Abstract
Topological centrality is a significant measure for characterising the relative importance of a node in a complex network. For directed networks that model dynamic processes, however, it is of more practical importance to quantify a vertex's ability to dominate (control or observe) the state of other vertices. In this paper, based on the determination of controllable and observable subspaces under the global minimum-cost condition, we introduce a novel direction-specific index, domination centrality, to assess the intervention capabilities of vertices in a directed network. Statistical studies demonstrate that the domination centrality is, to a great extent, encoded by the underlying network's degree distribution and that most network positions through which one can intervene in a system are vertices with high domination centrality rather than network hubs. To analyse the interaction and functional dependence between vertices when they are used to dominate a network, we define the domination similarity and detect significant functional modules in glossary and metabolic networks through clustering analysis. The experimental results provide strong evidence that our indices are effective and practical in accurately depicting the structure of directed networks.
Similar content being viewed by others
Introduction
Studies of the structure and function of complex networks can yield a variety of useful quantities or measures that capture particular features of social, biological and information-technology systems1. In this context, the concept of centrality addresses the most important or central vertices in a network. Despite the diversity of systems, several basic, universal measures of centrality have been developed to rank the vertices of a network according to their topological importance, including the vertex degree, betweenness2,3, closeness4, eigenvector5, subgraph6, PageRank7 and various types of random walks8,9. Although these measures have significantly enriched our understanding of many networks, our ultimate goal is to locate the most significant vertices that have the ability to dominate the networks.
Although the actual domination of complex networks has not yet been achieved at present, a necessary stepping stone is to understand the controllability and observability of complex networks, which has become a topic of active pursuit38,39,40,41,42. Based on control theory, Liu et al.10 have proposed an efficient methodology for identifying the minimum driver vertex set (MDS), the time-dependent control of which can guide the entire network to any desired final state11,12. The number of elements of the MDS, ND, is thus a key quantity of interest, as it characterises the cost of bringing the system under full control. The proposed maximum matching link set M can be used to assess and quantify structural controllability. On the other hand, a network is observable if its internal state can be determined from the given output vertex set, where observability depends on both the number and placement of the output vertices13. Liu et al.14 have adopted a graphical approach to determining the set of output vertices that are not only necessary but also sufficient for the observability of a complex network. Specifically, given a complex-networked dynamical system, the controllable subspace reflects the control capability of a vertex when we input a signal at that single vertex only and the observable subspace reflects the observation capability of a vertex when we measure the output from that single vertex only. Recently, Liu et al.15 and Wang et al.16 have further introduced the concept of control centrality to quantify the ability of a single vertex to control a directed weighted network. However, the use of only the control capability to quantify the vertex centrality is not comprehensive, as a vertex may directly intervene only in its downstream subspace from the viewpoint of controllability. For example, in figure 1, by inputting a signal at the vertex i, the state variables in the downstream system S2 can be controlled. This is the manner in which a vertex controls its downstream system, but this process embodies only one aspect of a vertex's power in dominating a system. If the state variables are looped back, the feedback signal can then control a system within itself. State feedback is self-related and helps to maintain stability in a system despite external changes. However, state feedback can be established only on the condition that all state variables are measurable. If not, the state variables must be estimated by utilising a state observer. In figure 1, the state observer obtains the state variables of the upstream system S1 by measuring the state of vertex i; the feedback loop can then be constructed to control S1. In this manner, a vertex can control its upstream system through feedback and this process reflects another aspect of a vertex's power in dominating a system. Therefore, the ability to examine the role that a vertex plays in both controlling the downstream subspace and observing the upstream subspace is an issue of significant practical interest in vertex centrality.
A schematic diagram illustrating the physical meaning of domination capability.
By inputting a signal at the vertex i, the state variables in the downstream system S2 can be controlled. By observing the state variables by measuring the state of vertex i, a feedback loop can be constructed to control the upstream system S1.
In this paper, we focus on the domination centrality (DC) index to assess the capabilities of vertices in directed networks. Intuitively, domination centrality includes two aspects: control capability and observation capability. Under the minimum-control-cost condition, for a single vertex, the control capability captures the dimension of the controllable subspace and quantifies the influence that can be exerted on the downstream subnetwork through this vertex. Similarly, the observation capability captures the dimension of the observable subspace and quantifies the intervention that can be exerted on the upstream subnetwork through this vertex. The purpose of emphasising the global minimum cost is to determine the responsibility and capability of each individual vertex cooperating with others in dominating the entire system. Mathematically, domination centrality is the harmonic mean of these two capabilities and represents the capability of a vertex synthetically. This approach is in good agreement with our original notion regarding the “power” of a vertex in dominating the entire network. Inspired by this general consideration, we perform statistical studies of the index DC for several types of real-world directed networks, including citation, metabolic, glossary and synthetic networks and analyse the underlying topological factors by which the distribution of DC is primarily determined. To uncover DC and functions of vertices, a clustering analysis is presented based on the intuitive assumption that vertices that control and observe the same subspaces tend to serve identical functions in a network. Our domination centrality index bridges the concepts of directed network topology and function by providing useful insights into the effect of the former on the latter from the viewpoint of cybernetics.
Results
Domination centrality
Consider the linear time-invariant dynamic system  , Y(t) = C·X(t) with the state vector
, Y(t) = C·X(t) with the state vector  , the adjacency matrix
, the adjacency matrix  , the input matrix
, the input matrix  , the control vector
, the control vector  , the output matrix
, the output matrix  and the output vector
and the output vector  . The underlying directed network of this system is denoted by G(A), with vertex set V and link set L. The rank of the n × nm controllability matrix QC ≡ [B, AB, A2B,…, An−1B], which is denoted by rank(QC), provides the dimension of the controllable subspace of the structural system (A, B, C)17,18. (A, B, C) is completely controllable12 iff rank (CC) = n. Analogously, the rank of the nr × n observability matrix QO ≡ [[C]T, [CA]T, [CA2]T, …, [CAn−1]T]T, which is denoted by rank(QO), provides the dimension of the observable subspace of this system. (A, B, C) is completely observable iff rank (QO) = n. Furthermore, the duality theorem12 indicates that system (A, B, C) is completely controllable if and only if system (AT, CT, BT) is completely observable and vice versa.
. The underlying directed network of this system is denoted by G(A), with vertex set V and link set L. The rank of the n × nm controllability matrix QC ≡ [B, AB, A2B,…, An−1B], which is denoted by rank(QC), provides the dimension of the controllable subspace of the structural system (A, B, C)17,18. (A, B, C) is completely controllable12 iff rank (CC) = n. Analogously, the rank of the nr × n observability matrix QO ≡ [[C]T, [CA]T, [CA2]T, …, [CAn−1]T]T, which is denoted by rank(QO), provides the dimension of the observable subspace of this system. (A, B, C) is completely observable iff rank (QO) = n. Furthermore, the duality theorem12 indicates that system (A, B, C) is completely controllable if and only if system (AT, CT, BT) is completely observable and vice versa.
Liu's Minimum Input Theorem10 states that the minimum number of driver vertices (ND) required to fully control a network G(A) is one if there is a perfect matching in G(A). Otherwise, it is equal to the number of unmatched vertices with respect to any maximum matching, ND = max{n − |M|,1}. A maximum matching is a link set  with maximum cardinality (size) and no two links in M may share a common starting vertex or a common ending vertex. A vertex is matched if it is an ending vertex of a link in M. |M| denotes the size of the maximum matching.
with maximum cardinality (size) and no two links in M may share a common starting vertex or a common ending vertex. A vertex is matched if it is an ending vertex of a link in M. |M| denotes the size of the maximum matching.
For a given maximum matching link set M of a directed network G(A), the minimum-control-cost configuration  carries the structural information of completely control16. CF is a spanning subnetwork of G(A), with vertex set V and link set
carries the structural information of completely control16. CF is a spanning subnetwork of G(A), with vertex set V and link set  . M is a stem-cycle disjoint cover of G(A) and indicates the directed routes along which the input control signals are transmitted. AL is the set of additional links that begin in vertices of stems (except the top vertices) and end in vertices of cycles. The n × n adjacent matrix A(M) is used to indicate the wiring diagram of the spanning subnetwork CF that corresponds to the maximum matching link set M of G(A). As an example, in figure 2(a), the red links are elements of a maximum matching. When vertices are connected by red links, the network thus constructed is composed of vertex-disjoint stems (two in shades of green) and cycles (four in shades of red); l3,7 and l4,9 are the additional links that connect the stems and cycles.
. M is a stem-cycle disjoint cover of G(A) and indicates the directed routes along which the input control signals are transmitted. AL is the set of additional links that begin in vertices of stems (except the top vertices) and end in vertices of cycles. The n × n adjacent matrix A(M) is used to indicate the wiring diagram of the spanning subnetwork CF that corresponds to the maximum matching link set M of G(A). As an example, in figure 2(a), the red links are elements of a maximum matching. When vertices are connected by red links, the network thus constructed is composed of vertex-disjoint stems (two in shades of green) and cycles (four in shades of red); l3,7 and l4,9 are the additional links that connect the stems and cycles.
A schematic diagram illustrating the domination centrality and the domination similarity of vertices in a directed network.
(a): A maximum matching M1 consisting of red links, forms a stem (in shades of green)-cycle (in shades of red) disjoint cover of the network. Additional links (AL) that connect the stems and cycles are highlighted by bold dashed lines. (b): The controllable subspace of vertex 1 is highlighted by a purple dotted line and its observable subspace is highlighted by a green dotted line. The domination centrality of vertex 1 is the harmonic mean of the size of these two subspaces. (c): Another maximum matching, M2, is given. (d): The controllable subspace of vertex 2 is highlighted by an orange dotted line and its observable subspace is highlighted by a blue dotted line. (e): The overlapping phenomenon of the controllable subspaces and the observable subspaces is depicted. The Jaccard similarity coefficients of the controllable subspaces and the observable subspaces are calculated and the arithmetic-geometric mean thereof is used to determine the domination similarity.
The controllability of a complex network concentrates on the interaction structure in which the pattern of influence may be known, but not the specific extent of influence. In response to unknown or uncertain edge weights, the controllability is used to uncover the generic properties of systems, independent of parameter values. The cactus is the most economical topology-structure pattern to propagate control influence, since the cactus is a minimal structure such that removing any link will render the structure uncontrollable. A maximum matching shows the important links by which we can construct the cactus structures efficiently in a complex system. Therefore, the maximum matching not only reveals the minimum driver set but also consists of a backbone of the key control routes, which are a stem-cycle cover of the original network. The minimum-control-cost configuration CF is just constructed for showing the backbone of the propagation of control influence.
To quantify the control capability of a single vertex i under the minimum-control-cost condition, B reduces to the vector b(i) with a single non-zero entry and A reduces to the matrix A(M). Then, the control capability of a single vertex i can be defined as

Lin's theorem11 has demonstrated that a linear control system (A, B) is structurally controllable if and only if the associated digraph G(A) can be spanned by cacti. A cactus is a subnetwork in the form of a distinct stem or a stem connected to several buds. A stem is simply an elementary path that originates from an input vertex. The initial (or terminal) vertex of a stem is known as the root (or top) of the stem. A bud is an elementary cycle with an additional link that ends, but does not begin, in a vertex of the cycle and the top vertex of the stem is not the initial vertex of any additional link. The network can be spanned by cacti using links of A(M). Thus, A(M) demonstrates the manner in which vertices control the entire network under the minimum-control-cost condition. When the vertex i is taken as an input vertex, the subspace that is accessible from vertex i in the spanning subnetwork CF is cactus-structured and structurally controllable. A vertex j is called accessible if there is at least one directed path that passes from the input vertex i to vertex j. For example, in figure 2(b), the accessible subspace of vertex 1 is highlighted in bold purple and spanned by the links of this CF in the form of a cactus.
We can therefore use the size of the accessible subspace of vertex i as an accurate measure of  . Thus, equation (1) can be represented by
. Thus, equation (1) can be represented by

where

is the set of vertices in the controllable subspace of vertex i.
By invoking the duality between controllability and observability in a linear system, it can be seen that the driver vertices in network G(A) for inputting signals are simply the output vertices for measurement in the transposed network G(AT), which is obtained by inverting the direction of all links. The network G(AT) is guaranteed to be observable by monitoring those output vertices. Thus, all our controllability conditions can be readily extended to the observability case. The link set MT that is obtained by inverting the direction of all links in M forms a maximum matching link set of G(AT). Thus, the minimum-observation-cost configuration (OF) can be defined as  , where AL′ is the set of additional links in G(AT) and AT(MT) can be used to indicate the wiring diagram of OF that corresponds to MT in G(AT). As an example, in figure 2(a), l12,18 is the only additional link in AL′.
, where AL′ is the set of additional links in G(AT) and AT(MT) can be used to indicate the wiring diagram of OF that corresponds to MT in G(AT). As an example, in figure 2(a), l12,18 is the only additional link in AL′.
To quantify the observation capability of a single vertex i under the minimum-observation-cost condition, the output matrix BT reduces to the vector (B(i))T with a single non-zero entry and AT reduces to the matrix AT(MT). Then, the observation capability can be represented by the size of the observable subspace OSi(MT) of vertex i in  and can be accurately measured as follows:
and can be accurately measured as follows:


where

Considering the role that a vertex plays in both controlling the downstream subspace and observing the upstream subspace, the domination centrality (DC) index for the assessment of the capabilities of vertices in directed networks can be synthetically defined as the harmonic mean of a vertex's control capability and observation capability. The domination centrality of vertex i is represented by

The DC index is used to detect the most powerful vertex through which we can not only control but also observe a network. Therefore, as the harmonic mean of the control capability and observation capability of the vertex, DC will be significant only when the control capability and observation capability attain high values simultaneously. In figure 2(b), for the given maximum matching M1, the controllable subspace of vertex 1, CS1(M1), is highlighted by a purple dotted line and has  and the observable subspace,
and the observable subspace,  , is highlighted by a green dotted line and has
, is highlighted by a green dotted line and has  ; thus, the domination centrality of vertex 1 is DC1 = 2/(1/10 + 1/7) ≈ 8.2 and vertex 1 is powerful in dominating the network. By contrast, vertex 14 has the highest value of control capability but a very small observation capability, meaning that DC14 = 2/(1/13 + 1/1) ≈ 1.9. Thus, vertex 1 has a stronger overall ability to dominate the network than does vertex 14. In the worst case, when a vertex i can only control and observe itself, DCi = 1.
; thus, the domination centrality of vertex 1 is DC1 = 2/(1/10 + 1/7) ≈ 8.2 and vertex 1 is powerful in dominating the network. By contrast, vertex 14 has the highest value of control capability but a very small observation capability, meaning that DC14 = 2/(1/13 + 1/1) ≈ 1.9. Thus, vertex 1 has a stronger overall ability to dominate the network than does vertex 14. In the worst case, when a vertex i can only control and observe itself, DCi = 1.
Furthermore, we note that there are multiple different maximum matchings (N! matchings for a complete connected network). Each one illustrates a unique manner in which vertices may control and observe the entire network under a minimum-cost condition. Therefore, in combination with other vertices, a vertex may play several different roles in dominating a network. Thus, we may ask this question: in all possible minimum-cost configurations, if two vertices can perform similar control and observation functions, does that fact indicate that they can also play similar functions in intervening in the system? To answer this question, the domination similarity (DS) is defined as

where agm(x, y) is the arithmetic-geometric mean29 of two positive real numbers x and y. We calculate the Jaccard similarity coefficient of the complete controllable subspaces of i and j to determine their control-function similarity. Meanwhile, the Jaccard similarity coefficients of the complete observable subspaces of i and j are calculated to determine their observation-function similarity. The complete controllable subspace  and the complete observable subspace
and the complete observable subspace  , where K is the number of different maximum matchings. JC(i, j) = Jaccard(CSi, CSj), JO(i, j) = Jaccard(OSi, OSj). agm(x, y) is a number between the geometric and arithmetic means of x and y; thus, DS(i, j) will be significant only when JC(i, j) and JO(i, j) attain high values simultaneously. Furthermore, in the case that there is a large difference between the two quantities, agm (x,y) yields a more reasonable result than the arithmetic or harmonic mean.
, where K is the number of different maximum matchings. JC(i, j) = Jaccard(CSi, CSj), JO(i, j) = Jaccard(OSi, OSj). agm(x, y) is a number between the geometric and arithmetic means of x and y; thus, DS(i, j) will be significant only when JC(i, j) and JO(i, j) attain high values simultaneously. Furthermore, in the case that there is a large difference between the two quantities, agm (x,y) yields a more reasonable result than the arithmetic or harmonic mean.
Figure 2 vividly illustrates this concept. M1 and M2 are two different maximum matchings of this toy network, with links highlighted in red in figure 2(a) and figure 2(c), respectively. We concentrate on the domination capabilities of vertices 1 and 2. In figure 2(b), for vertex 1, the controllable subspace CS1(M1) in CF(A(M1)) is indicated in purple and the observable subspace  in
in  is indicated in green. Similarly, the situation for vertex 2 is illustrated in figure 2(d), with orange and blue colours corresponding to M2. A great deal of information regarding the functions of these two vertices can be determined based on the overlapping of their controllable and observable subspaces, as shown in figure 2(e).
is indicated in green. Similarly, the situation for vertex 2 is illustrated in figure 2(d), with orange and blue colours corresponding to M2. A great deal of information regarding the functions of these two vertices can be determined based on the overlapping of their controllable and observable subspaces, as shown in figure 2(e).
Distribution of domination centrality
If a structural system can be shown to be controllable for almost all weight combinations10 and the dimension of the controllable subspace is stable, in the sense that for almost any set of system parameters, the dimension is equal to some maximal constant (the generic rank of the controllability matrix)19, all these properties also hold for observability14. Thus, to some extent, domination centrality and domination similarity can be calculated without assessing the link weights. This property is one of the greatest advantages of controllability-based topological measures: they are robust to uncertainty in link weights, which frequently arises in networks constructed from real data, such as biological networks.
In this section, we perform statistical studies of the domination centrality on several types of real-world directed networks, including citation, glossary, metabolic and synthetic scale-free networks, as summarised in table 1. We have manually reconstructed the global human enzyme-centric network based on data available in the August 2009 release of the Kyoto Encyclopedia of Genes and Genomes (KEGG)20. The citation and glossary networks are drawn from Pajek datasets and can be downloaded at http://vlado.fmf.uni-lj.si/pub/networks/data/. The synthetic scale-free networks were constructed using the method of Fan et al.21. In table 1, we provide the statistical values of the numbers of vertices (n), links (m) and minimum driver vertices (ND) for the original networks.
We first consider the distribution of the domination centrality. For a given network, any existing algorithm22,23 can be used to compute a maximum matching M. For this M, the domination centrality reveals the responsibility and capability of each individual vertex in controlling and observing the system with the global minimum cost. Figure 3 presents the distribution of the domination centrality for the synthetic scale-free networks listed in table 1. In double-logarithmic coordinates, the relation between the DC value and the probability P(DC) is nearly linear, suggesting the coexistence of a few powerful vertices and a large number of vertices that have little domination over the system's dynamics. However, we also must consider that even the most powerful vertex can dominate only a small local subspace within the entire system and thus, it is preferable to identify multiple collaborating vertices for the domination of the entire system. Therefore, cooperative relations among vertices are of significant concern. This is why we insist on measuring all vertices' capabilities in collaboratively dominating an entire network in a certain minimum-control-cost configuration.
To statistically explain which topological features determine the distribution of the domination centrality itself, we compare the DCs of each vertex in the real networks and their randomised counterparts (denoted as rand-ER and rand-Degree). A full randomisation procedure (rand-ER) turns the network into a fairly homogeneous, directed Erdo˝s-Rényi random network24. The domination centrality values in rand-ER (DCER) and the corresponding number of driver vertices ( ) change dramatically, as shown in table 2. For almost all networks, there is no correlation between DC and DCER, indicating that full randomisation eliminates the topological characteristics that influence domination centrality. We also apply a degree-preserving randomisation (rand-Degree)25, which leaves the in-degree, kin and the out-degree, kout, of each vertex unchanged but randomly selects which vertices link to each other. We find that this procedure does not significantly alter the number of driver vertices (
) change dramatically, as shown in table 2. For almost all networks, there is no correlation between DC and DCER, indicating that full randomisation eliminates the topological characteristics that influence domination centrality. We also apply a degree-preserving randomisation (rand-Degree)25, which leaves the in-degree, kin and the out-degree, kout, of each vertex unchanged but randomly selects which vertices link to each other. We find that this procedure does not significantly alter the number of driver vertices ( ) or the domination centrality (DCDgeree). For example, in figure 4(b, c, d), we present scatter plots of the DC values versus the in-degree, out-degree and degree in the Homo sapiens networks; the results for the real networks (green) are reasonably consistent with that for the rand-Degree counterparts (blue), whereas the results for the rand-ER counterparts (purple) are significantly different from the others.
) or the domination centrality (DCDgeree). For example, in figure 4(b, c, d), we present scatter plots of the DC values versus the in-degree, out-degree and degree in the Homo sapiens networks; the results for the real networks (green) are reasonably consistent with that for the rand-Degree counterparts (blue), whereas the results for the rand-ER counterparts (purple) are significantly different from the others.
A schematic diagram illustrating the domination centrality in Homo sapiens networks.
(a): The average values of domination centrality among low-, medium- and high-degree vertices. The scatter plots of the domination centrality versus the vertex in-degree, out-degree and degree are presented in panel (b), panel (c) and panel (d), respectively. The green, blue and purple plots represent the real network, rand-Degree network and rand-ER network, respectively.
In addition, we calculate the mean, the average of absolute deviation26 and the relative entropy27 for the distribution of the domination centrality in each real network and their random counterparts in table 3. Compared to the real networks, the rand-Degree counterparts yield similar mean values, similar averages of absolute deviation and small relative entropies. The same indices of the rand-ER counterparts differ significantly in comparison. From all these observations, we conclude that domination centrality is, to a great extent, encoded by the degree distribution of the underlying network.
Another interesting phenomenon observed in this study is that the hubs (vertices of high degree) do not tend to play more important roles in dominating a system. We divide the vertices into three groups of equal size according to their degree k (low, medium and high) and calculate the average values of DC among the low-degree, medium-degree and high-degree vertices. As table 2 demonstrates, for real networks and two random network models (Erdo˝s–Rényi24 and scale-free21,22,23,24,25), the average DC value of the set of low-degree vertices is not significantly lower than that of the set of hubs in each case. Figure 4(a) graphically represents the values for the Homo sapiens networks. In figure 4(b, c, d), as expected, in all cases, a low-degree vertex can also have a significant domination centrality. For a vertex with a degree equal to 1, either the control capability or the observation capability must also be equal to 1; thus, as the harmonic mean of these two capabilities, the domination centrality must be less than 2. Intuitively, a vertex with a degree of 1 must have either no downstream space it can control or no upstream space it can observe. This is the reason why the hubs are observed to attain slightly larger DC values than the low-degree vertices. To conclude, this experimental study demonstrates that there is no obvious correlation between the degree and the DC. This result is very useful in the following sense: the most effective method by which we can intervene in a system's dynamics is to identify vertices with great domination capability, which are not restricted to hubs alone.
Clustering analysis
In fact, a consensus among the topological criteria for measuring the functional similarity of vertices is often lacking in directed networks28. Ignoring the direction of the links may lead to partial or even misleading clustering results. Domination similarity is a direction-specific index and concentrates on quantifying the unique relation between the upstream and downstream subspaces of vertices in directed networks. Independent of the weights of the links used in the calculation, the domination similarity is a parameter-free index for analysing data with noise. With the global minimum-cost limitation, the domination similarity represents the ability of vertices to work synergistically with others and provides guidance for dominating a system using multiple vertices operating cooperatively at the minimum cost. In this section, we apply the DS index to detect and analyse functional modules in a glossary network and the enzyme-centric network of Homo sapiens.
We utilise the DS values as the input of the AP algorithm30 to identify the functional modules in the glossary network. In this case, we test the performance on a directed word network that has also been recently introduced by Newman31 and Boccaletti32. The network represents the connections among a set of technical terms, such as “Tree” and “Digraph”, contained in a glossary of network jargon. Vertices represent terms and a directed link from one vertex to another exists in the network iff the second term is used to describe the meaning of the first term. Because circular definitions are unhelpful and are normally avoided, most links in the network are not reciprocal. The statistics for this network are provided in table 1.
Figure 5 shows the modules identified in this network using our DS-based method. This method identifies nine modules in this case, which appear to correspond to the meaningful groups in understanding the relations among glossary terms. For instance, module 1, which is highlighted in red in the figures, deals with words that describe tree structure. Remarkably, as an upstream vertex, the term “Decision Tree” can be explained by its downstream terms in module 1 and these downstream vertices constitute the controllable subspace of the vertex “Decision Tree”. As a downstream vertex, the term “Tree” is the basic foundation for the formation of other upstream terms and these upstream vertices constitute the observable subspace of the vertex “Tree”. Module 9 contains the glossary terms derived from the fundamental term “Digraph” and provides an overview of the dominance of this term. Additionally, all other detected modules represent not only groups of terms with similar meanings but also the etymology of the network jargon. Thus, the DS-based method appears to identify meaningful structure in the network, of a type that could be useful in understanding the broader shapes of otherwise poorly understood systems.
We return now to the global human metabolic (enzyme-centric) network. The vertices in this network represent enzymes and there is a directed link from one enzyme to another if the product of a reaction catalysed by the first enzyme is used as the substrate of a reaction catalysed by the second. The statistics for the human enzyme-centric network are provided in table 1. There are 689 enzymes and 2382 directed links derived from 90 metabolic pathways. Metabolism is a vital cellular process and its malfunction is a major contributor to human disease33. Metabolic networks are complex and thus, systems-level computational approaches are required to elucidate and understand them. Here, we wish to discuss the interventional effect of enzymes in the metabolic system and detect the relevant modules.
We cluster the network into modules using the AP algorithm augmented by our DS index to identify the pathways that correspond to metabolic functions. In total, 63 modules are detected by our method. We measure the biological quality of the clustering result by means of Gene Ontology (GO) enrichment34 and use the tool GO TermFinder35 to compute the functional enrichment p-values of components with respect to their biological process annotations. In the results, 28 modules are annotated by GO terms with p-values ≤ 0.01 (the most significant p-value = 3.45E-16), which means that these modules represent significant biological functions in the metabolic system. In figure 6, certain representative modules are depicted with their corresponding GO terms and p-values. We note that not only dense subnetworks but also functional modules, with distinctive circle and path structures, are detected. These experiments provide compelling evidence that the DS is a meaningful and practical indicator in accurately depicting the structures of directed networks.
Discussion
The domination capability of a vertex reflects the vertex's ability to interfere in dynamical control processes in many directed complex systems. The key task is to explain the manner in which a vertex intervenes in its downstream and upstream spaces in the implementation of dynamical functions. In this paper, based on the determination of controllable and observable subspaces under the minimum-cost condition, we introduced the DC index to assess the capabilities of vertices in directed networks. The results of our statistical studies demonstrate that the domination centrality is, to a great extent, encoded by the degree distribution of the underlying network, yet there is no discernible correlation between the degree and DC of a vertex. This result provides guidelines for the selection of the most effective means through which we can intervene in a system's dynamics. Furthermore, to analyse the cooperative relations among vertices in the domination of an entire network, we defined the domination similarity and we were able to detect significant functional modules in glossary and metabolic networks through clustering. As direction-specific and parameter-free indexes, DC and DS are effective and practical in accurately depicting the structures of directed networks. In our future studies, we intend to investigate the most effective approach to intervening in the dynamical functions of complex systems through selected vertices.
Methods
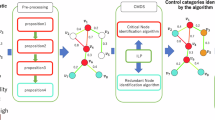
Enumerating all possible different maximum matching link sets Ms is infeasible when calculating DS, as in the worst case scenario, there may be an exponential number of them. However, we note that there are many “redundant” links in real networks that may never appear in any maximum matching. Based on their role in the Ms, links can be classified into three categories: “critical” links must appear in all Ms, “redundant” links may never appear in any one of them and “ordinary” links play roles in some, but not all, Ms10. In combination with the sparseness of real networks, we can approximate the control subspaces and observation subspaces of vertices via an optimisation routine. As shown in figure 7, we observe the beneficial phenomenon that in a real network, there always exists some consistent set of control subspaces and observation subspaces of vertices induced by different maximum matchings. This observation supports the feasibility of using a small number of maximum matchings to approximate the complete control and observation subspaces of vertices. A random optimisation can be performed rather quickly using a Markov sampling process.
The influence of the number of random samples among all maximum matchings on the domination-similarity result.
The growth rates of the sum of the complete control and observation subspaces as functions of the number of random samples of maximum matchings in the GlossTG, Homo sapiens and SmaGri networks are shown.
Algorithm for DS
Input network G(A)
θ = 1, τ = 1, t = 0
do
Markov random sampling to produce a maximum matching M




if |θ − τ|/τ ≤ ε t = t + 1 else t = 0
τ = θ
Whilet ≤ ψ
Calculate Domination Similarity
In every loop, we randomly produce a maximum matching M and update the complete control subspace CS and complete observation subspace OS of each vertex by merging the additional accessible vertices introduced in this M. CS s and OS s are added but never deleted throughout the entire procedure. θ is the sum of all CS s and OS s. If the rate of increase of θ is less than ε for ψ continuous loops, the random optimisation procedure terminates and we then calculate the Jaccard similarity coefficients of the CS s and OS s of two arbitrary vertices. The growth rates of θ during the random optimisation procedures for the GlossTG, Homo sapiens and SmaGri networks are presented in figure 7. Clear improvement in θ is achieved for 90, 7845 and 4527 maximum matchings in 247, 27995 and 15172 random samples for GlossTG, Homo sapiens and SmaGri, respectively. We note that the growth rate of θ rapidly decreases to nearly 0 as the sampling number increases. This observation supports the appropriateness of using only a certain number of Ms to approximate the domination similarities of vertices in real networks. We set ε = 0.000001 and ψ = 50 in the clustering-analysis case studies.
A Markov process, as described by Jia et al.36,37, performs unbiased random sampling among all maximum matchings and can be used to estimate the role of each vertex in controlling the network. This algorithm randomly chooses a vertex in a given M, enumerates all alternative maximum matchings that include all other elements except this vertex by removing all its links, then randomly chooses one of these alternative maximum matchings as the current M and repeats the process.
However, removing a vertex in a random sampling may not be effective for calculating DS. Usually, we identify a maximum matching in a bipartite graph and attempt to increase the matching size via an augmenting path that begins at a matched vertex, ends at an unmatched vertex and alternates between unmatched and matched links on the path. For example, in figure 8, a bipartite graph that is separated into the out and in sets is constructed in figure 8(b) for the network in figure 8(a). The red links are matched, the black dotted links are unmatched and the matched link set {l1,2, l2,3, l4,5} forms a maximum matching M1. Proceeding from this maximum matching, we randomly choose vertex 2 and leave the current matched vertices and links unchanged. Instead of removing all links of vertex 2, we delete only the matched link l1,2. Then, we can identify an augmenting path that begins at the relevant matched vertex 1 and ends at the presently unmatched vertex 2; in the figure, this path is indicated by a blue line. Finally, by alternating between unmatched and matched links on this blue path, we obtain a new maximum matching M2 in which the matching of vertex 2 has been replaced, as shown in figure 8(c). By contrast, because it removes all links of vertex 2, the method of Jia et al.37 cannot produce any new maximum matching from the given M1. Nevertheless, we use the Markov process defined by these authors to perform unbiased random sampling among all maximum matchings to estimate DS. The only difference is that we also enumerate all alternative maximum matchings that include all other elements except the matching link of the chosen vertex.
A schematic diagram illustrating the process of random sampling.
(a): The original network. (b): A bipartite graph separated into the out and in sets; the red link set {l1,2,l2,3,l4,5} forms a maximum matching, M1 and the blue path is an augmenting path when the matched link l1,2 is removed. (c): A new maximum matching M2 constructed by alternating the blue augmenting path.
In fact, the maximum matchings reveal the functions and the roles that the vertices and links play for controlling the whole network with minimum cost. Different combinations of ordinary links constitute different maximum matchings and produce different choices of minimum-control-cost configurations. Ordinary links are alternatives for constructing the backbone of the propagation of control influence. Therefore, we consider each combination of ordinary links (COL) to be one state. The set of ordinary links of a network is {l1,l2,…,lv}, where v is the number of ordinary links. The Markov chain can be characterised by a transition matrix P with the elements  , where Qi is the probability of ordinary link li being included in an M. The transition from state i to state j requires the choice of a matched link from an M, with a probability of
, where Qi is the probability of ordinary link li being included in an M. The transition from state i to state j requires the choice of a matched link from an M, with a probability of  ; the choice of a COL set that excludes li, with a probability of (1 − Qi); and the choice of a COL set that includes lj, with a probability of Qj. Clearly, Pi,j ≠ Pj,i; our algorithm is not guaranteed to choose each set of matched links with equal probability. For example, in figure 9(a), for the real network GlossTG with 164 COLs, we perform 239,000 iterations of our sampling algorithm and count the number of times that each COL is picked. We find that a few COLs are sampled many times, but it is very difficult to ensure that all COLs are sampled at least once. Thus, we adjust the transition matrix P and construct a new transition matrix P′ with the elements
; the choice of a COL set that excludes li, with a probability of (1 − Qi); and the choice of a COL set that includes lj, with a probability of Qj. Clearly, Pi,j ≠ Pj,i; our algorithm is not guaranteed to choose each set of matched links with equal probability. For example, in figure 9(a), for the real network GlossTG with 164 COLs, we perform 239,000 iterations of our sampling algorithm and count the number of times that each COL is picked. We find that a few COLs are sampled many times, but it is very difficult to ensure that all COLs are sampled at least once. Thus, we adjust the transition matrix P and construct a new transition matrix P′ with the elements  . If we can set
. If we can set  , then Pi,j ≡ Pj,i, meaning that the transition matrix P is symmetric and the steady-state distribution possesses equal probabilities for all states. However, Qj cannot be determined effectively; in practice,
, then Pi,j ≡ Pj,i, meaning that the transition matrix P is symmetric and the steady-state distribution possesses equal probabilities for all states. However, Qj cannot be determined effectively; in practice,  is used to approximate (1 − Qj), where uj is the average number of all alternative COLs that can be enumerated by removing lj in the first |M|2 iterations of our sampling procedure. Intuitively, if many alternative COLs can be enumerated by removing lj, then the probability of choosing lj from an M should be increased. With this modification, the sampling procedure becomes more efficient and the 164 COLs in GlossTG can be obtained within 20,000 iterations. As shown in figure 9(b), we perform our modified sampling algorithm 193,500 iterations and count the number of times that each COL is picked. The result demonstrates that this procedure provides a more even-handed random sampling among all maximum matchings.
is used to approximate (1 − Qj), where uj is the average number of all alternative COLs that can be enumerated by removing lj in the first |M|2 iterations of our sampling procedure. Intuitively, if many alternative COLs can be enumerated by removing lj, then the probability of choosing lj from an M should be increased. With this modification, the sampling procedure becomes more efficient and the 164 COLs in GlossTG can be obtained within 20,000 iterations. As shown in figure 9(b), we perform our modified sampling algorithm 193,500 iterations and count the number of times that each COL is picked. The result demonstrates that this procedure provides a more even-handed random sampling among all maximum matchings.
References
Newman, M. Networks: an introduction. (Oxford University Press, Oxford, 2009).
Freeman, L. C. A set of measures of centrality based on betweenness. Sociometry 40, 35–41 (1977).
Barthelemy, M. Betweenness centrality in large complex networks. Eur. Phys. J. B 38, 163–168 (2004).
Sabidussi, G. The centrality index of a graph. Psychometrika 31, 581–603 (1966).
Bonacich, P. & Lloyd, P. Eigenvector-like measures of centrality for asymmetric relations. Social Networks 23, 191–201 (2001).
Estrada, E. & Rodríguez-Velázquez, J. A. Subgraph centrality in complex networks. Phys. Rev. E 71, 056103 (2005).
Brin, S. & Page, L. The anatomy of a large-scale hypertextual Web search engine. Comput. networks and ISDN syst. 30, 107–117 (1998).
Delvenne, J.-C. & Libert, A.-S. Centrality measures and thermodynamic formalism for complex networks. Phys. Rev. E 83, 046117 (2011).
Fortunato, S. & Flammini, A. Random walks on directed networks: the case of PageRank. Int. J. Bifurcation Chaos 17, 2343–2353 (2007).
Liu, Y.-Y., Slotine, J.-J. & Barabási, A.-L. Controllability of complex networks. Nature 473, 167–173 (2011).
Lin, C.-T. Structural controllability. IEEE T. Automat. Contr. 19, 201–208 (1974).
Luenberger, D. Introduction to dynamic systems: theory, models and applications. (John Wiley & Sons, New York, 1979).
Hermann, R. & Krener, A. Nonlinear controllability and observability. IEEE T. Automat. Contr. 22, 728–740 (1977).
Liu, Y.-Y., Slotine, J.-J. & Barabási, A.-L. Observability of complex systems. Proc. Natl Acad. Sci. USA 110, 2460–2465 (2013).
Liu, Y.-Y., Slotine, J.-J. & Barabási, A.-L. Control centrality and hierarchical structure in complex networks. Plos one 7, e44459 (2012).
Wang, B., Gao, L. & Gao, Y. Control range: a controllability-based index for node significance in directed networks. J. Stat. Mech-Theory E. 2012, P04011 (2012).
Kalman, R. E. Mathematical description of linear dynamical systems. SIAM, Ser. A: Contr. 1, 152–192 (1963).
Poljak, S. On the generic dimension of controllable subspaces. IEEE T. Automat. Contr. 35, 367–369 (1990).
Hosoe, S. Determination of generic dimensions of controllable subspaces and its application. IEEE T. Automat. Contr. 25, 1192–1196 (1980).
Kanehisa, M. et al. From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Res. 34, D354–D357 (2006).
Chung, F. & Lu, L. Connected components in random graphs with given expected degree sequences. Ann. Comb. 6, 125–145 (2002).
Kuhn, H. W. The Hungarian method for the assignment problem. Nav. Res. Log. 2, 83–97 (1955).
Hopcroft, J. E. & Karp, R. M. An n∧5/2 algorithm for maximum matchings in bipartite graphs. SIAM J. on computing 2, 225–231 (1973).
Bollobás, B. Random graphs. (Cambridge university press, Cambridge, 2001).
Barabási, A.-L. & Albert, R. Emergence of scaling in random networks. Science 286, 509–512 (1999).
Ruppert, D. Statistics and data analysis for financial engineering. (Springer, New York, 2011).
Kullback, S. & Leibler, R. A. On information and sufficiency. Ann. Math. Stat. 22, 79–86 (1951).
Lancichinetti, A. & Fortunato, S. Community detection algorithms: a comparative analysis. Phys. Rev. E 80, 056117 (2009).
Borwein, J. M. & Borwein, P. B. The arithmetic-geometric mean and fast computation of elementary functions. SIAM rev. 26, 351–366 (1984).
Frey, B. J. & Dueck, D. Clustering by passing messages between data points. Science 315, 972–976 (2007).
Newman, M. E. The structure and function of complex networks. SIAM rev. 45, 167–256 (2003).
Boccaletti, S., Latora, V., Moreno, Y., Chavez, M. & Hwang, D.-U. Complex networks: Structure and dynamics. Phys. Rep. 424, 175–308 (2006).
Duarte, N. C. et al. Global reconstruction of the human metabolic network based on genomic and bibliomic data. Proc. Natl Acad. Sci. USA 104, 1777–1782 (2007).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Boyle, E. I. et al. GO:: TermFinder—open source software for accessing Gene Ontology information and finding significantly enriched Gene Ontology terms associated with a list of genes. Bioinformatics 20, 3710–3715 (2004).
Jia, T. et al. Emergence of bimodality in controlling complex networks. Nat. Commun. 4, 2002 (2013).
Jia, T. & Barabási, A.-L. Control Capacity and A Random Sampling Method in Exploring Controllability of Complex Networks. Sci. Rep. 3, 2354 (2013).
Wang, B., Gao, L., Gao, Y. et al. Maintain the structural controllability under malicious attacks on directed networks. Europhys. Letts. 101, 58003 (2013).
Yuan, Z., Zhao, C., Di, Z., Wang, W. X. & Lai, Y. C. Exact controllability of complex networks. Nat. Commun. 4, 2447 (2013).
Cornelius, S. P., Kath, W. L. & Motter, A. E. Realistic control of network dynamics. Nat. Commun. 4, 1942 (2013).
Sun, J. & Motter, A. E. Controllability transition and nonlocality in network control. Phys. Rev. Lett. 110, 208701 (2013).
Ruths, J. & Ruths, D. Control Profiles of Complex Networks. Science 343, 1373–1376 (2014).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant No. 91130006, 61303122, 61100157 & 61303118) and the Fundamental Research Funds for the Central Universities (Grant No. BDZ021404).
Author information
Authors and Affiliations
Contributions
B.-B.W. designed and performed experiments, analyzed data and wrote the paper; L.G. and Y.G. designed experiments and wrote the paper; Y.D. and Y.W. designed experiments and wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Wang, B., Gao, L., Gao, Y. et al. Controllability and observability analysis for vertex domination centrality in directed networks. Sci Rep 4, 5399 (2014). https://doi.org/10.1038/srep05399
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep05399
This article is cited by
-
Controllability and Its Applications to Biological Networks
Journal of Computer Science and Technology (2019)
-
A symbolic network-based nonlinear theory for dynamical systems observability
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.





 and
and  are shown.
are shown.




