Abstract
Rhodopsins are membrane-embedded photoreceptors found in all major taxonomic kingdoms using retinal as their chromophore. They play well-known functions in different biological systems, but their roles in fungi remain unknown. The filamentous fungus Fusarium fujikuroi contains two putative rhodopsins, CarO and OpsA. The gene carO is light-regulated and the predicted polypeptide contains all conserved residues required for proton pumping. We aimed to elucidate the expression and cellular location of the fungal rhodopsin CarO, its presumed proton-pumping activity and the possible effect of such function on F. fujikuroi growth. In electrophysiology experiments we confirmed that CarO is a green-light driven proton pump. Visualization of fluorescent CarO-YFP expressed in F. fujikuroi under control of its native promoter revealed higher accumulation in spores (conidia) produced by light-exposed mycelia. Germination analyses of conidia from carO− mutant and carO+ control strains showed a faster development of light-exposed carO− germlings. In conclusion, CarO is an active proton pump, abundant in light-formed conidia, whose activity slows down early hyphal development under light. Interestingly, CarO-related rhodopsins are typically found in plant-associated fungi, where green light dominates the phyllosphere. Our data provide the first reliable clue on a possible biological role of a fungal rhodopsin.
Similar content being viewed by others
Introduction
Light strongly influences the behaviour of microorganisms in their natural habitats. Various photoreceptors, able to detect blue, green, or red wavelengths in the visible spectrum1,2 provide filamentous fungi the ability to sense light and adapt accordingly. Among them stand out blue-light photoreceptors, such as the proteins of the White collar, Vivid and cryptochrome families3,4 which absorb light by flavin chromophores. In contrast, phytochromes are red light-sensing proteins that use the chromophore biliverdin4. Analyses of mutants have revealed the participation of these photoreceptors in the modulation of a variety of physiological and morphological responses in fungi1,5,6.
A further class of fungal photoreceptors comprises the green light sensing microbial rhodopsins7,8,9,10. These proteins exhibit a characteristic structure, consisting of seven transmembrane helices forming an interior pocket for the chromophore all-trans-retinal, which is covalently bound via a protonated Schiff-base to a lysine residue in helix-G. In light-driven rhodopsin-pumps a single ion is translocated per photocycle, which is triggered by light-induced all-trans to 13-cis retinal isomerization. Some residues of the binding pocket are well conserved across kingdoms, whereas conservation outside the pocket is more sparse11. According to differences in the binding pocket residues three classes of fungal rhodopsins are distinguished12: NR-like rhodopsins (from Neurospora rhodopsin), LR-like rhodopsins (from Leptosphaeria rhodopsin) and auxiliary opsin-related protein (ORP)-like rhodopsins. Genes for putative rhodopsins are ubiquitous in the genomes of fungal species13 and they are frequently up-regulated by light14,15,16. In chytridiomycetes rhodopsin is involved in the phototaxis of swimming zoospores17 and recently a unique gene fusion was described in this fungal group, which combines the action of rhodopsin and guanylyl cyclase in a single protein18. Some fungal rhodopsins have been purified and photochemically analysed, but their physiological roles are unknown19,20,21. Moreover, the analyses of their functions through targeted deletion of the encoding genes provided no clear clues so far3,22.
The filamentous fungus Fusarium fujikuroi, the causing agent of the bakanae-disease in rice, is of economic importance as a biotechnological source of gibberellins, growth-promoting plant hormones23. This fungus is also a research model for the production of these and other metabolites24, including the carotenoids and its genome has been recently sequenced25. The Fusarium genomes contain genes encoding for two different rhodopsins, called in F. fujikuroi OpsA15 and CarO26. Based on sequence similarity OpsA is a NR-like rhodopsin, alike to Nop-1 of N. crassa. The photocycle of Nop-1 is very slow and proton-pump activity was not detected in electrophysiological measurements27 and similar features are presumed for OpsA. In contrast, CarO is among the auxiliary ORP-like rhodopsins, which together with LR-like rhodopsins exhibit fast photocycles and possess all conserved amino acid residues required for proton pumping. LR exhibits proton-pumping function, but the auxiliary ORP-like rhodopsins were not investigated in this respect4,19.
Fungal light-driven proton pumps may be involved in the generation of pH gradients or in pH homeostasis, but they also may have signalling functions, as acidification of some cell compartments could be used to activate certain biochemical responses3,20. In F. fujikuroi, carO is located in a gene cluster with the genes needed for retinal synthesis, but the carO− mutant did not show any visible phenotype26. The opsA deletion mutants did not exhibit growth or morphological alterations either, but they showed minor changes in the expression of carotenoid biosynthesis genes4 and a minor regulatory alteration of these genes was also exhibited by the mutant of the gene encoding the retinal forming enzyme CarX28.
Here we combined electrophysiology and fluorescence microscopy analyses to detect pump activity and cellular localization of the CarO protein in the fungus F. fujikuroi, which reproduces asexually through the formation of spores called conidia. We analysed the effect of light on conidia germination and growth of a carO− mutant and a carO+ control strain, referred hereafter as CarO+ and CarO−. We show that CarO is a green light-activated proton pump mainly distributed in the plasma membrane of conidia, slowing down spore germination and early hyphal development in this fungus.
Results
CarO is a light-dependent H+ pump
CarO was predicted to be a light-activated proton pump due to the occurrence of conserved residues, which are required for proton pumping in related microbial rhodopsins10. We modelled the amino acid sequence of CarO to the structure of bacteriorhodopsin (BR; Fig. 1). As other microbial rhodopsins, this protein consists of 7 transmembrane domains (helix-A-G). Most importantly CarO contains proton donor and proton acceptor counterparts of BR-D96 and BR-D85 (D128 and D117, respectively). Also the lysine residue (BR-K216) required for covalent binding of the retinal chromophore via a protonated Schiff base is present in helix-G of CarO (K246). The occurrence of Trp or Glu in helix-D in position 146 is characteristic for ORP-like rhodopsins, while other microbial rhodopsins typically exhibit apolar residues in this position19.
Schematic representation of CarO::YFP.
Topo2 transmembrane protein display software and NCBI protein-Blast was used to define transmembrane sections in CarO (GenBank: CAD97459.1) in analogy to BR (SWISS Model62, template 4hwl.3.B). Conserved residues are highlighted in orange and related residues in yellow. Asp117 and Asp128 in helix-C, highlighted in red, represent the putative proton acceptor and donor, respectively (D85 and D96 in BR). The lysine (Lys246) in helix-G is also conserved and allows for covalent binding of retinal via protonated Schiff base. In helix-D the Trp148 (red font) is typical for auxiliary ORP-like rhodopsins, which contain Glu or Trp in this positions instead of Gly in BR and other related rhodopsins. A linker peptide (gray letters) was used to connect YFP to the C-terminus of the CarO protein.
In order to test the capability of CarO to act as an ion pump, we established a Flip-In T-REx 293 cell line expressing a fluorescent version of CarO, C-terminally tagged with enhanced yellow fluorescent protein (CarO::YFP; Fig. 1). As expected, after 15–24 hours activation of the promoter by tetracycline, the cells exhibited membrane-localized fluorescence (Fig. 2a). Localization of CarO::YFP in the cytoplasmic membrane supported the subsequent use of this cell line to test CarO activity in electrophysiological experiments.
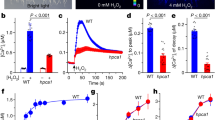
Electrophysiological analysis of CarO::YFP expressed in Flip-In T-Rex 293 cells.
(a) Wide field fluorescence image of Flip-In T-Rex 293 CarO::YFP cells used in patch-clamp experiments. White bar represents 10 μm. The fusion protein is mainly located in the plasma membrane but also in internal membranes. (b) Typical current traces of the ion pump CarO recorded in NaCl pH 7.4 at membrane potentials of +40 mV, 0 mV and −100 mV, respectively, as indicated. Cells were illuminated by a 561 nm diode pumped laser. The green bar represents illumination time. (c) I–V-relationship of the light-dependent normalized pump current of CarO::YFP measured at pH 7.4 corresponding to data shown in b. Note, that activity of CarO is voltage-dependent.
Patch-clamp measurements were performed in the whole-cell configuration in voltage clamp mode and cells were illuminated with green light (561 nm) emitted by a laser diode. Upon light activation CarO evoked an outward directed current, exhibiting the typical behaviour of ion-pumping rhodopsin (Fig. 2b10). After a fast rise (transient photocurrent) characterized by a small time constant (measured at 0 mV holding potential and pH 7.4; τon 0.3 ± 0.1 ms s.d., n = 8) the photocurrent decreased to a positive stationary value. Upon switching off light the pump current decayed in a biexponential manner exhibiting a fast (τoff_1 1.4 ± 0.26 ms s.d., n = 10) and a slower time constant (τoff_2 20 ± 3.8 ms s.d., n = 10). The photocurrent showed clear voltage-dependency (Fig. 2c), with larger pump activity at higher membrane potentials. The direction of charge transfer remained outward, when the electrochemical gradient was contrary to the charge-transfer direction (Fig. 2c). Accordingly, the values for the reversal potentials estimated by nonlinear fit of the I–V-plot (Supplementary Fig. S1) were much more negative than those calculated by Nernst equation at intracellular pH 7.4 and extracellular pH 5 (−150 mV vs. 139 mV), pH 7.4 (−222 mV vs. 0 mV) and pH 9 (−243 mV vs. −9.3 mV). In conclusion, CarO exhibits the behaviour expected for an active ion pump.
CarO pump activity depends on light intensity and wavelength
The activity of photoreceptors depends on the light-wavelength and according to former studies maximal efficiency of fungal rhodopsins is expected with green light. We recorded the pump currents at different wavelengths and plotted the data resulting in a bell-shaped action spectrum (Fig. 3a). The pumping activity of CarO::YFP was highly wavelength dependent with highest activity at around 520 nm (green light).
Light dependency of CarO::YFP.
(a) Action spectrum. Effect of light-wavelength on CarO::YFP activity (mean value and standard deviation of 7 cells). The light emitted by a 150W-XBO lamp was filtered through narrow-band filters. At every wavelength the same dose of photons per time and area (2 × 1016 photons s−1 mm−2) was used and values were corrected for eventual intensity loss (reference wavelength 560 nm). The action spectrum was essentially the same at different intracellular proton concentrations (pH 5, 7.4 and 9, Supplementary Fig. S2); therefore, for better signal to noise-ratio, pH of internal pipette solution was set to 5. (b) Dependency of the pump current amplitude on photon density. Mean normalized photocurrent and corresponding standard deviation (5 cells, 3 recordings each). Data were fitted by a Hill function (red curve).
Beside the light wavelength the pump activity also depended on the light intensity. We measured the pump current at 0 mV voltage potential (1 ms illumination time) while varying the photon-density emitted by a 561 nm-diode pumped solid state (DPSS)-laser (Fig. 3b). The pump exhibited typical saturation behaviour, which could be fitted with a standard Hill equation. Half maximal activity was reached at a photon density of 1.6 × 1017 photons s−1 mm−2. For patch-clamp experiments photon density was adjusted to 2.3 × 1017 photons s−1 mm−2, providing >60% pump-activity.
Sequence features of CarO suggested a pump function for protons (Fig. 14,26). If CarO was an outward directed proton pump, we would expect an increase or decrease in pump activity when the amount of intracellular protons is heightened or diminished, respectively. Therefore, we varied the pH of bath (Fig. 4a) and pipette (Fig. 4b) solution and investigated the influence of the proton gradient on the pump intensity. Indeed, when the pH of the bath-solution was reduced from 9 to 5 the mean photocurrent of CarO decreased. This circumstance was especially pronounced when the extracellular pH was set to pH 7.4 and the intracellular pH was increased from 5 to 9, which resulted in a remarkable loss in pump activity (Fig. 4b).
Pump-activity of CarO::YFP at various pHs and other ion concentrations.
Cells were illuminated by a 561 nm-laser. (a) Influence of external pH on pump-activity. The relative pump activity is shown, which was normalized to the pump current at pH 7.4 and 0 mV. pH of bath solution varied from pH 5 (black squares, 10 cells), to pH 7.4 (red circles, 25 cells) and pH 9 (blue triangles, 13 cells); intracellular pH was kept at 7.4. (b) Influence of internal pH on pump activity. Extracellular pH was maintained at pH 7.4. Mean area specific photocurrent at intracellular pH 9 (3 cells), 7.4 (8 cells) and 5 (10 cells), respectively. (c) Analysis of pump activity in absence of extracellular chloride or intracellular sodium. Mean area-specific pump current to membrane voltage relationship of CarO is given. Sodium chloride in standard solutions (red circles, 14 cells) was replaced either intracellularly by potassium chloride (black squares, 5 cells) or extracellularly by sodium sulfate (blue triangles, 7 cells), without any remarkable effect on pump activity.
Some microbial rhodopsins were reported to pump chloride29 or even sodium30 ions instead of protons. To test these possible activities of CarO, we changed the ion composition of either bath or pipette solution and measured the corresponding pump current. When sodium chloride was replaced extracellularly by sodium sulphate or intracellularly by potassium chloride, the pump activity was not affected (Fig. 4c), providing further support to CarO as a proton pump.
Gluconate enhances CarO pump activity
Gluconate is one of the standard compounds used in electrophysiological experiments, when chloride ions have to be substituted31. Unexpectedly, CarO pump activity was strongly influenced by gluconate (Fig. 5a). Indeed, the activity was enhanced about 3-fold in sodium gluconate at extracellular pH 5 compared to sodium chloride at neutral pH (7.4). When the pH was suddenly raised from 5 to 9 a further transient increase (up to ten fold compared to neutral sodium chloride) in pump activity could be observed, which relaxed within 10–15 min. After relaxation the I–V-curve for gluconate pH 9 was similar as for sodium chloride pH 7.4. In contrast, this transient pH-dependent increase in pump activity was not observed when sodium chloride in the pipette solution was exchanged for sodium gluconate, while at pH 5 and pH 7.4 the pump activity was about the same as with sodium chloride (Supplementary Fig. S4).
Influence of weak organic acids on the pump-activity of CarO::YFP.
I–V blot of the normalized light-driven pump current (linear fit of mean values and standard deviation of measurements recorded from at least 5 different cells) at pH 5 (black squares), pH 7.4 (red circles) and pH 9 (blue triangles), respectively, in extracellular presence of the weak organic acids gluconate (a), glutamate (b) and galacturonate (c). Note that in the presence of gluconate and glutamate at pH 5 the pump activity is enhanced, though the contra-directed gradient is increased. In case of gluconate at pH 9 a transient enhanced pump current (half triangles, pH 9*) is observed, which disappears in a time scale of over ten minutes (closed triangles).
In addition to gluconate we also tested the activity of CarO in the presence of other weak organic acids. With glutamate (Fig. 5b) the pump activity was noticeably increased at pH 5 compared to pH 7.4. However, no further increase was observed at pH 9; actually, the activity at pH 9 was even reduced compared to pH 7.4. In contrast, the behaviour of CarO in galacturonic acid (Fig. 5c) did not significantly differ from its behaviour in sodium chloride.
CarO is localized to membranes and accumulates in conidia
To gain more insights on its possible biological function, we aimed to find out in which membranes of the F. fujikuroi conidia and hyphae CarO is mainly localized. For this purpose we generated a F. fujikuroi strain expressing the fluorescent carO::yfp construct under control of its native light-regulated promoter. Successful transformation of the plasmid by protoplast electroporation and integration of the fusion gene into the F. fujikuroi genome was demonstrated by Southern blot analyses: a single band specific for carO was observed in wild type genomic DNA, while an additional band was present in the genomic DNA of the carO::yfp transformant (Fig. 6a). A hybridizing band was also found with an yfp-probe in the transformant, absent in the wild type (data not shown). Taken together, these results indicate that the carO::yfp construct was incorporated into the F. fujikuroi genome, but it did not replace the native carO gene.
Localization of CarO::YFP in F. fujikuroi.
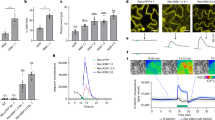
(a) Southern blot analysis of genomic DNA of F. fujikuroi FKMC1995 transformed with pHJA2-carO::YFP and selected for hygromycin resistance. DNA samples were digested with NcoI or BclI. The carO::yfp gene was introduced into the fungal genome and the native carO gene was not replaced. Complete blot is shown in the Supplementary Fig. S13. (b) Localization of CarO::YFP in F. fujikuroi conidia. The CarO::YFP fusion protein is mainly located in the plasma membrane but also in the membranes of inner organelles (vacuole, nucleus, ER). i–iii. Photomicrographs of conidia obtained by transmitted light microscopy (upper row) and corresponding confocal laser scanning microscopy (cLSM; lower row). The expression of carO::yfp in the conidia is induced by light. i) Light exposed (left: bicellular, right, monocellular), ii) dark exposed, iii) control (light exposed wild type). White bars represent 5 μm (i) and 2 μm (ii, iii), respectively. (c) Confocal laser scanning microphotograph of a yeast cell (S. cerevisiae DSY5) expressing CarO::YFP. The fusion protein is mainly located in the plasma membrane but in addition also in internal membranes (most likely ER). White bar represents 1 μm. (d) Localization of CarO::YFP in light-exposed F. fujikuroi hyphae 20 h after germination. The CarO::YFP fusion protein is heterogeneously distributed in the mycelia according to a dynamic process (Supplementary Fig. S6 and Movie 1). White bar represents 5 μm.
Fluorescence analysis of F. fujikuroi carO::yfp conidia and hyphae by confocal laser scanning microscopy confirmed the expression of CarO::YFP from the carO promoter, which is upregulated by light26. Accordingly, conidia produced by light-grown mycelia (abbreviated hereafter as LG conidia) exhibited strong YFP-mediated fluorescence (Fig. 6b.i), whereas, using the same microscope settings, the fluorescence of conidia from dark-grown mycelia (DG conidia) was notably weaker (Fig. 6b.ii). In contrast, only dim background fluorescence was observed in wild type LG or DG conidia (Fig. 6b.iii). CarO::YFP was mainly located in the plasma membrane, but it was also present in membranes of inner organelles, possibly corresponding to vacuoles, endoplasmic reticulum and nuclear envelope (Fig. 6b, d). A similar distribution was observed when carO::yfp cDNA was expressed in S. cerevisiae (Fig. 6c).
The expression of CarO::YFP was also analysed in young F. fujikuroi hyphae (<24 h after germination) developed in light or dark and germinated either from LG or DG conidia. Overall, fluorescence intensity was lower in young mycelia (<24 h after germination) than in conidia (Supplementary Fig. S5). Interestingly, the signal was not evenly found in the hyphae (representative example shown in Fig. 6d). For better understanding of this heterogeneous distribution, we recorded the fluorescence in young hyphae in the first 18 hours, starting 5 hours after spore seeding. We observed a highly dynamic localization for CarO. Though the rhodopsin remained concentrated in regions of the former conidia, it also extended alongside the hyphae. In addition CarO was newly produced and trafficked to plasma membrane and septae behind growth regions (ca. 50–100 μm behind the hyphal tip). In certain regions, often next to a branch, also the fungal vacuoles exhibited strong fluorescence in their membranes (Supplementary Fig. S6 and Movie 1).
CarO activity affects conidia germination
The abundance of CarO in conidia (Fig. 6b) led us to hypothesize that this rhodopsin might play a role in spore viability or germination. We observed that germination of F. fujikuroi conidia is supported by low pH (Supplementary Fig. S8). Therefore, we wanted to find out if CarO acidifies the medium and thereby supports spore germination.
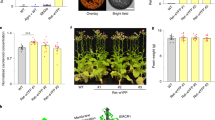
As carO expression is up-regulated by light, we compared the effect of light on growth and germination in different conditions of conidia from a strain with a frame shift mutation in the gene carO26 (CarO− strain), in parallel with an isogenic CarO+ control strain (Fig. 7a). LG conidia were spotted on solid media (N-source ammonium chloride; C-source glucose) and germinated either in light or darkness. To visualize acidification, we used media supplemented with bromocresol green (Supplementary Fig. S7).
Colony growth of F. fujikuroi carO− and carO+ strains in the dark and under light.
(a) carO+ and carO− strains were spotted in standard DA minimal medium (104 conidia per drop) and documented after 2 days incubation under light. White bar represents 5 mm. (b) Influence of carbon source on colony size. Inoculation was done as described in (a) but in DA media with varying additional carbon sources as indicated. Colonies were either grown in the light or in the dark. (c) Linear growth experiments in race tubes. carO+ and carO− strains were grown in standard DA minimal medium (104 conidia per drop; n = 4) and colony length was measured over a period of 9 days. Mean values are given with standard deviation (y-bars).
Acidification halos were noticeable in the medium surrounding the colonies in a degree that depended on the colony diameters. Taking this into consideration, we could not detect clear differences in the acidification patterns between CarO− and CarO+ strains. In another set of experiments, concentrated suspensions of conidia of both strains were grown in the dark and illuminated and pH of the medium was monitored submerging the pH sensor in the sample. Again, no differences were identified in pH values between both strains. These results suggest that CarO is not significantly involved in medium acidification.
However, when exposed to light, the mycelia of the CarO− strain grew faster than those of the CarO+ control strain in all tested conditions, while the difference was basically non-existent in darkness. Because of the stimulatory effect of gluconate described above, this organic acid was added in parallel with other carbon compounds, such as galacturonic acid, arabinose and xylose. The inhibition under light was especially clear in the standard minimal medium or upon addition of arabinose or xylose. Interestingly, the presence of gluconate led to a faster growth of the fungus irrespective of the carO mutation, making the difference in growth less apparent at the same incubation time (Supplementary Fig. S11). The stimulating effect of gluconate on growth velocity was particularly noticeable in the dark.
The slower growth under light became less apparent after more prolonged incubation, suggesting that the effect is achieved at the start of mycelial development, possibly at the level of conidia germination. Therefore we performed linear growth experiments and recorded the differences in colony size between the CarO+ and CarO− strains over a period of 9 days (Fig. 7c). LG CarO− colonies appeared earlier than LG CarO+ colonies, but no significant difference could be observed afterwards in the growth rate of both strains.
Since the difference in growth under light between the CarO− and CarO+ strains was only noticeable in short-term incubations, we wanted to know if CarO photoactivity delays the spore germination or if it slows down germling development. We harvested conidia from CarO+ and CarO− strains and let them germinate either in the dark or under illumination. For this purpose the conidia were distributed in 24-well plates (3 wells per strain and condition) coated with poly-D-lysin and growth of hyphae was documented by microscopy (Fig. 8; 1 picture/well).
Influence of CarO on conidia germination.
Conidia from light and dark exposed mycelia of the CarO+ and CarO− strains were seeded in liquid DA medium adjusted to pH 5 and germination was observed. Data from one single representative experiment are shown, but similar results were obtained in four independent experiments. (a) Representative images of germinated conidia of light exposed strains. (b) Length of germlings of CarO+ LL (n = 1476) and CarO− LL (n = 1413) conidia as indicated. The percentage of >40 μm germlings is higher in CarO− LL than in CarO+ LL, while the opposite is the case for <30 μm germlings. (c) Length of germlings of CarO+ DD (n = 1392) and CarO− DD (n = 1196) conidia as indicated. Only slight difference was observed between both strains incubated in the dark.
The size of fresh conidia was similar for CarO+ and CarO− strains irrespective of light exposition of the mycelia (CarO+: LG 9 ± 2.7 μm, n = 62; DG 8.3 ± 2.7 μm, n = 175. CarO−: LG 8.1 ± 2.9 μm, n = 224; DG 7.5 ± 3 μm, n = 231). Germlings produced by conidia from LG mycelia exhibited faster growth than those from DG mycelia. When conidia from DG mycelia were used in germination experiments, after 12 hours there was no clear difference in size of hyphae between CarO+ and CarO− strains, but light exposed conidia (CarO+ mean length 23.2 μm, n = 1147 vs. CarO− 25 μm, n = 1225) germinated slower than dark exposed conidia (CarO+ 31.9 μm, n = 1392 vs. CarO− 30.4 μm, n = 1196). Faster germination in the dark was also observed with spores from LG mycelia. However, in this case CarO+ germlings clearly exhibited smaller size than CarO− germlings (light exposed: 25.5 μm, n = 1476 vs. 39.5 μm, n = 1413; dark exposed: 31.9 μm, n = 1253 vs. 34.4 μm, n = 1168, respectively). Hyphal length varied considerably between different germlings, ranging from <7 μm to >160 μm. However, the distribution of germling length differed between CarO+ and CarO− strains and longer hyphae were found in CarO− mutant (Fig. 8). We also analysed the spore germination rate after 9 hours and found that the percentage of germinated conidia (>10 μm) was almost double for CarO− strain (64.9%) than for CarO+ strain (34.2%). Taking all data together, the results suggest a faster germination of CarO− conidia in the light compared to those of the control CarO+ strain.
Discussion
Fungal rhodopsins are widespread in the fungal kingdom3,13, but their biological functions, particularly how their light-dependent functions affect fungal growth and physiology, remain unknown. In this report we investigate the biological role of the auxiliary ORP-like rhodopsin CarO of the filamentous fungus F. fujikuroi. For this purpose, we used three different experimental approaches (a) the study of pump activity of heterologously expressed CarO through patch-clamp experiments, (b) the identification of in vivo location of YFP-tagged CarO and (c) the phenotypic comparison of a targeted CarO− mutant and an isogenic CarO+ strain. Taken together, the results provide solid clues on a possible role of CarO as a proton pump associated to germination of conidia.
The first approach showed that CarO is an ion pump rather than a photosensor, as expected from the conservation of residues needed for proton pumping in other rhodopsins (Fig. 1). While pumps must have rather fast photochemical cycles to maintain transmembrane ion gradients, the photosensors are characterized by their relatively slow turnover-rates. The photocycle of CarO has not been investigated, but recently it was shown that another auxiliary ORP-like rhodopsin, PhaeoRD2 from Phaeosphaeria nodorum, exhibits a very fast photocycle, hence providing support to a role of this group of rhodopsins as proton pumps. However, to our knowledge, no electrophysiology studies are available on auxiliary ORP-like rhodopsins.
To facilitate the generation of reliable electrophysiological data, we carried out patch-clamp experiments in mammalian cells expressing CarO::YFP. We chose this experimental model because patch-clamp experiments of fungal protoplasts are complex32,33,34 and high-quality F. fujikuroi protoplasts could only be obtained in a narrow time-window. In contrast, the use of a stable CarO::YFP-expressing mammal cell line allowed us to record abundant and reproducible data. The fusion protein was sufficiently expressed, targeted to plasma membranes and therefore suitable for the patch-clamp technique. The expression pattern was similar in F. fujikuroi, S. cerevisiae and 293 cells (Fig. 1 and Fig. 6), leading us to expect the same biochemical activity for the protein in different eukaryotic host systems, as was shown in former reports on other microbial rhodopsins10,33,35. The activity of YFP is not expected to influence the physiological properties of microbial rhodopsins as has been shown recently for an YFP-, mCherry- and myc-tagged ChR236.
Our results provide solid support to the activity of the ORP-like rhodopsin CarO as an outwardly directed, green light-driven proton pump. CarO exhibited bell-shaped action spectrum, similar to that found for other microbial rhodopsins37,38, with a maximal efficiency at around 520 nm (Fig. 3a). Active ion transport against the electrochemical gradient was demonstrated by the distinct shift of reversal potentials towards negative values when compared with the values calculated for passive transport by Nernst equation. Specificity for proton transport was clearly shown by the rise of outward-directed light-induced pump current with increasing proton gradient (Fig. 4a, b) and by the lack of effect of chloride or sodium ions on pump activity (Fig. 4c).
In general, pump characteristics of CarO were similar to those described for BR. As observed for BR and other microbial proton pumps35, pump activity increased with rising membrane potentials. In BR this voltage-dependent behaviour is due to the M-decay, which is slowed down at negative membrane potentials and represents the rate-limiting step of the pump cycle39. Our data on CarO reflect a similar situation, as upon switching off the light, the photocurrent of CarO decayed in a voltage-dependent, biexponential manner. While the fast closing time constant τoff_1 for CarO was in the same range with BR40, the slow closing time constant τoff_2 was smaller in CarO (20 ± 3.8 ms) than in BR (27.2 ± 2.2 ms). In our system the rise time constant (0.3 ± 0.1 ms) was limited by the opening time of the fast shutter (around 300 μs). Hence we could not determine, if activation of CarO occurred more rapidly than in BR. However, as the M-decay is rate-limiting in BR and obviously occurs faster in CarO, we assume that the whole photocycle of CarO is faster than in BR, an assumption that fits flash-photolysis data from the auxiliary ORP-like Rhodopsin PhaeoRD219. In PhaeoRD2 deprotonation of the Schiff base occurred very fast, resulting in reduced accumulation of the late red-shifted intermediate and thus, in a very fast photocycle. Fast photocycling of CarO was also supported by the observation that, in comparison to other rhodopsins, its pumping activity saturated at very high light intensities (Fig. 3b).
CarO activity declined with increasing extracellular proton concentration when the bath solution was based on the anions chloride, sulphate, or galacturonate. Since at low extracellular pHs the release of protons is impeded, this is the expected behaviour for proton pumping rhodopsins. In contrast, in the presence of the weak organic acids (WOAs) gluconate or glutamate the outward directed current was increased at pH 5 compared to neutral pH (Fig. 5). As this behaviour was mostly independent from the presence or absence of an WOA-gradient (Supplementary Fig. S4), the enhanced charge transfer cannot be explained by additional WOA pump function beside proton pump ability. A more reliable explanation is inferred from the proton transfer mechanism of BR and other microbial rhodopsins. Once the proton has been transferred from the Schiff base to the proton acceptor BR-D85, it is guided to the extracellular space via a cascade of proton acceptors with different pKas to the well conserved proton releasing site41,42. A fast pumping activity is expected to generate a “proton traffic jam”, i.e., an accumulation of protons nearby the releasing site, thus slowing down proton liberation to the extracellular medium43. The substances producing enhanced CarO pump activity exhibit pKa values around 4 (sodium gluconate pKa 3.86, sodium glutamate second pKa 4.07). Thus, at pH 5 those WOAs may act as buffers, supporting the release of protons from the pumping CarO protein. This would lead to a faster photocycle and thus to more efficient pump activity, as indicated by smaller time constants (Supplementary Fig. S3) and higher current amplitudes (Fig. 5).
Among the investigated WOAs, gluconate provoked an additional extraordinary effect on CarO activity in response to pH change. When CarO was subjected to gluconate solution at pH 5 and the pH gradient was turned to pH 9, the pump activity transiently increased to about 10-fold of that in standard bath solution. Seemingly the supporting effect of gluconate transiently remained at pH 9, which in turn resulted – now in combination with the driving force of the outward directed proton gradient – in a temporary faster photocycle (Supplementary Fig. S3). However, this supporting effect disappeared within a time scale of about 10 minutes. Strong pH-dependency of the photocycle was also observed in the related auxiliary ORP-like rhodopsin PhaeoRD2, whose photocycle was considerably faster at pH 5 than at neutral or basic pH19. Auxiliary ORP-like rhodopsins are presumed to interact with transducer proteins19. So, we may also speculate that gluconate could affect CarO conformation, or its interaction capacity. Interestingly gluconate does not only influence the pump activity of CarO: its presence results in a remarkable stimulation of mycelial growth of F. fujikuroi on solid medium (Fig. 7b, Supplementary Fig. S11). This behaviour is independent of the presence of a functional CarO protein and it is also noticeable in the dark and therefore has no connection with the stimulatory effect of gluconate on CarO pump activity. The ecological relevance of gluconate in the F. fujikuroi natural habitat is not clear. The bakanae disease is seed born and it is known that root associated bacteria produce gluconic acid44. Such bacteria were present in ten different rice root samples45, thus this compound might play a role as a chemical attractant supporting fungal growth.
Under standard laboratory conditions no visible phenotype was observed for the CarO− mutant26. Our fluorescence assays (Fig. 6) with a CarO::YFP fusion protein expressed under control of the carO promoter showed that the protein is accumulated mainly in conidia produced by light-exposed mycelia and predominantly in the cytoplasmic membrane. In young hyphae the protein was irregularly distributed in a highly dynamic fashion, with fluorescence appearing transiently in the cytoplasmic membrane in regions behind the hyphal tip and septae, or in the vacuoles. These variations in plasma membranes and in inner organelles could be consistent with a role in pH dependent signalling and pH homeostasis46. So, CarO activity could generate external and internal changes in pH, predictably affecting hyphal development. Proton-gradients were reported to be involved in cell polarity and tip growth of fungi47,48,49, although more recent communications contradict the pH-gradient based cell polarity46,50,51. However, the prevalence of CarO in the conidia suggests that it might play a role in germination, possibly through the generation of a local proton gradient. In support to this hypothesis, the formation of mycelia from conidial spots on the agar surface was delayed by illumination under certain culture conditions, an effect partially mitigated by the absence of a functional CarO protein (Figs. 7 and 8). Ambient pH can influence spore germination rates and germ tube lengths, as reported for the phytopathogen Penicillium expansum52. Nevertheless, F. fujikuroi acidifies the medium and favours conidia germination (Supplementary Fig. S8) independently of CarO activity (Supplementary Fig. S7). Thus, we hypothesize that CarO uses protons to generate a signal promoting the delay of germination. This could play a relevant role in the life cycle of the fungus. During rice pathogenesis (“bakane” disease), F. fujikuroi mycelia emerge from infected aerial plant tissues and form cottony conidiating masses. Airborne CarO-containing conidia may contaminate healthy seeds, but they may also reach the soaked fields and produce new infections through the roots of healthy plants. Under these conditions it may be advantageous to delay the germination, promoted by water availability, waiting for a closer localization to rice roots in the marsh soils, predictably in a darker environment. CarO could be also involved in the control of conidiation, as suggested for NOP-1 from N. crassa53,54. However, the CarO− mutant exhibits similar conidiation levels as the CarO+ control26. Moreover, the mutants of the NOP-1 homologous rhodopsin of F. fujikuroi, OpsA, are not affected in conidiation either15. The biological role of OpsA and its possible connection with CarO is currently under investigation.
Beside this role, CarO may also participate in the maintenance of the proton gradient across the plasma membrane, which is mainly used for nutrient uptake. In fungi typically an H+-ATPase is responsible for the maintenance of this gradient50. Thus, in light exposed mycelia the consumption of ATP would be reduced due to the proton transfer by the rhodopsin, a clear advantage for the fungus. This hypothesis is supported by the recent observation that green-light sensing microbial rhodopsins are frequent in the phyllosphere55. It is suggestive that green-light should be predominant in internal tissues of thin aerial plant structures, filtered by chlorophyll. Interestingly, phytopathogenic ascomycetes frequently possess several fungal rhodopsins (see Supplementary Tab. S1 and Supplementary Fig. S12), while some well-known non-pathogenic fungal models, as Neurospora crassa, do not. Furthermore, fungi with CarO orthologues in their genome are usually either plant pathogens or endophytes (Supplementary Tab. S2) suggesting an evolutionary advantage of this rhodopsin for fungi living in these habitats.
We cannot exclude a potential role of CarO in fungal pathogenesis and/or plant interaction. A pH regulatory network mediated by the specific transcription factor PacC plays a decisive role in phytopathogenesis by Fusarium oxysporum56 and a similar PacC protein is involved in pH control of transcriptional regulation in F. fujikuroi57. A possible connection between CarO activity and PacC regulation and/or phytopathogenesis cannot be discarded. However, we could not observe any stimulation of CarO activity in the presence of different cell-wall related carbon sources. In addition, our plant tissue invasion assays in tomato and apple (Supplementary Fig. S9) revealed no significant difference between the CarO+ and CarO− strains, but situation might be different in rice, its natural host.
Conclusion
The ORP-like fungal rhodopsin CarO from the fungus F. fujikuroi is a functional green-light driven proton pump, predominantly found in the cytoplasmic membrane of conidia. In the presence of ammonium chloride, conidia from a CarO deficient strain germinated faster under light than those of a control strain providing the first relevant phenotypic alteration observed for a rhodopsin mutant in a fungus. During rice pathogenesis, conidia are formed by mycelia emerging from aerial tissues of infected plants. Airborne conidia may contaminate seeds or infect new plants in the soil of soaked fields, presumably in partial darkness. A biological task of CarO might be to retard conidia-germination under light to improve the chance of infection. CarO exhibits a fast photocycle, saturates at high light intensities, making it potentially useful for the fungus to save energy under daylight. CarO orthologues prevail in phytopathogenic fungi, which inhabit niches where green light is particularly abundant.
Methods
Strains and culture conditions
The wild type strain FKMC1995 of Fusarium fujikuroi (Gibberella fujikuroi mating population C) was kindly provided by J.F. Leslie (Kansas State University Collection, Manhattan, KS, USA). The carO− strain (SF100) and its carO+ control (SF101) were formerly described (strains R3 and R2, respectively26). Light incubations were done below a battery of four fluorescent tubes (Philips TL-D 18W/840) at a distance of 58 cm, providing ca. 7 W m−2 white light or in an Peltier-cooled incubator (IPP110 Plus, Memmert, Germany) equipped with white light emitting diodes (LEDs, mixed 2700 K/5500 K) providing 6–10 W m−2. Dark grown cultures were protected in a box or by aluminum foil.
To obtain conidia, the F. fujikuroi strains were grown on CG media (10 g l−1 D+(-)glucose, 0.1 g l−1 NH4NO3, 1 g l−1 KH2PO4, 0.5 g l−1 MgSO4*7H2O and 16 g l−1 agar) or on DG media58 with 3 g l−1 glutamine or asparagine instead of NaNO3 (DGgln or DGasn media, respectively). Strains were incubated at 25–28°C for 7 days before conidia were harvested in water, filtered through a sterile Whatman paper or glass filter (Pore size 2, Robu, Germany) and counted. Conidia from dark grown mycelia were harvested in a darkened room.
For phenotypic characterization, SF100 and SF101 were grown on minimal media in the presence of different carbon sources. For this purpose, DA (dextrose ammonium) medium (30 g l−1 D+(-)glucose, 2 g l−1 NH4Cl, 0.5 g l−1 MgSO4*7H2O, 0.5 g l−1 KCl, 2 ml l−1 microelements solution of DG medium51 and 16 g l−1 agar, adjusted to pH 5.5 by 4.31 mM citrate/11.37 mM NaHPO4) was chosen. In some cases, DA-medium was augmented by 28 mM of respective carbon source as indicated while D-glucose was reduced to 25 g l−1. Plates were inoculated with 4 μl drops containing 104 fresh conidia and incubated for 3 days at 30°C. Differences in colony growth were documented under a Leica M125 stereoscopic microscope.
For the following experiments conidia from either light-exposed (L) or dark-grown (D) mycelia of SF100 and SF101 were seeded and thereafter plates were either grown in dark (D) or light (L). For better readability, when appropriate the letters L and D will be combined in couples to indicate conditions of (i) mycelia incubation and (ii) germination of the corresponding conidia, respectively, for the referred strain. E.g. CarO− DL means spores obtained from dark-grown mycelia of strain SF100 and germinated in the light. Race tubes (10 ml DA agar, pH 5, 25 cm length) were inoculated with 1 μl drop of 104 spores SF100 D, SF100 L, SF101 D, or SF101 L, respectively, placed on either side of the tube and colony extension was documented daily during a 9-day period.
Transformation and molecular biology techniques
To investigate the capability of CarO protein to act as a proton pump, we established a cell line stably expressing a heterologous protein consisting on CarO fused to the enhanced yellow fluorescent protein (YFP). For this purpose, a 5.5 kb plasmid, called pSCDcarO, was synthesized. A fragment consisting of carO cDNA, without the stop codon, was fused to an 18 bp adapter followed by a DNA sequence coding for YFP. The resulting fragment was called cDcarO::YFP and was surrounded by two XhoI restriction sites. CdcarO::YFP was cloned in pMA-RQ to yield pScarO. The resulting plasmid was digested with XhoI to release cDcarO::YFP, which was purified and cloned in two different vectors previously linearized by digesting with XhoI: pcDNA5/FRT/TO to give pcDNA/FRT/TO-cDcarO::YFP and pKS1-ST to give pKS1-ST-cDcarO::YFP. The former plasmid was transfected into the Flip-In T-Rex 293 cell line (Invitrogen) to get a cell line stably expressing the heterologous CarO::YFP protein. In prevention of CarO::YFP being misprocessed in mammalian cells, S. cerevisiae DSY-5 was transformed with pKS1-ST-cDcarO::YFP as described before31, providing a second model for heterologous expression of the CarO protein.
A F. fujikuroi strain expressing CarO fused to YFP was generated to study the in vivo localization of the putative proton pump. For this purpose, a 6.8 kb plasmid, called pScarO, was synthesized. pScarO was generated by cloning into pMA-RQ, previously linearized with XbaI, a 3.2 kb DNA fragment containing the carO gene without the codon stop, surrounded by its native promoter and terminator region (ca. 1 kb each). An 18 bp adapter followed by YFP coding sequence was fused to the carO gene. Two XbaI restriction sites were added to the ends of the fragment and allowed the generation of pScarO. Digestion of pScarO with XbaI released the 3.2 kb fragment, that was cloned into XbaI-linearized pHJA2 plasmid to yield pHJA2-carO::YFP. Transformation of F. fujikuroi protoplasts with pHJA2-carO::YFP generated a strain expressing CarO::YFP.
For Southern blot analyses, genomic DNA from the wild type and two transformants was extracted from filtered samples following Weinkove et al.59 and digested with NcoI and BclI. DNA was quantified using a Nanodrop ND-100 spectrophotometer (Nanodrop Technologies) and the same amounts of digested DNA were loaded per lane on a 0.8% agarose gel. After electrophoresis, DNA fragmentes were transferred to a Hybond-N membrane (GE Healthcare). As a probe, a 1 kb segment of the carO gene was obtained by polymerase chain reaction (PCR) with primer set 1 (5′-GTCAACCGTGAGCCATATCAG-3′ and 5′-GGGAAGATGCGCTCGTCAATG-3′). The probe was heated for 10 min at 100°C and incubated afterwards for 4 h at 37°C with 2 μl hexanucleotide mix (10x concentrated), 2 μl of a pool of nonradioactive dATP, dGTP, dTTP (0.5 mM), 2 μl of [α-32P]dCTP (1.1 × 105 GBq mmol−1, Perkin Elmer) and 2 U of the Klenow fragment of Escherichia coli DNA polymerase I. The labelled probe was purified with GFX DNA Purification kit (GE Healthcare). Other molecular biology techniques were achieved following Sambrook et al.60. The presence of yfp sequene in the transformants was also checked by PCR with primer set 2 (5′-GGTGAGCAAGGGCGAGGA-3′ and 5′- CAGCTCGTCCATGCCGTG-3′).
PCR reactions were carried out with the Expand High Fidelity PCR System (Roche, Mannheim, Germany). Initial denaturation was at 94°C for 2 min, followed by 35 cycles of 30 s at 94°C, 30 s at 54°C and N s at 72°C (N = 80 s with primer set 1 and 60 s with primer set 2) and a final elongation step at 72°C for 10 min.
Transformation was done by electroporation of protoplasts, adapting a former protocol61. After digestion with enzymatic solution, FKMC1995 protoplasts were washed in 0.7 M NaCl and 1.2 M Sorbitol. Afterwards, they were resuspended in 1.2 M Sorbitol to a final density of 108 protoplasts ml−1 and lineal pHJA2-carO::YFP was added yielding a final concentration of 40 ng μl−1. An aliquot of 100 μl of the protoplast/DNA-suspension was used in each electroporation (Multiporator, Eppendorf, Hamburg, Germany; cuvette providing 1 mm electrode distance, one pulse, voltage amplitude 6 kV/cm, pulse duration 200 μs). After electroporation the protoplast suspension was transferred to a tube containing 4 ml of top-agar, mixed and spread on Petri dishes containing 25 ml of regeneration medium. These Petri dishes were incubated 15 hours at 25°C to allow protoplast regeneration and expression of the hygromycin resistance cassette. Then, 140 μl of hygromycin B (50 mg ml−1) were added for selection of transformants and the Petri dishes were incubated at 25°C for 10–14 days.
Stable mammalian cell line
The plasmid pcDNA5/FRT/TO-cDcarO::YFP was transfected into Flip-In T-Rex 293 cells (Invitrogen) according to the manufactures manual. Flip-In T-Rex 293 cells were grown in Dulbecco's modified eagle medium (DMEM) supplemented with fetal calf serum (10%), L-glutamine (2 mM), penicillin (100 U ml−1), streptomycin (100 μg ml−1), zeocin (100 μg ml−1) and blasticidin (15 μg ml−1). After transfection the Flip-In T-Rex 293 cells containing pcDNA5/FRT/TO-CdcarO::YFP were selected by adding hygromycin (100 μg/ml) and blasticidin (15 μg/ml). One day before patch-clamp experiments the cells were seeded on glass coverslips and protein expression was induced by adding tetracycline (1–5 μg ml−1). In addition the medium was supplemented with trans-retinal (1 nM) to ensure the presence of the presumed rhodopsin chromophore.
Patch-Clamp Experiments
The patch-clamp setup was installed on a vibration-attenuated table (TMC, Peabody, USA). A 561 nm-DPSS laser (150 mW, Changchun New Industries Optogenetics Tech. Co., Ltd.) was used to trigger rhodopsins activity. The laser beam was coupled into the fluorescence beam path of an inverse microscope (Axiovert 200, Zeiss, Jena, Germany). Illumination-time was controlled by means of a fast electronic shutter (S3ZM2-NL, Vincent associates, Rochester, USA). The illuminated area Ai was calculated according to the formula Ai = [(field of view/magnification)/2]2 * π. In our experiments we used a 63 fold objective with a field of view of 23 mm.
For the recording of action spectra a 150W-xenon arc lamp equipped with shutter and filter holder (Lot-qd, Darmstadt, Germany) was used. Illumination with different wavelengths at constant photon density of (2 × 1016 photons s−1 mm−2) was reached with narrowband interference filters (420, 450, 470, 490, 510, 532, 560, 580, 600, 620 and 640 nm; bandwidth 10 nm, Andovor Corporation, Salem, USA) in combination with neutral density filters. Light energy was measured by means of laser-powermeter (Coherent Labmax/Power Max PS10Q). The order of filters during measurements was varied to prevent that illumination by certain wavelengths could influence the subsequent reaction of the protein at other wavelength, which would lead to a distorted spectrum.
Whole-cell currents were recorded with an Axopatch 200B amplifier coupled to a DigiData 1440 interface (Molecular Devices Corporation, Union City, USA), low-pass filtered at 5 kHz and digitized at a sampling rate of 100 kHz. Data recordings were controlled by the Software Clampex 10.3 (Molecular Devices Co.). Data analysis was performed using Clampfit 10.3 (Molecular Devices Co.) and Origin 8.5 (OriginLab Corporation, Northampton, USA). Patch-Clamp experiments were performed using a perfusion-chamber described in detail before32. Pipettes (GB150F-8P, Scientific-Instruments, Hofheim, Germany) with a tip opening diameter of 1.5 μm exhibited a resistance of 3–5 MΩ in standard bath solution. Typically, a voltage step protocol reaching from −120 mV to +40 mV was applied. Compensation current for every membrane potential was recorded 3 times and sweeps were averaged.
Standard bath solution contained 140 mM NaCl, 2 mM MgCl2, 2 mM CaCl2 and 10 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES) pH 7.4. For pH screening, 2-(N-morpholino)ethanesulfonic acid (TRIS) or 2-(N-morpholino)ethanesulfonic acid (MES) was used instead of HEPES and the pH of the bath solution was adjusted to 5 or 9, respectively, by means of HCl, acetic acid, NaOH, or the hydroxid of the particular cation. CarO::YFP activity was investigated in the presence of different ions. For this purpose NaCl was substituted by the same amount of sodium gluconate, sodium glutamate, sodium galacturonate or by 90 mM sodium sulphate. Pipette solution contained 110 mM NaCl or NaGluconate as indicated, 2 mM MgCl2, 10 mM EGTA and 10 mM HEPES pH 7.4. In some experiments the pH was adjusted to 5 or 9, using MES or TRIS, respectively.
Microscopy
Hyphae and yeasts were microscopically investigated in 8-well Labtek II chambers coated with 0.01% Poly-D-lysine (125 μl per well, incubation >24 h at 37°C). 5 × 103 F. fujikuroi conidia were seeded in 500 μl DGgln medium and incubated for 16–17 h at 28°C. S. cerevisiae DSY5 pKS1-ST-carO::YFP yeasts were grown in liquid yeast extract peptone dextrose (YPD) medium supplemented with 60 μg/ml G418 and seeded to the Labtek II chamber immediately before fixation. Probes were fixed for 10 minutes by adding 200 μl 4% formaldehyde in phosphate buffered saline (PBS, pH 7.4) and stored in PBS at 4°C. Probes were analysed using a confocal laser scanning microscope (Leica SP2, Wetzlar, Germany) or a fluorescence microscope (Olympus IX71, Hamburg, Germany) equipped with an electron multiplying charge-coupled device (EMCCD) camera (Andor, Belfast, UK). For the documentation of CarO::YFP distribution during germination an inverse fluorescence microscope (Axiovert200) equipped with a CMOS camera (Zyla 5.5, Andor, Belfast, UK) controlled by the software Micromanager (University of California, San Francisco, USA) was used. Every 2–5 min a picture was recorded and images were processed using imageJ and combined in a movie (Supplementary Movie 1).
Germination assays were performed in 24-well plates, previously coated with 0.01% Poly-D-Lysine. 500 μl of DA liquid media were inoculated with 5 × 104 fresh conidia per well. Germination of conidia was investigated using the Axiovert200 and documented 0, 3, 7, 9 and 12 hours (light-exposed germlings only) after inoculation. Dark exposed germlings were only fixed and analysed after 12 h. The length of germlings was determined using ImageJ software (1.49b).
References
Corrochano, L. M. Fungal photoreceptors: sensory molecules for fungal development and behaviour. Photochem. Photobiol. Sci. 6, 725–736 (2007).
Bayram, Ö., Braus, G. H., Fischer, R. & Rodríguez-Romero, J. Spotlight on Aspergillus nidulans photosensory systems. Fungal Genet. Biol. 47, 900–908 (2010).
Idnurm, A., Verma, S. & Corrochano, L. M. A glimpse into the basis of vision in the kingdom Mycota. Fungal Genet. Biol. 47, 881–892 (2010).
Avalos, J. & Estrada, A. F. Regulation by light in Fusarium. Fungal Genet. Biol. 47, 930–938 (2010).
Herrera-Estrella, A. & Horwitz, B. A. Looking through the eyes of fungi: molecular genetics of photoreception. Mol. Microbiol. 64, 5–15 (2007).
Corrochano, L. M. & Avalos, J. [Light sensing]. Cellular and molecular biology of filamentous fungi [Borkovich K., & Ebbole D. J. (eds.)] [417–441] (ASM Press, Washington, 2010).
Klare, J. P., Chizhov, I. & Engelhard, M. Microbial rhodopsins: scaffolds for ion pumps, channels and sensors. Results Probl. Cell Differ. 45, 73–122 (2008).
Spudich, J. L. & Jung, K.-H. [Microbial rhodopsins: phylogenetic and functional diversity]. Handbook of photosensory receptors [Briggs W. R., & Spudich J. L. (eds.)] [1–23] (VILEY-VCH, Weinheim, 2005).
Grote, M., Engelhard, M. & Hegemann, P. Of ion pumps, sensors and channels -Perspectives on microbial rhodopsins between science and history. Biochim. Biophys. Acta - Bioenergetics 1837, 533–545 (2013).
Ernst, O. P. et al. Microbial and animal rhodopsins: structures, functions and molecular mechanisms. Chem. Rev. 114, 126–163 (2014).
Sharma, A. K., Spudich, J. L. & Doolittle, W. F. Microbial rhodopsins: functional versatility and genetic mobility. Trends Microbiol. 14, 463–469 (2006).
Brown, L. S. & Jung, K. H. Bacteriorhodopsin-like proteins of eubacteria and fungi: the extent of conservation of the haloarchaeal proton-pumping mechanism. Photochem. Photobiol. Sci. 5, 538–546 (2006).
Brown, L. S. Fungal rhodopsins and opsin-related proteins: eukaryotic homologues of bacteriorhodopsin with unknown functions. Photochem. Photobiol. Sci. 3, 555–565 (2004).
Kihara, J., Tanaka, N., Ueno, M. & Arase, S. Cloning and expression analysis of two opsin-like genes in the phytopathogenic fungus Bipolaris oryzae. FEMS Microbiol. Lett. 295, 289–294 (2009).
Estrada, A. F. & Avalos, J. Regulation and targeted mutation of opsA, coding for the NOP-1 opsin orthologue in Fusarium fujikuroi. J. Mol. Biol. 387, 59–73 (2009).
Estrada, A. F. et al. Ustilago maydis accumulates β-carotene at levels determined by a retinal-forming carotenoid oxygenase. Fungal Genet. Biol. 46, 803–813 (2009).
Saranak, J. & Foster, K. W. Rhodopsin guides fungal phototaxis. Nature 387, 465–466 (1997).
Avelar, G. M. et al. A rhodopsin-guanylyl cyclase gene fusion functions in visual perception in a fungus. Curr. Biol. 24, 1234–1240 (2014).
Fan, Y., Solomon, P., Oliver, R. P. & Brown, L. S. Photochemical characterization of a novel fungal rhodopsin from Phaeosphaeria nodorum. Biochim. Biophys. Acta - Bioenergetics 1807, 1457–1466 (2011).
Waschuk, S. A., Bezerra, A. G., Shi, L. & Brown, L. S. Leptosphaeria rhodopsin: Bacteriorhodopsin-like proton pump from a eukaryote. Proc. Natl. Acad. Sci. U. S. A. 102, 6879–6883 (2005).
Brown, L. S. [Proton-pumping microbial rhodopsins – ubiquitous structurally simple helpers of respiration and photosynthesis]. The structural basis of biological energy generation [Hohmann-Marriott M. F. (ed.)] [1–20] (Springer, Dordrecht, 2014).
Idnurm, A. & Heitman, J. Light controls growth and development via a conserved pathway in the fungal kingdom. PLoS Biol. 3, e95 (2005).
Rademacher, W. [Gibberilins]. Fungal Biotechnology [Timm A. (ed.)] [193–205] (Chapmann & Hall, Weinheim, 1997).
Avalos, J., Cerdá-Olmedo, E., Reyes, F. & Barrero, A. F. Gibberellins and other metabolites of Fusarium fujikuroi and related fungi. Curr. Org. Chem. 11, 721–737 (2007).
Wiemann, P. et al. Deciphering the cryptic genome: genome-wide analyses of the rice pathogen Fusarium fujikuroi reveal complex regulation of secondary metabolism and novel metabolites. PLoS Pathog. 9, e1003475 (2013).
Prado, M. M., Prado-Cabrero, A., Fernández-Martín, R. & Avalos, J. A gene of the opsin family in the carotenoid gene cluster of Fusarium fujikuroi. Curr. Genet. 46, 47–58 (2004).
Chow, B. Y. et al. High-performance genetically targetable optical neural silencing by light-driven proton pumps. Nature 463, 98–102 (2010).
Thewes, S., Prado-Cabrero, A., Prado, M. M., Tudzynski, B. & Avalos, J. Characterization of a gene in the car cluster of Fusarium fujikuroi that codes for a protein of the carotenoid oxygenase family. Mol. Genet. Genomics 274, 217–228 (2005).
Bamberg, E., Tittor, J. & Oesterhelt, D. Light-driven proton or chloride pumping by halorhodopsin. Proc. Natl. Acad. Sci. U. S. A. 90, 639–643 (1993).
Inoue, K. et al. A light-driven sodium ion pump in marine bacteria. Nat. Commun. 4, 1678 (2013).
Walz, W. Patch-clamp analysis: advanced techniques. 2nd edn. (Humana Press, Totowa, 2007).
Terpitz, U. et al. Dielectric analysis and multi-cell electrofusion of the yeast Pichia pastoris for electrophysiological studies. J. Membr. Biol. 245, 815–826 (2012).
Terpitz, U. et al. Electrofused giant protoplasts of Saccharomyces cerevisiae as a novel system for electrophysiological studies on membrane proteins. Biochim. Biophys. Acta. - Biomembranes 1778, 1493–1500 (2008).
Garrill, A. & Davies, J. M. Patch clamping fungal membranes - a new perspective on ion-transport. Mycol. Res. 98, 257–263 (1994).
Nagel, G., Mockel, B., Büldt, G. & Bamberg, E. Functional expression of bacteriorhodopsin in oocytes allows direct measurement of voltage dependence of light induced H+ pumping. FEBS Lett. 377, 263–266 (1995).
Nikolic, K. et al. Photocycles of channelrhodopsin-2. Photochem. Photobiol. 85, 400–411 (2009).
Kleinlogel, S. et al. A gene-fusion strategy for stoichiometric and co-localized expression of light-gated membrane proteins. Nat. Methods 8, 1083–1088 (2011).
Nagel, G. et al. Channelrhodopsin-2, a directly light-gated cation-selective membrane channel. Proc. Natl. Acad. Sci. U. S. A. 100, 13940–13945 (2003).
Geibel, S. et al. The voltage-dependent proton pumping in bacteriorhodopsin is characterized by optoelectric behavior. Biophys. J. 81, 2059–2068 (2001).
Geibel, S., Lörinczi, E., Bamberg, E. & Friedrich, T. Voltage dependence of proton pumping by bacteriorhodopsin mutants with altered lifetime of the M intermediate. PLoS ONE 8, e73338 (2013).
Dencher, N. A., Sass, H. J. & Büldt, G. Water and bacteriorhodopsin: structure, dynamics and function. Biochim. Biophys. Acta - Bioenergetics 1460, 192–203 (2000).
Lanyi, J. K. Proton transfers in the bacteriorhodopsin photocycle. Biochim. Biophys. Acta - Bioenergetics 1757, 1012–1018 (2006).
Kikukawa, T. et al. [Photo-induced proton transfers of microbial rhodopsins]. Molecular photochemistry - various aspects [Saha S. (ed.)] [89–108] (InTech, Rijeka, 2012).
Rodríguez, H., González, T., Goire, I. & Bashan, Y. Gluconic acid production and phosphate solubilization by the plant growth-promoting bacterium Azospirillum spp. Naturwissenschaften 91, 552–555 (2004).
Hardoim, P. R. et al. Rice root-associated bacteria: insights into community structures across 10 cultivars. FEMS Microbiol. Ecol. 77, 154–164 (2011).
Bagar, T., Altenbach, K., Read, N. D. & Bencina, M. Live-cell imaging and measurement of intracellular pH in filamentous fungi using a genetically encoded ratiometric probe. Eukaryot. Cell 8, 703–712 (2009).
Jelitto, T. C. Confocal ratio imaging of cytoplasmic pH during germ tube growth and appressorium induction by Magnaporthe grisea. New Phytologist 144, 499–506 (1999).
Robson, G. D. et al. Polarized growth of fungal hyphae is defined by an alkaline pH gradient. Fungal Genet. Biol. 20, 289–298 (1996).
Turian, G. Polarized acidification at germ tube outgrowth from fungal spores (Morchella ascospores, Neurospora conidia). Bot. Helv. 93, 27–32 (1983).
Hesse, S. J. A., Ruijter, G. J. G., Dijkema, C. & Visser, J. Intracellular pH homeostasis in the filamentous fungus Aspergillus niger. Eur. J. Biochem. 269, 3485–3494 (2002).
Valkonen, M., Penttila, M. & Bencina, M. Intracellular pH responses in the industrially important fungus Trichoderma reesei. Fungal Genet. Biol. 70, 86–93 (2014).
Li, B. Q., Lai, T. F., Qin, G. Z. & Tian, S. P. Ambient pH stress inhibits spore germination of Penicillium expansum by impairing protein synthesis and folding: a proteomic-based study. J. Proteome Res. 9, 298–307 (2010).
Bieszke, J. A., Li, L. & Borkovich, K. A. The fungal opsin gene nop-1 is negatively-regulated by a component of the blue light sensing pathway and influences conidiation-specific gene expression in Neurospora crassa. Curr. Genet. 52, 149–157 (2007).
Olmedo, M. et al. A role in the regulation of transcription by light for RCO-1 and RCM-1, the Neurospora homologs of the yeast Tup1–Ssn6 repressor. Fungal Genet. Biol. 47, 939–952 (2010).
Atamna-Ismaeel, N. et al. Microbial rhodopsins on leaf surfaces of terrestrial plants. Environ. Microbiol. 14, 140–146 (2012).
Caracuel, Z. et al. The pH signalling transcription factor PacC controls virulence in the plant pathogen Fusarium oxysporum. Mol. Microbiol. 48, 765–779 (2003).
Wiemann, P. et al. Biosynthesis of the red pigment bikaverin in Fusarium fujikuroi: genes, their function and regulation. Mol. Microbiol. 72, 931–946 (2009).
Avalos, J., Casadesús, J. & Cerdá-Olmedo, E. Gibberella fujikuroi mutants obtained with UV-radiation and N-Methyl-N'-Nitro-N-Nitrosoguanidine. Appl. Environ. Microbiol. 49, 187–191 (1985).
Weinkove, D. et al. Mutants of Phycomyces with decreased gallic acid content. Fungal Genet. Biol. 25, 196–203 (1998).
Sambrook, J. & Russell, D. W. Molecular Cloning : a Laboratory Manual. 3rd edn. (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 2001).
Proctor, R. H., Hohn, T. M. & McCormick, S. P. Restoration of wild-type virulence to Tri5 disruption mutants of Gibberella zeae via gene reversion and mutant complementation. Microbiology 143, 2583–2591 (1997).
Biasini, M. et al. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 42, W252–W258 (2014).
Acknowledgements
We thank A. Adam, D. Lühders, N. Müller, A. Geringer and M. Castrillo for great assistance in the lab. We are grateful that this work was supported by the German Research foundation (DFG, TE-832-4/1), the Bavarian Research Alliance (Travel costs, BayIntAn_Uni_Würzburg_2012_39), the Spanish Government (projects EEBB-I-12-0582, BIO2009-11131 and BIO2012-39716) and the Andalusian Government (projects P07-CVI-02813 and CTS-6638). Spanish grants included support from the European Union (European Regional Development Fund [ERDF]).
Author information
Authors and Affiliations
Contributions
U.T. and J.A. designed research; J.G., M.B. and U.T. performed research; J.G., M.B. and U.T. analysed data; U.T. and J.A. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Supplementary Information
Germination of F.fujikuroi CarO::YFP LG conidia
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
García-Martínez, J., Brunk, M., Avalos, J. et al. The CarO rhodopsin of the fungus Fusarium fujikuroi is a light-driven proton pump that retards spore germination. Sci Rep 5, 7798 (2015). https://doi.org/10.1038/srep07798
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07798
This article is cited by
-
Light in the box—photobiological examination chamber with light trap ventilation system for studying fungal surface cultures illustrated with Metarhizium brunneum and Beauveria brongniartii
Fungal Biology and Biotechnology (2023)
-
Recent advances in metabolic regulation and bioengineering of gibberellic acid biosynthesis in Fusarium fujikuroi
World Journal of Microbiology and Biotechnology (2022)
-
Use of red, far-red, and near-infrared light in imaging of yeasts and filamentous fungi
Applied Microbiology and Biotechnology (2022)
-
Functional expression of the eukaryotic proton pump rhodopsin OmR2 in Escherichia coli and its photochemical characterization
Scientific Reports (2021)
-
Superaufgelöste Mikroskopie: Pilze unter Beobachtung
BIOspektrum (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.