Abstract
The plants effect in subsurface flow constructed wetlands (SSF-CWs) is controversial, especially at low temperatures. Consequently, several SSF-CWs planted with Iris pseudacorus (CWI) or Typha orientalis Presl. (CWT) and several unplanted ones (CWC) were set up and fed with secondary effluent of sewage treatment plant during the winter in Eastern China. The 16S rDNA Illumina Miseq sequencing analysis indicated the positive effects of I. pseudacorus on the bacterial community richness and diversity in the substrate. Moreover, the community compositions of the bacteria involved with denitrification presented a significant difference in the three systems. Additionally, higher relative abundances of nitrifying bacteria (0.4140%, 0.2402% and 0.4318% for Nitrosomonas, Nitrosospira and Nitrospira, respectively) were recorded in CWI compared with CWT (0.2074%, 0.0648% and 0.0181%, respectively) and CWC (0.3013%, 0.1107% and 0.1185%, respectively). Meanwhile, the average removal rates of NH4+-N and TN in CWI showed a prominent advantage compared to CWC, but no distinct advantage was found in CWT. The hardy plant I. pseudacorus, which still had active root oxygen release in cold temperatures, positively affected the abundance of nitrifying bacteria in the substrate, and accordingly was supposed to contribute to a comparatively high nitrogen removal efficiency of the system during the winter.
Similar content being viewed by others
Introduction
During the past several decades, constructed wetlands (CWs), which are an economical and environmentally friendly wastewater treatment technology, have been widely used for removing pollutants from a variety of wastewaters including domestic wastewater, agricultural wastewater, industrial wastewater, acid mine drainage, etc. around the world1,2,3,4,5. They are generally categorized into two basic types: free water surface system (FWS-CWs) and subsurface flow system (SSF-CWs). In recent years, SSF-CWs have attracted more and more studies and applications in China and Europe, due to its decreased demand for land and high efficiency for various pollutants1. Researches on the mechanism of pollutants removal in CWs have revealed that it is a fairly complex process which involves a variety of removal pathways, including microbial degradation, plant uptake, substrate filtration and adsorption, precipitation, sedimentation and volatilization3,6,7,8.
Microorganisms have been universally realized as an original and critical factor in the wetland purification process8,9,10. Biodegradation of organic compounds in wastewater is primarily due to some heterotrophic and autotrophic bacteria, for example, methane oxidizing bacteria (methanotrophs) and sulfate-reducing bacteria (SRB). Moreover, nitrification-denitrification and anaerobic ammonium oxidation (anammox) are usually considered to be the two key links for nitrogen removal from wastewater in wetlands. Ammonia oxidizing bacteria (AOB), mainly belonging to β-proteobacteria and γ-proteobacteria, determined the rate-limiting step for nitrification that converts ammonium to nitrite with oxygen as the final electron acceptor, while a wider variety of bacterial phylogenetic groups contributed to the denitrification process8,11.
Plants have several properties that make them an essential component in wetlands, but their exact effect is complex and controversial. Apart from absorbing and storing the nutrients from wastewater, the presence of macrophytes is believed to have a close relationship with the increased microbial density, activity and diversity in wetlands6,7,12,13. On the one hand, plant roots can provide a substratum for attached microorganism growth1,8. On the other hand, the plant litter can serve as carbon source for heterotrophic bacteria6,13,14. In addition, the emerging plants growing in SSF-CWs can also work as oxygen transfer passage for aerobic microorganism in the substrate6,15,16. However, several studies posited that plants had little statistical effect on the community abundance or structure of the overall bacteria or particular microbial functional groups17,18,19,20. In fact, the interaction between the macrophyte and microorganism community depends on a variety of factors such as plant species, vegetation growth status, season and even random chance17,20,21,22. Therefore, the effects of different plants on microorganism groups in CWs, especially in the cold season, need deeper exploration. Further study on the complicated process occurring in CWs would be beneficial to optimizing design parameters and improving the overall performance in engineering application practices.
In the present research, yellow flag (Iris pseudacorus) and oriental cattail (Typha orientalis Presl.) were chosen. The two different species have different physiological properties and growth characteristics; for example, the yellow flag can remain active, but the acrial part of the oriental cattail will wither away in the winter. Furthermore, to investigate the microorganism community in the substrate, the V3-V4 regions of the bacteria 16S rDNA were sequenced via an Illumina MiSeq 2500 platform. Because of its higher integrity and broader range of applications23,24, 16S rDNA Illumina Miseq sequencing has recently been used frequently to study the microbial diversity in various environments25,26,27,28. Recently, this technology is also used for the analysis of microorganism communities in CWs29,30,31,32,33,34.
Results
Overall performance
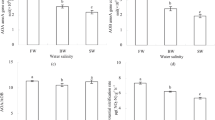
The influent characteristics and the time for water sample determination were described in the second section (Experimental Design and Operation). The overall performance of the two types of SSF-CWs is shown in Fig. 1. CWI had the highest removal rates of NH4+-N (80.50%), TN (47.50%) and COD (81.07%) and was superior to the control, which had removal rates of 72.25% (NH4+-N), 39.33% (TN) and 73.57% (COD), respectively. For CWT, only NH4+-N removal (75.75%) presented a slight advantage compared to the control, while the removal rates of TN (34.17%) and COD (71.43%) were the lowest among the three systems. The removal efficiency of NO3−-N in the three systems was consistent at approximately 33.00%. NO2−-N was not detected in the effluent.
Growth and physiological characteristics of the plants
Plant biomasses and nitrogen accumulations in the plants were determined during Stage III of the experiment (Table 1). The dry weight of the I. pseudacorus shoot increased slightly from 226.75 gDW·m−2 to 249.35 gDW·m−2, although no significant difference was observed. Conversely, the biomass of the T. orientalis Presl. shoot declined sharply from 135.24 gDW·m−2 to 50.91 gDW·m−2 for the dormancy of aboveground part. However, for the roots biomasses, both of species rose indistinctively. The I. pseudacorus shoot and T. orientalis Presl. shoot increased from 633.47 gDW·m−2 to 681.83 gDW·m−2 and 552.31 gDW·m−2 to 562.86 gDW·m−2, respectively. With the change in the plant biomasses, the nitrogen accumulation in the plants changed accordingly. An increase of 0.42 g·m−2, 0.38 g·m−2 and 0.06 g·m−2 and a decrease of 0.46 g·m−2 were recorded for the I. pseudacorus shoot, I. pseudacorus root, T. orientalis Presl. root and T. orientalis Presl. shoot, respectively. The rate of root radial oxygen loss (ROL) and the root vitality in the two plants were also measured (Table 1). The rate of ROL in I. pseudacorus was 5.95 μmolO2·g−1Root·h−1 and showed a striking difference compared to the T. orientalis Presl., which recorded as 1.73 μmolO2·g−1Root·h−1. Similarly, the root vitality of the I. pseudacorus root was 347.48 μgTTC·g−1Root·h−1 while it was 56.44 μgTTC·g−1Root·h−1 in the T. orientalis Presl. root.
Richness and diversity of the microbial communities
Three 16S rDNA libraries were established based on the Miseq Illumina sequencing of the three microorganism samples (Table 2). With 95% similarity, 4732, 3209 and 2991 Operational Taxonomic Units (OTUs) were clustered with a Good’s Coverage of 0.96, 0.97 and 0.98 in CWI, CWT and CWC, respectively. The total numbers of OTUs estimated by the ACE estimator were 8550, 7527 and 5401. The Shannon indexes were 6.57, 5.75 and 5.36 in CWI, CWT and CWC, respectively. With 97% similarity, 10393, 9118 and 7475 OTUs were clustered with a Good’s Coverage of 0.91, 0.90 and 0.92 respectively. The ACE estimators were 34531.54, 29659.38, 25433.24, and the Shannon indexes were 7.83, 7.87 and 7.28, respectively.
Microbial community composition
In addition to the richness and diversity of the microbial communities, the community composition in the substrate also plays a critical role in the pollutants removal in CWs. The abundance of different phyla of bacteria in the three types of SSF-CWs is shown in Fig. 2. A total of 24 identifiable phyla, in which Proteobacteria and Bacteroidetes were the two dominant species, were detected in the three systems. In CWI, Proteobacteria and Bacteroidetes accounted for 51.16% and 33.91%, respectively, followed by Cyanobacteria (6.05%) and Verrucomicrobia (4.23%). In CWT, Proteobacteria and Bacteroidetes accounted for 72.73% and 22.59%, respectively. In the control, Proteobacteria (64.29%) and Bacteroidetes (27.26%) constituted the primary phyla, and Cyanobacteria (3.39%) and Verrucomicrobia (2.55%) followed.
The bacterial composition of the three systems at the class level is shown in Fig. 3. Overall, 52, 42 and 50 classes were observed, with 13.80%, 5.26% and 5.71% of the total reads in each sample being undistinguishable at the present taxonomic level in CWI, CWT and CWC, respectively. The order of the primary classes was β-proteobacteria (26.77%) > Flavobacteria (14.13%) > γ-proteobacteria (13.57%) > Sphingobacteria (5.55%) > Chloroplast (4.93%) > δ-proteobacteria (4.48%) > α-proteobacteria (2.30%) in CWI. In CWT, it was β-proteobacteria (46.26%) > γ-proteobacteria (21.39%) > Flavobacteria (16.35%) > Sphingobacteria (2.12%) > α-proteobacteria (1.88%) > δ-proteobacteria (1.07%). In CWC, they mainly included γ-proteobacteria (35.72%), β-proteobacteria (18.51%), Flavobacteria (15.58%), Sphingobacteria (4.95%), δ-proteobacteria (3.91%), α-proteobacteria (2.68%) and ε-proteobacteria (2.18%).
The primary genera (relative abundance >0.50%) with the addition of three nitrifying bacteria (Nitrosomonas, Nitrosospira and Nitrospira) in the three systems are shown in Table 3. A total of 40 genera were listed, and they mainly belonged to two classes, β-proteobacteria (17 genera) and γ-proteobacteria (10 genera). The bacterial community composition showed a significant difference at the genera level among the three systems. In CWI, Flavobacterium (18.28%), Albidiferax (10.02%) and Ohtaekwangia (7.72%) constituted the three dominant genera. In CWT, they were Deefgea (22.21%), Flavobacterium (19.47%), Albidiferax (14.08%) and Halomonas (12.21%). In CWC, the two dominant genera composed of Halomonas (27.81%) and Flavobacterium (17.62%). Among the 40 listed genera, there were at least 8 genera reported to involve in denitrification. They included Azospira, Dechloromonas, Aeromonas, Shewanella, Halomonas, Pseudomonas, Arcobacter and Flavobacterium. The compositions of these denitrifying bacteria were also different among the three systems. In CWI, the order of their relative abundances was Flavobacterium (18.28%) > Halomonas (2.91%) > Aeromonas (1.81%) > Pseudomonas (1.68%) > Dechloromonas (1.29%) > Azospira (0.67%) > Shewanella (0.56%) > Arcobacter (0.24%). In CWT, there were richer Flavobacterium (19.47%) and Halomonas (12.21%). In CWC, the dominant denitrifying bacteria were Halomonas (27.81%) and Flavobacterium (17.62%), followed by Aeromonas (3.22%) and Arcobacter (2.99%). In addition, Nitrosomonas, Nitrosospira and Nitrospira were also observed in the three systems, although their relative abundances were not rich. CWI presented the highest proportion of nitrifying bacteria, and the relative abundances were 0.4140% (Nitrosomonas), 0.2402% (Nitrosospira) and 0.4318% (Nitrospira), respectively. The lowest percentage of the three nitrifiers was observed in CWT, which recorded as 0.2074% (Nitrosomonas), 0.0648% (Nitrosospira) and 0.0181% (Nitrospira), respectively. In CWC, their abundances were 0.3013% (Nitrosomonas), 0.1107% (Nitrosospira) and 0.1185% (Nitrospira), respectively.
Discussions
It was widely acknowledged that the transformation and removal of nitrogen in CWs were mainly attributed to nitrification and denitrification, followed by vegetation assimilation, substrate adsorption and the other pathways3,4,6,7,8,35. Because the sorption of ammonia by gravel or sand was not the primary means for ammonia removal22, only the nitrogen contents in the water and plants were determined in this research. Our results showed that the nitrogen accumulation in the vegetation was very limited during the variables determination stage (Stage III). This result was related to the weak growth of I. pseudacorus and the dormant aboveground part of T. orientalis during the cold season. Before the variable determination, the current systems had operated normally for 6 months, which was considered a long enough time for the microbial community in the substrate to reach a steady state13,36,37. Compared to traditional molecular methods, 16S rDNA Illumina Miseq sequencing is a more effective approach for studying microbial diversity due to its higher throughput and integrity as well as its broader application range23,24. PCR-DGGE is often restricted by an insufficient resolution to characterize the microbial communities in the complex samples13, while 454-pyrosequencing has a low throughput and high error rate and is costly24. In the present study, approximately ten thousand OTUs were identified in each sample, which were clustered with 97% similarity. This result was 4–5 times higher than that in several previous studies on this issue22,38.
Many researchers supported that the microorganism composition was closely related to the vegetable species in wetlands15,17,21,22,35,39. To reveal if there are any significant differences in the bacterial communities among the three types of CWs in the present research, a detailed comparison was carried out at different taxonomic levels. Firstly, the results indicated that Proteobacteria was the dominant phyla followed by Bacteroidetes in the three systems. This finding was consistent with a plenty of studies that reported Proteobacteria as the dominant phylum in various microorganism samples from the substrate or rhizosphere in wetlands22,38,40,41. Proteobacteria, which displayed a remarkably high level of bacterial metabolic diversity involved in global carbon, nitrogen and sulfur cycling, played an important role in pollutants removal40,42. Further analysis at the class level showed an obvious difference in bacterial community compositions among the three systems. The bacterial community mainly consisted of β-proteobacteria, Flavobacteria and γ-proteobacteria in CWI, while it was highly enriched with β-proteobacteria in CWT and the dominant class was γ-proteobacteria in CWC. Due to the amount of different influential factors, such as wetland types, substrate characteristics, wastewater components, operational parameters, environmental conditions and the different estimations by different approaches used in previous CW studies, the dominant class in CWs has been a long debated subject13,43,44. The present results suggested that the different plants with different growth characteristics and physiological properties had a considerable effect on the bacterial community compositions in CWs.
According to the list for genera in which at least one member had been characterized as a denitrifying strain reported by Heylen, et al.45 and Philippot, et al.46, it was found that a considerable proportion of the bacterial genera detected in the present research had a close relationship to denitrification. The compositions of denitrifying bacteria presented a remarkable difference among the three systems. The system grown with a hardy plant showed superior evenness of denitrifying bacteria, while certain genus such as Flavobacterium or Halomonas presented a fairly high abundance in the freezing-sensitive plant system and the unplanted system. However, it seemed that all the three types of SSF-CWs had adequate denitrification capability.
The difference in the overall efficiency of nitrogen removal among the three systems might result from the nitrification process. The bacteria involved in nitrification mainly consisted of two groups, the ammonium-oxidizing bacteria (AOB), which converted ammonium to nitrite, and the nitrite-oxidizing bacteria (NOB), which converted nitrite to nitrate11. A higher ratio of NOB to AOB populations was considered indicative of a higher nitrification capacity and more complete ammonia oxidation47. In this research, higher relative abundance of nitrifying bacteria and higher ratio of NOB to AOB populations accompanied by higher removal efficiency of NH4+-N were recorded in CWI compared to CWT and CWC. This phenomenon could be attributed to more oxygen being released from the hardy plant I. pseudacorus6,15. Zhong, et al.11 also reported that enhanced levels of oxygen and nitrite favoured the growth of NOB in CWs. In the present experiment, Nitrosomonas and Nitrosospira were the two dominant AOB lineages. This finding was consistent with a previous study in a vertical flow CW by Tietz, et al.48. This study also found that Nitrosomonas in the three systems was more abundant than Nitrosospira. The result might be related to the high influent NH4+-N concentration, which contributed to the formation of a community dominated by Nitrosomonas, because Nitrosomonas had a lower substrate affinity but a higher maximum activity than Nitrosospira49,50. Moreover, Nitrospira was the only NOB detected in the present systems. These results suggested that the three nitrifying bacteria, Nitrosomonas, Nitrosospira and Nitrospira, shared the most responsibility for nitrification.
The diversity of the microbial composition suggested a diversity of the functional characteristics that were relevant to pollutant removal in wastewater51. Any difference in microbial functional groups might lead directly to the difference in the purification performance of the CWs11. Alpha diversity analysis suggested that the presence of a hardy plant induced higher diversity and evenness of bacterial community composition at different taxonomic levels ranging from class to genus. This might be related to the multiple influencing mechanisms of plants on microorganisms such as complicating the attachment surface, providing organic carbon, as well as changing the oxidation-reduction conditions in the wetlands1,6,15. The results revealed by the ACE estimator and Shannon index indicated that the system grown with hardy plant had higher microbial richness. This finding was consistent with several earlier studies22,38,52,53. The oxygen released from plant roots could affect the microorganism richness in the substrate39. Previous studies have shown that the bacterial communities were grouped according to an oxygen concentration gradient40,54,55. In the current study, the two plants grown in the CWs have different physiological properties and growth characteristics. During the experiment period, I. pseudacorus remained active, but the acrial part of T. orientalis withered away. The rate of root radial oxygen loss (ROL) was also higher in CWI compared to CWT. This might contribute to a higher richness of aerobic microorganism in CWI. Moreover, the carbon source supply is also an important factor that influences the microbial communities in the substrate6,8. The senescent or dead plant could release a variety of labile organic compounds such as amino acids, sugars, volatile fatty acids, etc.56. In this research, T. orientalis, a dormant plant during the winter, showed a decline in shoot biomass because of the senescence of leaves during the determination period. Chen et al.38,57,58 suggested that the enhanced litter decomposition might contribute to increased microbial richness in the substrate of the CW. However, in this study, the effect of plant litter on denitrification efficiency enhancement was limited in the T. orientalis system. This could be because quite a large proportion of plant litter, such as highly crystalline lignocellulose, was recalcitrant and the complete degradation of litter needed a relatively long time7,59. Therefore, the present results suggested a positive effect of the hardy plant on the nitrogen removal in SSF-CWs during the winter, while the freezing-sensitive plant showed slight differences on the pollutants removal compared to the control.
In addition, some other factors may also have significant impact on the contaminants removal in CWs. Plant root exudates can also be used as the dissolved organic matter for the metabolism of heterotrophic bacteria13. In the current research, due to the limitations of experimental conditions, the root vitality was measured as a proxy indicator. The results seem to indicate that the presence of I. pseudacorus have more active effects towards the microbial community compositions in the substrate. Moreover, the root activity and growth status of the macrophytes in CWs also influence the richness and enzyme activity of microbial populations10,52,60. Wang, et al.61 reported that plant ROL played an important role in nutrient removal by significantly affecting microbial abundance and activity in CWs. Additionally, anammox was also recognized as another approach for nitrogen removal apart from nitrification-denitrification62. However, the anammox bacteria, placed in the order Brocadiales in the phylum Planctomycetes, has not been observed in this study. Besides, the coexistence of some autotrophic bacteria such as Cyanobacteria in wetlands was also reported benefit to the pollutants degradation, because they could provide the heterotrophic bacteria with oxygen, which played a similar and complementary role in addition to the macrophytes to a certain degree22,63. Furthermore, some archaea with specific activity of ammonia oxidation also play an indispensable role in nitrogen removal processes64,65. All these factors should be taken into consideration in further researches.
In conclusion, 16S rDNA Illumina Miseq sequencing revealed detailed information about the interaction between the plant and the bacterial community composition in SSF-CWs. The hardy plant I. pseudacorus had a positive effect on the bacterial abundance and eventually on the community composition in the substrate. The presence of a hardy plant with active root oxygen release enhanced the relative abundance of nitrifying bacteria in the substrate, and consequently was supposed to contribute to the increase in the nitrogen removal efficiency of the system during the winter. However, some other factors, such as the bacteria population and activity, the presence of Cyanobacteria and ammonia-oxidizing archaea may also have an important influence on the nitrogen removal in SSF-CWs. Hence, further researches will be needed to understand detailed mechanisms referring to the biodegradation process of pollutants in CWs.
Methods
Experimental Design and Operation
To simulate subsurface flow constructed wetlands (SSF-CWs), four outdoor mesocosms were planted with I. pseudacorus, four were planted with T. orientalis Presl., both at an initial density of 30 plants per m2, and four controls without plants were built in Huai’an, Jiangsu province, China (33.3°N, 119.0°E). The dimensions of each mesocosm were (0.8 m)2 × π × 0.75 m, and each bed consisted of three layers: 100 mm deep rough sand (1–2 mm in diameter) on the top, 100 mm deep gravel (10–20 mm in diameter, porosity of 0.45) in the middle, and 550 mm deep gravel (30–50 mm in diameter, porosity of 0.55) at the bottom (Fig. S1).
The operation of the SSF-CWs was divided into three stages. (1) Stage I (From June 10th, 2014 to Sept 10th, 2014) was for plants growth. The mesocosms were fed continuously with natural water until the plant shoots grew long enough. (2) Stage II (From Sept 10th, 2014 to Dec 10th, 2014) was for microbial establishment. The mesocosms were fed continuously with wastewater with appropriate artificial modification on the base of a secondary effluent from a sewage treatment plant for domestic wastewater. The pH of the influent water was 7.12–7.88, the average hydraulic loading rate (HLR) was 187.5 mm·d−1 and the hydraulic retention time (HRT) was 4 days. The concentrations of dissolved oxygen (DO), chemical oxygen demand (COD), ammonia nitrogen (NH4+-N), nitrite nitrogen (NO2−-N), nitrate nitrogen (NO3−-N) and total nitrogen (TN) in the influent water were 9.2–9.8 mg·L−1, 55.0–65.0 mg·L−1, 9.5–12.5 mg·L−1, 0.3–0.6 mg·L−1, 4.5–5.5 mg·L−1 and 18.0–24.0 mg·L−1, respectively. (3) Stage III (From Dec 10th, 2014 to Feb 10th, 2015) was for variables determination. The operation scheme was the same as Stage II. The water temperature was 4.3–6.7 °C during Stage III.
Water Sampling and Analysis
The water temperature in the mesocosms was recorded by the Temperature and Illuminance Data Logger (HOBO Pendant UA-002-08, Onset, USA). The pH was determined by a portable Multi-parameter Water Quality Meter (U-52, HORIBA, Japan). Dissolved oxygen (DO) was monitored in situ using DO electrodes (HQ40d-53 LED, HACH, USA). The concentrations of NH4+-N, NO2−-N, NO3−-N, TN and COD were determined with a Water Quality Analyzing System (DRB200 and DR2800, HACH, USA). All variables were analysed according to standard analytical procedures66.
Plant Sampling and Analysis
Plant samples were harvested and separated into roots and shoots, dried at 65 °C to a constant weight, and then ground into powder. The N content was determined by an Elemental analyzer (CHN-O-Rapid, Heraeus, Germany) at the beginning and end of the experiment67. The rate of root radial oxygen loss (ROL) was measured using the titanium (III) citrate buffer method68,69. The root vitality was quantified via the triphenyl tetrazolium chloride (TTC) method70.
Microbial Sampling and Analysis
Preparation of microbial samples
As shown in Fig. S1, microorganism sampling points of each system was divided into three layers according to the substrate. Each layer comprised four sampling points. The four points formed a circle with a radius of 0.4 m around the system center, and the connections between each point and the system center formed a cross. 100 g of sand from the top layer and 100 g of gravel from the middle layer were obtained by a cylindrical sampler with 2.5 cm diameter, while 200 g of gravel were obtained from the bottom layer by a sampling scoop on Feb 2nd 2015. Then, the samples in each system were mixed well and vigorously shaken at 200 rpm for 3 h in sterile glass bottles to isolate the biofilm from the substrate surface. After centrifuging twice (6,000 × g, 15 min), the precipitate was collected for subsequent DNA extraction, 16S rDNA PCR amplification and Illumina MiSeq sequencing.
DNA extraction and PCR amplification
DNA was extracted from the samples using a QIAamp Fast DNA Stool Mini Kit (QIAGEN, Chatsworth, CA, USA) according to the manufacturer’s instructions. DNA yields were determined using a SpectraMax 190 (Molecular Devices, California, USA), and the integrity was evaluated via 1.0% agarose gel electrophoresis. Then, DNA was diluted to 1 ng·μL−1 in sterile water. The universal primer sets 341F (5′-CCTAYGGGRBGCASCAG-3′) and 785R (5′-GACTACHVGGGTATCTAATCC-3′) were used for amplification of the V3-V4 regions of 16S rDNA. A 10 ng template, 0.5 μL of forward primers and 0.5 μL of reverse primers were added into a 25 μL reaction system for the PCR reaction. Thermal cycling consisted of denaturation at 94 °C for 3 min, which was followed by 30 cycles of 94 °C for 10 s, 55 °C for 15 s and 72 °C for 30 s, and finally held at 72 °C for 7 min. AmpureBeads (Beckman Coulter, Inc., CA, USA) were used for the PCR product purification.
16S rDNA Illumina MiSeq sequencing
The sequencing libraries were constructed with an NEB Next Ultra DNA Library Prep Kit for Illumina (New England Biolabs Inc., Boston, MA, USA), and then a Qubit 2.0 Fluorometer (Life Invitrogen, Inc., Carlsbad, CA, USA) was used to assess the libraries quality. Then, the libraries were sequenced on an Illumina MiSeq 2500 platform.
Quality filtering, OTUs picking, annotation and diversity analysis
The FASTX-Toolkit (version 0.0.14) and Mothur program (version 1.34.0) were used for the analysis, merging and quality filtering of the raw data sequences. All of the reads were quality filtered using an average quality value of 20 (Q20) during demultiplexing. Short reads (length < 40 bp) and chimeras were excluded. Reads were clustered by degree similarity levels with the UCLUST program (version 1.2.22q). Sequences with ≥ 97% similarity were assigned to the same genus. A RDP classifier (version 2.2) was used to annotate the taxonomic information. The Mothur program was used for alpha diversity analysis. ACE estimated the richness of the species, while the Shannon index estimated the diversity of the species.
Statistical Analysis
All statistical tests were performed with the statistical program SPSS 17.0 (SPSS Inc. Chicago, USA). The data were analysed with one-way analysis of variance to compare the performance of each mesocosm. Duncan’s test was performed to detect the statistical significance of differences (p > 0.05) between the mean values of the treatments.
Additional Information
How to cite this article: Wang, P. et al. A Hardy Plant Facilitates Nitrogen Removal via Microbial Communities in Subsurface Flow Constructed Wetlands in Winter. Sci. Rep. 6, 33600; doi: 10.1038/srep33600 (2016).
References
Vymazal, J. Constructed Wetlands for Wastewater Treatment: Five Decades of Experience. Environ Sci Technol 45, 61–69 (2011).
Vymazal, J. Constructed wetlands for treatment of industrial wastewaters: A review. Ecological Engineering 73, 724–751 (2014).
Wu, S., Kuschk, P., Brix, H., Vymazal, J. & Dong, R. Development of constructed wetlands in performance intensifications for wastewater treatment: a nitrogen and organic matter targeted review. Water Res 57, 40–55 (2014).
Wu, S. et al. Treatment of industrial effluents in constructed wetlands: challenges, operational strategies and overall performance. Environ Pollut 201, 107–120 (2015).
Zhang, D. Q. et al. Application of constructed wetlands for wastewater treatment in tropical and subtropical regions (2000–2013). J Environ Sci (China) 30, 30–46 (2015).
Faulwetter, J. L. et al. Microbial processes influencing performance of treatment wetlands: A review. Ecological Engineering 35, 987–1004 (2009).
Kadlec, R. & Wallace, S. Treatment Wetlands, 2nd (CRC Press, 2009).
Meng, P., Pei, H., Hu, W., Shao, Y. & Li, Z. How to increase microbial degradation in constructed wetlands: influencing factors and improvement measures. Bioresour Technol 157, 316–326 (2014).
Wu, Y., Li, T. & Yang, L. Mechanisms of removing pollutants from aqueous solutions by microorganisms and their aggregates: a review. Bioresour Technol 107, 10–18 (2012).
Cui, L., Ouyang, Y., Gu, W., Yang, W. & Xu, Q. Evaluation of nutrient removal efficiency and microbial enzyme activity in a baffled subsurface-flow constructed wetland system. Bioresour Technol 146, 656–662 (2013).
Zhong, F. et al. Bacterial community analysis by PCR-DGGE and 454-pyrosequencing of horizontal subsurface flow constructed wetlands with front aeration. Appl Microbiol Biotechnol 99, 1499–1512 (2015).
Brix, H. & Arias, C. A. Danish guidelines for small-scale constructed wetland systems for onsite treatment of domestic sewage. Water Sci & Technol 51, 1–9 (2005).
Truu, M., Juhanson, J. & Truu, J. Microbial biomass, activity and community composition in constructed wetlands. Sci Total Environ 407, 3958–3971 (2009).
Vymazal, J. & Kröpfelová, L. Removal of organics in constructed wetlands with horizontal sub-surface flow: a review of the field experience. Sci Total Environ 407, 3911–3922 (2009).
Faulwetter, J. L., Burr, M. D., Parker, A. E., Stein, O. R. & Camper, A. K. Influence of season and plant species on the abundance and diversity of sulfate reducing bacteria and ammonia oxidizing bacteria in constructed wetland microcosms. Microb Ecol 65, 111–127 (2013).
Shelef, O., Gross, A. & Rachmilevitch, S. Role of Plants in a Constructed Wetland: Current and New Perspectives. Water 5, 405–419 (2013).
Ahn, C., Gillevet, P. M. & Sikaroodi, M. Molecular characterization of microbial communities in treatment microcosm wetlands as influenced by macrophytes and phosphorus loading. Ecological Indicators 7, 852–863 (2007).
DeJournett, T. D., Arnold, W. A. & LaPara, T. M. The characterization and quantification of methanotrophic bacterial populations in constructed wetland sediments using PCR targeting 16S rRNA gene fragments. Applied Soil Ecology 35, 648–659 (2007).
Gorra, R., Coci, M., Ambrosoli, R. & Laanbroek, H. J. Effects of substratum on the diversity and stability of ammonia-oxidizing communities in a constructed wetland used for wastewater treatment. J Appl Microbiol 103, 1442–1452 (2007).
Baptista, J. C., Davenport, R. J., Donnelly, T. & Curtis, T. P. The microbial diversity of laboratory-scale wetlands appears to be randomly assembled. Water Res 42, 3182–3190 (2008).
Knief, C., Ramette, A., Frances, L., Alonso-Blanco, C. & Vorholt, J. A. Site and plant species are important determinants of the Methylobacterium community composition in the plant phyllosphere. ISME J 4, 719–728 (2010).
Wang, Q. et al. Microbial abundance and community in subsurface flow constructed wetland microcosms: role of plant presence. Environ Sci Pollut Res Int, 1–10 (2015).
Glenn, T. C. Field guide to next-generation DNA sequencers. Mol Ecol Resour 11, 759–769 (2011).
Liu, L. et al. Comparison of next-generation sequencing systems. J Biomed Biotechnol 2012, 1–11 (2012).
Staley, C. et al. Application of Illumina next-generation sequencing to characterize the bacterial community of the Upper Mississippi River. J Appl Microbiol 115, 1147–1158 (2013).
Moreau, M. M., Eades, S. C., Reinemeyer, C. R., Fugaro, M. N. & Onishi, J. C. Illumina sequencing of the V4 hypervariable region 16S rRNA gene reveals extensive changes in bacterial communities in the cecum following carbohydrate oral infusion and development of early-stage acute laminitis in the horse. Vet Microbiol 168, 436–441 (2014).
Zhang, J., Sun, Q. L., Zeng, Z. G., Chen, S. & Sun, L. Microbial diversity in the deep-sea sediments of Iheya North and Iheya Ridge, Okinawa Trough. Microbiol Res 177, 43–52 (2015).
Walujkar, S. A. et al. Characterization of bacterial community shift in human Ulcerative Colitis patients revealed by Illumina based 16S rRNA gene amplicon sequencing. Gut Pathogens 6, 1–11 (2014).
Ligi, T. et al. Characterization of bacterial communities in soil and sediment of a created riverine wetland complex using high-throughput 16S rRNA amplicon sequencing. Ecological Engineering 72, 56–66 (2014).
Guan, W., Yin, M., He, T. & Xie, S. Influence of substrate type on microbial community structure in vertical-flow constructed wetland treating polluted river water. Environ Sci Pollut Res Int 22, 16202–16209 (2015).
He, T., Guan, W., Luan, Z. & Xie, S. Spatiotemporal variation of bacterial and archaeal communities in a pilot-scale constructed wetland for surface water treatment. Appl Microbiol Biotechnol (2015).
Martinez-Lavanchy, P. M. et al. Microbial Toluene Removal in Hypoxic Model Constructed Wetlands Occurs Predominantly via the Ring Monooxygenation Pathway. Appl Environ Microbiol 81, 6241–6252 (2015).
Wu, J. et al. Biological mechanisms associated with triazophos (TAP) removal by horizontal subsurface flow constructed wetlands (HSFCW). Sci Total Environ 553, 13–19 (2016).
Zhao, X., Yang, J., Bai, S., Ma, F. & Wang, L. Microbial population dynamics in response to bioaugmentation in a constructed wetland system under 10 degrees C. Bioresour Technol 205, 166–173 (2016).
Meng, P., Hu, W., Pei, H., Hou, Q. & Ji, Y. Effect of different plant species on nutrient removal and rhizospheric microorganisms distribution in horizontal-flow constructed wetlands. Environ Technol 35, 808–816 (2014).
Weber, K. P. & Legge, R. L. Dynamics in the bacterial community-level physiological profiles and hydrological characteristics of constructed wetland mesocosms during start-up. Ecological Engineering 37, 666–677 (2011).
Samso, R. & Garcia, J. Bacteria distribution and dynamics in constructed wetlands based on modelling results. Sci Total Environ 461–462, 430–440 (2013).
Chen, Y. et al. Effects of plant biomass on bacterial community structure in constructed wetlands used for tertiary wastewater treatment. Ecological Engineering 84, 38–45 (2015).
Ruiz-Rueda, O., Hallin, S. & Baneras, L. Structure and function of denitrifying and nitrifying bacterial communities in relation to the plant species in a constructed wetland. FEMS Microbiol Ecol 67, 308–319 (2009).
Ansola, G., Arroyo, P. & Saenz de Miera, L. E. Characterisation of the soil bacterial community structure and composition of natural and constructed wetlands. Sci Total Environ 473–474, 63–71 (2014).
Zhang, J. et al. Comparisons of microbial abundance and community among different plant species in constructed wetlands in summer. Ecological Engineering 82, 376–380 (2015).
Kersters, K. et al. In The Prokaryotes Vol. 5 (eds Dwarkin, M. et al.) 3–37 (Springer, 2006).
Ibekwe, A. M., Lyon, S. R., Leddy, M. & Jacobson-Meyers, M. Impact of plant density and microbial composition on water quality from a free water surface constructed wetland. J Appl Microbiol 102, 921–936 (2007).
Calheiros, C. S. et al. Bacterial community dynamics in horizontal flow constructed wetlands with different plants for high salinity industrial wastewater polishing. Water Res 44, 5032–5038 (2010).
Heylen, K. et al. Cultivation of denitrifying bacteria: optimization of isolation conditions and diversity study. Appl Environ Microbiol 72, 2637–2643 (2006).
Philippot, L., Hallin, S. & Schloter, M. Ecology of Denitrifying Prokaryotes in Agricultural Soil. Advances in Agronomy 96, 249–305 (2007).
Wittebolle, L., Vervaeren, H., Verstraete, W. & Boon, N. Quantifying community dynamics of nitrifiers in functionally stable reactors. Appl Environ Microbiol 74, 286–293 (2008).
Tietz, A., Hornek, R., Langergraber, G., Kreuzinger, N. & Haberl, R. Diversity of ammonia oxidising bacteria in a vertical flow constructed wetland. Water Science & Technology 56, 241 (2007).
Juretschko, S. et al. Combined molecular and conventional analyses of nitrifying bacterium diversity in activated sludge: Nitrosococcus mobilis and Nitrospira-like bacteria as dominant populations. Appl Environ Microb 64, 3042–3051 (1998).
Schramm, A. et al. Structure and function of a nitrifying biofilm as determined by in situ hybridization and the use of microelectrodes. Appl Environ Microb 62, 4641–4647 (1996).
Saunders, A. M., Larsen, P. & Nielsen, P. H. Comparison of nutrient-removing microbial communities in activated sludge from full-scale MBRs and conventional plants. Water Sci Technol 68, 366–371 (2013).
Salvato, M. et al. Wetland plants, micro-organisms and enzymatic activities interrelations in treating N polluted water. Ecological Engineering 47, 36–43 (2012).
Menon, R., Jackson, C. R. & Holland, M. M. The Influence of Vegetation on Microbial Enzyme Activity and Bacterial Community Structure in Freshwater Constructed Wetland Sediments. Wetlands 33, 365–378 (2013).
Armstrong, W., Armstrong, J. & Beckett, P. In Constructed Wetlands in Water Pollution Control (eds Cooper, P. F. & Findlater, B. C. ) 41–52 (Pergamon Press, 1990).
Luederitz, V., Eckert, E., Lange-Weber, M., Lange, A. & Gersberg, R. Nutrient removal efficiency and resource economics of vertical flow and horizontal flow constructed wetlands. Ecological Engineering 18, 157–171 (2001).
Vymazal, J. Removal of nutrients in various types of constructed wetlands. Sci Total Environ 380, 48–65 (2007).
Chen, Y. et al. Effects of dissolved oxygen on extracellular enzymes activities and transformation of carbon sources from plant biomass: implications for denitrification in constructed wetlands. Bioresour Technol 102, 2433–2440 (2011).
Chen, Y. et al. Effects of cattail biomass on sulfate removal and carbon sources competition in subsurface-flow constructed wetlands treating secondary effluent. Water Res 59, 1–10 (2014).
Hume, N., Fleming, M. & Horne, A. Plant carbohydrate limitation on nitrate reduction in wetland microcosms. Water Res 36, 577–584 (2002).
Huang, L. et al. Correlation among soil microorganisms, soil enzyme activities, and removal rates of pollutants in three constructed wetlands purifying micro-polluted river water. Ecological Engineering 46, 98–106 (2012).
Wang, Q. et al. Effect of plant harvesting on the performance of constructed wetlands during winter: radial oxygen loss and microbial characteristics. Environ Sci Pollut Res Int 22, 7476–7484 (2015).
Zhu, G., Jetten, M. S., Kuschk, P., Ettwig, K. F. & Yin, C. Potential roles of anaerobic ammonium and methane oxidation in the nitrogen cycle of wetland ecosystems. Appl Microbiol Biotechnol 86, 1043–1055 (2010).
Subashchandrabose, S. R., Ramakrishnan, B., Megharaj, M., Venkateswarlu, K. & Naidu, R. Consortia of cyanobacteria/microalgae and bacteria: biotechnological potential. Biotechnol Adv 29, 896–907 (2011).
Fan, L.-F., Chen, H.-J., Hsieh, H.-L., Lin, H.-J. & Tang, S.-L. Comparing abundance, composition and environmental influences on prokaryotic ammonia oxidizers in two subtropical constructed wetlands. Ecological Engineering 90, 336–346 (2016).
Zhou, L., Wang, S., Zou, Y., Xia, C. & Zhu, G. Species, Abundance and Function of Ammonia-oxidizing Archaea in Inland Waters across China. Sci Rep 5, 15969 (2015).
APHA (American Public Health Association, 1998).
Toet, S., Bouwman, M., Cevaal, A. & Verhoeven, J. T. A. Nutrient Removal Through Autumn Harvest ofPhragmites australisandThypha latifoliaShoots in Relation to Nutrient Loading in a Wetland System Used for Polishing Sewage Treatment Plant Effluent. Journal of Environmental Science and Health, Part A 40, 1133–1156 (2005).
Wu, C., Ye, Z., Shu, W., Zhu, Y. & Wong, M. Arsenic accumulation and speciation in rice are affected by root aeration and variation of genotypes. J Exp Bot 62, 2889–2898 (2011).
Kludze, H. K., DeLaune, R. D. & Petric Jr, W. H. A Colorimetric Method for Assaying Dissolved Oxygen Loss from Container-Grown Rice Roots. Agorn. J. 86, 483–487 (1994).
Li, H., Sun, Q., Zhao, S. & Zhang, W. Principles and techniques of plant physiological biochemical experiment (Higher Education Press, 2000).
Acknowledgements
This research was supported by the Major Science and Technology Program for Water Pollution Control and Treatment (No. 2012ZX07204-004, No. 2014ZX07204-005-002). We would like to thank Genergy Biotechnology Limited Corporation (Shanghai, China) for the 16S rDNA Illumina Miseq analysis.
Author information
Authors and Affiliations
Contributions
D.Z. and S.A. designed the experiment, P.W., X.Z., J.Z. and H.Z. performed the experiment, Z.Z. and N.J. did the statistical analysis, P.W. wrote the first draft of the manuscript, H.Z. and X.L. provided many valuable views, comments and suggestions for the accomplishment of the revised paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wang, P., Zhang, H., Zuo, J. et al. A Hardy Plant Facilitates Nitrogen Removal via Microbial Communities in Subsurface Flow Constructed Wetlands in Winter. Sci Rep 6, 33600 (2016). https://doi.org/10.1038/srep33600
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep33600
This article is cited by
-
Improving the removal efficiency of nitrogen and organics in vertical-flow constructed wetlands: the correlation of substrate, aeration and microbial activity
Environmental Science and Pollution Research (2022)
-
Urban Wastewater Treatment by Pilot-Scale Vertical Subsurface Flow Constructed Wetland Planted with Typha latifolia and Phragmite australis Under Arid Climate
Water, Air, & Soil Pollution (2022)
-
Nitrogen removal performance and bacterial communities in zeolite trickling filter under different influent C/N ratios
Environmental Science and Pollution Research (2021)
-
Metagenomics analysis of rhizospheric bacterial communities of Saccharum arundinaceum growing on organometallic sludge of sugarcane molasses-based distillery
3 Biotech (2020)
-
Enhanced nitrogen removal in filled-and-drained vertical flow constructed wetlands: microbial responses to aeration mode and carbon source
Environmental Science and Pollution Research (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.