Abstract
The grease matrix was originally introduced as a microcrystal-carrier for serial femtosecond crystallography and has been expanded to applications for various types of proteins, including membrane proteins. However, the grease-based matrix has limited application for oil-sensitive proteins. Here we introduce a grease-free, water-based hyaluronic acid matrix. Applications for proteinase K and lysozyme proteins were able to produce electron density maps at 2.3-Å resolution.
Similar content being viewed by others
Introduction
Using femtosecond X-ray pulses, serial femtosecond crystallography (SFX) offers a route to overcome radiation damage to small protein crystals via the “diffraction-before-destruction” approach1,2,3,4,5,6. This has expanded the window to obtain room temperature structures of proteins7,8,9,10,11,12,13. Liquid jet injection of small protein crystals is often exploited for the serial sample loading14. Continuous flow at relative high speed (~10 m/sec) of the liquid-jet injectors consumes 10~100 mg of the sample (5~6 hours/a full data set), which may not be ideal for X-ray free-electron lasers (XFELs) with low repetition rates (30~120 Hz at current facilities). Micro-extrusion of specimens using viscous media such as the lipidic cubic phase (LCP)12, grease13, and Vaseline (petroleum jelly)15 can maintain a stable stream at a low flow rate of 0.02~0.5 μl/min, which helps to reduce sample consumption. On the other hand, the viscous media tends to produce stronger X-ray scatterings that increase noise levels. Efforts to discover a sample delivery medium with lower background scattering has been ongoing, and recent demonstrations using agarose media are notable in that it has lower background scattering compared to the LCP media16. However, in general, viscous media tends to cause cracking and dissolution of protein crystals due to various physical or chemical events such as osmotic shock arising from the properties of viscous media.
Here we introduce a crystal carrier of water-based hyaluronic acid which is a good alternative to the grease matrix, to expand its application to grease-sensitive proteins. A highly viscous hydrophilic polymer was chosen from commercially available polymers. The practicality of the hyaluronic acid matrix as a protein carrier for SFX was demonstrated using two different types of proteins. The relatively low background scattering was notable in comparison to the grease matrix.
Results & Discussion
Data collection and structure determination
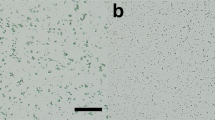
We carried out SFX experiments using femtosecond X-ray pulses from the SPring-8 Angstrom Compact Free Electron Laser (SACLA)6. The X-ray wavelength was kept at 1.77 Å (7 keV). At first, the background scatterings from three crystal matrix carriers, a mineral oil-based AZ grease13, a synthetic grease Super Lube17,18 and a hyaluronic acid aqueous solution, were compared. Grease generated diffuse scatterings in the resolution range of 4–5 Å (Fig. 1a,b). AZ grease gave a diffraction ring at ~14-Å resolution. For the Super Lube grease, we observed a diffraction ring at ~4.8-Å resolution in about 30% of all diffraction images (Supplementary Fig. 1). Weaker background scattering was noted when using hyaluronic acid compared with those of greases (Fig. 1c,d). In this study, using two carriers, synthetic grease Super Lube and hyaluronic acid matrices, we investigated proteinase K (5–10 μm) and lysozyme (7–10 μm) crystals (Supplementary Fig. 2) to demonstrate their general applicability as crystal carriers. A flow rate of 0.48 μl/min was used for all samples. The grease and hyaluronic acid matrices formed a stable flow for all protein samples (Supplementary Fig. 3).
(a) Mineral oil-based AZ grease, (b) Super Lube synthetic grease, and (c) hyaluronic acid. Resolution at the edges corresponds to ~2.3 Å (dashed circle). (d) The average background scattering intensities of ~2,000 images from each matrix. AZ grease, Super Lube synthetic grease and hyaluronic acid are depicted by the black, blue and green lines, respectively.
With the SACLA running at a 30 Hz repetition rate, we were able to collect ~100,000 diffraction patterns in approximately 1 hour. In total about 30 μl of the sample volume was used with the crystal number density of 6.7 × 107 crystals/ml (Table 1). We successfully indexed and integrated 21,000–27,000 patterns for each of the proteinase K and lysozyme crystals (space group P43212). The crystals yielded data sets at 2.3-Å resolution with a completeness of 100% and an Rsplit19 ranging from 8.5% to 9.7%. We determined and refined the crystal structures of proteinase K (Protein Data Bank (PDB) ID: 5B1D for grease and 5B1E for hyaluronic acid) and lysozyme (5B1F for grease and 5B1G for hyaluronic acid) at 2.3-Å resolution. Clear electron density maps of proteinase K and lysozyme were able to be observed (examples are shown here for proteinase K, Fig. 2).
Close-up views of the proteinase K structures for (a) Super Lube synthetic grease (PDB: 5B1D) and (b) hyaluronic acid (PDB: 5B1E) with 2Fo − Fc electron density maps contoured at the 1.0σ level (coloured gray) and the anomalous difference Fourier maps contoured at the 3.5σ level (coloured magenta). Bound calcium ion is depicted as a green sphere. These figures were drawn with PyMol (http://www.pymol.org).
Sample preparation
Using the two matrix carriers, Super Lube grease and hyaluronic acid, we successfully collected the data sets for proteinase K and lysozyme at 2.3-Å resolution (Table 1). A 12% (w/v) hyaluronic acid matrix prepared by mixing 24% (w/v) hyaluronic acid aqueous solution with an equal volume of the supernatant solution in the crystal suspension solution was used for the lysozyme crystals. Optimizing the hyaluronic acid solution buffer is important to prevent any damage to the crystals. We have observed that before adding protein crystals it is essential to mix the hyaluronic acid aqueous solution with the supernatant solution or the crystal harvest solution, which helps to avoid osmotic shock to the crystals when mixing with the medium. The hyaluronic acid solution was saturated with the supernatant solution or the crystal harvest solution, and then protein crystals were added, which helps to avoid potential osmotic shock to the crystals. The unit-cell axes of the lysozyme crystals for the grease matrix were slightly shorter than those for the hyaluronic acid matrix (Table 1). Dehydration of protein crystals might have been induced during the sample preparation process of the water-free grease matrix. In such cases, a water-based medium can be helpful for preventing the contraction of the unit cell in the SFX experiments.
In comparison with LCP12, grease13 and Vaseline (petroleum jelly)15, water-based media such as hyaluronic acid and agarose16 produce lower background scattering noise; however, the agarose medium requires heat treatment at temperatures higher than 80 °C. The sample preparation in our technique can be performed by simply mixing with hyaluronic acid medium. In SFX, the grease matrix may not always be useful, because some proteins are damaged while being mixed and soaked in them. In the matrix technique using viscous media, the first step is to find a carrier for the protein crystals of interest that is suitable for data collection at room temperature. For SFX experiments, it is important to provide a wide repertoire of carrier media for a wide variety of proteins. Currently we are studying other crystal matrix carriers with low background scattering. For example, hydroxyethyl cellulose medium appears to be a good candidate.
Background scattering & column diameter
The Super Lube grease tended to give a stronger background scattering in the resolution range of 4–5 Å than the hyaluronic acid (Fig. 1b,d). However, there was no noticeable difference in the data collection statistics or the electron density maps between the two carriers (Table 1 and Fig. 2). Statistics of I/σ(I) for proteinase K showed higher intensity values for the hyaluronic acid matrix at resolutions ranging from ~8 to ~3 Å in comparison to the grease matrix (Supplementary Fig. 4). However, it is reversed on the border of around 3 Å resolution, because water-based matrix gives a slightly higher background scattering in the resolution range of ~3.5–2.5 Å compared to the grease matrix (Fig. 1). To date, we have performed SFX experiments with the grease matrix using more than ten soluble proteins and three membrane proteins. However, we observed dissolution of crystals for one soluble and one membrane protein samples in the matrix. We do not yet have the data sets from these samples, but we confirmed that water-based matrix is useful for all these oil-sensitive crystals in SFX. These results suggest that the Super Lube grease has potential as a versatile matrix carrier, but the hyaluronic acid matrix would enable SFX experiments for grease-sensitive protein crystals.
Untreated Super Lube grease extruded through a 110-μm-i.d. needle tended to produce a larger-diameter grease column (approximately ~210 μm) about the size of the outer diameter (o.d.) of the needle, and similar to the mineral oil-based AZ grease13. By grinding the Super Lube grease for 30–60 min, the grease produced a sample column diameter of ~110 μm (Supplementary Fig. 2a). On the other hand, the hyaluronic acid matrix was extruded as a continuous column with a diameter of 110–130 μm through a 110-μm-i.d. needle (Supplementary Fig. 2b). A sample column with a smaller diameter contributes to reduce sample consumption and background noise from the matrices.
Recently, using the grease matrix technique, Yamashita and coworkers have demonstrated a single isomorphous replacement with anomalous scattering (SIRAS) phasing for Hg-derivatized luciferin-regenerating enzyme17. In addition, we have successfully determined the structure of native lysozyme with single-wavelength anomalous diffraction (SAD) by utilizing the anomalous signal of sulfur and chlorine18. One of the major challenges for phasing in SFX is to improve the signal-to-noise ratio. In this study, we could observe a weak anomalous scattering signal from the calcium atom in the proteinase K structures (Fig. 2). The anomalous difference Fourier maps showed that the signal from the calcium atom is stronger with the hyaluronic acid matrix than with the grease matrix. Furthermore, in the crystal structure for the hyaluronic acid matrix, we could observe anomalous signal from sulfur atoms (e.g. the sulfur atom of Cys178, Fig. 2b), which was not discernible when using the grease matrix. This technique using the matrices with low background scattering noise will contribute significantly to measuring weak anomalous signals for de novo phasing from SFX data.
In summary, using the hyaluronic acid matrix as a general carrier of protein microcrystals for serial sample loading in SFX, we successfully obtained the room-temperature structures at 2.3-Å resolution of two proteins in 5–10 μm microcrystals using less than 1 mg of sample. Oil- and water-based crystal carriers are complementary and their application to a wide variety of proteins is essential to firmly establish SFX. Recently, using viscous carrier media, synchrotron-based serial crystallography data collection at room temperature has also been demonstrated15,20. In the immediate future, the sample loading technique with a viscous medium which helps to reduce sample consumption will become more important in serial millisecond crystallography using synchrotron radiation. SFX has provided new opportunities for time-resolved studies of light-driven structural changes and chemical dynamics21,22,23. Matrix carriers with a stable sample flow and small diameter sample column should be applicable for time-resolved studies using pump-probe techniques.
Methods
Sample preparation
Proteinase K from Engyodontium album (No. P2308, Sigma) was crystalized by mixing a 1:1 ratio of 40 mg/ml protein solution in 20 mM MES–NaOH (pH 6.5) and a precipitant solution composed of 0.5 M NaNO3, 0.1 M CaCl2, 0.1 M MES–NaOH (pH 6.5). Microcrystals were produced by incubation for 5–10 min at 18 °C. A 1.0-ml sample of crystallization solution was centrifuged at 20 °C and 3,000 g for 3 min, and then the supernatant solution was removed. The crystals of proteinase K were suspended in 1.0 ml of the crystallization reagent. The crystal suspensions were filtered through a mesh (pore size, 30 μm). Lysozyme crystals were prepared as described previously13. Proteinase K and lysozyme samples were adjusted to a number density of 6.7 × 107crystals/ml. The samples were then stored at 18 °C for proteinase K and 4 °C for lysozyme.
Synthetic grease Super Lube (No. 21030, Synco Chemical Co.) was ground using a mortar for 30–60 min. The crystals were mixed with the ground grease using the same procedure reported by Sugahara et al.13. For hyaluronic acid (No. H5388, Sigma), protein microcrystals were prepared according to the following procedures. After a 100-μl sample of storage solution was centrifuged for 10 sec, a 90-μl aliquot of supernatant solution was removed. For proteinase K crystals, an 8.0-μl aliquot of the crystal solution was dispensed into 72 μl of 12% (w/v) hyaluronic acid aqueous solution on a glass slide and then mixed with a spatula. For lysozyme crystals, a hyaluronic acid matrix was prepared by mixing a 1:1 ratio of 24% (w/v) hyaluronic acid aqueous solution and the supernatant solution of crystallization solution. An 8.0-μl aliquot of the crystal solution was dispensed into 72 μl of the hyaluronic acid solution. An aliquot (30 μl) of the sample was extruded into a 100-μl syringe (No. 1710, Hamilton).
Data collection
We carried out the experiments using femtosecond X-ray pulses from the SPring-8 Angstrom Compact Free Electron Laser (SACLA)6. The X-ray wavelength was kept at 1.77 Å (7 keV) with a pulse energy of ~ 200 μJ. Each X-ray pulse delivered ~ 7 × 1010 photons within a 10-fs duration (FWHM) to the samples with a matrix. Data were collected using focused X-ray beams of 1.5 × 1.5 μm2 by Kirkpatrick-Baez mirrors24. The crystals in the matrix were serially loaded using a syringe injector installed in a helium ambiance, diffraction chamber. The experiments were carried out using a Diverse Application Platform for Hard X-ray Diffraction in SACLA (DAPHNIS)25 at BL326. The microcrystals embedded in grease or hyaluronic acid matrix were kept at a temperature of approximately 20 °C. The sample chamber was kept at a temperature of ~ 26 °C and humidity greater than 80%. Each matrix with randomly oriented crystals was extruded through a syringe needle with an inner diameter (i.d.) of 110 μm (outer diameter (o.d.), 210 μm; No. 7803-05, Hamilton). The sample flow rate was 0.48 μl/min. Diffraction patterns were collected using a custom-built multiport CCD27.
Background intensity determination
The background intensity was determined by a procedure similar to that used in16. First, the average (m) and standard deviation (s) of each detector pixel over images were calculated. To remove intensity contributions due to Bragg spots from protein crystals, pixels brighter than m + 3s were rejected. Remaining pixels were averaged again to yield a “clean” background image. This image was radially averaged by the resolution calculated from the detector metrology. When plotting Fig. 1d, datasets were scaled so that values at the highest resolution shell became the same.
Structure determination
Diffraction patterns were processed by Cheetah28 adapted for the SACLA data acquisition system29. Each pattern with more than 20 spots was accepted as a hit, and indexed and integrated using CrystFEL19. Diffraction peak positions were determined using the built-in Zaefferer algorithm and passed on to DirAx30 for indexing. Monte Carlo integrated intensities from CrystFEL were converted to MTZ format. The structures were determined by the molecular replacement method using Molrep31 with search models (PDB: 5AVJ for proteinase K and 3WUL for lysozyme). Manual model revision was performed using Coot32. The program Phenix33 was used for structure refinement. Details of the data collection and refinement statistics are summarized in Table 1.
Additional Information
Accession codes: The coordinates and structure factors have been deposited in the Protein Data Bank under the accession code 5B1D and 5B1E for proteinase K and 5B1F and 5B1G for lysozyme. Diffraction images have been deposited to CXIDB under ID #33 (lysozyme in grease) and #41 (others).
How to cite this article: Sugahara, M. et al. Oil-free hyaluronic acid matrix for serial femtosecond crystallography. Sci. Rep. 6, 24484; doi: 10.1038/srep24484 (2016).
References
Chapman, H. N. et al. Femtosecond X-ray protein nanocrystallography. Nature 470, 73–77 (2011).
Neutze, R., Wouts, R., van der Spoel, D., Weckert, E. & Hajdu, J. Potential for biomolecular imaging with femtosecond X-ray pulses. Nature 406, 752–757 (2000).
Emma, P. et al. First lasing and operation of an ångstrom-wavelength free-electron laser. Nat. Photonics 4, 641–647 (2010).
Schlichting, I. & Miao, J. Emerging opportunities in structural biology with X-ray free-electron lasers. J. Curr. Opin. Struct. Biol. 22, 613–626 (2012).
Barty, A. et al. Self-terminating diffraction gates femtosecond X-ray nanocrystallography measurements. Nat. Photonics 6, 35–40 (2012).
Ishikawa, T. et al. A compact X-ray free-electron laser emitting in the sub-ångström region. Nat. Photonics 6, 540–544 (2012).
Boutet, S. et al. High-resolution protein structure determination by serial femtosecond crystallography. Science 337, 362–364 (2012).
Barends, T. R. M. et al. De novo protein crystal structure determination from X-ray free-electron laser data. Nature 505, 244–247 (2014).
Johansson, L. C. et al. Lipidic phase membrane protein serial femtosecond crystallography. Nat. Methods 9, 263–265 (2012).
Redecke, L. et al. Natively inhibited Trypanosoma brucei cathepsin B structure determined by using an X-ray laser. Science 339, 227–230 (2013).
Liu, W. et al. Serial femtosecond crystallography of G protein–coupled receptors. Science 342, 1521–1524 (2013).
Weierstall, U. et al. Lipidic cubic phase injector facilitates membrane protein serial femtosecond crystallography. Nat. Commun. 5, 3309 (2014).
Sugahara, M. et al. Grease matrix as a versatile carrier of proteins for serial crystallography. Nat. Methods 12, 61–63 (2015).
DePonte, D. P. et al. Gas dynamic virtual nozzle for generation of microscopic droplet streams. J. Phys. D Appl. Phys. 41, 195505 (2008).
Botha, S. et al. Room-temperature serial crystallography at synchrotron X-ray sources using slowly flowing free-standing high-viscosity microstreams. Acta Crystallogr. D 71, 387–397 (2015).
Conrad, C. E. et al. A novel inert crystal delivery medium for serial femtosecond crystallography. IUCrJ 2, 421–430 (2015).
Yamashita, K. et al. An isomorphous replacement method for efficient de novo phasing for serial femtosecond crystallography. Sci. Rep. 5, 14017 (2015).
Nakane, T. et al. Native sulfur/chlorine SAD phasing for serial femtosecond crystallography. Acta Crystallogr. D 71, 2519– 2525 (2015).
White, T. A. et al. CrystFEL: a software suite for snapshot serial crystallography. J. Appl. Crystallogr. 45, 335–341 (2012).
Nogly, P. et al. Lipidic cubic phase serial millisecond crystallography using synchrotron radiation. IUCrJ 2, 168–176 (2015).
Kern, J. et al. Taking snapshots of photosynthetic water oxidation using femtosecond X-ray diffraction and spectroscopy. Nat. Commun. 5, 4371 (2014).
Kupitz, C. et al. Serial time-resolved crystallography of photosystem II using a femtosecond X-ray laser. Nature 513, 261–265 (2014).
Tenboer, J. et al. Time-resolved serial crystallography captures high-resolution intermediates of photoactive yellow protein. Science 346, 1242–1246 (2014).
Yumoto, H. et al. Focusing of X-ray free-electron laser pulses with reflective optics. Nat. Photonics 7, 43–47 (2013).
Tono, K. et al. Diverse application platform for hard X-ray diffraction in SACLA (DAPHNIS): application to serial protein crystallography using an X-ray free-electron laser. J. Synchrotron Rad. 22, 532–537 (2015).
Tono, K. et al. Beamline, experimental stations and photon beam diagnostics for the hard x-ray free electron laser of SACLA. New J. Phys. 15, 083035 (2013).
Kameshima, T. et al. Development of an X-ray pixel detector with multi-port charge-coupled device for X-ray free-electron laser experiments. Rev. Sci. Instrum. 85, 033110 (2014).
Barty, A. et al. Cheetah: software for high-throughput reduction and analysis of serial femtosecond X-ray diffraction data. J. Appl. Crystallogr. 47, 1118–1131 (2014).
Joti, Y., Kameshima, T., Yamaga, M., Sugimoto, T. & Okada, K. Data acquisition system for X-ray free-electron laser experiments at SACLA. J. Synchrotron Rad. 22. 571–576 (2015).
Duisenberg, A. J. M. Indexing in single-crystal diffractometry with an obstinate list of reflections. J. Appl. Crystallogr. 25, 92–96 (1992).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221. (2010).
Acknowledgements
The XFEL experiments were carried out at the BL3 of SACLA with the approval of the Japan Synchrotron Radiation Research Institute (JASRI) (proposal nos 2014A8032, 2014B8050, 2015A8048 and 2015A8026). This work was supported by the X-ray Free-Electron Laser Priority Strategy Program (MEXT), partly by a Grant-in-Aid for Scientific Research from the Japan Society for the Promotion of Science (KAKENHI No. 25650026), partly by the Research Acceleration Program of the Japan Science and Technology Agency and partly by the Platform for Drug Discovery, Informatics, and Structural Life Science (MEXT). C.S. is supported by NRF through the SRC (NRF-2015R1A5A1009962). The authors thank Dr. Jun Kobayashi for preparation of the grease matrix and the SACLA beamline staff for technical assistance. We are grateful for the computational support from SACLA HPC system and Mini-K super computer system. We thank A. Nisbet for his editing assistance.
Author information
Authors and Affiliations
Contributions
M.Sugahara. introduced the oil-free hyaluronic acid matrix extrusion scheme, M.Suzuki, T.M., S.Inoue, F.Y., E.N., R.T., C.S. and M.Sugahara performed the data collection, M.Sugahara. and M.Suzuki performed data processing and solved the structure, M.Sugahara, E.N. and K.N. prepared the microcrystals, T.N. and O.N. developed the data processing pipeline and performed the background scattering data analysis, K.T., Y.J., T.K., T.H. and M.Y. developed the DAPHNIS and detectors, M.Sugahara, C.S., M.Suzuki, T.M., S.Inoue and T.N. wrote the manuscript with input from all of the coauthors and S.Iwata coordinated the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Sugahara, M., Song, C., Suzuki, M. et al. Oil-free hyaluronic acid matrix for serial femtosecond crystallography. Sci Rep 6, 24484 (2016). https://doi.org/10.1038/srep24484
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep24484
This article is cited by
-
Beef tallow injection matrix for serial crystallography
Scientific Reports (2022)
-
Viscosity-adjustable grease matrices for serial nanocrystallography
Scientific Reports (2020)
-
Shortening injection matrix for serial crystallography
Scientific Reports (2020)
-
Polyacrylamide injection matrix for serial femtosecond crystallography
Scientific Reports (2019)
-
Microfluidic sample delivery for serial crystallography using XFELs
Analytical and Bioanalytical Chemistry (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.