Abstract
Gout is a common disease resulting from hyperuricemia. Recently, a genome-wide association study identified an association between gout and a single nucleotide polymorphism (SNP) rs2188380, located on an intergenic region between MYL2 and CUX2 on chromosome 12. However, other genes around rs2188380 could possibly be gout susceptibility genes. Therefore, we performed a fine-mapping study of the MYL2-CUX2 region. From 8,595 SNPs in the MYL2-CUX2 region, 9 tag SNPs were selected and genotyping of 1,048 male gout patients and 1,334 male controls was performed by TaqMan method. Eight SNPs showed significant associations with gout after Bonferroni correction. rs671 (Glu504Lys) of ALDH2 had the most significant association with gout (P = 1.7 × 10−18, odds ratio = 0.53). After adjustment for rs671, the other 8 SNPs no longer showed a significant association with gout, while the significant association of rs671 remained. rs671 has been reportedly associated with alcohol drinking behavior and it is well-known that alcohol drinking elevates serum uric acid levels. These data suggest that rs671, a common functional SNP of ALDH2, is a genuine gout-associated SNP in the MYL2-CUX2 locus and that “A” allele (Lys) of rs671 plays a protective role in the development of gout.
Similar content being viewed by others
Introduction
Gout is a common disease resulting from hyperuricemia and causes acute arthritis. Previous genetic and functional analyses revealed that ABCG2 dysfunctional variants caused gout1,2,3 due to decreased urate excretion in gut4 and kidney5. Genome-wide association studies (GWASs) of gout also showed genome-wide significant associations with ABCG2 and GLUT96,7,8. Recently, we revealed for the first time that the following 3 loci were associated with gout at the genome-wide significance level: rs1260326 of GCKR, rs4073582 of CNIH-2 and rs2188380 of MYL2-CUX28. Among them, 2 SNPs are located in gene regions: rs1260326 is a nonsynonymous single nucleotide polymorphism (SNP) (Leu446Pro) of GCKR and rs4073582 is an intronic SNP of CNIH-2. On the other hand, rs2188380 is located on an intergenic region between MYL2 and CUX28. Additionally, we detected many SNPs showing significant associations with gout across the chromosome 12q24 region which were in strong linkage disequilibrium (LD) with rs2188380. MYL2 encodes a regulatory light chain associated with cardiac myosin β (or slow) heavy chain and an association between MYL2 variant and high-density lipoprotein cholesterol metabolism was previously reported9. CUX2 regulates cell-cycle progression10 and plays important roles in neural progenitor development in the central nervous system10,11. An association between CUX2 and type 1 diabetes has also been reported12. However, there is a possibility that the other genes around rs2188380 of MYL2-CUX2 can be gout susceptibility genes. Therefore, we performed fine-mapping of the MYL2-CUX2 region and a further association analysis of gout.
Results
From 8,595 SNPs in the MYL2-CUX2 region within 10 Mb across rs2188380, 45 SNPs in LD (r2 ≥ 0.3) with rs2188380 were selected (Supplementary Figure S1). Among these 45 SNPs and rs2188380, 9 tag SNPs were selected for association analysis (Fig. 1 and Supplementary Table S1). Genotyping results of the 9 tag SNPs for 1,048 gout patients and 1,334 controls were shown in Table 1. The call rates for the 9 SNPs were more than 95.0%. All the SNPs in the control group were in Hardy-Weinberg equilibrium (P > 0.05). Except for rs2555004, the other 8 SNPs showed significant associations at P < 5.6 × 10−3 (=0.05/9) with the Bonferroni correction and rs671 (Glu504Lys) of aldehyde dehydrogenase 2 (ALDH2) had the most significant association with gout (P = 1.7 × 10−18; odds ratio [OR] = 0.53; 95% confidence interval [CI]: 0.46–0.61, Table 1 and Supplementary Figure S2A).
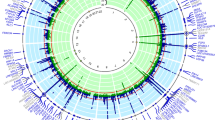
Linkage disequilibrium heat map of 46 SNPs.
We examined the linkage disequilibrium (LD) between each pair of 46 SNPs and searched the SNPs that were tagging other SNPs with strong LD (r2 ≥ 0.8). The 9 tag SNPs (rs7978484, rs16940688, rs2071629, rs2188380, rs11065783, rs3809297, rs4766566, rs671 and rs2555004), which are shown in bold and boxes, were selected for association analysis.
Next, the multivariate logistic regression analyses were performed to evaluate whether there was an additional association signal after the adjustment for the most significantly associated SNP rs671. We set the significance threshold as α = 6.3 × 10−3 (=0.05/8) with the Bonferroni correction. While the significant association of rs671 remained, the other 8 SNPs no longer showed a significant association with gout after the adjustment for rs671 (Table 2 and Supplementary Figure S2B). Among rs671 and 6 tagged SNPs (rs3782886, rs11066015, rs4646776, rs11066132, rs2074356 and rs11066280) shown in Supplementary Table S2, rs671 is the most promising functional SNP because only rs671 is a nonsynonymous variant (Glu504Lys). rs4766566 had a nominally significant association (P = 0.032), but did not pass the significant threshold for multiple hypothesis testing (Table 2). Additionally, the OR for rs4766566 became closer to 1.0 after the adjustment for rs671 (from 0.59 to 0.82; Tables 1 and 2). These data suggest that rs671 (Glu504Lys) of ALDH2 is a genuine gout-associated SNP in the MYL2-CUX2 locus.
It is well-known that individuals with rs671 heterozygotes (A/G; Lys/Glu) have only 6.25% of enzyme activity of those with normal ALDH2 (G/G; Glu/Glu) and those with homozygotes (A/A; Lys/Lys) show almost no activity13. Therefore, it is expected that dominant model (G/G v.s. A/G or A/A) is the most likely genetic model for the association between rs671 and gout. Actually, the statistical significance of the association between rs671 and gout was improved by applying the dominant model (P = 2.9 × 10−21; OR = 0.44; 95% CI: 0.37–0.52) compared with the result from the allelic model (P = 1.7 × 10−18; OR = 0.53; 95% CI: 0.46–0.61; Table 1). These findings indicate that “A” allele (Lys) of rs671 plays a protective role in the development of gout.
Discussion
In this study, among 9 tag SNPs selected from 8,595 SNPs in the MYL2-CUX2 locus, only the association of rs671 of ALDH2 remained significant after the adjustment for each SNP with the Bonferroni correction (Table 2). Together with the fact that rs671 (Glu504Lys) is a well-known dysfunctional SNP, we therefore concluded that rs671 is a genuine gout-associated SNP. Indeed, a previous Japanese study with 180 gout cases and 49 controls has indicated that the frequency of homozygotes (A/A; Lys/Lys) of rs671 was lower in gout patients than in controls14. The fine-mapping study for the associational signal on chromosome 12 identified by our GWAS reached the consistent result with the previous finding showing the association between ALDH2 and gout14.
ALDH2 is a crucial enzyme in the alcohol metabolism. Alcohol is oxidized to acetaldehyde by alcohol dehydrogenase and acetaldehyde is further metabolized to acetate by aldehyde dehydrogenase15, which largely depends on ALDH2. rs671 (Glu504Lys), a common missense SNP of ALDH2 gene, severely decreases the activity of ALDH2 enzyme13. When acetate is metabolized to acetyl-coenzyme A, adenosine triphosphate (ATP) hydrolyzes to adenosine monophosphate (AMP) which is ultimately metabolized to uric acid. This alcohol metabolism is one of the reasons why alcohol drinking elevates serum uric acid (SUA) levels. Thus, the association between rs671 and gout is partly due to alcohol drinking.
The allele frequencies of rs671 of ALDH2 differ among populations: the 504Lys allele (“A” allele) is common in East Asians including Japanese, but quite rare in other populations such as European and African descendants16. Therefore, it is reasonable that this SNP has not been detected in the previous GWASs of gout in Europeans and African Americans due to its low frequency. We showed that rs671 of ALDH2 is an influential genetic factor for Japanese as the other 4 previously reported loci8 (ABCG2, SLC2A9, GCKR and CNIH-2) of gout and further investigations in East Asian populations will be able to warrant these findings.
In summary, Glu504Lys polymorphism (rs671), a common dysfunctional SNP of ALDH2, is identified as a genuine gout-associated polymorphism in the MYL2-CUX2 locus and “A” allele (Lys) of rs671 plays a protective role in the development of gout.
Methods
Study participants
This study was approved by the institutional ethical committees (National Defense Medical College and Nagoya University) and all procedures involved in this study were performed in accordance with the Declaration of Helsinki with written informed consent from each subject.
1,048 gout cases were assigned from the Japanese male outpatients at the gout clinics of Kyoto Industrial Health Association (Kyoto, Japan), or Ryougoku East Gate Clinic (Tokyo, Japan). All patients were clinically diagnosed as primary gout according to the criteria established by the American College of Rheumatology17. Patients with inherited metabolism disorders including Lesch–Nyhan syndrome were excluded. For the control group, 1,334 Japanese males with normal SUA (≤7.0 mg/dl) and without a history of gout were collected from the participants in the Shizuoka area in the Japan Multi-Institutional Collaborative Cohort Study (J-MICC Study)18,19. The details of participants in this study are shown in Supplementary Table S3.
Selection of SNPs
First, 8,595 SNPs within 10 Mb across rs2188380 were selected using HapMap phase III JPT samples (http://hapmap.ncbi.nlm.nih.gov/)20; then, the pairwise LD was calculated between rs2188380 and the 8,595 SNPs (Supplementary Figure S1). After 8,550 SNPs in weak LD were excluded, the other 45 SNPs showing moderate to strong LD (r2 ≥ 0.3) with rs2188380 remained. Next, we examined the LD between each pair of these 46 SNPs (Fig. 1) and searched for the SNPs that were tagging other SNPs with strong LD (r2 ≥ 0.8). Finally, in addition to rs2188380, we selected 8 SNPs (rs7978484, rs16940688, rs2071629, rs11065783, rs3809297, rs4766566, rs671 and rs2555004) for association analysis (Supplementary Table S1).
Genetic analysis
Genomic DNA was extracted from whole peripheral blood cells21. Genotyping of the 8 SNPs was performed by the TaqMan method (Life Technologies Corporation, Carlsbad, CA USA) with a LightCycler 480 (Roche Diagnostics, Mannheim, Germany)22. To confirm their genotypes, DNA sequencing analyses were performed with the primers shown in Supplementary Table S4. Direct sequencing was performed with a 3130xl Genetic Analyzer (Life Technologies Corporation)22. The deviation from Hardy-Weinberg equilibrium in control samples was evaluated by chi-square test using the software R (version 3.1.1) (http://www.r-project.org/).
Statistical analyses
The associations between SNPs and gout were examined with logistic regression analyses. For the robustness of the statistical test, random re-sampling methods with computer simulation are often applied23,24. In this study, the permutation test24 was used for random re-sampling in a case-control study with replacement for 1,000,000 times and the robustness of statistics was confirmed. All the logistic regression analyses and chi-square tests were performed with SPSS v.22.0J (IBM Japan Inc., Tokyo, Japan) and the software R (version 3.1.1) (http://www.r-project.org/). We examined the pairwise LD using PLINK v1.0725.
Additional Information
How to cite this article: Sakiyama, M. et al. Identification of rs671, a common variant of ALDH2, as a gout susceptibility locus. Sci. Rep. 6, 25360; doi: 10.1038/srep25360 (2016).
References
Woodward, O. M. et al. Identification of a urate transporter, ABCG2, with a common functional polymorphism causing gout. Proc. Natl. Acad. Sci. USA 106, 10338–10342 (2009).
Matsuo, H. et al. Common defects of ABCG2, a high-capacity urate exporter, cause gout: a function-based genetic analysis in a Japanese population. Sci. Transl. Med. 1, 5ra11 (2009).
Matsuo, H. et al. Common dysfunctional variants in ABCG2 are a major cause of early-onset gout. Sci. Rep. 3, 2014 (2013).
Ichida, K. et al. Decreased extra-renal urate excretion is a common cause of hyperuricemia. Nat. Commun. 3, 764 (2012).
Matsuo, H. et al. ABCG2 dysfunction causes hyperuricemia due to both renal urate underexcretion and renal urate overload. Sci. Rep. 4, 3755 (2014).
Sulem, P. et al. Identification of low-frequency variants associated with gout and serum uric acid levels. Nat. Genet. 43, 1127–1130 (2011).
Köttgen, A. et al. Genome-wide association analyses identify 18 new loci associated with serum urate concentrations. Nat. Genet. 45, 145–154 (2013).
Matsuo, H. et al. Genome-wide association study of clinically defined gout identifies multiple risk loci and its association with clinical subtypes. Ann. Rheum. Dis. 75, 652–659 (2016).
Kim, Y. J. et al. Large-scale genome-wide association studies in East Asians identify new genetic loci influencing metabolic traits. Nat. Genet. 43, 990–995 (2011).
Iulianella, A., Sharma, M., Durnin, M., Vanden Heuvel, G. B. & Trainor, P. A. Cux2 (Cutl2) integrates neural progenitor development with cell-cycle progression during spinal cord neurogenesis. Development 135, 729–741 (2008).
Franco, S. J. et al. Fate-restricted neural progenitors in the mammalian cerebral cortex. Science 337, 746–749 (2012).
Huang, J., Ellinghaus, D., Franke, A., Howie, B. & Li, Y. 1000 Genomes-based imputation identifies novel and refined associations for the Wellcome Trust Case Control Consortium phase 1 Data. Eur. J. Hum. Genet. 20, 801–805 (2012).
Matsuo, K. et al. Alcohol dehydrogenase 2 His47Arg polymorphism influences drinking habit independently of aldehyde dehydrogenase 2 Glu487Lys polymorphism: analysis of 2,299 Japanese subjects. Cancer Epidemiol. Biomarkers Prev. 15, 1009–1013 (2006).
Yamanaka, H. et al. Analysis of the genotypes for aldehyde dehydrogenase 2 in Japanese patients with primary gout. Adv. Exp. Med. Biol. 370, 53–56 (1994).
Takeuchi, F. et al. Confirmation of ALDH2 as a major locus of drinking behavior and of its variants regulating multiple metabolic phenotypes in a Japanese population. Circ. J. 75, 911–918 (2011).
Goedde, H. W. et al. Distribution of ADH2 and ALDH2 genotypes in different populations. Hum. Genet. 88, 344–346 (1992).
Wallace, S. L. et al. Preliminary criteria for the classification of the acute arthritis of primary gout. Arthritis Rheum. 20, 895–900 (1977).
Hamajima, N. & J-MICC Study Group. The Japan Multi-Institutional Collaborative Cohort Study (J-MICC Study) to detect gene-environment interactions for cancer. Asian Pac. J. Cancer Prev. 8, 317–323 (2007).
Asai, Y. et al. Baseline data of Shizuoka area in the Japan Multi-Institutional Collaborative Cohort Study (J-MICC Study). Nagoya J. Med. Sci. 71, 137–144 (2009).
International HapMap, C. The International HapMap Project. Nature 426, 789–796 (2003).
Matsuo, H. et al. Mutations in glucose transporter 9 gene SLC2A9 cause renal hypouricemia. Am. J. Hum. Genet. 83, 744–751 (2008).
Sakiyama, M. et al. Common variant of leucine-rich repeat-containing 16A (LRRC16A) gene is associated with gout susceptibility. Hum. Cell 27, 1–4 (2014).
Li, J. et al. Identification of high-quality cancer prognostic markers and metastasis network modules. Nat. Commun. 1, 34 (2010).
Efron, B. & Tibshirani, R. J. In An Introduction to the Bootstrap (eds Cox, D. R. et al.) 202–219 (Chapman & Hall, 1993).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Acknowledgements
We would like to thank all the participants for their generous involvement in this study. Our sincere gratitude also to the members of the Japan Multi-Institutional Collaborative Cohort Study (J-MICC Study) Shizuoka Field for supporting the study. We are indebted to J. Abe, K. Gotanda, S. Shimizu, Y. Takada, T. Chiba, T. Higashino, M. Kawaguchi, Y. Kawamura, H. Ogata, A. Akashi, Y. Tanahashi, H. Nakashima and Y. Sakurai, National Defense Medical College, for genetic analysis and helpful discussion and to A. Tokumasu, Ryougoku East Gate Clinic and to M. Naito, K. Wakai and N. Hamajima, Nagoya University, for sample collection. We especially thank the following for enlightening discussion: T. Hosoya, Jikei University School of Medicine and K. Ichida, Tokyo University of Pharmacy and Life Sciences. This study was supported by grants from the Ministry of Education, Culture, Sports, Science and Technology (MEXT) of Japan including the MEXT KAKENHI (Grant numbers 221S0002, 221S0001, 25293145, 22689021, 25670307), the Ministry of Health, Labour and Welfare of Japan, the Ministry of Defense of Japan, the Japan Society for the Promotion of Science, the Kawano Masanori Memorial Foundation for Promotion of Pediatrics and the Gout Research Foundation of Japan.
Author information
Authors and Affiliations
Contributions
M.S., H.M., H.N. and A.N. conceived and designed this study. S.K., R.O., H.O. and T.S. collected samples and analyzed clinical data. M.S., H.M., K.Y. and A.N. performed genetic analysis. M.S., H.N. and T.N. performed statistical analyses. K.Y. and N.S. provided intellectual input and assisted with the preparation of the manuscript. M.S., H.M. and H.N. wrote the manuscript. M.S. and H.M. contributed equally to this work.
Ethics declarations
Competing interests
Yes, there is potential competing interest: H.M. and N.S. have a patent pending based on the work reported in this paper.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Sakiyama, M., Matsuo, H., Nakaoka, H. et al. Identification of rs671, a common variant of ALDH2, as a gout susceptibility locus. Sci Rep 6, 25360 (2016). https://doi.org/10.1038/srep25360
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep25360
This article is cited by
-
Risk of gout among Taiwanese adults with ALDH-2 rs671 polymorphism according to BMI and alcohol intake
Arthritis Research & Therapy (2021)
-
Identification of the 12q24 locus associated with fish intake frequency by genome-wide meta-analysis in Japanese populations
Genes & Nutrition (2019)
-
An update on the genetics of hyperuricaemia and gout
Nature Reviews Rheumatology (2018)
-
Independent effects of ADH1B and ALDH2 common dysfunctional variants on gout risk
Scientific Reports (2017)
-
A missense single nucleotide polymorphism in the ALDH2 gene, rs671, is associated with hip fracture
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.