Abstract
The temporal dynamics of desert soil microbial communities are poorly understood. Given the implications for ecosystem functioning under a global change scenario, a better understanding of desert microbial community stability is crucial. Here, we sampled soils in the central Namib Desert on sixteen different occasions over a one-year period. Using Illumina-based amplicon sequencing of the 16S rRNA gene, we found that α-diversity (richness) was more variable at a given sampling date (spatial variability) than over the course of one year (temporal variability). Community composition remained essentially unchanged across the first 10 months, indicating that spatial sampling might be more important than temporal sampling when assessing β-diversity patterns in desert soils. However, a major shift in microbial community composition was found following a single precipitation event. This shift in composition was associated with a rapid increase in CO2 respiration and productivity, supporting the view that desert soil microbial communities respond rapidly to re-wetting and that this response may be the result of both taxon-specific selection and changes in the availability or accessibility of organic substrates. Recovery to quasi pre-disturbance community composition was achieved within one month after rainfall.
Similar content being viewed by others
Introduction
A major goal in microbial community ecology is to understand the factors that underlie observed patterns in species distribution. As such, investigating changes in microbial community composition and how these changes relate to physical distance among communities has been the focus of considerable research (e.g. refs 1, 2, 3). Spatial changes in microbial community composition have been linked to four major processes: selection, drift, dispersal and mutation4.
In addition to varying in space, microbial communities are also dynamic over time5. In soils, studies have shown that community composition can change across days6, seasons7 and years8, as a result of both biotic and abiotic processes. However, most investigations on the temporal variability of soil microbial communities have focused on Northern hemisphere temperate environments7,9, where such soils are commonly characterised by high primary productivity, a major driver for compositional variation10. Clearly, more ecologically diverse communities should be studied in order to better understand temporal patterns in microbial diversity.
Deserts constitute the largest biome on Earth, covering around 20% of the global land surface11. Hot desert ecosystems are characterized by scant, erratic and low precipitation inputs, high temperatures with extreme seasonal and diurnal fluctuations in temperature, low nutrient status, high levels of incident ultraviolet (UV) radiation and physical disturbance12. However, despite these environmental constraints, deserts are globally significant, as they sustain ca. 6% of the human population, host many endemic plants and animals and store almost one third of total terrestrial carbon13,14. Nevertheless, the temporal variation of microbial communities has seldom been investigated in deserts3,15,16.
Combining Illumina MiSeq amplicon sequencing of replicated surface-soil samples, chlorophyll a (chla) content, CO2 respiration rates and environmental (climate and soil chemistry) data, we investigated the temporal variability of bacterial and archaeal communities and their associated activities and productivities, over a one-year period in the Namib Desert gravel plains. Because deserts are low nutrient and physically stressful environments with comparatively small species pool sizes17, we hypothesize that the rate of change in α-diversity over time should be lower relative to more benign edaphic ecosystems. We also expect a low β-diversity turnover, which will be accompanied by small changes in activity and productivity. Whether significant temporal shifts in either α- or β-diversity do occur, we hypothesize that these shifts should be linked to changes in soil conditions.
Results and Discussion
We investigated the temporal variability of edaphic microbial communities (bacteria and archaea) in the Namib Desert gravel plains over a one-year period. Macro- and micro-environmental conditions and soil nutrient status data were collected throughout the period. We also linked this information to shifts in microbial activity (CO2 respiration rates) and productivity (chla content).
Soil chemical analysis results showed low nutrient levels (Supp. Table 1), comparable to those previously found in Namib Desert soils18,19,20,21. Soil chemical properties remained stable throughout the year (Fig. 1a), with no significant differences in any of the parameters measured (excluding soil moisture content) according to sampling date or season. Air temperature and relative humidity showed a similar pattern (data not shown). Rainfall during the sampling period was restricted to a single precipitation event (38 mm) on day 325. Rainfall in the Namib Desert is typically highly stochastic, both on temporal and spatial scales22. As expected, the soil moisture contents responded to the single precipitation event by a 100-fold increase between day 298 (30 days pre-rainfall) and day 328 (three days post-rainfall) (Supp. Table 1, Supp. Figure 1).
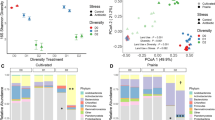
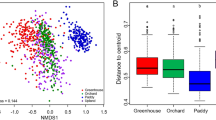
Non-metric multidimensional scaling ordination plots.
(a) Soil chemistry profiles (Euclidean distances with standardized data), excluding moisture content. To appreciate the changes induced by the addition of soil moisture content in the grouping of the samples see Supp. Figure 1. (b) Sequencing data (Bray-Curtis distances after Hellinger transformation). D328, microbial communities 3 days after rainfall; D355, microbial communities 30 days after rainfall. Samples that are closer together are more similar in soil chemistry or microbial community composition.
Respiration rates were low for most of the year (mean 0.022 mg CO2-C g−1 soil hour−1; SE = 0.001), but were ca. 4-fold higher 3 days after rainfall (mean 0.079 mg CO2-C g−1 soil hour−1; SE = 0.015) and remained well above average on day 355 (mean 0.065 mg CO2-C g−1 soil hour−1; SE = 0.007), 30 days after the rainfall event. Rainfall induced CO2 pulses and increases in other microbial processes, resulting from resurrection of microbial communities and changing accessibility to organic substrates, have been shown to occur in Mediterranean23,24 and desert soil environments25, both of which have long periods of drought interspaced with precipitation events. The cause of this phenomenon, which has become known as the “Birch effect” (see ref. 23 and references therein), is not completely understood, but the likely contributory biophysical conditions are: (1) drying and rewetting shatters soil aggregates and exposes previously unavailable organic substrates for decomposition; (2) microorganisms killed by soil drying are decomposed on rewetting to release their nutrients; (3) there is a spontaneous rapid increase in microbial biomass in response to the availability of water perhaps associated with the breaking of dormancy; and (4) there is a microbial hypo-osmotic stress response. These mechanisms are not mutually exclusive26. Irrespective of the mechanisms and whether they operate simultaneously, with the projected intensification of intra-annual precipitation in desert systems27, our results suggest that these environments may become increasing sources of net CO2 output, potentially having a positive (and possibly detrimental) feedback on atmospheric CO2-linked global change processes.
Chla content concentrations (a proxy for productivity) were positively correlated with respiration rates (Spearman p = 0.5, P < 0.05). The average chla concentration in pre-rainfall communities was 0.13 mg chla mg-¹ soil, but 1.96 mg chla mg−¹ soil for post-rainfall communities. Positive relationships between microbial activity, productivity and moisture content have been shown across different soil types28. The increase in soil moisture is probably altering conditions for desert soil microbial communities, mobilizing nutrients and salts and stimulating primary productivity by cyanobacterial populations13. Indeed, cyanobacteria have been reported to require minimum soil moisture level of 15% to be photosynthetically active29.
Amplicon sequencing analysis of the 102 samples collected over 12 months yielded a total of 2170 OTUs (97% identity cut-off) from 33456 sequences. The number of OTUs per sample (α-diversity) ranged from 125 to 208, values similar to those reported for Atacama Desert soils30. We do not expect this variability to be related to sequencing depth, as all samples were rarefied to the same number of reads (328). While we do not have a complete justification for the relatively low number of reads, similar results have been obtained in other nutrient-limited environments31,32,33. Additionally, one focus of the study is on β-diversity and it has been shown that shallow sequencing captures similar β-diversity patterns compared to deep sequencing34. This is because β-diversity is less affected by the number of samples reads, as in the case of α-diversity.
No significant correlations between α-diversity and any of the environmental variables were observed (not shown). A possible explanation for this observation is that α-diversity in these soils may be driven by unmeasured biotic and abiotic variables. As hypothesized, despite some differences between samples collected at a given time, the temporal variability of α-diversity was very low (Fig. 2), indicating that spatial variability in richness may exceed temporal variability in desert soil microbial communities. This is not an unexpected result, as soils are typically spatially heterogeneous, but is not in agreement with results of temporal studies performed in other terrestrial9 and aquatic35 ecosystems. The contrast in α-diversity patterns between this and previous studies is probably related primarily to the aridity, because in the absence of water, transport of microorganisms and nutrients will be limited.
The slope value (0.39) of the species-time relationship, an index of temporal turnover, was within the lower range (0.24–0.61) found for microbial communities in a meta-analysis5. This suggests that the taxa present at this location do not change appreciably over time, in accordance with previous results36. This low turnover could be a product of slow growth rates overall, the low degree of variability in soil properties, low regional species pool size, or perhaps because of the high degree of dormancy in these microbial communities37.
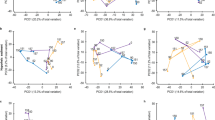
As for α-diversity, β-diversity values (Bray-Curtis dissimilarities ca. 0.63) did not change significantly over time (Supp. Figure 2), even after the rainfall event and the consequent increase in soil moisture and productivity. An increase in β-diversity with rising moisture38 and productivity10 is frequently observed in microbial communities. A possible reason for the absence of a similar pattern in these communities might be that rainfall acted as a selection factor affecting more members of the community, which may result in lower β-diversity values39. Furthermore, the insignificant time-decay relationship confirmed that the microbial communities remained largely unchanged for 10 of the 12 months (Fig. 3), from the beginning of the study until the last sampling point before the rainfall event. Microbial community shifts commonly result from changes in environmental variables40, which remained relatively constant in the Namib Desert during 10 months of the 12-month sampling period. Altogether, these results suggest that environmental filtering, as a consequence of low water content and nutrient concentrations, is critical in shaping these microbial communities.
Temporal decay curves for microbial communities.
The blue line denotes the regression for communities sampled before the rainfall event. The red line denotes a separate regression for all sampling points. Both lines were obtained using the generalized dissimilarity model of Millar et al.58. Only the slope of the solid red line, including the samples after the rainfall event, is significantly less than zero.
The dataset indicated that the single rainfall event greatly affected the soil microbial communities. A nonmetric multidimensional scaling plot showed differences in community structure between pre- and post-rainfall samples (Fig. 1b). PERMANOVA: P < 0.05 in all pairwise comparison including D328), which resulted in a significant decrease in the microbial community similarity with time over the full year (Fig. 3). This is in line with a recent study that showed that microbial communities from the Namib Desert responded rapidly to (intense) water inputs18. Furthermore, post-rainfall samples formed two separated clusters (Fig. 1b). The first cluster included samples obtained 3 days after the precipitation event (day 328), whereas the second corresponded to those collected 30 days after the precipitation event (day 355). Interestingly, the latter communities (day 355) were more similar to those from pre-rainfall samples than to those from 3 days post-rainfall (D328), suggesting that the microbial community structure tended to revert to the pre-disturbance status, a sign of microbial community resilience41. The unique clustering of the microbial communities 3 days after the precipitation event was driven both by the detection of 137 new taxa (Supp. Figure 3) and by changes in the relative abundance of taxa previously identified (see below). Many rare taxa that are below the detection limit can become detectable when dominant species are reduced in their relative abundance. Alternatively, this high short-term community turnover may be a consequence of stochastic dispersal from the surrounding environment42.
A total of 19 bacterial phyla and 2 archaeal phyla were identified in all samples. Microbial communities were dominated by Bacteroidetes (29% of all sequences), Proteobacteria (23%, mostly Alphaproteobacteria (17%) and Betaproteobacteria (3%)) and Actinobacteria (22%); followed by Firmicutes (4%), Acidobacteria (4%), Chloroflexi (4%) and Verrucomicrobia (3%). Archaea (mainly phylum Crenarchaeota) contributed 2% of the sequences. Similar results have been found in other studies17,43,44, indicating that common members of these phyla are probably well adapted to survive and possibly thrive in desert soils20. The relative abundance of Proteobacteria (mainly Betaproteobacteria, Supp. Figure 4) and Actinobacteria increased after the single rainfall event (Fig. 4). Previous studies have also observed that Actinobacteria9 and Betaproteobacteria45 tend to dominate with increasing moisture. This may reflect the nutritional preferences of these two groups, as members of the Actinobacteria and Betaproteobacteria seem to respond positively to carbon availability46, which is thought to increase with soil moisture47. Nevertheless Bacteroidetes, also proposed as a copiotrophic group46, showed the opposite trend in our data, decreasing with moisture (Fig. 4). Bacteroidetes were also shown to decrease in biocrust communities following a simulated rainfall event43. Overall, these apparently contradictory results can be explained in light of recent investigations showing that physiological traits and not phylogeny are better predictors of moisture preferences48.
Taxonomic distribution, phylum level, of bacterial OTUs (97% cutoff).
Affiliation was performed using the Ribosomal Database Project Classifier with a confidence threshold of 80%. Pre-rainfall, microbial communities before the rainfall event; D328, microbial communities 3 days after rainfall; D355, microbial communities 30 days after rainfall.
In summary, we have demonstrated low temporal variability over a year period in desert soil microbial communities. Although a major shift in microbial community composition was induced by a precipitation event. The precipitation-induced shifts in microbial community composition increased CO2 respiration and productivity, demonstrating that desert soil microbial communities can respond to water addition. The major community shift after wetting accompanied by the occurrence of increased soil respiration rates and chla concentrations is consistent with a multi-mechanisms response, involving both taxon-specific selection and changes in the availability or accessibility of organic substrates. In general, the variability in microbial communities within the sampling location (spatial variability) at a given time was higher than the intra-annual temporal variability, suggesting that sampling more sites (preferably at different locations) might be more important than frequent sampling if the aim is to describe compositional changes in desert soil microbial communities and their implications for ecosystem functioning. Nevertheless, collecting time series of soil microbial communities is important to further understanding community stability, given ongoing global climate change disturbances.
Materials and Methods
Study site description and sampling strategy
Soil samples were collected between 08:00 and 10:00 (am) from the gravel plains in the central Namib Desert (S23° 33.302', E15° 3.288) near Gobabeb over a 12-month period (from 04 May 2012 to 28 April 2013). The study area (8100 m2) consisted of 81 (10 × 10 m) plots on a south-facing 2º slope.
Using a random number generator, eight plots were sampled during each of the 16 sampling campaigns (days 0, 4, 12, 28, 42, 57, 88, 118, 138, 178, 198, 238, 268, 298, 328, 355). From each plot, surface soil samples (0–10 cm) were collected using a 1-m2 grid divided into 16 quadrats. These samples were pooled and homogenized to obtain a single sample. Soil samples were maintained at −20 °C in sterile Whirl-Pak bags (Nasco International, Fort Atkinson, WI, USA). A total of 128 samples were collected (16 sampling campaigns each consisting of 8 samples).
Macroenvironmental variables
Air temperature and relative humidity data were obtained from the Gobabeb Land Surface Temperature (LST) weather station using thermal infrared satellite measurements. The station was established by the Karlsruhe Institute of Technology7 in the central Namib Desert gravel plains (23°33′S, 15°03′E), located 400 m above sea level (350 m west of the study site). The station instruments were mounted at 2 m, measuring air temperature and relative humidity at 1-minute intervals. Rainfall data was obtained from the Gobabeb Research and Training Centre located 1.5 km east of the study area (http://www.gobabebtrc.org/).
Soil physicochemical characterisation
Analysis of soil samples was conducted at the Soil Science Laboratory of the University of Pretoria, South Africa in accordance with the standard procedures (SSSA, 1996). Prior to analysis, soil samples were sieved (2 mm) and dried overnight at 50 °C.
The slurry technique was used to measure pH (1:3 soil/deionised water) with a Crison Bench pH meter (Crison Instruments, Barcelona, Spain) after allowing soil to settle for 30 min. Soil N-NO3-, N-NH4+ and cation exchange capacity (CEC) were determined by extraction (2M KCl) with subsequent titration. Total P was measured using the P Bray method and total C was measured using the Walkley-Black method. Ammonium acetate extraction was used to measure salt concentrations (K+, Ca2+, Mg2+, Na+), analysed using inductively coupled plasma atomic emission spectroscopy (ICP-OES; Spectro Genesis, Spectro Analytical Instruments GmbH, Germany). Moisture content was determined by oven drying 10 g of soil at 60 °C for 48 h and comparing the weight of soil pre- and post-drying.
Soil respiration (decomposition)
Between 8.5 and 10.5 g soil were incubated for up to 10 days in a conductiometric respirometer (Nordgren, 1988) at 20˚C in the dark and CO2 production measured at hourly intervals. CO2 production data were used to estimate soil respiration rate by linear regression of the CO2 over the first 2–3 days. Linear regression was significant (P < 0.05) in all cases. The mean and standard errors were calculated for all replicates after the replicates that gave the largest and smallest estimates had been removed. In no case was the mean estimate dependent on fewer than three replicates. The extreme replicates were removed because observable fragments of organic matter or salt encrustations led to anomalously high or unreproducibly low respiration rates, which in neither case produced statistically significant linear regressions of CO2 production with time.
Chlorophyll a content (productivity)
Chla extraction was performed on frozen (−20 °C) soil samples using a modification of the method described by Richie et al. Briefly, 3 g of soil was added to a sterile glass vial containing 8 ml 90% ethanol and incubated at 72 °C for 10 min and subsequently sonicated at 100% amplitude for 2 min. The supernatant was passed through a 0.2 mm filter into a glass test tube and the absorbance measured at 665 nm. Background fluorescence was determined by measuring absorbance at 750 nm. Chla absorbance was calculated according to the formula

where: E665 = Absorbance at 665 nm, E750 = Absorbance at 750 nm, V = volume of extraction solution, λ = 11.9035 the cyanobacterial-specific coefficient and gSoil corresponds to the mass of soil49.
DNA extraction and 16S rRNA gene amplicon barcoded sequencing
Metagenomic DNA was extracted from 0.25 g of soil using the PowerSoil®- htp 96-Well Soil DNA Isolation (MoBio, Carlsbad, CA, USA) following manufacturer’s instructions. In total, DNA was extracted from 128 samples according to the methods used by the Earth Microbiome Project (http://www.Earthmicrobiome.Org/emp-standard-protocols/dna-extraction-protocol/). Sample sequencing was performed using bacterial/archaeal 515f and 806r error-correcting barcoded primers as described previously50. Illumina MiSeq sequence data was screened using the QIIME toolkit51 with the following parameters: quality score >25, sequence length >200 and <400, maximum homopolymer of 6, 0 maximum ambiguous bases and 0 mismatched bases in the primers and barcodes. Sequence counts before and after quality filtering were 348687 and 124597, respectively. Following chimera removal (USEARCH 6.152), 115102 sequences remained. OTUs were picked at the 97% identity level using UCLUST52 with the default settings. Phylogenetic identification was performed against the most recent Greengenes database (gg_13_5)53. After removal of singletons, samples were rarefied to 328 reads, which represents the lowest number of sequences obtained for a single sample. In total 26 samples, which failed quality control, were excluded from further analysis, yielding a total of 102 samples.
Data analysis
Biotic and abiotic data sets were analysed using the R environment for statistical computing (version 3.0.354) using vegan55 and custom scripts. Differences in abiotic data were assessed using Wilcoxon-Mann-Whitney post hoc tests after ensuring that an overall Kruskal-Wallis test was significant (P < 0.05). Microbial community similarity was calculated using the Bray-Curtis index and visualised using non-metric multidimensional scaling (NMDS). Permutational multivariate analysis of variance (PERMANOVA) was used to test for statistically significant variance between groups56.
Richness trend analysis was performed using generalized least squares (GLS) with AR1 temporal autocorrelation of errors57. Distance-decay relationships between microbial community composition and temporal distance were plotted using the generalized dissimilarity model of Millar et al.58. Species-time relationship (STR) was constructed in R by calculating richness using the moving window approach of White et al.59. This approach was used to allow comparison with the results from microbial communities reported in shade et al.5. Analyses were performed on Hellinger transformed sequence data and on standardized environmental data.
Additional Information
Accession codes: The sequence data generated in this study were deposited in the NCBI Sequence Read Archive and are available under the project number: SRP066886.
How to cite this article: Armstrong, A. et al. Temporal dynamics of hot desert microbial communities reveal structural and functional responses to water input. Sci. Rep. 6, 34434; doi: 10.1038/srep34434 (2016).
References
Soininen, J., Lennon, J. J. & Hillebrand, H. A multivariate analysis of beta diversity across organisms and environments. Ecology 88, 2830–2838, doi: 10.1890/06-1730.1 (2007).
Martiny, J. B. H., Eisen, J. A., Penn, K., Allison, S. D. & Horner-Devine, M. C. Drivers of bacterial beta-diversity depend on spatial scale. Proc Nat Acad Sci USA 108, 7850–7854, doi: 10.1073/pnas.1016308108 (2011).
Bell, C., McIntyre, N., Cox, S., Tissue, D. & Zak, J. Soil microbial responses to temporal variations of moisture and temperature in a Chihuahuan Desert Grassland. Microb Ecol 56, 153–167, doi: 10.1007/s00248-007-9333 (2008).
Hanson, C. A., Fuhrman, J. A., Horner-Devine, M. C. & Martiny, J. B. H. Beyond biogeographic patterns: processes shaping the microbial landscape. Nat Rev Microbiol 10, 497–506, doi: 10.1038/nrmicro2795 (2012).
Shade, A., Gregory Caporaso, J., Handelsman, J., Knight, R. & Fierer, N. A meta-analysis of changes in bacterial and archaeal communities with time. ISME J 7, 1493–1506, doi: 10.1038/ismej.2013.54 (2013).
Noll, M., Matthies, D., Frenzel, P., Derakshani, M. & Liesack, W. Succession of bacterial community structure and diversity in a paddy soil oxygen gradient. Environ Microbiol 7, 382–395, doi: 10.1111/j.1462-2920.2004.00700 (2005).
Rasche, F. et al. Seasonality and resource availability control bacterial and archaeal communities in soils of a temperate beech forest. ISME J 5, 389–402, doi: 10.1038/ismej.2010.138 (2011).
Moore-Kucera, J. & Dick, R. P. PLFA profiling of microbial community structure and seasonal shifts in soils of a Douglas-fir chronosequence. Micro Ecol 55, 500–511, doi: 10.1007/s00248-007-9295-1 (2008).
Lauber, C. L., Ramirez, K. S., Aanderud, Z., Lennon, J. & Fierer, N. Temporal variability in soil microbial communities across land-use types. ISME J 7, 1641–1650, doi: 10.1038/ismej.2013.50 (2013).
Langenheder, S., Berga, M., Ostman, O. & Szekely, A. J. Temporal variation of beta-diversity and assembly mechanisms in a bacterial metacommunity. ISME J 6, 1107–1114, doi: 10.1038/ismej.2011.177 (2012).
Middleton, N. & Thomas, D. World Atlas of Desertification, 2nd edn. London, UK: UNEP (1997).
Lester, E. D., Satomi, M. & Ponce, A. Microflora of extreme arid Atacama Desert soils. Soil Biol Biochem 39, 704–708, doi: 10.1016/j.soilbio.2006.09.020 (2007).
Pointing, S. B. & Belnap, J. Microbial colonization and controls in dryland systems. Nat Rev Microbiol, 10, 551–562, doi: 10.1038/nrmicro2831 (2012).
Makhalanyane, T. P. et al. Microbial ecology of hot desert edaphic systems. FEMS Microbiol Rev 39, 203–221, doi: 10.1093/femsre/fuu011 (2015).
Saul-Tcherkas, V., Unc, A. & Steinberger, Y. Soil Microbial Diversity in the Vicinity of Desert Shrubs. Micro Ecol 65, 689–699, doi: 10.1007/s00248-012-0141-8 (2013).
Sher, Y., Zaady, E. & Nejidat, A. Spatial and temporal diversity and abundance of ammonia oxidizers in semi-arid and arid soils: indications for a differential seasonal effect on archaeal and bacterial ammonia oxidizers. FEMS Microbiol Ecol 86, 544–556, doi: 10.1111/1574-6941.12180 (2013).
Fierer, N. et al. Cross-biome metagenomic analyses of soil microbial communities and their functional attributes. Proc Nat Acad Sci USA 109, 21390–21395, doi: 10.1073/pnas.1215210110 (2012).
Frossard, A., Ramond, J. B., Seely, M. & Cowan, D. A. Water regime history drives responses of soil Namib Desert microbial communities to wetting events. Sci Rep 5, 571–578, doi: 10.1038/srep12263 (2015).
Gombeer, S., Ramond, J. B., Eckardt, F. D., Seely, M. & Cowan, D. A. The influence of surface soil physicochemistry on the edaphic bacterial communities in contrasting terrain types of the Central Namib Desert. Geobiology 13, 494–505, doi: 10.1111/gbi.12144 (2015).
Makhalanyane, T. P. et al. Evidence of species recruitment and development of hot desert hypolithic communities. Environ Microbiol Rep. 5, 219–224 (2013).
Stomeo, F. et al. Hypolithic and soil microbial community assembly along an aridity gradient in the Namib Desert. Extremophiles 17, 329–337 (2013).
Eckardt, F. D. et al. The nature of moisture at Gobabeb, in the central Namib Desert. J Arid Environ 93, 7–19, doi: 10.1016/j.jaridenv.2012.01.011 (2013).
Jarvis, P. et al. Drying and wetting of Mediterranean soils stimulates decomposition and carbon dioxide emission: The "Birch effect". Tree Physiol, 27, 929–940 (2007).
Placella, S. A., Brodie, E. L. & Firestone, M. K. Rainfall-induced carbon dioxide pulses result from sequential resuscitation of phylogenetically clustered microbial groups. Proc Nat Acad Sci USA 109, 10931–10936, doi: 10.1073/pnas.1204306109 (2012).
Talmon, Y., Sternberg, M. & Grünzweig, J. M. Impact of rainfall manipulations and biotic controls on soil respiration in Mediterranean and desert ecosystems along an aridity gradient. Global Change Biol 17, 1108–1118, doi: 10.1111/j.1365-2486.2010.02285 (2011).
Dungait, J. A. J., Hopkins, D. W., Gregory, A. S. & Whitmore, A. P. Soil organic matter turnover is governed by accessibility not recalcitrance. Global Change Biol 18, 1781–1796, doi: 10.1111/j.1365-2486.2012.02665 (2012).
[IPCC]. Intergovernmental Panel on Climate Change 2007. Climate Change 2007. The Physical Science Basis. Summary for Policymakers New York Cambridge University Press. (2007).
Manzoni, S., Schimel, J. P. & Porporato, A. Responses of soil microbial communities to water stress: Results from a meta-analysis. Ecolog 93, 930–938, doi: 10.1890/11-0026.1 (2012).
Tracy, C. R. et al. Microclimate and limits to photosynthesis in a diverse community of hypolithic cyanobacteria in northern Australia. Environ Microbiol 12, 592–607, doi: 10.1111/j.1462-2920.2009.02098 (2010).
Crits-Christoph, A. et al. Colonization patterns of soil microbial communities in the Atacama Desert. Microbiome 1, doi: 10.1186/2049-2618-1-28 (2013).
Zablocki, O. et al. High-level diversity of tailed Phages, eukaryote-associated viruses and virophage-like elements in the metaviromes of Antarctic soils. App Environ Microbiol 80, 6888–6897, doi: 10.1128/AEM.01525-14 (2014).
Lee, C. K., Barbier, B. A., Bottos, E. M., McDonald, I. R. & Cary, S. C. The inter-valley soil comparative survey: the ecology of Dry Valley edaphic microbial communities. ISME J 6, 1046–1057, doi: 10.1038/ismej.2011.170 (2012).
Makhalanyane, T. P. et al. Evidence for successional development in Antarctic hypolithic bacterial communities. ISME J, 7, 2080–2090, doi: 10.1038/ismej.2013.94 (2013).
Mariadassou, M., Pichon, S. & Ebert, D. Microbial ecosystems are dominated by specialist taxa. Ecol Lett 18, 974–982, doi: 10.1111/ele.12478 (2015).
Portillo, M. C., Anderson, S. P. & Fierer, N. Temporal variability in the diversity and composition of stream bacterioplankton communities. Environ Microbiol 14, 2417–2428, doi: 10.1111/j.1462-2920.2012.02785 (2012).
Fierer, N. & Jackson, R. B. The diversity and biogeography of soil bacterial communities. Proc Nat Acad Sci USA 103, 626–631, doi: 10.1073/pnas.0507535103 (2006).
Lennon, J. T. & Jones, S. E. Microbial seed banks: the ecological and evolutionary implications of dormancy. Nat Rev Microbiol 9, 119–130, doi: 10.1038/nrmicro2504 (2011).
Zimmerman, N. B. & Vitousek, P. M. Fungal endophyte communities reflect environmental structuring across a Hawaiian landscape. Proc Nat Acad Sci USA 109, 13022–13027, doi: 10.1073/pnas.1209872109 (2012).
Zhou, J. et al. Stochasticity, succession and environmental perturbations in a fluidic ecosystem. Proc Nat Acad Sci USA 111, E836–E845, doi: 10.1073/pnas.1324044111 (2014).
van der Gast, C. J., Ager, D. & Lilley, A. K. Temporal scaling of bacterial taxa is influenced by both stochastic and deterministic ecological factors. Environ Microbiol 10, 1411–1418, doi: 10.1111/j.1462-2920.2007.01550 (2008).
Allison, S. D. & Martiny, J. B. H. Resistance, resilience and redundancy in microbial communities. Proc Nat Acad Sci USA 105, 11512–11519, doi: 10.1073/pnas.0801925105 (2008).
Ferrenberg, S. et al. Changes in assembly processes in soil bacterial communities following a wildfire disturbance. ISME J 7, 1102–1111, doi: 10.1038/ismej.2013.11 (2013).
Angel, R. & Conrad, R. Elucidating the microbial resuscitation cascade in biological soil crusts following a simulated rain event. Environ Microbiol 15, 2799–2815, doi: 10.1111/1462-2920.12140 (2013).
Valverde, A., Makhalanyane, T. P., Seely, M. & Cowan, D. A. Cyanobacteria drive community composition and functionality in rock-soil interface communities. Mol Ecol 24, 812–821, doi: 10.1111/mec.13068 (2015).
Lennon, J. T., Aanderud, Z. T., Lehmkuhl, B. K. & Schoolmaster Jr, D. R. Mapping the niche space of soil microorganisms using taxonomy and traits. Ecology 93, 1867–1879, doi: 10.1890/11-1745.1 (2012).
Fierer, N., Bradford, M. A. & Jackson, R. B. Toward an ecological classification of soil bacteria. Ecology, 88, 1354–1364, doi: 10.1890/05-1839 (2007).
Knorr, W., Prentice, I. C., House, J. I. & Holland, E. A. Long-term sensitivity of soil carbon turnover to warming. Nature, 433, 298–301, doi: 10.1038/nature03226 (2005).
Evans, S. E. & Wallenstein, M. D. Climate change alters ecological strategies of soil bacteria. Ecol Let 17, 155–164, doi: 10.1111/ele.12206 (2014).
Ritchie, R. J. Consistent sets of spectrophotometric chlorophyll equations for acetone, methanol and ethanol solvents. Photosynth Res 89, 27–41, doi: 10.1007/s11120-006-9065-9 (2006).
Gilbert, J. A., Jansson, J. K. & Knight, R. The Earth Microbiome project: successes and aspirations. BMC Biol 12, 1 (2014).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat Met 7, 335–336, doi: 10.1038/nmeth.f.303 (2010).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461, doi: 10.1093/bioinformatics/btq461 (2010).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol, 72, 5069–5072, doi: 10.1128/aem.03006-05 (2006).
R Development Core Team. (2013). R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, http://www.R-project.org
Oksanen, J. et al. Vegan: Community Ecology Package http://CRAN.R-project.org/package=vegan (2013).
Anderson, M. J. A new method for non-parametric multivariate analysis of variance. Austral Ecol 26, 32–46, doi: 10.1111/j.1442-9993.2001.01070 (2001).
Dornelas, M. et al. Quantifying temporal change in biodiversity: challenges and opportunities. Proc R Soc B Biol 280, doi: 10.1098/rspb.2012.1931 (2013).
Millar, R. B., Anderson, M. J. & Tolimieri, N. Much ado about nothings: using zero similarity points in distance-decay curves. Ecology 92, 1717–1722 (2011).
White, E. P. et al. A comparison of the species-time relationship across ecosystems and taxonomic groups. Oikos 112, 185–195 (2006).
Acknowledgements
We gratefully acknowledge financial support from the National Research Foundation of South Africa (grant no. 81779 and TTK2008052000003), the Research Council of Norway (grant No. 180352) and the University of the Western Cape. Partial support was also provided under the Laboratory Directed Research and Development Program at PNNL, a multiprogram national laboratory operated by Battelle for the U.S. Department of Energy under contract DE-AC05-76RL01830. We would also like to thank the Earth Microbiome Project (www.earthmicrobiome.org) for sequencing of the samples.
Author information
Authors and Affiliations
Contributions
J.-B.R., A.V. and D.A.C. designed research. A.A., J.-B.R. and T.P.M. performed research. A.A., T.P.M. and A.V. analysed the microbial data. T.J.A. and D.H. analysed CO2 data. D.A.C., D.W.H., M.S. and M.I.T. contributed reagent/materials/analysis tools. J.K.J., was responsible for DNA extraction and sequencing through the Earth Microbiome Project. A.V. and A.A. wrote the manuscript. All authors commented on the manuscript at all stages.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Armstrong, A., Valverde, A., Ramond, JB. et al. Temporal dynamics of hot desert microbial communities reveal structural and functional responses to water input. Sci Rep 6, 34434 (2016). https://doi.org/10.1038/srep34434
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep34434
This article is cited by
-
Microbial growth under drought is confined to distinct taxa and modified by potential future climate conditions
Nature Communications (2023)
-
Use of an automated respirometer for in situ chemical oxidation (ISCO) activator type and concentration selection
Environmental Science and Pollution Research (2022)
-
Differences in Precipitation Regime Shape Microbial Community Composition and Functional Potential in Namib Desert Soils
Microbial Ecology (2022)
-
Biogeographical survey of soil microbiomes across sub-Saharan Africa: structure, drivers, and predicted climate-driven changes
Microbiome (2022)
-
Agricultural management and cultivation period alter soil enzymatic activity and bacterial diversity in litchi (Litchi chinensis Sonn.) orchards
Botanical Studies (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.