Abstract
Complex III in C. glutamicum has an unusual di-heme cyt. c1 and it co-purifies with complex IV in a supercomplex. Here, we investigated the kinetics of electron transfer within this supercomplex and in the cyt. aa3 alone (cyt. bc1 was removed genetically). In the reaction of the reduced cyt. aa3 with O2, we identified the same sequence of events as with other A-type oxidases. However, even though this reaction is associated with proton uptake, no pH dependence was observed in the kinetics. For the cyt. bc1-cyt. aa3 supercomplex, we observed that electrons from the c-hemes were transferred to CuA with time constants 0.1–1 ms. The b-hemes were oxidized with a time constant of 6.5 ms, indicating that this electron transfer is rate-limiting for the overall quinol oxidation/O2 reduction activity (~210 e−/s). Furthermore, electron transfer from externally added cyt. c to cyt. aa3 was significantly faster upon removal of cyt. bc1 from the supercomplex, suggesting that one of the c-hemes occupies a position near CuA. In conclusion, isolation of the III-IV-supercomplex allowed us to investigate the kinetics of electron transfer from the b-hemes, via the di-heme cyt. c1 and heme a to the heme a3-CuB catalytic site of cyt. aa3.
Similar content being viewed by others
Introduction
The respiratory chain in aerobic organisms is composed of a number of membrane-bound protein complexes through which electrons, originating from the oxidation of organic compounds, are transferred to finally reach O2. The free energy released in this process is employed to establish a proton electrochemical gradient across the membrane, which is used to synthesize ATP by the F1FO ATP synthase or for secondary transmembrane transport. In mitochondria, Complex III (the cytochrome (cyt.) bc1-complex) of the respiratory chain links the two-electron oxidation of quinol (QH2) to the one-electron reduction of water-soluble cyt. c in the respiratory chain:

where the subscripts N and P refer to the more negative and positive sides of the membrane, respectively. Reduced cyt. c delivers electrons to Complex IV (cytochrome c oxidase, CytcO), which catalyzes the reduction of dioxygen to water:

Part of the respiratory-chain enzymes in mitochondria are organized in so-called supercomplexes1,2,3,4,5,6,7,8,9,10,11. There are also reports of supercomplexes in bacteria, for example in Paracoccus denitrificans. In this bacterium, depending on the detergent used, supercomplexes composed of respiratory-enzyme complexes III-IV or I-III-IV at variable stoichiometries were identified1,12,13,14. Furthermore, in several bacterial systems electrons could be transferred directly between complexes III and IV via a membrane-anchored cyt. c15,16,17.
Corynebacterium glutamicum is a rod-shaped, Gram positive soil bacterium, which harbors two different terminal oxidases; an aa3-type CytcO and a bd-type menaquinol oxidase18,19,20. The cyt. aa3 in C. glutamicum is a four-subunit protein complex, comprising subunits CtaD, C, E and F. Mass spectrometric analyses of the purified CytcO revealed that instead of heme a, the C. glutamicum contains heme as in the active site. Furthermore, the C. glutamicum CytcO harbors an extra charged amino-acid cluster near the cyt. c-binding domain of subunit II (CtaC), which was suggested to interact with the second cyt. c of the cyt. bc1 complex18.
Complex III in C. glutamicum is a three-subunit protein, containing cyt. c1 (QcrC), the Rieske iron-sulfur protein (QcrA) and cyt. b (QcrB) (Fig. 1A)21,22. The QcrC subunit contains two CXXCH heme-binding motifs, suggesting that this protein complex contains two c-type hemes19,22, hence referred to as the di-heme c1 cyt. bc1 complex. Furthermore, C. glutamicum contains no other c-type hemes, which suggested that the second heme c in cyt. bc1 shuttles electrons between complexes III and IV that form a tight supercomplex21. Such a supercomplex was isolated, its shape was determined using electron microscopy23 and it was shown to exhibit quinol-oxidase activity20:
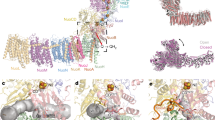
Schematic model of the C. glutamicum cyt. bc1-CytcO supercomplex and a homology model of subunit I in CytcO.
(A) Cyt. bc1 (in blue) and CytcO (in green), represented as a supercomplex. The electron-transfer pathway is indicated by black bold arrows. Reduction of O2 at the heme a3-CuB catalytic site is accompanied by proton uptake from solution. The Figure is adapted from19. (B,C) A structural model of subunit I of CytcO from C. glutamicum (B,C, top and side view, respectively), based on the structure of aa3 (PDB ID 1M56) from R. sphaeroides39. Water molecules found in the structure of the R. sphaeroides CytcO are shown in the model to indicate the location of the D proton pathway (some residues defining this pathway are shown in green, Glu267 in magenta). Tyr269 is presumably part of the catalytic site. The putative K-pathway residue Lys341 (equivalent to Lys362 in R. sphaeroides CytcO) is shown as a yellow stick.

The involvement of the second cyt. c of cyt. bc1 in electron transfer between cyt. c1 and CuA, the electron acceptor of CytcO (see below), is also supported by mutagenesis data20. Quinol-oxidase activity was also found for the cyt. bc1-aa3 supercomplex from Mycobacterium smegmatis24, a bacterium also devoid of soluble cyt. c.
The mechanism of the cyt. bc1 complex involves a Q-cycle in which the net reaction results in oxidation of menaquinol and reduction of cyt. c (see Fig. 1A), linked to proton uptake from the N side and release to the P side of the membrane. The process is initiated by binding of a menaquinol at the quinone-binding site, QP, located near the low-potential heme bL (see Fig. 1A)25. In the next step, one electron is transferred from the menaquinol, via the Rieske protein [2Fe–2S] cluster, to cyt. c1 while one electron is transferred via heme bL and heme bH to a menaquinone bound at a second quinone-binding site, QN. This bifurcated electron transfer yields reduced cyt. c1, a semireduced menaquinone at the QN-site and release of two protons to the P-side of the membrane. After binding a second menaquinol at the QP-site the same process is repeated. The doubly reduced menaquinone at the QN site picks up two protons from the N-side of the membrane to form a menaquinol that is released into the membrane.
Cytochrome c oxidase, which belongs to a large family of enzymes called the heme-copper oxidases, catalyzes oxidation of cyt. c and reduction of O2 to H2O. Here, we refer to the C. glutamicum cytochrome aa3 as a CytcO even though in the cyt. bc1-cyt. aa3 supercomplex it receives electrons from quinol, via the cyt. bc1 complex and not from (a water-soluble) cyt. c. The heme-copper oxidases are classified according to sequence, phylogenetic and structural analyses into three main classes, A, B and C26,27. Subunit I, the core subunit shared by all of the three oxidase types, contains a low-spin heme group and the catalytic site, which is composed of a copper ion, CuB and a high-spin heme (for review on structure and function of the CytcOs, see28,29,30,31,32,33,34,35,36,37). The fourth redox-active site, CuA, is found in subunit II. The most studied CytcOs are those from bovine heart mitochondria and the bacterial aa3 CytcOs from Paracoccus denitrificans and Rhodobacter sphaeroides, which all belong to the A-class. In these CytcOs, electrons delivered by cyt. c are transferred consecutively to the CuA site, heme a and finally to the binuclear center composed of heme a3 and CuB. In these bacterial CytcOs protons are transferred to the catalytic site through two pathways denoted by the letters D and K after conserved residues Asp132 and Lys362, respectively (numbering refers to the R. sphaeroides aa3-type CytcO).
Internal electron and proton-transfer reactions in CytcOs from several organisms have been studied in the past (see e.g.30,31,34,37). An experimental technique that yields particularly detailed information about the sequence and rates of these reactions is the so-called flow-flash technique, in which the oxidative part of a reaction cycle (single turnover) of the enzyme is monitored. In this approach, the oxidase is first fully reduced by four electrons (see dashed line in Fig. 2) and incubated under an atmosphere of carbon monoxide, which binds to heme a3 at the catalytic site. The CytcO-CO complex is then rapidly mixed with O2-containing solution followed in time by light-induced dissociation of the blocking CO ligand, which allows O2 from the surrounding medium to bind. Initially, the reduced CytcO (the state is called R) binds O2 to heme a3 with a time constant of ~10 μs (at 1 mM O2) yielding the ferrous heme a3-O2 state (state A) (see Fig. 2). Next, an electron is transferred from heme a to the catalytic site and the O=O bond is cleaved, resulting in formation of a ferryl intermediate that is called “peroxy” (PR) for historical reasons (τ ≅ 30 μs). A proton is then taken up to the catalytic site resulting in formation of the ferryl state, F, with a time constant of ~100 μs at pH 7. At the same time, an electron is transferred from CuA to heme a in a small fraction of the population (not shown in Fig. 2). Finally, the last electron is transferred from the CuA-heme a equilibrium to the catalytic site forming the oxidized CytcO (state O) with a time constant of ~1 ms at pH 7 (the time constants are those observed with the R. sphaeroides CytcO38).
Schematic illustration of the reactions in CytcO studied in this work.
The four red circles represent the redox-active metal sites as indicated (filled - reduced). During preparation of the sample, the oxidized CytcO (O) is reduced by 4 electrons under an atmosphere of pure N2 (anaerobic reduction) to yield the fully reduced CytcO (state R). The sample is then incubated under an atmosphere of CO, which binds to heme a3 at the catalytic site forming R-CO (see dashed line). The reaction studied in this work (indicated by a solid line) is started by photo-dissociation of the CO ligand (laser flash), yielding the reduced CytcO (R), which allows O2 to bind to heme a3 forming state A with a time constant of ~10 μs at 1 mM O2. An electron is then transferred from heme a to the catalytic site forming the “peroxy” state called PR with a time constant of ~30 μs. Next, a proton is taken up from solution forming the ferryl state F with a time constant of ~100 μs. In most oxidases studied to date, at the same time the electron at CuA equilibrates with heme a. This electron transfer is not indicated in the figure because in the C. glutamicum CytcO the equilibrium is shifted towards reduced CuA. In the final step of the reaction the electron from CuA (or the CuA-heme a equilibrium) is transferred to the catalytic site forming the oxidized CytcO (state O) in 1–2 ms. In cyt. bc1-CytcO supercomplex, as soon as CuA is partly oxidized (τ ≅ 100 μs), electrons are transferred from the cyt. bc1 complex to the CytcO. Further electron transfer from the c and b hemes occurs over a slower time scale. For simplicity, all these electron transfers are indicated schematically in the lower part of this scheme (the time constants are given in Fig. 8). The pumped protons are not shown.
In the present study we used the flow-flash technique to investigate the reaction of the purified C. glutamicum CytcO, as well as the cyt. bc1-CytcO supercomplex with O2. The data indicate rapid electron transfer from the cyt. bc1-complex to the CytcO, suggesting a functional supercomplex in which the additional cyt. c of the cyt. bc1 complex acts as an electron bridge between the two respiratory-enzyme complexes. Furthermore, a comparison of the TMPD/cyt. c-driven O2-reduction activities with the cyt. bc1-CytcO complex and with CytcO alone, indicate that the second heme c of the cyt. bc1 complex does bind at the surface of the CytcO, presumably near CuA. We also studied the electron-transfer kinetics in purified CytcO and found that reaction steps linked to proton uptake during O2 reduction displayed pH-independent kinetics, suggesting differences in the pKa of residues involved in proton transfer, compared to other A-type CytcOs.
Results
Sequence Alignment and Homology Modeling
To analyze the structural characteristics of CytcO from C. glutamicum, we performed homology modeling of the highly conserved subunit I with the three-dimensional structure of that from the R. sphaeroides aa3-type CytcO39 using the SwissModel program40,41,42 (Fig. 1B,C). The two protein sequences are ~40% identical and ~60% similar. On the basis of the sequence itself and this model, the C. glutamicum protein is identified as an A1-type CytcO. It holds the active site tyrosine (Tyr269 in C. glutamicum)21 and Glu267 in the D pathway (Glu286 in R. sphaeroides). Furthermore, in the C. glutamicum CytcO an Asp residue (Asp116) is found at the same location as Asp132, the entry point of the D pathway in the R. sphaeroides CytcO. Other residues that are discussed below are Asn123 and Asn189 in the C. glutamicum CytcO (Asn139 and Asn207, respectively, in the R. sphaeroides CytcO).
Purification and multiple turnover activity
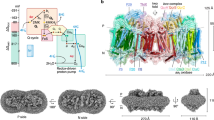
We have purified the cyt. bc1-aa3 supercomplex from strain ΔC-DSt and the aa3 oxidase from the cyt. bc1 deficient strain ΔQ-DSt. The quality of the preparations was assessed using SDS gel electrophoresis (Figure S1) and dithionite-reduced minus ferricyanide-oxidized difference spectra of the two samples (Figure S2). As expected, both spectra showed the heme a signature (605 nm) and the supercomplex preparation displayed additional peaks for b-heme (560 nm) and c-heme (550 nm) (Figure S2). The approximately similar height of the three alpha peaks indicates the presence of a 1:1 complex of cyt. bc1 (2 heme b and 2 heme c) and cytochrome aa3 (two heme a). The reduced CO-bound minus reduced difference spectrum (Figure S3) displayed the characteristic features of CO binding to heme a3.
Next, the quinol-oxidase activity of the purified supercomplex was measured by following O2 consumption after addition of a pre-reduced menaquinone. This process involves quinol oxidation by the cyt. bc1 complex followed by electron transfer to CytcO, where oxygen is reduced to water. The measured quinol-oxidase activity for the purified supercomplex was 210 ± 20 e−/s (SD, n = 4 measurements) (normalized to the total CytcO), i.e. in the same range as the published value20. The activity dropped rapidly upon flash freezing and thawing the preparation. Therefore, the sample was kept at 4 °C, where no activity loss was observed in the time frame between purification and functional studies (typically ~1 day, but the preparation was stable up to 7 days, Figure S4).
CytcO activity of the purified supercomplexes and pure CytcO (without cyt. bc1) was also measured using ascorbate as electron source and either TMPD, or TMPD and water-soluble cyt. c as electron mediators (Fig. 3). For the cyt. bc1-CytcO, the TMPD activity was 90 ± 10 e−/s and it increased to 130 ± 10 e−/s (SD, n = 3) upon addition of free cyt. c, i.e. both rates were lower than the coupled quinol activity. For pure CytcO, the activity increased from 160 ± 10 e−/s to 440 ± 20 e−/s (SD, n = 3) upon addition of cyt. c, thus displaying a much greater stimulation by the soluble electron carrier.
O2-reduction activity upon addition of an electron donor to CytcO.
The measurements were done with either CytcO or the cyt. bc1-CytcO super complex using ascorbate as electron donor and, TMPD and cyt. c as electron mediators. The activity was determined by measuring the O2-reduction rate (e−/s/CytcO). Conditions: 100 mM Tris-HCl pH 7.5, 100 mM NaCl, 2 mM MgSO4, 0.015% (w/v) DDM, 2 mM sodium ascorbate, 0.2 mM TMPD, with or without 24 μM cyt.c.
Kinetics of CO rebinding after flash photolysis
CO binds with high affinity to heme a3 in the binuclear site in the reduced CytcO. Upon illumination with a short laser flash, CO dissociates instantly, but in the absence of oxygen rebinds to the binuclear site (CO recombination). The CO-recombination kinetics was biphasic where the relative contribution of the two components varied slightly between the purified CytcO, the cyt. bc1-CytcO complex and the intact membrane (Fig. 4A,B). This observation indicates the presence of two CytcO populations, also in the native membrane (see Discussion). The time constants for the fast and the slow phases were about the same for all samples (pure CytcO, the purified cyt. bc1-CytcO supercomplex and whole cells), i.e., 11 ± 3 ms and 130 ± 30 ms (SD, n = 15) (at 1 mM CO), respectively. The kinetic difference spectra of the fast and the slow components were similar (Fig. 4C,D).
CO-recombination kinetics and kinetic difference spectra.
Absorbance changes at 445 nm (A) and 605 nm (B) associated with light-induced CO dissociation and recombination with detergent-purified CytcO, cyt. bc1-CytcO and with the ΔC-DSt whole cells. All traces were scaled to yield the same absorbance changes at t = 0. A laser artifact at t = 0 has been removed for clarity. Kinetic difference spectra, i.e. the amplitude of the absorbance changes of the two kinetic components (slow and fast components, empty and filled circles, respectively), for CytcO (C) and the cyt. bc1-CytcO complex (D). Note the different ordinate scales before and after the break on the abscissa, respectively. The black line represents the reduced minus CO-bound difference spectrum. Experimental conditions: 100 mM Tris-HCl at pH 7.5, 1 μM PMS, 5 mM sodium ascorbate. For the cyt. bc1-CytcO supercomplex, 1 mM sodium dithionite was added to reduce the b-hemes.
The apparent binding affinity for CO to the binuclear center of the oxidase was determined by measuring the observed CO-recombination rates for the slower kinetic phase at different CO-concentrations, for both the oxidase alone and the purified supercomplex (Figure S5). The second-order rate constants, determined from a linear fit to the data in Figure S5, were 7.6 ± 0.2 *103 M−1s−1 and 9.4 ± 0.2 *103 M−1s−1 for the CytcO and cyt. bc1-CytcO complex, respectively. The rate of the faster component was CO-concentration independent.
Single-turnover measurements
Figure 5 shows absorbance changes after flash-induced dissociation of the CO ligand from the reduced cyt. bc1-CytcO supercomplex in the presence of O2. At 605 nm (Fig. 5A), three kinetic phases were resolved. The initial decrease in absorbance, with a time constant of ∼25 μs [The standard deviation of the time constants was typically 10% of the measured values (n = 7–17), except for the P → F reaction for which the standard deviation was 20% of the measured values (for both the pure CytcO and the supercomplex)], is attributed to oxidation of heme a, i.e. electron transfer from heme a to heme a3, which yields the PR state at the catalytic site (formation of state A was not resolved, see below). This component is also seen at 445 nm (initial decrease, Fig. 5B). At 605 nm, the decrease is followed by a small increase in absorbance in the time range 0.05–0.2 ms that is attributed to re-reduction of heme a with a time constant of 120 μs concomitant with the PR → F reaction at the catalytic site. The slowest absorbance decrease (τ ≅ 1.7 ms) at 605 nm is associated with oxidation of the CytcO (F → O), also seen at 445 nm (Fig. 5B). At 830 nm, oxidation of CuA is observed as an increase in absorbance. The small initial increase in absorbance is associated with oxidation of CuA during the PR → F transition, i.e. τ ≅ 120 μs while the major oxidation component displayed a time constant of ~1.7 ms (Fig. 5D).
Absorbance changes associated with the reaction of the reduced cyt. bc1-CytcO complex and CytcO with O2.
Absorbance changes were recorded over time at 605 nm (A), 445 nm (B) both associated with oxidation of a-hemes and at 445 nm with the pure CytcO (C) (this trace was multiplied by a factor of two to yield approximately the same absorbance changes as those seen in (B). The absorbance changes at 830 nm (D) are associated with oxidation of CuA, at 550 nm (E) oxidation of cyt. c and at 563 nm (F) oxidation of the b-hemes. The CO ligand was dissociated at t = 0. Experimental conditions: 100 mM Tris-HCl at pH 7.5, 4 mM ascorbate, 1 μM PMS, ~900 μM O2. In addition, dithionite was added to fully reduce the cyt. bc1 complex. The CytcO concentration was ~1 μM. A laser artifact at t = 0 has been removed for clarity. The filled circle in (B) represents the extrapolated initial absorbance after the laser flash.
The absorbance changes at 550 nm (Fig. 5E) are mainly attributed to c-heme absorption, where a decrease in absorbance is associated with oxidation of the hemes. Two kinetic components with time constants of ~120 μs and ~1.7 ms, respectively, were observed, i.e. concomitant with electron transfer from CuA to heme a during the PR → F reaction and during the F → O reaction. Finally, at 563 nm (Fig. 5F) after the unresolved initial drop in absorbance (presumably associated with a small absorbance contribution from CO dissociation), a further decrease in absorbance associated with oxidation of heme b (τ ≅ 6.5 ms) was observed, i.e. oxidation of heme b occurred after oxidation of the CytcO and the c heme (compare panels D-F in Fig. 5, after the break on the abscissa).
The concentration of reacting CytcO (i.e. CytcO from which the CO ligand is dissociated) was estimated from the change in absorbance at t = 0 at 445 nm (Fig. 5B), which yields ~0.15 μM CytcO (using an absorption coefficient (ε) of 82 mM−1cm−1 43). The absorbance at t = 0+, i.e. just after CO dissociation corresponds to that of reduced CytcO. Consequently, the decrease in absorbance from this point until t ≅ 10 ms, when the reaction is essentially over, corresponds to the amount oxidized CytcO (~0.10 μM, using ε = 164 mM−1cm−1 44). Consequently, ~0.05 μM (~35%) of the reacting CytcO becomes re-reduced by the cyt. bc1 complex. From the absorbance changes at 550 nm and 563 nm we estimate that ~0.025 μM (ε = 19.1 mM−1cm−1 20) heme b and ~0.028 μM (ε = 22 mM−1cm−1 20) heme c, respectively, become oxidized, which together account for ~0.05 μM CytcO that is re-reduced during the experiment. It should be noted that these estimations are only approximate because we were not able to accurately resolve O2 binding to the reduced heme a3 in all samples. In part, this problem is attributed to the requirement to add dithionite in order to fully reduce the cyt. bc1-CytcO supercomplex. Because dithionite reduces O2 directly during mixing, the O2 concentration was lowered before initiation of the reaction of CytcO with O2 thereby slowing the R → A reaction. Consequently, it was difficult to resolve the associated absorbance changes from those associated with the next, A → PR transition.
The single-turnover reaction was also studied with purified CytcO (Fig. 5C). Here, heme a was oxidized with a time constant of ~21 μs (R → PR), followed in time by re-reduction from CuA with a time constant of ~90 μs (PR → F) and oxidation with a time constant of ~1.3 ms (F → O). These time constants were almost the same as those observed with the cyt. bc1-CytcO supercomplex. No changes in absorbance were observed at 550 nm nor 563 nm for the purified oxidase (data not shown). The end absorbance level at 445 nm was slightly lower with pure CytcO than with CytcO that is part of the supercomplex (c.f. Fig. 5B,C), which means that the former was more oxidized than the latter. This observation presumably reflects the re-reduction of CytcO by the cyt. bc1 complex in the latter. However, as mentioned above, it was difficult to quantify the relative absorbance differences in the two samples because we were unable to confidently scale the two different traces to each other due to unresolved absorbance changes associated with O2 binding.
Binding of water-soluble cyt. c to e.g. the bovine heart CytcOs is highly dependent on the salt concentration, reflecting electrostatic interactions of the two proteins45. In order to investigate whether or not the interactions of CytcO with cyt. c in the supercomplex could be disrupted, we studied the reaction with O2 at increasing ionic strengths (addition of KCl). As seen in Figure S6, the amplitude of the absorbance changes associated with the F → O reaction decreased slightly (indicating more re-reduction of CytcO) rather than increasing with increased ionic strength (we expect more oxidation of CytcO upon cyt. c dissociation), which indicates that the cyt. c-CytcO interactions were not disrupted at high salt concentrations.
Net Proton Uptake from Solution
The protons required for the reduction of O2 to water are taken up from the medium, a process that can be monitored during flow-flash experiments in the absence of buffer by use of a pH-sensitive dye. Using phenol red, we studied proton uptake during reaction of the reduced cyt. bc1-CytcO supercomplex with O2, following absorbance changes at 560 nm. As seen in Fig. 6, the absorbance increased over time, which indicates net proton uptake during the reaction. The process displayed two components with time constants of 130 μs (~25% of the total absorbance change) and 1.9 ms, i.e. they coincided with the PR → F and F → O reactions, respectively. Upon addition of buffer the dye signal was quenched and only a very small decrease in absorbance associated with oxidation of b-hemes was observed (c.f. Fig. 6).
Absorbance changes of the pH dye phenol red, associated with proton uptake during O2 reduction.
The reaction of the reduced cyt. bc1-CytcO complex with O2 was initiated by a laser flash at t = 0 (see Fig. 5). Absorbance changes were monitored over time at 560 nm in the absence and presence of buffer, respectively. Experimental conditions: 0.05% DDM, 100 μM EDTA, 40 μM Phenol red at pH 7.8 2 mM ascorbate, 0.2 μM Hexamminerutheniumchloride, ~900 μM O2 and either 100 mM KCl or 100 mM HEPES. The CytcO concentration was ~5 μM. A laser artifact at t = 0 has been removed for clarity.
pH-dependence
Reactions steps that are associated with proton uptake from solution (e.g. the F → O step of the reaction of reduced CytcO with O2) often display pH dependent rates30,46,47. We therefore investigated the pH dependence of the reaction with O2 of the reduced cyt. bc1-CytcO supercomplex and CytcO alone (Fig. 7A,B). At 445 nm, for the supercomplex, only a slight pH dependence was observed for the F → O reaction rate. As the total amplitude of the oxidation also changed, the results presumably reflect a pH-dependence in the extent of re-reduction by the cyt. bc1 complex, i.e. “down-stream” steps of the reaction. Also with the pure CytcO, the F → O reaction rate was pH independent and all kinetic components displayed essentially the same amplitudes in the measured pH range (c.f. data at pH 7.5 and 8.5 in Fig. 7B).
pH dependence of the reaction of cyt. bc1-CytcO (A) and CytcO (B) with O2. Kinetic data at representative pH-values are shown (color codes as indicated in the graph) and the time constants are summarized in the inset to panel B (measurements were also performed at pH 6 with the CytcO). The CO ligand was dissociated at t = 0. Experimental conditions: ~1 μM CytcO, 10–100 mM Tris-HCl (see Materials and Methods), 4 mM ascorbate, 1 μM PMS and 100 μM dithionite. The pH was set as described in the Materials and Methods section and measured after mixing using a pH meter. A laser artifact at t = 0 has been removed for clarity.
Na+ -dependence
Because we did not observe any significant pH-dependence in the reaction rates with O2, we also investigated the Na+ -concentration dependence to test the possibility that the CytcO transports Na+ (see refs 48 and 49). As can be seen in Figure S7, the addition of Na+ had a slight effect on the time constant of the A → PR reaction, i.e. electron transfer from heme a to the catalytic site (inset Figure S7), but this reaction step is not linked to pumping in other A-type oxidases. No effect on the kinetics of the F → O reaction was observed.
Discussion
Purification and Activity
The purified cyt. bc1-CytcO complex displayed quinol-oxidase activity, i.e. electrons were transferred first from quinol to the cyt. bc1 complex and then, via the two c-hemes20, to the CytcO, which reduces oxygen to water. The activity was ~210 e−/s, which is in good agreement with previously published results20. The data are also consistent with those obtained for the cyt. bc1-CytcO supercomplex from Mycobacterium smegmatis24, which exhibited quinol-oxidase activity even at high detergent concentrations, supporting the presence of the cyt. bc1-CytcO supercomplex. Both the C. glutamicum cyt. bc1-CytcO and CytcO preparations exhibited TMPD-driven O2-reduction activities. In both cases the activity increased upon addition of soluble cyt. c. However, the increase was larger for the pure CytcO (a factor of ~2.8) than for the supercomplex (a factor of ~1.4) (but see data with the M. smegmatis24). This observation suggests that in C. glutamicum cyt. c is more accessible for binding to the CytcO upon removal of the cyt. bc1 complex, which indicates that in the supercomplex one of the c-hemes is located near the CytcO electron entry point (i.e. presumably near CuA). Thus, the results indicate that pure CytcO (i.e. with the cyt. bc1 removed) is capable of binding soluble cyt. c in spite of the presence of an extra loop of charged amino acids located at the cyt. c binding site18 (this binding can only occur in the mutant where the cyt. bc1 complex is removed, i.e. not in vivo). Formation of a stable cyt. bc1-CytcO complex that is capable of transferring electrons directly from cyt. bc1 to CytcO is also consistent with the rapid electron transfer from the b and c-hemes to CytcO (see below).
We note that the TMPD and cyt. c-oxidation activities measured here are higher than those presented previously18,20. The discrepancy is presumably due to differences in experimental conditions. While in the earlier studies the rate was derived from changes in the concentration of reduced electron donor, here the data was obtained by measuring the O2-reduction rate at a constant concentration of the reduced electron donor (with excess ascorbate).
Ligand Binding to CytcO
Results from earlier studies with e.g. the bovine heart oxidase indicate that upon pulsed illumination the CO ligand dissociates from heme a3 and binds transiently to CuB before it dissociates into solution50:

Dissociation of CO from heme a3 in the dark is very slow (k−1 ≅ 0.03 s−1), but upon illumination the ligand moves to CuB in << 10 ns, if the light intensity of the pulse is strong enough (such as in this study). The dissociation rate constant from CuB, k−2, is ~7·105 s−1 with the bovine heart CytcO50. Recombination of CO occurs via CuB with a second-order process (k2 ≅ 1·108 M−1s−1). The rate for internal CO transfer from CuB to heme a3, k1, is ~103 s−1. The observed rate of CO recombination is approximately given by the fraction of CuB with bound CO (middle state in scheme 1) multiplied by the rate of CO transfer from CuB to heme a3,

With the rate constants given above, we obtain kobs ≅ 0.12 × 1000 s−1 = 120 s−1 (τ ≅ 8 ms) at 1 mM CO.
The CO-recombination kinetics measured with the C. glutamicum CytcO was biphasic (see Fig. 4A). The kinetic difference spectra of the two components with the purified supercomplex and the oxidase alone were similar (see Fig. 4C,D), indicating that both components are associated with CO binding to heme a3 after light-induced dissociation. Consequently, the data indicate the presence of two populations of CytcO with different CO-recombination rates to heme a3. The smaller (~25%) CytcO population, displayed a CO-concentration independent rate constant of ~90 s−1 (also the relative amplitude was independent on the CO concentration). Assuming the model in Eq. 4, this observation indicates that for this population the ratio k2/(k2 + k−2) is equal to ~1 at all CO concentrations used in this study and that k1 = 90 s−1. Alternatively, after photolysis from heme a3 and binding to CuB, the CO ligand did not equilibrate with solution in this population.
The slower, major CO-recombination component displayed a CO-concentration dependent rate constant of 6.7 s−1 at 1 mM CO. Assuming that k1 has the same value of 90 s−1 for the two CytcO populations, the ratio k2/(k2 + k−2) ≅ 0.07. Thus, the difference between the two CytcO populations reflecting the two time constants could be explained by different values of k2/(k2 + k−2), i.e. by differences in CO binding to CuB. Because CO-recombination was biphasic both in whole cells and in the detergent-purified samples, the presence of the two components is not an artifact caused by the purification of the supercomplex or the oxidase. Instead, we speculate that the two components reflect two CytcO populations that are present in the native membrane and that could differ, for example, in the local structure of CuB resulting in different CO-binding affinities. Most likely these two CytcO populations would display different reactivity towards the natural ligand and electron acceptor, O2. If the relative fraction of these two populations would be modulated by the cell, this mechanism could be used to regulate electron transfer through the respiratory chain.
Reaction with O2
The four-electron reduction of O2 to H2O takes place in a number of distinct kinetic steps in which the CytcO is gradually oxidized and O2 is reduced. We identified the kinetic components on the basis of a comparison to data obtained earlier with other well-studied oxidases38. With the pure CytcO we observed electron transfer from heme a to the catalytic site and formation of the “peroxy” state, PR, with a time constant of ~21 μs. Formation of the next, ferryl intermediate (F) displayed a time constant of 90 μs. Finally, the CytcO was oxidized forming the oxidized state (O) with a time constant of 1.3 ms. All these time constants are essentially the same as those observed previously with e.g. the well-studied CytcO from bovine heart or R. sphaeroides. Consequently, the differences in CO-binding kinetics between the earlier studied A-type oxidases and C. glutamicum CytcO are apparently not reflected in the kinetics of O2 binding and reduction.
For the cyt. bc1-CytcO complex, all time constants for the different reaction steps were similar to those observed with the pure CytcO, however, there were also notable differences reflecting intra-complex electron transfer. During the PR → F reaction (τ ≅ 120 μs) an electron is transferred from CuA to heme a, which leaves CuA oxidized allowing electron transfer from cyt. c of the cyt. bc1 complex to CuA. This electron transfer is rate-limited by proton uptake51 and does not occur at the same rate as when photochemically injected into CuA52. As seen at 550 nm (Fig. 5E), cyt. c was partially oxidized over a time scale of ~100 μs, which indicates that CuA was re-reduced concomitantly with the CuA-to-heme a electron transfer. This interpretation is also supported by the very small extent of net CuA oxidation (observed at 830 nm) over the 100-μs time scale (Fig. 5D, see the small increase in absorbance). In the next step of the reaction, F → O (τ ≅ 1.7 ms), the fourth electron is transferred to the catalytic site, which allows further electron transfer from heme c to the CytcO, reflected in a further decrease in absorbance at 550 nm over the time scale of the F → O reaction in CytcO. The absorbance changes at 550 nm reflect oxidation of cyt. c, but we could not determine the degree of oxidation of each of the two cyt. cs separately. Most likely these two cyt. cs are oxidized simultaneously, but not necessarily to the same degree. In the summarizing Fig. 8 we indicate that both cyt. cs are oxidized over time scales of 100 μs and 1.7 ms (approximated by 2 ms in the figure).
A schematic picture showing the approximate time constants for the electron-transfer reactions.
Note that the positions of the heme groups in this picture are not compatible with the X-ray structure (e.g.39); they have been adjusted for clarity. Because the data cannot discriminate between the two heme cs or the two heme bs, we have indicated the oxidation time constants for each type of heme without specifying which heme that is oxidized. Redox reactions of the quinones are not observed in these experiments.
We also observed further electron transfer from heme b (absorbance decrease at 563 nm, Fig. 5F), which reflects electron transfer from heme b to the CytcO, but this electron transfer significantly lags behind (τ ≅ 6.5 ms) that of the F → O reaction (τ ≅ 1.7 ms). As described above for cyt. c, also for heme b oxidation we could not discriminate between hemes bL or bH and conclude only that heme b of the cyt. bc1 is oxidized over the 6.5-ms time scale. As outlined in the Results section, the amounts of oxidized heme c and heme b approximately equal the amount of CytcO that becomes re-reduced during or after reaction with O2. Furthermore, the electron-transfer time constant from heme b to CytcO (τ ≅ 6.5 ms) is approximately compatible with the overall quinol-oxidation/O2 reduction turnover rate of the cyt. bc1-CytcO supercomplex (~210 s−1). This rapid electron transfer with a time constant of ~6.5 ms corresponds to a maximum electron-transfer rate over a distance of ~25 Å53. Even though the distances connecting the heme cs with their partners are not known for the C. glutamicum cyt. bc1-CytcO complex, this distance estimation is consistent with that between heme bL and the iron-sulfur cluster in cyt. bc1 from e.g. S. cerevisiae25.
To investigate the functional stability of the cyt. bc1-CytcO supercomplex we investigated the reaction with O2 upon increasing the ionic strength (Figure S6). Neither the amplitudes nor rates of the observed absorbance changes at 445 nm were significantly altered even at the highest KCl concentration of 1.5 M. These results suggest that the purified supercomplex is stable and remains functionally intact. Furthermore, the interactions between one of the c hemes of the cyt. bc1 complex and CytcO seem more stable than those observed for the water-soluble cyt. c and CytcO from e.g. bovine heart as in the latter case the 1:1 cyt. c-CytcO complex dissociates at ionic strength above ~300 mM45.
pH (in)dependence of the Reaction with O2
As seen in Fig. 7 we did not observe any significant pH-dependence in the kinetics of the F → O reaction, neither for the pure CytcO nor for the cyt. bc1-CytcO supercomplex. Slight differences were observed for the two samples (see inset to Fig. 7), which may be attributed to binding of cyt. bc1 to CytcO in the supercomplex. The observation that the F → O rate is essentially pH-independent is surprising given that the reaction is associated with proton uptake (see Fig. 6) and therefore expected to display pH-dependent kinetics, as in other oxidases30,46,47. One possible explanation for pH-independent kinetics of a reaction step that is linked to proton uptake is that the pKa in this pH dependence may be outside of the accessible pH range. Alternatively, the proton uptake may not be part of the rate-limiting step, however, this explanation is less likely based on results from earlier studies with other A- and B-type oxidases where proton uptake is rate limiting51,54,55,56,57.
In earlier studies with the R. sphaeroides CytcO it has been observed that the PR → F rate is essentially pH-independent up to pH ~9 and then decreases with increasing pH with an apparent pKa of 9.458. The F → O rate in R. sphaeroides CytcO displayed a more complex pH dependence and was found to titrate with two pKas of ~9 and <6, respectively30. The pKa around 9 was attributed to residue Glu286 within the D proton pathway, which is conserved in the C. glutamicum CytcO (Glu267). In the R. sphaeroides CytcO, replacement of e.g. Asn139 or Asn207 by Asp resulted in an increase in the Glu286 pKa; for example in the Asn139Asp variant the pKa increased to a value above the accessible pH range46,59,60. In other words, even though the reaction is associated with proton uptake in these structural variants, it did not display a pH-dependent kinetics. In the C. glutamicum CytcO, many but not all residues “below” the Glu267 in the D pathway are conserved. The data with the R. sphaeroides structural variants show that very small changes in the D pathway structure, also at a distance from the Glu, may yield pH-independent kinetics. Thus, considering the differences in the environment of Glu267 in the C. glutamicum CytcO it is possible that it’s pKa is tuned to adopt a value that is higher than that of Glu286 in the R. sphaeroides CytcO. However, more experiments with single point mutations in the D pathway are necessary to confirm this speculation.
Concluding Remarks
The reaction of the purified reduced CytcO with O2 displayed the same sequence of electron transfers as that observed previously with other bacterial and mitochondrial A-type CytcOs. However, in contrast to data obtained with the other oxidases, none of the reaction steps associated with proton uptake displayed any pH dependent rates, which is explained in terms of an elevated apparent pKa of Glu267. The data indicate that one of the c hemes of cyt. bc1 is bound near CuA, at the same site where externally added water-soluble cyt. c would bind. Furthermore, the data indicate that the interaction of this heme c with CytcO is strong in the cyt. bc1-CytcO supercomplex and insensitive to changes in ionic strength. The second heme c of the cyt. bc1-CytcO supercomplex provides a link for direct electron transfer between cyt. bc1 and CuA, the electron acceptor of CytcO (see Fig. 8), which was also indicated from earlier mutagenesis data20. Electron transfer from heme c to the CytcO occurred over time scales of approximately 100 μs–2 ms, while re-reduction of heme c by heme b displayed a time constant of ~6.5 ms, which suggest that this reaction may be rate limiting for the overall quinol-oxidation O2-reduction turnover rate. In conclusion, the isolation of a stable supercomplex from C. glutamicum allowed us to investigate the kinetics of electron transfer all the way from heme b in the cyt. bc1 complex to the catalytic site of CytcO, via the bridging c hemes.
Materials and Methods
If not stated otherwise, the chemicals were purchased from Sigma-Aldrich.
Corynebacterium glutamicum strains
The C. glutamicum strains used for purification of CytcO and the cyt. bc1-CytcO supercomplex were described before20. The ΔC-DSt strain refers to the 13032ΔctaD strain transformed with the pJC1-ctaDSt plasmid (KanR), which serves as an expression plasmid for Strep-tagged CtaD; ctaD is expressed from its native promoter and contains 10 additional codons at the 3′-end (AAWSHPQFEK)). The ΔQ-DSt strain refers to the 13032ΔqcrCAB strain transformed with the pJC1-ctaDSt plasmid.
Culture Conditions
The cells were cultivated at 30 °C in all steps. Single C. glutamicum colonies were picked from BHI-Agar plates (33 g/l brain heart infusion broth, 15 g/l agar agar, 20 g/l D-(+)-glucose, 25 mg/l kanamycin) and inoculated into 10 ml BHI culture medium (33 g/l brain heart infusion broth, 20 g/l D-(+)-glucose, 25 mg/l kanamycin) at 220 rpm. After overnight growth the pre-culture was inoculated into 500 ml CGXII medium20 in a 2 l Erlenmeyer flask (at 160 rpm). After the optical density at 600 nm (OD600) reached between 25 and 30, the cells were diluted 1:20 into 2 l of CGXII medium into a 5 l baffled Erlenmeyer flask (at 130 rpm) and cultivated to an OD600 of 15–17 before harvest.
Membrane Preparation
The cells were harvested using a Beckman centrifuge equipped with the JLA 8.1000 rotor at 7,500 rpm (~10,000 × g) for 30 min. The cells were homogenized in 4 ml cell lysis buffer (100 mM Tris-HCl pH 7.5, 5 mM MgSO4, some crystals phenylmethanesulfonyl fluoride, some crystals DNaseI (Roche)) per 1 g of cells (wet weight) and passed through a cell disrupter 4 times at 40 kPsi (Constant Systems). Cell debris was collected by centrifugation at 27,000 × g for 20 min. (Type 45Ti rotor, Beckman) and subsequently the membranes were collected by ultracentrifugation at 150,000 × g for 90 min (Type 45Ti rotor, Beckman).
Protein Purification
The purification was done essentially as described in ref. 20 with minor modifications. Briefly, the membranes were mixed with solubilization buffer (100 mM Tris-HCl pH 7.5, 100 mM NaCl, 2 mM MgSO4, 50 mg/l avidin (iba lifescience), 1% (w/v) DDM (GLYCON Biochemicals) at a protein concentration of 5 mg/ml and incubated at 4 °C for 45 min under slow stirring. Unsolubilized material was removed by ultracentrifugation (180,000 × g, 20 min, 4 °C). The supernatant was collected and concentrated in an Amicon Ultra 15 ml filter spin tube with 100 kDa cutoff until the volume was around 5 ml. The concentrated supernatant was diluted tenfold in solubilization buffer without DDM (yielding a final DDM concentration of 0.1% (w/v) and concentrated again to reach a volume <15 ml. The concentrated supernatant was applied to a Gravity flow Strep-Tactin Superflow column (bed volume 5 ml, iba lifescience). Subsequently, the column was washed 3 times with 0.5 column volumes of washing buffer (100 mM Tris-HCl pH7.5, 100 mM NaCl, 2 mM MgSO4, 0.015% (w/v) DDM) and protein was eluted with up to 3 column volumes elution buffer (100 mM Tris-HCl pH 7.5, 100 mM NaCl, 2 mM MgSO4, 0.015% (w/v) DDM, 2.5 mM D-desthiobiotin) and concentrated as described above. The protein samples (CytcO alone green colored, bc1-CytcO brown colored) were stored at 4 °C.
The CytcO alone, as well as the cyt. bc1-CytcO supercomplex, could be purified from membranes originating from the ΔQ-DSt and the ΔC-DSt C. glutamicum strains, respectively, both having a Strep-tag on subunit I of the CytcO. The SDS-PAGE for the two purifications (supplementary Figure S1) shows that the eluate of the supercomplex purification contains the three core subunits of CytcO (CtaD, C and E), along with the three subunits of the bc1-complex (QcrC, A and B), as well as some additional subunits, which were also co-purified with the supercomplex20. Accordingly, the eluate for the oxidase purification from ΔQ-DSt membranes contained only the three core subunits of the CytcO.
Dithionite-reduced minus ferricyanide-oxidized difference spectra were recorded at room temperature using a Cary100 UV-Vis Spectrophotometer. The CytcO concentration was determined from the reduced minus oxidized difference spectrum using the absorption coefficient Δε600–630 = 3.2 mM−1cm−1 20.
Quinone Reduction
Instead of ubiquinone, C. glutamicum, as a Gram-positive bacterium, employs menaquinones as an electron carrier in the membrane61. To prepare reduced quinol, 3.7 mg 2,3-dimethyl-[1,4]naphthoquinone (Rare Chemicals GmbH) was dissolved in 1 ml N2-saturated anhydrous cyclohexane to yield a 20 mM solution. The solution was mixed with 5 ml N2-saturated 1 M sodium dithionite solution (in H2O) and shaken vigorously. After phase separation, the organic phase containing the reduced 2,3-dimethyl-[1,4]naphthoquinol was removed and transferred to a 15 ml Falcon tube (all steps performed under a stream of N2). The cyclohexane was evaporated under an N2 stream, while the sample was kept at ~40 °C in a water bath. Subsequently, the reduced quinol was dissolved in N2-saturated, acidified ethanol (ethanol with 10 mM HCl), aliquoted and flash frozen in liquid nitrogen and stored at −20 °C.
Determination of enzyme activities
Oxygen consumption in multiple turnover experiments was measured using a Clark-type oxygraph (Hansatech). The TMPD (N,N,N′,N′-tetramethyl-p-phenylenediamine)-oxidase activity was measured in 100 mM Tris-HCl (pH 7.5), 100 mM NaCl, 0.015% (w/v) DDM, 0.2 mM TMPD, 2 mM sodium ascorbate. Both TMPD and ascorbate were added before the sample and background oxygen consumption was measured and the reaction was started by adding the protein sample. Cytochrome c driven oxidase activity was measured as above in the presence of ascorbate and TMPD, but bovine heart cyt. c (25 μM final concentration) was also supplied before addition of the sample. Quinol-driven oxidase activity was measured in the same buffer (100 mM Tris-HCl (pH 7.5), 100 mM NaCl, 0.015% (w/v) DDM), but 20 μl of the reduced quinol solution (see above) was added before the protein sample. Background oxygen consumption was measured, followed by addition of the protein sample and monitoring the enzymatic oxygen consumption. When measuring the CytcO activity with ascorbate/TMPD/cyt. c as a substrate, this activity was insignificant. However, in the presence of quinol the background O2-reduction rate increased significantly, due to quinol auto-oxidation and at most it reached 70% of the rate measured after addition of the supercomplex.
Flash Photolysis and Flow-flash Experiments
The purified samples (typical protein concentrations were in the range of 2–3 μM and 1–1.5 μM for bc1-CytcO and CytcO, respectively) were transferred into a Thunberg cuvette and the atmosphere was exchanged for N2 on the vacuum line. The sample was reduced by the addition of 1 μM phenazine methosulfate and 5 mM sodium ascorbate from the sidearm of the cuvette. In order to achieve complete reduction of all the redox-active centers in samples containing supercomplexes, 0.5–1 mM sodium dithionite (Merck Millipore) was added and the reduction state was confirmed spectrophotometrically. After complete reduction, the atmosphere was exchanged for CO on a vacuum line. To measure the pH dependence of the reaction of the reduced CytcO or cyt. bc1-CytcO with O2, the sample was prepared in 10 mM Tris-HCl, the atmosphere was replaced consecutively by N2 and then CO and the sample was reduced by 4 mM ascorbate, 1 μM PMS and 100 μM dithionite at pH 7.5. Samples at different, higher pH values were prepared by adding various volumes of 1 M Tris-HCl buffer at pH 10 followed by incubation for at least 1 hour. Alternatively, the sample in 10 mM Tris-HCl at pH 7.5, was reduced with 4 mM ascorbate, 1 μM PMS and 100 μM dithionite (under CO atmosphere) and then mixed with an O2-saturated solution at different pH values containing 100 mM buffer. The buffering capacity of the O2 solution was much higher (100 mM) than that of the enzyme solution (10 mM) so the final pH after mixing was determined by the former. The pH after mixing was measured using a pH meter.
The CO rebinding/recombination kinetics to the catalytic site was measured as a change in absorbance over time at several wavelengths after the photolysis by a ~10-ns laser flash (λ = 532 nm, Nd-YAG laser, Quantel; the flash-photolysis/flow-flash setup was purchased from Applied Photophysics UK). The time resolution of the set-up was ~10−7 s. The absorbance changes were then fitted to an exponential decay function using the ProK software from Applied Photophysics U.K.
In order to determine the CO concentration dependence of the CO-recombination, 1 ml of a protein sample was transferred into a Thunberg cuvette and treated as described above, except that after reduction it was left under a nitrogen atmosphere. The sample was covered with a layer of paraffin oil and CO was added in small aliquots of CO-saturated buffer (100 mM Tris-HCl (pH 7.5), 100 mM NaCl, 0.015% (w/v) DDM) with a gas-tight Hamilton syringe through the paraffin layer. The CO-recombination kinetics was studied after each CO addition using a flash-photolysis set-up (Applied Photophysics, UK).
In flow-flash experiments, the reduced and CO-blocked protein sample was mixed 1:3 with oxygen-saturated buffer (~1.2 mM O2) in a flow-flash setup (Applied Photophysics, UK.). About 200 ms after mixing (mixing time <10 ms) with the oxygenated buffer, CO was dissociated from the catalytic site by a short laser pulse (~10-ns laser flash (λ = 532 nm, Nd YAG-laser, Quantel). Changes in absorbance were recorded over time at different wavelengths. The data were fitted to a kinetic model using the ProK software from Applied Photophysics, UK.
Additional Information
How to cite this article: Graf, S. et al. Rapid Electron Transfer within the III-IV Supercomplex in Corynebacterium glutamicum. Sci. Rep. 6, 34098; doi: 10.1038/srep34098 (2016).
References
Schägger, H. Respiratory chain supercomplexes. IUBMB Life 52, 119–128 (2002).
Stuart, R. A. Supercomplex organization of the oxidative phosphorylation enzymes in yeast mitochondria. J. Bioenerg. Biomembr. 40, 411–417 (2008).
Heinemeyer, J., Braun, H. P., Boekema, E. J. & Kouřil, R. A structural model of the cytochrome c reductase/oxidase supercomplex from yeast mitochondria. J. Biol. Chem. 282, 12240–12248 (2007).
Winge, D. R. Sealing the Mitochondrial Respirasome. Molecular and Cellular Biology 32, 2647–2652 (2012).
Mileykovskaya, E. et al. Arrangement of the respiratory chain complexes in Saccharomyces cerevisiae supercomplex III2IV2 revealed by single particle cryo-electron microscopy. J. Biol. Chem. 287, 23095–23103, doi: 10.1074/jbc.M112.367888 (2012).
Genova, M. L. & Lenaz, G. Functional role of mitochondrial respiratory supercomplexes. Biochimica et Biophysica Acta - Bioenergetics 1837, 427–443, doi: 10.1016/j.bbabio.2013.11.002 (2014).
Acin-Perez, R. & Enriquez, J. A. The function of the respiratory supercomplexes: The plasticity model. Biochimica et Biophysica Acta - Bioenergetics 1837, 444–450, doi: 10.1016/j.bbabio.2013.12.009 (2014).
Boumans, H., Grivell, L. A. & Berden, J. A. The respiratory chain in yeast behaves as a single functional unit. J. Biol. Chem. 273, 4872–4877, doi: 10.1074/jbc.273.9.4872 (1998).
Althoff, T., Mills, D. J., Popot, J. L. & Kühlbrandt, W. Arrangement of electron transport chain components in bovine mitochondrial supercomplex I 1 III 2 IV 1. EMBO Journal 30, 4652–4664, doi: 10.1038/emboj.2011.324 (2011).
Dudkina, N. V., Folea, I. M. & Boekema, E. J. Towards structural and functional characterization of photosynthetic and mitochondrial supercomplexes. Micron 72, 39–51, doi: 10.1016/j.micron.2015.03.002 (2015).
Cruciat, C. M., Brunner, S., Baumann, F., Neupert, W. & Stuart, R. A. The cytochrome bc1 and cytochrome c oxidase complexes associate to form a single supracomplex in yeast mitochondria. J. Biol. Chem. 275, 18093–18098, doi: 10.1074/jbc.M001901200 (2000).
Berry, E. A. & Trumpower, B. L. Isolation of ubiquinol oxidase from Paracoccus denitrificans and resolution into cytochrome bc1 and cytochrome c-aa3 complexes. J. Biol. Chem. 260, 2458–2467 (1985).
Sone, N., Sekimachi, M. & Kutoh, E. Identification and properties of a quinol oxidase super-complex composed of a bc1 complex and cytochrome oxidase in the thermophilic bacterium PS3. J. Biol. Chem. 262, 15386–15391 (1987).
Stroh, A. et al. Assembly of Respiratory Complexes I, III and IV into NADH Oxidase Supercomplex Stabilizes Complex I in Paracoccus denitrificans. J. Biol. Chem. 279, 5000–5007, doi: 10.1074/jbc.M309505200 (2004).
Reincke, B. et al. Heterologous expression of soluble fragments of cytochrome c552 acting as electron donor to the Paracoccus denitrificans cytochrome c oxidase. Biochimica et Biophysica Acta - Bioenergetics 1411, 114–120, doi: 10.1016/s0005-2728(99)00037-7 (1999).
Myllykallio, H., Drepper, F., Mathis, P. & Daldal, F. Membrane-anchored cytochrome c(y) mediated microsecond time range electron transfer from the cytochrome bc1 complex to the reaction center in Rhodobacter capsulatus. Biochemistry 37, 5501–5510, doi: 10.1021/bi973123d (1998).
Daldal, F. et al. Mobile cytochrome c2 and membrane-anchored cytochrome cy are both efficient electron donors to the cbb3- and aa3-type cytochrome c oxidases during respiratory growth of Rhodobacter sphaeroides. J. Bacteriol. 183, 2013–2024, doi: 10.1128/jb.183.6.2013-2024.2001 (2001).
Sakamoto, J. et al. Cytochrome c oxidase contains an extra charged amino acid cluster in a new type of respiratory chain in the amino-acid-producing Gram-positive bacterium Corynebacterium glutamicum. Microbiology 147, 2865–2871 (2001).
Bott, M. & Niebisch, A. The respiratory chain of Corynebacterium glutamicum. Journal of Biotechnology 104, 129–153, doi: 10.1016/s0168-1656(03)00144-5 (2003).
Niebisch, A. & Bott, M. Purification of a cytochrome bc1-aa3 supercomplex with quinol oxidase activity from Corynebacterium glutamicum: Identification of a fourth subunit of cytochrome aa3 oxidase and mutational analysis of diheme cytochrome c1 . J. Biol. Chem. 278, 4339–4346, doi: 10.1074/jbc.M210499200 (2003).
Niebisch, A. & Bott, M. Molecular analysis of the cytochrome bc1-aa3 branch of the Corynebacterium glutamicum respiratory chain containing an unusual diheme cytochrome c1. Archives of Microbiology 175, 282–294, doi: 10.1007/s002030100262 (2001).
Sone, N. et al. A novel hydrophobic diheme c-type cytochrome. Purification from Corynebacterium glutamicum and analysis of the QcrCBA operon encoding three subunit proteins of a putative cytochrome reductase complex. Biochimica et Biophysica Acta - Bioenergetics 1503, 279–290, doi: 10.1016/s0005-2728(00)00205-x (2001).
Kao, W.-C. et al. The obligate respiratory supercomplex from Actinobacteria. Biochimica et Biophysica Acta (BBA) - Bioenergetics, 10.1016/j.bbabio.2016.07.009.
Megehee, J. A., Hosler, J. P. & Lundrigan, M. D. Evidence for a cytochrome bcc-aa3 interaction in the respiratory chain of Mycobacterium smegmatis. Microbiology 152, 823–829, doi: 10.1099/mic.0.28723-0 (2006).
Hunte, C., Koepke, J., Lange, C., Roßmanith, T. & Michel, H. Structure at 2.3 Å resolution of the cytochrome bc1 complex from the yeast Saccharomyces cerevisiae co-crystallized with an antibody Fv fragment. Structure 8, 669–684, doi: 10.1016/S0969-2126(00)00152-0 (2000).
Hemp, J. & Gennis, R. B. Diversity of the heme-copper superfamily in archaea: insights from genomics and structural modeling. Results Probl Cell Differ 45, 1–31 (2008).
Pereira, M. M., Santana, M. & Teixeira, M. A novel scenario for the evolution of haem-copper oxygen reductases. Biochim. Biophys. Acta-Bioenerg. 1505, 185–208 (2001).
Hosler, J. P., Ferguson-Miller, S. & Mills, D. A. Energy transduction: Proton transfer through the respiratory complexes. Annual Review of Biochemistry 75, 165–187 (2006).
Yoshikawa, S. et al. Proton pumping mechanism of bovine heart cytochrome c oxidase. Biochimica et Biophysica Acta - Bioenergetics 1757, 1110–1116 (2006).
Namslauer, A. & Brzezinski, P. Structural elements involved in electron-coupled proton transfer in cytochrome c oxidase. FEBS Lett 567, 103–110 (2004).
Brzezinski, P. & Gennis, R. B. Cytochrome c oxidase: exciting progress and remaining mysteries. J. Bioenerg. Biomembr. 40, 521–531 (2008).
Brzezinski, P. & Ädelroth, P. Design principles of proton-pumping haem-copper oxidases. Curr Opin Struct Biol 16, 465–472 (2006).
Richter, O. M. H. & Ludwig, B. Electron transfer and energy transduction in the terminal part of the respiratory chain - Lessons from bacterial model systems. Biochimica et Biophysica Acta - Bioenergetics 1787, 626–634 (2009).
Belevich, I. & Verkhovsky, M. I. Molecular mechanism of proton translocation by cytochrome c oxidase. Antioxid Redox Signal 10, 1–29 (2008).
Ferguson-Miller, S., Hiser, C. & Liu, J. Gating and regulation of the cytochrome c oxidase proton pump. Biochimica et Biophysica Acta - Bioenergetics 1817, 489–494 (2012).
Rich, P. R. & Maréchal, A. Functions of the hydrophilic channels in protonmotive cytochrome c oxidase. Journal of the Royal Society Interface 10, 183–196 (2013).
Kaila, V. R. I., Verkhovsky, M. I. & Wikström, M. Proton-coupled electron transfer in cytochrome oxidase. Chem. Rev. 110, 7062–7081 (2010).
Ädelroth, P., Ek, M. & Brzezinski, P. Factors Determining Electron-Transfer Rates in Cytochrome c Oxidase: Investigation of the Oxygen Reaction in the R. sphaeroides and Bovine Enzymes. Biochim. Biophys. Acta 1367, 107–117 (1998).
Svensson-Ek, M. et al. The X-ray Crystal Structures of Wild-Type and EQ(I-286) Mutant Cytochrome c Oxidases from Rhodobacter sphaeroides. J. Mol. Biol. 321, 329–339 (2002).
Biasini, M. et al. SWISS-MODEL: Modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Research 42, W252–W258, doi: 10.1093/nar/gku340 (2014).
Arnold, K., Bordoli, L., Kopp, J. & Schwede, T. The SWISS-MODEL workspace: A web-based environment for protein structure homology modelling. Bioinformatics 22, 195–201, doi: 10.1093/bioinformatics/bti770 (2006).
Guex, N., Peitsch, M. C. & Schwede, T. Automated comparative protein structure modeling with SWISS-MODEL and Swiss-PdbViewer: A historical perspective. Electrophoresis 30, S162–S173, doi: 10.1002/elps.200900140 (2009).
Vanneste, W. H. The stoichiometry and absorption spectra of components a and a-3 in cytochrome c oxidase. Biochemistry 5, 838–848 (1966).
van Gelder, B. F. On cytochrome c oxidase. I. The extinction coefficients of cytochrome a and cytochrome a3. Biochimica Et Biophysica Acta 118, 36–46 (1966).
Michel, B. & Bosshard, H. R. Spectroscopic analysis of the interaction between cytochrome c and cytochrome c oxidase. J. Biol. Chem. 259, 10085–10091 (1984).
Brzezinski, P. & Johansson, A. L. Variable proton-pumping stoichiometry in structural variants of cytochrome c oxidase. Biochimica et Biophysica Acta - Bioenergetics 1797, 710–723 (2010).
Von Ballmoos, C., Gennis, R. B., Ädelroth, P. & Brzezinski, P. Kinetic design of the respiratory oxidases. Proc. Natl. Acad. Sci. USA 108, 11057–11062 (2011).
Muntyan, M. S. et al. Cytochrome cbb3 of Thioalkalivibrio is a Na+−pumping cytochrome oxidase. Proc. Natl. Acad. Sci. USA 112, 7695–7700, doi: 10.1073/pnas.1417071112 (2015).
Park, C., Moon, J. Y., Cokic, P. & Webster, D. A. Na+−translocating cytochrome bo terminal oxidase from Vitreoscilla: Some parameters of its Na+ pumping and orientation in synthetic vesicles. Biochemistry 35, 11895–11900, doi: 10.1021/bi9530503 (1996).
Einarsdóttir, Ó. et al. Photodissociation and recombination of carbonmonoxy cytochrome oxidase: dynamics from picoseconds to kiloseconds. Biochemistry 32, 12013–12024 (1993).
Karpefors, M., Ädelroth, P., Zhen, Y., Ferguson-Miller, S. & Brzezinski, P. Proton uptake controls electron transfer in cytochrome c oxidase. Proc. Natl. Acad. Sci. USA 95, 13606–13611 (1998).
Wang, K. F. et al. Definition of the interaction domain for cytochrome c on cytochrome c oxidase - II. Rapid kinetic analysis of electron transfer from cytochrome c to Rhodobacter sphaeroides cytochrome oxidase surface mutants. J. Biol. Chem. 274, 38042–38050 (1999).
Gray, H. B. & Winkler, J. R. Long-range electron transfer. Proc. Natl. Acad. Sci. USA 102, 3534–3539, doi: 10.1073/pnas.0408029102 (2005).
Von Ballmoos, C. et al. Mutation of a single residue in the ba3 oxidase specifically impairs protonation of the pump site. Proc. Natl. Acad. Sci. USA 112, 3397–3402, doi: 10.1073/pnas.1422434112 (2015).
Vilhjálmsdóttir, J., Johansson, A. L. & Brzezinski, P. Structural Changes and Proton Transfer in Cytochrome c Oxidase. Scientific Reports 5, doi: 10.1038/srep12047 (2015).
Salomonsson, L., Faxén, K., Ädelroth, P. & Brzezinski, P. The timing of proton migration in membrane-reconstituted cytochrome c oxidase. Proc. Natl. Acad. Sci. USA 102, 17624–17629 (2005).
Aagaard, A. & Brzezinski, P. Zinc ions inhibit oxidation of cytochrome c oxidase by oxygen. FEBS Lett. 494, 157–160 (2001).
Namslauer, A., Aagaard, A., Katsonouri, A. & Brzezinski, P. Intramolecular proton-transfer reactions in a membrane-bound proton pump: the effect of pH on the peroxy to ferryl transition in cytochrome c oxidase. Biochemsitry 42, 1488–1498 (2003).
Han, D. et al. Replacing Asn207 by Aspartate at the Neck of the D Channel in the aa3-Type Cytochrome c Oxidase from Rhodobacter sphaeroides Results in Decoupling the Proton Pump. Biochemistry 45, 14064–14074 (2006).
Namslauer, A., Pawate, A. S., Gennis, R. B. & Brzezinski, P. Redox-coupled proton translocation in biological systems: Proton shuttling in cytochrome c oxidase. Proc. Natl. Acad. Sci. USA 100, 15543–15547 (2003).
Kanzaki, T., Sugiyama, Y., Kitano, K., Ashida, Y. & Imada, I. Quinones of brevibacterium. Biochimica et Biophysica Acta (BBA)/Lipids and Lipid Metabolism 348, 162–165, doi: 10.1016/0005-2760(74)90102-7 (1974).
Acknowledgements
We would like to thank Dr. Emelie Svahn for technical assistance. This study was supported by grants from the Knut and Alice Wallenberg Foundation (KAW 2013.0006) and Swedish Research Council (to PB, CvB, PÄ). Support was also obtained from the Excellence Initiative of the German Federal and State Governments (EXC 294 BIOSS to CH) and the COST European Cooperation in Science and Technology, CM1306, “Understanding Movement and Mechanism in Molecular Machines.
Author information
Authors and Affiliations
Contributions
P.B., C.v.B. and P.Ä. planned the research; S.G. and O.F. performed the experiments; M.B. constructed the strains; P.B., S.G., C.v.B. and P.Ä. wrote the manuscript; W.-C.K. and C.H. assisted in development of methods for enzyme purification.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Graf, S., Fedotovskaya, O., Kao, WC. et al. Rapid Electron Transfer within the III-IV Supercomplex in Corynebacterium glutamicum. Sci Rep 6, 34098 (2016). https://doi.org/10.1038/srep34098
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep34098
This article is cited by
-
Structural basis for safe and efficient energy conversion in a respiratory supercomplex
Nature Communications (2022)
-
Cryo-EM structure of the yeast respiratory supercomplex
Nature Structural & Molecular Biology (2019)
-
Bacterial periplasmic nitrate and trimethylamine-N-oxide respiration coupled to menaquinol-cytochrome c reductase (Qcr): Implications for electrogenic reduction of alternative electron acceptors
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.