Abstract
The role of [-2]proPSA (p2PSA) based diagnostic tests for the detection of aggressive prostate cancer (PCa) has not been fully evaluated. We conducted a meta-analysis to evaluate the diagnostic performance of p2PSA/free PSA (%p2PSA) and prostate health index (Phi) tests for PCa and to evaluate their ability in discriminating between aggressive and non-aggressive PCa. A total of 16 articles were included in this meta-analysis. For the detection of PCa, the pooled sensitivity, specificity and AUC were 0.86 (95% CI, 0.84–0.87), 0.40 (95% CI, 0.39–042) and 0.72 (95% CI, 0.67–0.77) for %p2PSA respectively and were 0.85 (95% CI, 0.83–0.86), 0.45 (95% CI, 0.44–0.47) and 0.70 (95% CI = 0.65–0.74) for Phi, respectively. In addition, the sensitivity for discriminating PCa between higher Gleason score (≥7) and lower Gleason score (<7) was 0.96 (95% CI, 0.93–0.98) and 0.90 (95% CI, 0.87–0.92) for %p2PSA and Phi respectively and the specificity was low, only 0.09 (95% CI, 0.06–0.12) and 0.17 (95% CI, 0.14–0.19) for %p2PSA and Phi, respectively. Phi and %p2PSA have a high diagnostic accuracy rates and can be used in PCa diagnosis. Phi and %p2PSA may be useful as tumor markers in predicating patients harboring more aggressive disease and guiding biopsy decisions.
Similar content being viewed by others
Introduction
Prostate Cancer (PCa) is one of the most commonly diagnosed malignant genitourinary tumors among men in United States and other Western countries1. To date, prostate-specific antigen (PSA) is the most widely used and successful serum marker to detect PCa, however, the clinical value of PSA for early detection of PCa has been questioned because of its poor specificity, especially in men with serum PSA levels in the 2–10 ng/ml range2. It is well-known that PSA testing leads to overdiagnosis and overtreatment of indolent PCa that does not evolve into aggressive life-threatening cancers3. Thus, significant effort has been placed on the development of new serum markers that could complement PSA for early aggressive PCa diagnosis.
It has been suggested that measurement of the precursor PSA isoform [−2] proPSA (p2PSA) and its derivatives might offer improvement of PCa detection in men with a total PSA (tPSA) level between 2 and 10 ng/ml4. The isoform p2PSA is a promising marker for PCa detection and compared to other isoforms, measurement of the p2PSA to free PSA (%p2PSA) and Prostate Health Index (Phi), which is calculated using the formula (p2PSA/fPSA × √tPSA) can better distinguish between PCa and non-PCa than tPSA and fPSA/tPSA (%fPSA)5,6,7. Values for %p2PSA and Phi were significantly higher in patients with PCa than patients with benign prostatic hyperplasia (BPH) and chronic histologic prostatic inflammation8,9.
Previous studies have also shown that %p2PSA and Phi were significantly associated with an increased probability of detecting aggressive PCa, as indicated by a Gleason score ≥75,8. Both %p2PSA and Phi gained attention as potentially very promising markers, that is, to be able to help predict aggressive PCa within a tPSA range of 2–10 ng/ml10,11,12. But the ability of these two markers to distinguish between aggressive PCa and non-aggressive PCa remains unclear. Here, a systematic review was performed to evaluate the emerging role of %p2PSA and Phi in diagnosing PCa and the ability of these markers to distinguish patients with and without aggressive PCa.
Results
Description of included studies
As shown in Figure 1, we identified 323 papers from a literature search and excluded 299 irrelevant articles. The remaining 24 potentially relevant studies were further assessed. Eight studies were further excluded because they contained duplicated populations6,8,13,14,15,16,17,18. Although the study by Lazzeri et al.6 was excluded due to partially duplicated data with the study of Lughezzani et al.19, the data on aggressive and non-aggressive PCa were available and extracted.
A total of 16 papers were included in the meta-analysis5,7,10,11,19,20,21,22,23,24,25,26,27,28,29,30, 13 papers reported on %P2PSA and 12 papers reported on Phi. In addition, the study by Jansen7 included two different populations which were treated as two data sets and the study by Scattoni20 contained initial and repeat biopsy groups which were treated as two separate studies. Finally, 14 studies were reported on %P2PSA and Phi. The detailed characteristics of each study are presented in Table 1. All studies presented the results of sensitivity and specificity and most studies presented the results of ROC curve. The cutoff points were reported in seven studies for %p2PSA and nine studies for Phi. The data on the patients with and without aggressive PCa were extracted from four and five studies for %p2PSA and Phi, respectively.
Overall meta-analyses for %p2PSA and Phi
A total of 6,279 patients (2,722 PCa and 3,557 non-PCa) and 5,856 patients (2,474 PCa and 3,382 non-PCa) were included in the pooled analyses for %p2PSA (Supplementary table 1) and Phi (Supplementary table 2), respectively. The pooled sensitivity and specificity were 0.86 (95% CI, 0.84–0.87) and 0.40 (95% CI, 0.39–042) for %p2PSA respectively and were 0.85 (95% CI, 0.83–0.86) and 0.45 (95% CI, 0.44–0.47) for Phi, respectively (Figure 2).
The ROC space showed a curvilinear trend of points and Spearman's correlation coefficient was 0.705 (P = 0.005) for %p2PSA and 0.744 (P = 0.002) for Phi. It was suggested that there was the existence of a threshold. Due to the presence of significant heterogeneity for sensitivity (I2 = 90.4%, χ2 = 135.15, P < 0.001), specificity (I2 = 96.1%, χ2 = 329.39, P < 0.001) for %p2PSA; and sensitivity (I2 = 91.5%, χ2 = 152.79, P < 0.001), specificity (I2 = 93.1%, χ2 = 188.66, P < 0.001) and diagnostic OR (Cochran-Q = 24.25, P = 0.0289) for Phi; the diagnostic indices were calculated using a random effect model. We used a summary SROC curve to aggregate data and obtained a symmetrical curve with an AUC of 0.72 (95% CI = 0.67–0.77) and 0.70 (95% CI = 0.65–0.74) for %p2PSA and Phi, respectively (Supplementary Figure 1).
Subgroup analysis
Significant heterogeneity for Phi (I2 = 46.4%, P = 0.0289) and low heterogeneity (I2 = 29.1%, p = 0.1448) were observed. Therefore, further subgroup analyses were performed to explore the heterogeneity on data stratified by ethnicity group, study design and QUADAS score. Subgroup analyses showed that a retrospective study design (I2 = 61.6%, p = 0.0231) and low QUADAS score (I2 = 63.7%, p = 0.0171) were responsible for the heterogeneity of %p2PSA (Table 2); and the prospective study design (I2 = 51.3%, p = 0.0368), Caucasian group (I2 = 52.5%, p = 0.0168) and low QUADAS score (I2 = 55.9%, p = 0.0201) were responsible for the heterogeneity of Phi (Table 3).
The ability of %p2PSA and Phi to discriminate between aggressive and non-aggressive PCa
A Total of 728 cases of PCa (292 with Gleason score ≥ 7 versus 436 with Gleason score < 7) and 1,157 cases of PCa (431 with Gleason score ≥ 7 versus 726 with Gleason score < 7) were included in the pooled analysis of %p2PSA (Supplementary table 3) and Phi (Supplementary table 4).
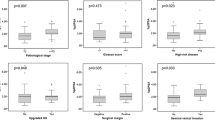
The pooled sensitivity for the detection of PCa with higher Gleason score (≥7) was 0.96 (95% CI, 0.93–0.98) and 0.90 (95% CI, 0.87–0.92), while the specificity was low, only 0.09 (95% CI, 0.06–0.12) and 0.17 (95% CI, 0.14–0.19) for %p2PSA and Phi respectively (Figure 3). The pooled AUC of SROC was 0.54 (95% CI,−0.52–1.61) and 0.67 (95% CI, 0.57–0.77) for %p2PSA and Phi respectively (Supplementary Figure 1). The accuracy of %p2PSA and Phi in discriminating aggressive PCa with non-aggressive PCa is summarized in Table 4.
Assessment of publication bias
We analyzed possible publication bias by generating funnel plots and the shape of the funnel plots was symmetrical in the diagnosis of PCa suggesting the absence of publication bias. We also used the Deeks' funnel plot asymmetry tests (t = 1.86, P = 0.09 for %p2PSA and t = 1.37, P = 0.20 for Phi), results suggested that there was no publication bias in the meta-analysis (Figure 4).
Discussion
Despite the wide and successful use of PSA blood testing, its use as a serum marker for PCa is far from ideal, due to the lack of specificity; it has only 20% specificity31. This review synthesized the current knowledge about early diagnosis of PCa with %p2PSA and Phi determination. The present meta-analysis indicated that %p2PSA and Phi had a high cancer detection rate. Most importantly, %p2PSA and Phi may be useful as tumor markers for high risk PCa detection.
PSA cannot accurately distinguish between benign and malignant pathology in the tPSA range of 2–10 ng/ml. In addition, more indolent PCas were detected, which will not develop into clinical significant PCas32. New markers that can accurately detect as well as differentiate patients with aggressive and non-aggressive PCa are needed. According to preliminary investigations and observational studies, %p2PSA and Phi may improve discrimination between men with and without PCa and they are associated with aggressiveness of PCa, as indicated by Gleason score6,10,11,12. But some studies did not demonstrate a relationship between Gleason score and p2PSA level7. So we carried out a meta-analysis to further confirm the ability of %p2PSA and Phi in PCa patient's diagnosis and derive a more precise estimation of their ability to predicate aggressive PCa.
To date, only one systematic review has been conducted to assess the ability of %p2PSA and Phi to diagnose PCa. Both %p2PSA and Phi improved the accuracy of PCa detection compared to PSA and %fPSA, particularly with the PSA range of 2–10 ng/ml33. However, only the papers published before December 2011 were included and subgroup analysis or meta-regression were not performed to explore the sources of heterogeneity. In addition, the diagnostic accuracy of %p2PSA and Phi for differentiating aggressive PCa from non-aggressive PCa has not yet been summarized. In our meta-analysis, articles published in 2012, 2013 and 2014 were also included. To the best of our knowledge, this was the first attempt to synthesize existing literature to evaluate the diagnostic accuracy of %p2PSA and Phi for aggressive PCa.
Continued emergence of promising data indicate that %p2PSA and Phi are better parameters than tPSA and fPSA for the detection of PCa and aggressive PCa27. Based on synthesis of the results in our meta-analysis, the diagnostic test may be particularly useful in patients with diagnostically challenging PSA scores ranging 2–10 ng/ml, as sensitivity reaches 85%–86% and specificity reaches 40%–45% for %p2PSA and Phi respectively. As shown previously, the PSA test had varying sensitivity and poor specificity, only about 20% for specificity31. Our results showed that %p2PSA and Phi test have potential to complement PSA screening due to their high specificity. We also observed that %p2PSA and Phi have nearly the same effect in diagnosing PCa. Both %p2PSA and Phi showed a high accuracy for detecting PCa (AUC of 0.72 and 0.70 respectively). SROC of our systematic review showed a curvilinear relationship between sensitivity and 1-specificity and the Spearman's correlation coefficient was 0.705 (P = 0.005) and 0.744 (P = 0.002) for %p2PSA and Phi respectively, which may be due to a threshold effect.
The detection of aggressive PCa is the main concern of current discussions on the usefulness of PCa detection markers. Recent studies have shown that higher %p2PSA and Phi values were associated with increased probability of detecting Gleason score ≥ 7 PCa and were higher in aggressive cancers, so %p2PSA and Phi may be able to predict more aggressive PCa on prostatic biopsies5. Phi provided the best discrimination between aggressive and non-aggressive PCa than other markers5. According to our pooled results, the sensitivity for discriminating between aggressive and non-aggressive PCa was very high, 96% and 90% for %p2PSA and Phi respectively, which would miss a small number of aggressive PCa, but the specificity was relatively low, only 9% and 17% for %p2PSA and Phi respectively. The AUC for Phi (0.67) exceeded that of the %p2PSA (0.54) in discriminating aggressive versus non-aggressive PCa. Increased Phi levels and %p2PSA might be helpful in identifying patients harboring more aggressive forms of the disease. Some cutoffs were used for %p2PSA and Phi, at a threshold of 1.2 and 25.5 for %p2PSA and Phi respectively, about 4% of aggressive cancers would have been missed8. And these cutoffs did not allow significant improvement in differentiation between Gleason score 4 + 3 = 7 and 3 + 4 = 7 cancers5. Phi values did not differ with age and race11,24, this suggests that Phi may be more prosperous to be used in a wider scope of all men, irrespective of race, age and background in predicting aggressive PCa. A limited number of studies were included, therefore it was difficult to draw a definitive conclusion for their ability to discriminate. Hence, further research should aim at determining the potential aggressiveness of PCa in patients with high %p2PSA and Phi values.
When interpreting the results, the heterogeneity caused by the different optimum cutoff points determining whether a test is positive or negative must be taken into consideration, since there is actually no consensus regarding the most appropriate level, in part, due to differences in study design and methodology. The cut-off points were only reported in 7 and 9 studies for %p2PSA and Phi respectively and the cut-off values were quite different for all the studies. Increased Phi values were associated with increasing PCa detection rate, when the phi values were <25, 25–34.9, 35–54.9 and ≥55, the PCa detection rates were 11%,18.1%, 32.7% and 52.1% respectively11. The full clinical potential and the threshold of %p2PSA and Phi for diagnosing PCa and identification of aggressive disease need to be determined. If the maximum number of aggressive PCas were to be detected, it might be acceptable to not detect some of the less aggressive, clinically insignificant PCas.
The present meta-analysis has a number of limitations that must be taken into account. First, the preanalytic and analytic management of blood samples were different in the studies such as the case when archived serum was used in some studies and different calibrations were used in the literatures, including Hybritech and WHO standardization. PSA and fPSA were measured using Hybritech Tandem assays except for one study. Second, the main bias of all the studies was that the inclusion criteria was based on the risk of having PCa, the final decision on whether to biopsy or not was based on elevated PSA values and not based on the %p2PSA or Phi, which would cause selection bias. Third, only one study was a randomized study, differences in study designs including prospective, retrospective and mixed, population characteristics, cutoff points and the number of cores obtained in prostate biopsies (rang, 6–24) may contribute to the heterogeneity. Fourth, another limitation was potential publication bias, since yet unpublished studies and non-English studies were excluded, however, we did not confirm the presence of publication bias. We should also note that the number of studies included for analysis for the purpose of discriminating aggressive and non-aggressive PCa was relatively low. Due to limited literatures, subgroup-analyses and meta-regression were not performed. Finally, the best strategy to correlate %p2PSA and phi with aggressiveness is to perform a correlation between preoperative %p2PSA, phi and whole gland sample after radical prostatectomy; unfortunately only the paper by Guazzoni et al12 focused on this topic. Due to the pathological difference between prostate biopsy and sample after radical prostatectomy, we did not include this study into our meta-analysis.
Due to the limitations mentioned above, the results of this meta-analysis should be interpreted with care. Nevertheless, our study was conducted at an appropriate time, because enough data was available to use meta-analytical methods. And we provided the most up-to-date information on this topic. However, it is necessary to conduct the large multi-center randomized-controlled studies or prospective trials with similar methodology to evaluate the ability of %p2PSA and Phi in predicting aggressive PCa in the future. In addition, similar comparison outcomes should be based on the same preanalytic and analytic management for blood samples, same calibration technology (Hybritech or WHO), same TRUS prostate biopsy techniques and biopsy cores or based on the pathology of prostatectomy specimen.
In conclusion, our meta-analysis suggested that both %p2PSA and Phi have high diagnostic accuracy rates and can be used in early PCa diagnosis. In addition, %p2PSA and Phi might be potential biomarkers in predicating aggressive PCa. In our opinion, if the data will be confirmed by further larger sample size studies, the two biomarkers might be helpful in guiding biopsy decisions.
Methods
Literature search and selection criteria
Our meta-analysis was performed following the guidelines concerning Preferred Reporting Items for Systematic reviews and Meta-Analysis (PRISMA) statement34 (Supplementary Table 5). A systematic literature search on PubMed was conducted for articles published from January 2000 to March 2014. The strategy was to use and combine the following terms included in their titles, abstracts, or keywords lists: “prostatic neoplasm”, “prostate cancer”, “p2PSA”, “[−2]proPSA”, “[−2]proenzyme prostate specific antigen”, “Prostate Health Index”, “Phi”, “diagnosis”, “sensitivity“ and “specificity”. Additionally, we reviewed the reference list of each relevant paper by hand. Two researchers (W Wang and M Wang) independently reviewed all identified titles, abstracts and manuscripts to determine if a study was suitable for the meta-analysis.
Studies that met the following criteria were included in the meta-analysis: (1) case-control or cohort design; (2) diagnostic test about %p2PSA or Phi for PCa; (3) histological results were based on prostate biopsy; (4) data on sensitivity and specificity could be extracted. Non-English language papers or studies with only abstracts available for information extraction were excluded. When more than one article was published based on the same patient population or same project, only the most recent or complete report was used, to avoid overlapping between cohorts.
Data extraction and quality assessment
We collected information from each eligible study and entered the data into a structured database including: last name of the first author, year of publication, country of origin, study design, ethnicity, age range of participants, the tPSA and fPSA range, sample size, indication for biopsy, type of biopsy used, the number of biopsy-confirmed PCa and the values of true positive (TP), false-positive (FP), false negative (FN), true negative (TN), summary receiver operating curve (SROC) and cut-off points if available. Of these studies, Gleason score ≥7 was defined as high Gleason score and aggressive and Gleason score <7 was defined as low Gleason score and non-aggressive. The study quality was assessed with Quality Assessment of Diagnostic Accuracy Studies (QUADAS)35,36. Two authors extracted the data and undertook the quality assessment independently. If disagreements appeared, a third author helped to resolve the problem.
Data analysis and presentation
Based on the collected data, 2 × 2 contingency tables were created to calculate the indices of diagnostic validity, the main outcome data included pooled sensitivity (Se) and specificity (Sp), each value was determined with 95% confidence intervals. A Forest plot was used to display the results of the meta-analysis. Methodological heterogeneity was assessed during selection.
We determined the existence of a threshold effect, which is the primary cause of heterogeneity in test accuracy studies, defined as the use of different cut-offs or thresholds in different studies to determine a positive test result. The analysis of the diagnostic threshold was assessed using the graphic representation of “Se” vs “1-Sp” on a receiver operating characteristic (ROC) plane and also by calculating the Spearman correlation coefficient. The ROC plane provides a graphic representation of the pairs of Se and Sp; characteristically its points show a typical pattern of “shoulder arm” if the threshold effect exists. The results were synthesized and represented graphically in a forest plot. If there was evidence of a threshold effect, the studies were combined to create a SROC and we calculated an additional measurement of accuracy of the technique (Q*) and obtained the area under the curve (AUC). Possible sources of heterogeneity were assessed through chi-square, the null hypothesis that the studies are homogeneous was rejected with a p value of <0.05 and heterogeneity was quantified using the I2 statistic which was interpreted as low (25%–50%), moderate (51%–75%) and high (>75%) levels of heterogeneity. The Mantel-Haenszel fixed effect model was used if heterogeneity was not found; otherwise, the Dersimonian-Laird random effect model was used. In addition, subgroup analyses were also conducted by collection of sera (prospective, retrospective), Ethnicities (Caucasian and Asian) and QUADAS score. Potential publication bias was examined using Deeks' funnel plots37, with P < 0.05 for the slope coefficient indicating significant asymmetry. All meta-analyses were performed using MetaDisc (Version 1.4)38 and STATA (Version 10.0).
References
Sokoll, L. J. et al. Proenzyme psa for the early detection of prostate cancer in the 2.5–4.0 ng/ml total psa range: preliminary analysis. Urology 61, 274–6 (2003).
Schroder, F. H. & Roobol, M. J. Defining the optimal prostate-specific antigen threshold for the diagnosis of prostate cancer. Curr Opin Urol 19, 227–31 (2009).
Hori, S., Blanchet, J. S. & McLoughlin, J. From prostate-specific antigen (PSA) to precursor PSA (proPSA) isoforms: a review of the emerging role of proPSAs in the detection and management of early prostate cancer. BJU Int 112, 717–28 (2013).
Khan, M. A. et al. Evaluation of proprostate specific antigen for early detection of prostate cancer in men with a total prostate specific antigen range of 4.0 to 10.0 ng/ml. J Urol 170, 723–6 (2003).
Stephan, C. et al. Multicenter evaluation of [−2]proprostate-specific antigen and the prostate health index for detecting prostate cancer. Clin Chem 59, 306–14 (2013).
Lazzeri, M. et al. Serum index test %[−2]proPSA and Prostate Health Index are more accurate than prostate specific antigen and %fPSA in predicting a positive repeat prostate biopsy. J Urol 188, 1137–43 (2012).
Jansen, F. H. et al. Prostate-specific antigen (PSA) isoform p2PSA in combination with total PSA and free PSA improves diagnostic accuracy in prostate cancer detection. Eur Urol 57, 921–7 (2010).
Lazzeri, M. et al. Clinical performance of serum prostate-specific antigen isoform [−2]proPSA (p2PSA) and its derivatives, %p2PSA and the prostate health index (PHI), in men with a family history of prostate cancer: results from a multicentre European study, the PROMEtheuS project. BJU Int 112, 313–21 (2013).
Lazzeri, M. et al. Relationship of chronic histologic prostatic inflammation in biopsy specimens with serum isoform [−2]proPSA (p2PSA), %p2PSA and prostate health index in men with a total prostate-specific antigen of 4–10 ng/ml and normal digital rectal examination. Urology 83, 606–12 (2014).
Guazzoni, G. et al. Prostate-specific antigen (PSA) isoform p2PSA significantly improves the prediction of prostate cancer at initial extended prostate biopsies in patients with total PSA between 2.0 and 10 ng/ml: results of a prospective study in a clinical setting. Eur Urol 60, 214–22 (2011).
Catalona, W. J. et al. A multicenter study of [−2]pro-prostate specific antigen combined with prostate specific antigen and free prostate specific antigen for prostate cancer detection in the 2.0 to 10.0 ng/ml prostate specific antigen range. J Urol 185, 1650–5 (2011).
Guazzoni, G. et al. Preoperative prostate-specific antigen isoform p2PSA and its derivatives, %p2PSA and prostate health index, predict pathologic outcomes in patients undergoing radical prostatectomy for prostate cancer. Eur Urol 61, 455–66 (2012).
Stephan, C. et al. Comparative assessment of urinary prostate cancer antigen 3 and TMPRSS2:ERG gene fusion with the serum [-2]proprostate-specific antigen-based prostate health index for detection of prostate cancer. Clin Chem 59, 280–8 (2013).
Ferro, M. et al. Predicting prostate biopsy outcome: prostate health index (phi) and prostate cancer antigen 3 (PCA3) are useful biomarkers. Clin Chim Acta 413, 1274–8 (2012).
Mikolajczyk, S. D. et al. Proenzyme forms of prostate-specific antigen in serum improve the detection of prostate cancer. Clin Chem 50, 1017–25 (2004).
Loeb, S. et al. Prospective multicenter evaluation of the Beckman Coulter Prostate Health Index using WHO calibration. J Urol 189, 1702–6 (2013).
Lughezzani, G. et al. Multicenter European External Validation of a Prostate Health Index-based Nomogram for Predicting Prostate Cancer at Extended Biopsy. Eur Urol (2013).
Fossati, N. et al. Clinical performance of serum isoform [−2]proPSA (p2PSA) and its derivatives, namely %p2PSA and PHI (Prostate Health Index) in men younger than 60 years of age: results from a multicentric European study. BJU Int (2014).
Lughezzani, G. et al. Development and internal validation of a Prostate Health Index based nomogram for predicting prostate cancer at extended biopsy. J Urol 188, 1144–50 (2012).
Scattoni, V. et al. Head-to-head comparison of prostate health index and urinary PCA3 for predicting cancer at initial or repeat biopsy. J Urol 190, 496–501 (2013).
Perdona, S. et al. Prostate health index (phi) and prostate cancer antigen 3 (PCA3) significantly improve diagnostic accuracy in patients undergoing prostate biopsy. Prostate 73, 227–35 (2013).
Ferro, M. et al. Prostate Health Index (Phi) and Prostate Cancer Antigen 3 (PCA3) significantly improve prostate cancer detection at initial biopsy in a total PSA range of 2–10 ng/ml. PLoS One 8, e67687 (2013).
Lazzeri, M. et al. Serum isoform [−2]proPSA derivatives significantly improve prediction of prostate cancer at initial biopsy in a total PSA range of 2–10 ng/ml: a multicentric European study. Eur Urol 63, 986–94 (2013).
Ito, K. et al. Diagnostic significance of [−2]pro-PSA and prostate dimension-adjusted PSA-related indices in men with total PSA in the 2.0-10.0 ng/mL range. World J Urol 31, 305–11 (2013).
Le, B. V. et al. [−2]Proenzyme prostate specific antigen is more accurate than total and free prostate specific antigen in differentiating prostate cancer from benign disease in a prospective prostate cancer screening study. J Urol 183, 1355–9 (2010).
Sokoll, L. J. et al. A prospective, multicenter, National Cancer Institute Early Detection Research Network study of [−2]proPSA: improving prostate cancer detection and correlating with cancer aggressiveness. Cancer Epidemiol Biomarkers Prev 19, 1193–200 (2010).
Stephan, C. et al. A [−2]proPSA-based artificial neural network significantly improves differentiation between prostate cancer and benign prostatic diseases. Prostate 69, 198–207 (2009).
Sokoll, L. J. et al. [−2]proenzyme prostate specific antigen for prostate cancer detection: a national cancer institute early detection research network validation study. J Urol 180, 539–43; discussion 543 (2008).
Catalona, W. J. et al. Serum pro prostate specific antigen improves cancer detection compared to free and complexed prostate specific antigen in men with prostate specific antigen 2 to 4 ng/ml. J Urol 170, 2181–5 (2003).
Ng, C. F. et al. The Prostate Health Index in predicting initial prostate biopsy outcomes in Asian men with prostate-specific antigen levels of 4–10 ng/mL. Int Urol Nephrol (2013).
van Leeuwen, P. J. et al. Prostate cancer mortality in screen and clinically detected prostate cancer: estimating the screening benefit. Eur J Cancer 46, 377–83 (2010).
Mikolajczyk, S. D., Marks, L. S., Partin, A. W. & Rittenhouse, H. G. Free prostate-specific antigen in serum is becoming more complex. Urology 59, 797–802 (2002).
Filella, X. & Gimenez, N. Evaluation of [−2] proPSA and Prostate Health Index (phi) for the detection of prostate cancer: a systematic review and meta-analysis. Clin Chem Lab Med 51, 729–39 (2013).
Liberati, A. et al. The PRISMA statement for reporting systematic reviews and meta-analyses of studies that evaluate healthcare interventions: explanation and elaboration. BMJ 339, b2700 (2009).
Whiting, P., Rutjes, A. W., Reitsma, J. B., Bossuyt, P. M. & Kleijnen, J. The development of QUADAS: a tool for the quality assessment of studies of diagnostic accuracy included in systematic reviews. BMC Med Res Methodol 3, 25 (2003).
Whiting, P. et al. Development and validation of methods for assessing the quality of diagnostic accuracy studies. Health Technol Assess 8,1–234 (2004).
Deeks, J. J., Macaskill, P. & Irwig, L. The performance of tests of publication bias and other sample size effects in systematic reviews of diagnostic test accuracy was assessed. J Clin Epidemiol 58, 882–93 (2005).
Zamora, J., Abraira, V., Muriel, A., Khan, K. & Coomarasamy, A. Meta-DiSc: a software for meta-analysis of test accuracy data. BMC Med Res Methodol 6, 31 (2006).
Acknowledgements
No funding was used for this project.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: Y.T., J.F.X. Performed the experiments: W.Y.W., M.L.W. Analyzed the data: L.W., M.L.W. Contributed reagents/material/analysis tools: T.S.A., L.W., Wrote the manuscript: W.Y.W., M.L.W., Reference collection and data management: Y.T., T.S.A.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary data
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. The images in this article are included in the article's Creative Commons license, unless indicated otherwise in the image credit; if the image is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the image. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Wang, W., Wang, M., Wang, L. et al. Diagnostic ability of %p2PSA and prostate health index for aggressive prostate cancer: a meta-analysis. Sci Rep 4, 5012 (2014). https://doi.org/10.1038/srep05012
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep05012
This article is cited by
-
Relationship between Proclarix and the Aggressiveness of Prostate Cancer
Molecular Diagnosis & Therapy (2023)
-
Glycosylation in cancer: its application as a biomarker and recent advances of analytical techniques
Glycoconjugate Journal (2022)
-
Evaluation of prostate health index in predicting bone metastasis of prostate cancer before bone scanning
International Urology and Nephrology (2022)
-
Usefulness of the prostate health index in predicting the presence and aggressiveness of prostate cancer among Korean men: a prospective observational study
BMC Urology (2021)
-
Clinical utility and cost modelling of the phi test to triage referrals into image-based diagnostic services for suspected prostate cancer: the PRIM (Phi to RefIne Mri) study
BMC Medicine (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.