation between LOC146880 and KPNA2 expression. (c) High LOC146880 expression associated with poor overall survival (left: training set; right: validation set). (d) Lower methylation of cg12562461 in tumor than in adjacent non-tumor tissues. (e) Lower methylation of cg12562461 in high than in low LOC146880 expression tumors. (f) Higher expression of ENST00000439577 and RCC2 in tumor than in adjacent non-tumor tissues. (g) A positive correlation between ENST00000439577 and RCC2 expression. (h) High ENST00000439577 expression associated with poor overall survival (left: training set; right: validation set). Data are expressed as mean ± s.d.; *P < 0.05; **P < 0.01; ***P < 0.001.
Functional analysis of LOC146880 in NSCLC cell lines
To examine the biological relevance of LOC146880 in lung cancer, we analyzed baseline expression of LOC146880 in three NSCLC (A549, PC9 and H1299) and one normal cell lines (Beas2B). Our analysis showed that LOC146880 expression was high in A549 and PC9 compared to Beas2B, but low in H1299 (see Supplementary Fig. S3). Based on these results, we selected A549 and PC9 for our knockdown experiments. We made a small interfering RNA against LOC146880, siRNA-LOC146880, and transfected the siRNA into A549 and PC9 cell lines. Transfected cells showed significant decreases in LOC146880 expression (Fig. 3a). As a downstream molecule, KPNA2 expression were also declined in A549 and PC9 at both mRNA and protein levels (Fig. 3a). Moreover, low expression of LOC146880 inhibited both A549 and PC9 proliferation (Fig. 3b). Furthermore, knockdown of LOC146880 by siRNA significantly inhibited cell migration (Fig. 3c) and invasion (Fig. 3d) in A549 and PC9. Flow cytometry analysis showed that lowering LOC146880 expression slowed down cell cycle progression at the G2/M phase (Fig. 3e), suggesting that declined LOC146880 expression may put a break on cell cycle progression resulting in suppression of cell proliferation and migration. Taken together, our findings indicate that LOC146880 may function as an oncogenic transcript in NSCLC.
Functional analysis of LOC146880 expression in NSCLC cell lines.
(a) Declined expression of LOC146880 and KPNA2 after siRNA-LOC146880 transfection. Declined expression of KPNA2 proteins after siRNA-LOC146880 transfection. (b) Declined A549/PC9 cell proliferation after lowering the expression of LOC146880. (c) Declined A549/PC9 cell migration after lowering the expression of LOC146880. (d) Declined A549/PC9 cell invasion after lowering the expression of LOC147880. (e) Reduced cell cycle progression measured by the C6 flow cytometry after lowering the expression of LOC146880. (f) A promoter region of LOC146880 contains a CpG site which was predicted to be a transcription factor SP1 binding site. A LOC146880 fragment (−623/+123) was cloned into the pGL3-basic vector. The vector containing the LOC146880 (−623/+123) construct displayed 15-fold increases in promoter activities compared with the PGL3-basic vector. LOC146880 methylation is lower in tumors than in normal tissues. Data are expressed as mean ± s.d.; *P < 0.05, **P < 0.01. NC, Negative Control.
Based on the prediction by TRANSFAC® (http://www.gene-regulation.com/pub/databases.html), LOC146880 promoter contains one CpG site which is located in the transcription factor (SP1) binding site, suggesting that methylation in the region may block the SP1 binding to the promoter, resulting in suppression of LOC146880 expression. To test this possibility, We inserted a LOC146880 promoter construct (−623/+123) into a plasmid (pGL3), and transfected it into HEK293T, A549 and H1299 cell lines which were co-transfacted with a SP1 overexpression vector. Compared to the control (pGL3-basic), LOC146880 transfected cells (pGL3-LOC146880) displayed significant promoter activities, suggesting the existence of SP1 activation in the LOC146880 promoter (Fig. 3f).
Baseline expression of ENST00000439577 was measured in three NSCLC (A549, H1975 and H1299) and one normal cell lines (Beas2B). The analysis showed that ENST00000439577 expression was high in A549, H1299 and H1975 compared to Beas2B (see Supplementary Fig. S4). A549 and H1299 cell lines were selected for testing the effects of ENST00000439577. We made a small interfering RNA against ENST00000439577, siRNA-ENST00000439577, and transfected the siRNA into A549 and H1299 cell lines. After transfection, cells showed significant declines in ENST00000439577 expression. RCC2 expression was also reduced in A549 and H1299 (Fig. 4a). Low expression of ENST00000439577 inhibited both A549 and H1299 proliferation (Fig. 4b). Furthermore, knockdown of ENST00000439577 by siRNA significantly inhibited cell migration (Fig. 4c) and invasion (Fig. 4d) in A549 and H1299. Flow cytometry analysis indicated that cell cycle pregresion was slightly reduced at the G2/M phase in both cell lines after lowering ENST00000439577 expression, although the changes were not statistically signigicant (Fig. 4e). We also analyzed CpG site methylation in the promoter of ENST00000439577, and found no significant correlation between methylation and expression.
Functional analysis of ENST00000439577 expression in NSCLC cell lines A549 and H1299.
(a) Declined expression of ENST00000439577 and RCC2 after siRNA-ENST00000439577 transfection. (b) Reduced A549/H1299 cell proliferation after lowering the expression of ENST00000439577. (c) Reduced A549/H1299 cell migration after lowering the expression of ENST00000439577. (d) Reduced A549/H1299 cell invasion after lowering the expression of ENST00000439577. (e) Cell cycle analysis by the C6 flow cytometry after lowering the expression of ENST00000439577. Data are expressed as mean ± s.d.; *P < 0.05. NC, Negative Control.
Discussion
With the rapid improvement in our understanding of the human genome from nucleotide sequences to functional regulation, we now recognize that changes in epigenetic regulation, including histone modifications, DNA methylation and non-coding RNA expression, play key roles in the development and progression of human cancer. High-throughput technology allows us to analyze genome-wide changes of DNA methylation and gene expression in cancer, but the challenge is how to effectively use the massive high-dimensional data to understand the process of tumorigenesis.
Although much research on DNA methylation and gene expression has been directed towards the understanding of protein-coding genes and their functions, evidence also suggests that we should integrate multiple layers of epigenetic regulation to reveal the well-orchestrated regulatory network that is beyond the production and activities of proteins. In this study, we performed genome-wide analyses of lncRNA, mRNA expression and DNA methylation, and integrated the two platforms of high-dimensional data to evaluate concurrent changes between epigenetic regulation and gene expression in NSCLC. Integrating the expression profiles of both coding and non-coding RNAs with genome-wide DNA methylation may also aid our understanding of the interactions between epigenetic modifications and gene expression in lung cancer. Furthermore, this integrated approach may help to discover and develop novel epigenetic biomarkers for diagnosis, prognosis and treatment of lung cancer.
In our study, we discovered two novel lncRNAs whose expression were significantly elevated in NSCLC, and high expression were associated with poor overall survival as well as with two possible onco-proteins. High expression of LOC146880 was associated with high expression of KPNA2, and suppressing LOC146880 expression resulted in declines in KPNA2 expression in NSCLC cell lines. KPNA2 belonges to the karyopherin α family, which has been reported as a key nucleocytoplasmic transport protein in the translocation of several cancer-related proteins. KPNA2 may play critical roles in carcinogenesis, cell differentiation, transcription regulation and immune response. Recently studies have demonstracted that high expression of KPNA2 is associated with poor prognosis of esophageal squamous cell carcinoma12, epithelial ovarian carcinamos13, gastric cancer14, prostate cancer15, upper tract urothelial carcinoma16 and breast cancer17. Teng et al. found that KPNA2 was involved in DNA damage repair through mediating the NBS1 subcellular location and functions of the MRE11-RAD50-NBS1 complex in tumorigenesis18. It was also suggested that silencing KPNA2 could decrease the nuclear translocation of POU class 5 homeobox 1 (Oct4), suppressing the proliferation of lung cancer cells19. We speculate that LOC146880 may play a critical role in tumorigenesis through the regulation of KPNA2 expression. In our analysis, we found not only LOC146880 expression in NSCLC being associated with patient survival outcomes, but also changes of its expression in lung tumor cells resulting in downregulation of KPNA2 as well as suppression of cell proliferation, migration and invasion. In our investigation, we also found that the transcription activity of LOC146880 was affected by promoter methylation.
ENST00000439577 and RCC2 were another pair of correlated lncRNA/mRNA expression associated with NSCLC. High ENST00000439577 expression was associated with high expression of RCC2, as well as with poor overall survival of NSCLC patients. We knocked down the expression of ENST00000439577 using siRNA, and found decreases in the expression of RCC2 in NSCLC cell lines, which resulted in inhibition of cell proliferation, migration and invasion. Yenjerla et al. reported that TD-60 (also known as RCC2) was a mitotic centromere-associated protein and an essetial regulator of cell cycle progression through G1, S, and G2 phases into mitosis20,21. Moreover, TD-60 displayed high guanine exchange factor (GEF) activity for the Ras-related protein RalA, and regulated the kinetochore-microtubule stability in early mitosis22. Subsequent studies revealed that Ral GEFs palyed an important role during the Ras-induced tumourigenesis via activating the Ral GTPases23. Furthermore, down-regulation of RalA could reactivate p53 protein expression and stability, and up-regulate the expression of p53-downstream molecule p21 in NSCLC cell lines H460 that harbored mutant K-Ras and wild-type p5324. Guin et al. found that RalA/RalB, as the downstream effector of RAS, might be important for tumor metastasis and invasion of NSCLC, but, however, their mechanisms remained unknown25. According to our results, we speculate that high expression of ENST00000439577 could up-regulate RCC2 expression, which may promote the activity of RalA, stimulating the growth and invasion of lung tumour cells. Consequently, lncRNA ENST00000439577, which influences the Ral signalling pathways via RCC2, may be a potential therapeutic target for the treatment of lung cancer.
Although we used a limited discovery set to identify lncRNAs and methylation loci, we were able to validate our microarray results in additional specimens. This process demonstrates that microarray is quite efficient in identifying potential molecules involved in lung cancer. In addition to the integrated analysis of lncRNAs and mRNAs expression, we also investigated the correlation of lncRNAs expression and their methylation in lung cancer. All the analyses indicate that our approach to integrate different microarray analyses is clinically and biologically relevant and that our findings of expression of coding and non-coding RNAs and methylation loci may help to develop useful biomarkers for prognosis and treatment of lung cancer. More studies are needed to further explore and integrate different high-dimensional data. These approaches will help to generate valuable insights into the molecular mechanisms of lung tumorigenesis and progression.
Methods
Study patients
NSCLC patients (n = 490) were recruited from the Tianjin Medical University Cancer Hospital (TMUCH) between May 2006 and July 2011. These patients were divided into two groups: one training set including patients recruited before December 31 2008, one validation set including patients recruited after January 1, 2009. Informed consents were obtained from all the subjects. Fresh tumor samples and adjacent non-tumor tissues were collected from the patients during their surgery. Twelve pairs of tumor and adjacent non-tumor samples were used for microarray analyses of DNA methylation and gene expression, and 217 tumor samples from the training set and 185 tumor samples from the validation set were analyzed for expression of lncRNAs using qPCR, respectively, and 126 tumor samples were analyzed for validation of methylation using pyrosequencing. Patients’ characteristics are shown in Supplementary files (see Supplementary Table S1). All the patients were followed from surgery (earliest in May 2006) to August 2013. The study was approved both by the Medical Ethical Review Committees at TMUCH and Shanghai Jiao Tong University School of Public Health. These experimental protocols were carried out in accordance to with the approved guidelines of TMUCH and Shanghai Jiao Tong University School of Public Health.
Microarray of gene expression
Expression of lncRNAs and mRNAs were measured using a SBC microarray chip (Shanghai Biotechnology Company, Shanghai, China), which was based on the Agilent eArray platform with additional probes to lncRNAs. The chip detects 48,011 lncRNA and 41,712 mRNA transcripts. Data were extracted using the Feature Extraction software 10.7 (Agilent technologies, Santa Clara, CA). Raw data were normalized with the Quantile algorithm, Gene Spring Software 11.0 (Agilent technologies, Santa Clara, CA).
Microarray of DNA methylation
DNA methylation was measured in 12 paires of tissue samples using the Illumina HumanMethylation450 BeadChip (Illumina, San Diego, CA). The chip analysis of DNA methylation has been described elsewhere26.
DNA and RNA extraction
Genomic DNA and total RNAs were isolated from tissue samples using the QIAamp DNA Mini Kit (Qiagen, Hilden, Germany) and TRIzol reagents (Invitrogen, Grand Island, NY), respectively, according to the manufacturer’s protocols. DNA and RNA concentrations were quantified with UV light using a Nanodrop ND-8000 spectrophotometer (Nanodrop Technologies, Wilmington, DE).
Reverse-transcriptase quantitative PCR (RT-qPCR)
Total RNA was reversely transcribed to cDNA using the PrimerScript RT Kit with the gDNA Eraser (Takara, Dalian, China) in accordance with the manufacturer’s instructions. Beta-actin was used as an internal control. For data analysis, the qPCR results were quantified as fold changes between Beta-actin and target genes using the formula 2−△△Ct.
DNA extraction and pyrosequencing analysis of DNA methylation
Bisulfite pyrosequencing was performed to validate our microarray results in tumor samples. Genomic DNA was bisulfite-converted using the EpiTect plus DNA Bisulfite Kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol. The treated DNA samples were PCR-amplified and fragmented. The processed samples were then precipitated, suspended and sequenced in the Pyro Mark Q96 system (Qiagen, Hilden, Germany). Primer sequences and PCR conditions are provided in Supplementary files (see Supplementary Table S2). The primers were designed to include CpG sites within 0.5 kb of the transcription start site. A methylation level >15% was considered positive for methylation. Any levels equal to or lower than that could not be easily distinguished from the background, and therefore were regarded as negative for methylation.
Cell culture
Normal cell line HEK293T, Beas2B and NSCLC cell lines A549, H1299, H1975 and PC9 were purchased from the Shanghai Cell Bank, Type Culture Collection Committee of Chinese Academy of Science (CAS, Shanghai, China) and were authenticated using short tandem repeat profiling by the cell bank in 2015. These cells were maintained in RPMI-1640 medium supplement with 10% fetal calf serum (Gibco, Grand Island, NY) at 37 °C in a humid atmosphere containing 5% CO2 according to the recommendation by CAS. We only used cells within 6 months from being thawed for the current study and recorded the cell morphology and doubling times regularly to ensure the maintenance of phenotypes.
Cell transfection experiments
To knockdown the expression of LOC146880 or ENST00000439577, siRNA-LOC146880 or siRNA-ENST00000439577 were transfected into the NSCLC cell lines using the Lipofectamine® 2000 Reagent (Invitrogen, Grand Island, NY) following the manufacturer’s instructions. We designed three specific siRNAs for knockdown of each target of interest and mixed them for transfection. The siRNAs information is shown in Supplementary Table S3. For comparison, siRNA-negative control was also transfected under the same condition.
Luciferase report assay
To determine if LOC146880 expression was regulated by promoter methylation, a 746 bp PCR product containing the LOC146880 promoter (from −623 to +123 relative to the transcription start site) was cloned into the pGL3 luciferase report vector (Promega, Madison, WI) between the restriction sits KpnI and HindIII. HEK293T and NSCLC cell lines (H1299 and A549) were seeded onto 24-well plates at 5 × 104 cells per well for 24 hours prior to transfection. The cells were transfected either with 0.5 mg pGL3-LOC146880 (−623/+123) or pGL3-basic (control) using the Lipofectamine® 2000 Reagent according to the manufacturer’s instructions. PRL-TK plasmid (10 ng) containing the Renilla luciferase gene (Promega, Madison, WI) was co-transfected to standardize transfection efficiency. The relative luminescence was measured 24 hours post-transfection using the Dual-Luciferase Reporter Assay System (Promega, Madison, WI). The experiments were performed in triplicate.
Assays of cell proliferation, wound healing, trans-well mobility and cell cycle progression
These assays have been described elsewhere27.
Western blot assay
Cultured cells were lysed on ice for 30 minutes in 1x RIPA Lysis Buffer supplemented with a protease inhibitor cocktail (Millipore, Billerica, MA). Protein concentrations of the cell lysates were measured with the BCA Protein Assay Kit (Sangon Biotech, Shanghai, China). Denatured proteins (40 μg) were electrophoresed on 10% SDS-PAGE gel and were electrotransferred from the gel to PDVF membranes (Millipore, Billerica, MA). After blocking with 5% non-fat milk in a Tris buffered saline with 0.05% Tween-20 for one hour at room temperature, the membranes were incubated overnight at 4 °C with primary antibodies against KPNA2 (Abcam, Cambridge, MA) at a dilution of 1:500, and β-actin 1:1000 (Santa Cruz Biotechnology, Dallas, TX). Secondary antibodies were added at concentrations of 1:4000. ECL Kit (Millipore, Billerica, MA) was used to visualize protein expression.
Microarray data analysis
Expression data were evaluated for reliability with reference to internal controls. Raw data were normalized with Quantile algorithm, Gene Spring 11.0 (Agilent technologies, Santa Clara, CA). Limma package in R (3.2.2 version) was used to determine differential expression of lncRNA and mRNA between matched tumor and non-tumor samples, followed by the Benjamin-Hochberg FDR correction. Correlations between lncRNAs and mRNAs were analyzed using R (3.2.2 version). LncRNAs were annotated for their protein-coding targets based on 1) their proximities to the protein coding genes (cis-targets) and 2) sequence homologies determined by BLAST (trans-targets). Heatmap was generated to show expression patterns. Principal component analysis (PCA) was performed to assess sample clustering. Analysis of functional enrichment on selected genes was performed using the Ingenuity Pathway Analysis (IPA) (http://www.ingenuity.com; September 2015).
Correlations between lncRNA expression and DNA methylation were analyzed for the methylation sites located within the lncRNA genes or corresponding to the TSS1500 regions. To show the patterns of these relationships, correlation coefficients with p values less than or equal to 0.01 were selected for plot. For each chromosome, the correlation’s P values, correlation coefficients and distances to TSS were plotted.
Statistical analysis
SPSS software version 16.0 was used for statistical analysis. Student’s t-test was performed to compare changes in expression. Log-rank test and Kaplan-Meier survival plot were used to evaluate associations of overall survival with methylation or expression of lncRNAs or coding genes. Cox proportional hazards regression was employed for survival analysis with adjustment for covariates. All statistical tests were two-tailed. Overall survival was calculated from the date of surgery to the date of death or last live contact.
Additional Information
How to cite this article: Feng, N. et al. Analysis of Microarray Data on Gene Expression and Methylation to Identify Long Non-coding RNAs in Non-small Cell Lung Cancer. Sci. Rep. 6, 37233; doi: 10.1038/srep37233 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Ferlay, J. et al. GLOBOCAN 2012 v1.0, Cancer Incidence and Mortality Worldwide: IARC CancerBase No. 11 [Internet]. Lyon, France: International Agency for Research on Cancer. (2013) Available at: http://globocan.iarc.fr. (Accessed: 4th November 2015).
Shames, D. S. et al. A Genome-Wide Screen for Promoter Methylation in Lung Cancer Identifies Novel Methylation Markers for Multiple Malignancies. PLoS Med. 3, e486 (2006).
Anglim, P. P. et al. Identification of a panel of sensitive and specific DNA methylation markers for squamous cell lung cancer. Mol Cancer. 7, 62 (2008).
Heller, G., Zielinski, C. C. & Zöchbauer-Müller, S. Lung cancer: From single-gene methylation to methylome profiling. Cancer Metastasis Rev. 29, 95–107 (2010).
Walter, K. et al. DNA methylation profiling defines clinically relevant biological subsets of non-small cell lung cancer. Clin Cancer Res. 18, 2360–2373 (2012).
Kim, B., Kang, S., Jeong, G., Park, S. B. & Kim, S. J. Identification and Comparison of Aberrant Key Regulatory Networks in Breast, Colon, Liver, Lung, and Stomach Cancers through Methylome Database Analysis. PLoS One. 9, e97818 (2014).
Rhee, J. K. et al. Integrated analysis of genome-wide DNA methylation and gene expression profiles in molecular subtypes of breast cancer. Nucleic Acids Res. 41, 8464–8474 (2013).
Yu, Y. P. et al. Whole-genome methylation sequencing reveals distinct impact of differential methylations on gene transcription in prostate cancer. Am J Pathol. 183, 1960–1970 (2013).
Li, J., Ching, T., Huang, S. & Garmire, L. X. Using epigenomics data to predict gene expression in lung cancer. BMC Bioinformatics. 16 (Suppl 5), S10 (2015).
Ponting, C. P., Oliver, P. L. & Reik, W. Evolution and functions of long noncoding RNAs. Cell. 136, 629–641 (2009).
Faghihi, M. A. & Wahlestedt, C. Regulatory roles of natural antisense transcripts. Nat Rev Mol Cell Biol. 10, 637–643 (2009).
Sakai, M. et al. Significance of karyopherin-{alpha} 2 (KPNA2) expression in esophageal squamous cell carcinoma. Anticancer Res. 30, 851–856 (2010).
Zheng, M. et al. Overexpression of karyopherin-2 in epithelial ovarian cancer and correlation with poor prognosis. Obstet Gynecol. 116, 884–891 (2010).
Altan, B. et al. Nuclear karyopherin-α2 expression in primary lesions and metastatic lymph nodes was associated with poor prognosis and progression in gastric cancer. Carcinogenesis. 34, 2314–2321 (2013).
Grupp, K. et al. High nuclear karyopherin α 2 expression is a strong and independent predictor of biochemical recurrence in prostate cancer patients treated by radical prostatectomy. Mod Pathol. 27, 96–106 (2014).
Shi, B. et al. High expression of KPNA2 defines poor prognosis in patients with upper tract urothelial carcinoma treated with radical nephroureterectomy. BMC Cancer. 15, 380 (2015).
Alshareeda, A. T. et al. KPNA2 is a nuclear export protein that contributes to aberrant localisation of key proteins and poor prognosis of breast cancer. Br J Cancer. 112, 1929–1937 (2015).
Teng, S. C., Wu, K. J., Tseng, S. F., Wong, C. W. & Kao, L. Importin KPNA2, NBS1, DNA repair and tumorigenesis. J Mol Hist. 37, 293–299 (2006).
Li, X. L. et al. Downregulation of KPNA2 in non-small-cell lung cancer is associated with Oct4 expression. J Transl Med. 11, 232 (2013).
Yenjerla, M., Panopoulos, A., Reynaud, C., Fotedar, R. & Margolis, R. L. TD-60 is required for interphase cell cycle progression. Cell Cycle. 12, 837–841 (2013).
Grigera, P. R. et al. Mass spectrometric analysis identifies a cortactin-RCC2/TD60 interaction in mitotic cells. J Proteomics. 75, 2153–2159 (2012).
Papini, D. et al. TD-60 links RalA GTPase function to the CPC in mitosis. Nat Commun. 6, 7678 (2015).
Bodemann, B. O. & White, M. A. Ral GTPases and cancer: linchpin support of the tumorigenic platform. Nat Rev Cancer. 8, 133–140 (2008).
Tecleab, A., Zhang, X. & Sebti, S. M. Ral GTPase down-regulation stabilizes and reactivates p53 to inhibit malignant transformation. J Biol Chem. 289, 31296–31309 (2014).
Guin, S. & Theodorescu, D. The RAS-RAL axis in cancer: evidence for mutation-specific selectivity in non-small cell lung cancer. Acta Pharmacol Sin. 36, 291–297 (2015).
Touleimat, N. & Tost, J. Complete pipeline for Infinium(®) Human Methylation 450K BeadChip data processing using subset quantile normalization for accurate DNA methylation estimation. Epigenomics. 4, 325–341 (2012).
Liu, B. et al. Mir-195 suppresses non-small cell lung cancer by targeting CHEK1. Oncotarget. 6, 9445–9456 (2015).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant No. 81573231); the Natural Science Foundation of Shanghai (Grant No. 15ZR1424400); and the Translational Medicine Foundation of Shanghai Jiao Tong University medical college (Grant No. 15ZH1001).
Author information
Authors and Affiliations
Contributions
B.Q. conceived and designed the study; N.F. and T.C. performed the experiments, the contribution is equal; N.F., T.C., X.Z., H.Y., L.G. and B.Q. analyzed and interpreted the data; Y.W., B.L., M.Z. and M.G. collected subjects and clinical data; N.F., Y.W., H.L., X.Z. and O.S. performed experiments in vitro. B.Q., N.F., Z.Z., H.Y. and L.G. wrote and revised the manuscript. All authors reviewed and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Feng, N., Ching, T., Wang, Y. et al. Analysis of Microarray Data on Gene Expression and Methylation to Identify Long Non-coding RNAs in Non-small Cell Lung Cancer. Sci Rep 6, 37233 (2016). https://doi.org/10.1038/srep37233
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep37233
This article is cited by
-
Mutual interaction of lncRNAs and epigenetics: focusing on cancer
Egyptian Journal of Medical Human Genetics (2023)
-
A SNP-mediated lncRNA (LOC146880) and microRNA (miR-539-5p) interaction and its potential impact on the NSCLC risk
Journal of Experimental & Clinical Cancer Research (2020)
-
Methylation and transcriptome analysis reveal lung adenocarcinoma-specific diagnostic biomarkers
Journal of Translational Medicine (2019)
-
Epigenetics of lung cancer: a translational perspective
Cellular Oncology (2019)
-
Long non-coding RNAs involved in autophagy regulation
Cell Death & Disease (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.