Abstract
Experimental models that recapitulate mutational landscapes of human cancers are needed to decipher the rapidly expanding data on human somatic mutations. We demonstrate that mutation patterns in immortalised cell lines derived from primary murine embryonic fibroblasts (MEFs) exposed in vitro to carcinogens recapitulate key features of mutational signatures observed in human cancers. In experiments with several cancer-causing agents we obtained high genome-wide concordance between human tumour mutation data and in vitro data with respect to predominant substitution types, strand bias and sequence context. Moreover, we found signature mutations in well-studied human cancer driver genes. To explore endogenous mutagenesis, we used MEFs ectopically expressing activation-induced cytidine deaminase (AID) and observed an excess of AID signature mutations in immortalised cell lines compared to their non-transgenic counterparts. MEF immortalisation is thus a simple and powerful strategy for modelling cancer mutation landscapes that facilitates the interpretation of human tumour genome-wide sequencing data.
Similar content being viewed by others
Introduction
High-throughput sequencing has shown that cancer genomes are riddled with somatic alterations, with numerous tumour types harbouring hundreds to thousands of mutations. Bioinformatic analyses of this massive catalogue in search of recurring mutations have substantially contributed to the identification of most of the genes functionally impaired in cancer development1. The genome-wide human cancer sequencing data also provide a powerful resource for investigating the nature of mutagenic insults that give rise to mutations in human population2,3. However, what is lacking in order to make optimal use of this resource for such a purpose is a suitable database of experimentally induced mutations to test inferences that come from inspecting human mutation patterns. It is well known that mutagenic factors, whether chemical or enzymatic, mutate DNA in characteristic ways, thereby revealing clues to their identities. This principle was elegantly demonstrated decades before the advent of new sequencing technologies in a wide variety of assays4,5. These assays, however, had one feature in common that limited their scope. Mutations were typically scored in a single gene (allowing clonal selection), or at best, in a discrete number of specific genes. Although data from such experiments have been fundamentally important to biology, the tests were not designed to recapitulate or interpret the more complex mutation profiles generated from genome-wide data. Genome-wide sequencing of cells exposed to sources of mutation in a controlled fashion will now allow more comprehensive experimental investigations of the mutagenic activities of human carcinogens.
This study aims to determine whether a simple experimental system using in vitro immortalisation of normal mammalian cells would generate genome-wide mutation data relevant to human tumours. Immortalisation of primary cells, notably murine embryonic fibroblasts (MEFs), has been used extensively as a powerful in vitro model for exploring genetic control of cellular homeostasis and its disruption in disease6. Recent research in this area has shown that various molecular pathways that control cellular senescence and become circumvented in vitro to allow cell immortalisation are cancer gene pathways, including oncogenes and tumour suppressor genes known to be mutated in human cancer7,8,9. Encouragingly, we showed in previous work that when carcinogens are applied to Hupki MEFs (MEFs carrying normal human p53 sequences embedded in the Trp53 gene, human p53 knock-in) prior to senescence, emerging clonal cell lines harbour TP53 gene signature mutations characteristic of the carcinogens and consistent with TP53 mutations detected in human cancers from exposed patient cohorts10,11,12.
In the present study, we expanded this approach by assessing exome-wide mutation patterns in the Hupki MEF immortalisation assay ( Supplementary Fig. 1 ). We sequenced the exome of immortalised MEF lines established from primary cultures exposed to well-known carcinogens and compared the mutation profiles obtained in these assays to those observed in genome-wide data from human tumours with related aetiologies. Mutation signatures derived from these assays were also compared to currently known signatures in human cancers2. Since the MEF in vitro immortalisation process has parallels with the conversion of normal cells to tumour cells in vivo, we also investigated alterations in specific driver genes that could provide mechanistic clues to molecular events governing senescence bypass and immortalisation. Finally, in a proof-of-concept experiment devised to explore an endogenous process proposed to contribute to the human mutation load, we examined the effect of activation-induced cytidine deaminase transgene (AID-Tg) expression on the pattern of base substitutions that accumulate during MEF immortalisation13.
Results
Genome-wide mutation spectra from immortalised MEF cell lines
Genomic DNA isolated from primary MEFs and from immortalised cell lines derived from MEF cultures exposed to aristolochic acid (AA), ultraviolet light subclass C (UVC), the alkylating agent N-methyl-N'-nitro-N-nitrosoguanidine (MNNG), the tobacco mutagen benzo(a)pyrene (BaP), or unexposed cultures (see Supplementary Table 1 ), were subjected to genome-wide mutation profiling by whole-exome sequencing (WES) (see Supplementary Fig. 1 for assay overview and Methods for WES data processing and analysis). Two cell lines per exposure category were investigated. As shown in Fig. 1 , the patterns of mutations found in the MEF cell lines were in marked concordance with those observed in human tumours with aetiologies related to the mutagens tested and were as expected from previous knowledge on the mutagenic properties of these particular exposures. In the cell lines derived from AA-exposed cultures, the most frequent type of mutation was A:T > T:A as in urinary tract urothelial cancers (UTUC) from AA exposed patients ( Fig. 1a ) and a significant strand-bias towards the non-transcribed strand was observed for A > T ( Table 1 ), in keeping with previous reports on human UTUC from AA-exposed patients14,15. In cell lines from BaP-exposed cultures, the most frequent type of mutation was C:G > A:T with a strand bias towards the non-transcribed strand for G > T as is seen in lung cancers (Lung_Ca) from heavy smokers ( Fig. 1b and Table 1 ). In cell lines from MNNG-exposed cells, the most frequent mutation type was C:G > T:A with no significant strand-bias ( Fig. 1c and Table 1 ), consistent with the alkylating properties of this agent and with the pattern observed in brain tumours from patients treated with the alkylating agent temozolomide. In the cell lines from UVC-exposed cultures, the most frequent type of mutation was C:G > T:A, as in skin squamous cell carcinomas (Skin_SCC) and with a strand-bias of borderline significance ( Fig. 1d and Table 1 ).
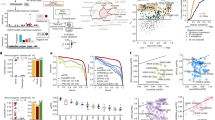
Mutation patterns derived from exome data obtained from MEF immortalised cell lines.
Mutation type distributions (a–f) and sequence context (g–l) of single base substitutions. For each treatment condition, data are shown for two independent immortalised cell lines and for a set of human tumours related to the tested condition. In (a–f), the percentage of each substitution type is shown with the total number of mutations indicated in parentheses. In (g–l), heat maps of mutation sequence context are shown. The percentage of each substitution type within a triplet sequence context is colour-coded according to the percent values. Highly abundant mutations are represented in red and low abundance mutations are in yellow. (a,g) Aristolochic-acid treatment (two left panels) and upper urinary tract human tumours (right panel) from patients exposed to AA. (b,h) Benzo(a)pyrene treatment (two left panels) and lung adenocarcinomas (right panel) from heavy smokers. (c,i) N-methyl-N'-nitro-N-nitrosoguanidine treatment (two left panels) and human recurrent glioblastoma treated with temozolomide (right panel). (d,j) UVC treatment (two left panels) and human skin squamous cell carcinomas (right panel) (COSMIC v65). (e,k) AID transgene (two panels). (f,l) Data from four independent cell lines obtained by spontaneous immortalisation of the Hupki MEF primary cells (Spont, no treatment).
In addition to exogenous exposures, we assessed the effect of an endogenous mutagenic process by analysing immortalised MEFs harbouring a transgene expressing activation-induced cytidine deaminase (AID) (see Methods). In these cell lines, referred to as HxAID-Tg, a predominance of C:G > T:A transitions was observed ( Fig. 1e ), as expected from experimental studies on the mutagenic properties of AID13,16. Finally, four immortalised MEF cell lines from untreated cultures were analysed to determine underlying mutagenesis in this model. Interestingly, the most predominant mutation type was C:G > G:C ( Fig. 1f ) in all four cell lines, as has been observed previously in the Trp53 gene of immortalised MEFs17.
The sequence context of mutations is an important feature of mutation patterns because many mutagenic agents and processes exhibit a preferred base context. We analysed the 5′ and 3′ base context of mutations in all conditions described above. As shown in Fig. 1 , a previously described preferred sequence context for each specific exposure was recapitulated in the MEF assay. Indeed, A:T > T:A mutations occurred predominantly within a 5′-CAG-3′ motif, as in the selected human set ( Fig. 1g ) and as reported in other published series14,15,18,19,20. For BaP exposure, C > A mutations occurred most frequently in 5′-CCN-3′ triplets (corresponding to 5′-NGG-3′ for the complementary G > T), as in the human lung tumour dataset ( Fig. 1h ). In cell lines from MNNG cultures, C > T transitions with a C or T in 3′ and any base in 5′ were the most frequent (corresponding to 5′-(G/A)GN-3′ for the complementary G > A mutations), observed also in recurrent glioblastomas of temozolomide-treated patients ( Fig. 1i ). These mutations occurred mainly outside CpG sites as expected. In the cell lines derived from UVC treated MEF cultures, C > T changes within a 5′-(C/T)CN-3′ motif were the major events as seen in human skin SCC ( Fig. 1j ). This context is expected from the published literature on UV mutagenesis, which describes the highly characteristic alterations at pyrimidine dimers induced by UV exposure. Interestingly, the frequent C > G mutations found in the spontaneous lines showed a preferred sequence context for 5′-GCC-3′ a signature that was also present, although much less prominently, in most of the other cell lines ( Fig. 1l ). Finally, in the spontaneously immortalised lines from HxAID-Tg MEFs ( Fig. 1k ) the predominant C > T changes were most frequently observed in a 5′-GC(A/C/T)-3′ sequence context, followed by 5′-AC(A/C/T)-3′. These findings match the preferred contexts previously demonstrated for AID activity21,22. The most frequent single base substitutions (SBS) observed in human cancers and in mammalian evolution are C > T transitions at 5′-NCG-3′ sites (CpGs). These mutations occur following spontaneous deamination of 5-methyl-cytosines and result in C > T transitions23. In the cell lines analysed here, CpGs accounted for 25–30% of the C > T mutations in the cell lines derived from AA, BaP and untreated MEF cultures, but for less than 15% of C > T mutations in the MNNG, UVC or HxAID-Tg cell lines, which is consistent with the treatment-specific sequence context of C > T transitions in these latter cell lines.
Two types of statistical analyses were then applied to these data ( Fig. 2 ). Firstly, principal component analysis (PCA) was performed to assess whether global mutation patterns obtained in the MEF immortalisation assays can distinguish between cell lines obtained from different treatments/conditions. Using percent frequency values of the six mutation types in their triplet sequence context (amounting to a total of 96 variables), the two first components were able to discriminate the replicate cell lines according to each specific treatment condition ( Fig. 2a ). When including the human cancer datasets in the exposure model, we observed a good concordance with the mouse datasets for most conditions, with the exception of the UVC treatment which showed a broader confidence interval ( Fig. 2b ). Secondly, the method used by Alexandrov et al.24, to extract signatures was adapted and applied to the 14 cell line data (see Methods). Although this method is optimized for large datasets, it could identify six signatures that corresponded to the six experimental conditions ( Supplementary Fig. 2 ) and were concordant with the mutation patterns shown in Fig. 1 . The comparison of the MEF experimental signatures with the 27 human-cancer derived signatures reported by Alexandrov et al.2, showed high similarity between the MNNG signature and Signature 11 (temolozomide), similarity between the BaP signature and Signature 4 (smoking) and between AID signature and Signature 19 (not identified) ( Fig. 2c ). No similarity was found for the AA signature (patients with AA-associated tumours were not analysed by Alexandrov et al.), or for the signature observed in the spontaneous immortalised cell lines.
Analysis of mutation signatures derived from exome data obtained from MEF immortalised cell lines.
(a) Principal component analysis (PCA) of WES data using mutation signatures. PCA was computed using as input the frequency matrices of sequence context mutations (96 variables) from cell lines immortalised following exposure of primary Hupki MEFs to a carcinogen (AA, BaP, MNNG or UVC), from Hupki MEFs carrying the AID transgene (HxAID-Tg) or from Hupki MEF-derived cell lines that immortalised spontaneously (Spont). Each sample is plotted considering the value of the first and second principal components (PC1 and PC2). The percentage of variance explained by each component is indicated within brackets in each axis. A 95% confidence ellipsis is drawn for each experimental condition and the empty squares indicate the respective centre of gravity. Cells and samples are represented by round and squared solid symbols, respectively. (b) Same as in (a) but with the human tumour datasets (same as shown in Figure 1) added to the input and labelled by arrows. HxAID-Tg samples are omitted in (b) as no corresponding relevant tumour data were identified. AAN_UTUC, aristolochic acid nephropathy-related upper urinary tract urothelial carcinoma; GBM_TZM, glioblastoma after temozolomide treatment; Lung_Ca, lung carcinoma; Skin_SCC, skin squamous cell carcinoma. (c) Graphical representation of the similarity distance of each of the six MEF signatures (front-back axis) to each of the 27 human cancer signatures2 (horizontal axis, 1A through U2). The vertical axis measures the similarity of signatures between the two systems, expressed as negative log(tan(angle), see Methods. Negative values below the x-z plane correspond to angles >45° and represent dissimilarity and are thus not shown.
These results show that mutation patterns obtained in the MEF immortalisation assay are specific to the exposure and can reveal carcinogen-specific signatures that are relevant to human cancers.
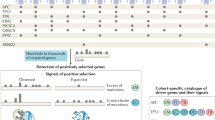
Driver gene mutation status in immortalised MEF cell lines
The cell lines from carcinogen exposure experiments chosen here for WES studies harbour TP53 mutations that arose during immortalisation of the primary cells. To investigate whether other cancer driver genes were recurrently affected during the senescence bypass/immortalisation process, we analysed the mutation status of other established or putative cancer drivers, including all those defined as oncogenes or tumour suppressor genes according to the “20/20 rule” formulated by Vogelstein et al.1, as well as genes encoding regulators of the epigenome that have been described as a newly emerging class of cancer driver genes1,25,26. Non-synonymous and truncating mutations found in these selected driver genes are detailed in Supplementary Dataset 1 and graphically summarized in Supplementary Dataset 2 . A number of genes in these functional classes were found altered by mutations characteristic of the exposure that cells underwent prior to immortalisation. Although most genes were mutated only in one line, the Ep400, Dnmt1, Kdm6b, Kmt2d, Arid1b and Arid2 genes were mutated in at least two lines. Ep400 and Kmt2d in particular were mutated in four cell lines. Ep400, a regulator of cellular senescence within the p53-p21 axis27, was affected by a truncating mutation in one line and Kmt2d carried three mutations in important functional domains. The most unequivocal driver gene mutations were two activating Ras missense mutations, highly recurrent in human cancers: the (c.A182T/p.Q61L) Hras1 mutation in one AA cell line corresponding to the HRAS mutation previously associated with exposure to AA in humans and animal models14,18,28,29,30 and an activating mutation in the Kras oncogene (c.A182G/p.Q61R) identified in one of the UVC lines. Overall, these observations suggest that the MEF immortalisation assay captures and selects for driver gene mutations relevant to cancer biology and are in keeping with extensive literature on the impact of cancer-related genes on senescence bypass, immortalisation and transformation of MEFs9,31.
Discussion
In this report we show that mutations acquired in MEFs during establishment in culture and studied at the exome level reveal patterns relevant to human cancers. While the MEF immortalisation assay protocol has been shown previously to recapitulate TP53 mutation patterns in the context of specific carcinogen exposures10,11, we demonstrate here that this assay is highly suitable as a selection strategy to obtain a cell population harbouring a suite of base substitutions relevant to exome-wide mutation data derived from human cancers. In principle, one single immortalised cell line provides information to identify a mutation signature, whereas many cell lines would be necessary when interrogating a single gene such as TP53. In practice, of course, WES on multiple cell line replicates per exposure or condition is warranted and will be called for in extended studies in the future to generate highly robust mutation signatures. The scope of overlap between mutation patterns in human datasets and immortalised MEF lines includes: (a) the global distribution of mutation types, (b) the accumulation of mutations on the non-transcribed strand (strand bias) for treatments with carcinogens known to elicit transcription-coupled DNA repair and (c) the sequence context of the dominant mutation type. Thus, using four carcinogens with well-known mutagenic properties, the predominant mutation signatures we found with this model for the four tested carcinogens were the ones expected for these mutagenic agents. Although we analysed only two cell lines for each carcinogen, the expected signatures were evident in single cell lines and were highly reproducible between the two cell lines. The human tumour datasets used for comparison with our in vitro data were selected from publicly available data and our selection was based on whether the suspected aetiologies of the tumour sets were linked to the carcinogen tested in MEFs. The most striking matching condition was the AA treatment. In both human and in vitro MEF data, over 50% of mutations were A > T transversions, with a significant strand bias of 2:1 and a sequence context dominated by 5′-CAG-3′. The aetiology of the tumours included in the human set has been clearly associated with the AA exposure14. Since AA is a potent carcinogen that mainly causes A > T transversions, enrichment of these somatic mutations in exposed individuals is likely to reflect the insult of AA exposure. The AA signature has not been found in any other cancer type so far32. In the case of the in vitro WES data from cell lines arising from MNNG-exposed cultures, the human set chosen for comparison consisted of patients treated with the drug temozolomide, which, like MNNG, is an alkylating agent. The global mutation type distribution was strikingly similar between the mouse and human data and very distinct from primary tumours of the same type but not exposed to temozolomide ( Supplementary Fig. 3 ). The MNNG signature was also very similar to the temozolomide signature derived from another set of temozolomide-exposed patients reported previously2. This signature is thus highly specific for alkylating agents such as MNNG and temozolomide. With respect to other exposures, it is clear that human tumour development typically involves various mutational mixtures and selection processes, resulting in a complex picture of mutation signatures. These considerations may explain why the tumour data from lungs of heavy smokers differ from the in vitro data from BaP exposure with respect to the less prominent mutation types. Tobacco smoke contains a highly complex mixture of carcinogens. Nevertheless, the BaP signature derived from the in vitro assay exhibited similarity with the human tumour-derived smoking signature reported previously2, suggesting that BaP and possibly other smoke components that have similar mutagenic properties constitute one of the main carcinogenic insults responsible for the smoking signature observed in human tumours. The dataset from tumours associated with UV exposure was the most distant from the signature obtained in vitro, although the expected C > T mutations within a 5′-(C/T)CN-3′ context were prominent in both sets. There are several possible explanations, such as the technical aspects of the experimental procedure in vitro, or the mutagenic activities of sunlight compared to UVC alone. These results are reflected in the principal component analysis that showed the closest relationship between human and MEF data for AA and BaP and a more distant relationship for UVC. In addition to exogenous exposures, the spontaneous decay of DNA is a well-known cause of the human mutation load33 and the deregulation of endogenous cellular enzymes that accelerate the accumulation of sequence changes is becoming of increasing interest to cancer biologists. It is a considerable challenge, however, to determine the relative contributions of different DNA metabolism pathways to genetic alterations observed in human cancers and to understand the factors that may result in the deregulation of normal processes governing DNA integrity. Recently the APOBEC/AID families of cytidine deaminases have come under scrutiny because of their potential roles in cancer as endogenous sources of mutation in various cancer types2,34,35. AID, which is normally expressed in B-lymphocytes, has been proposed as a possible source of mutagenic activity in the development of various inflammation-associated cancer types when expressed inappropriately36,37. An early investigation on mutation patterns produced by AID in a single reporter gene showed a strong C > T mutation signature as anticipated16 and there are now many studies exploring the impact of ectopic AID expression on cancer development13. Here we compared the sequence changes during immortalisation of MEFs harbouring a constitutively active AID transgene with MEFs that did not carry the transgene. The AID signature mutation was easily captured by this strategy, providing a proof-of-concept demonstration of the applicability of this approach to investigating endogenous mutagenesis. Interestingly, the in vitro AID signature showed some similarity with one of the signatures found in pilocytic astrocytoma2,34,35, but was not represented in other cancer types and had no similarity to two previously reported APOBEC signatures2,34,35. The full role of AID in shaping mutation patterns in humans remains to be investigated.
Surprisingly, the analysis of spontaneously immortalised cell lines from untreated cultures showed a strikingly high frequency of C > G mutations in the 5′-GCC-3′ sequence context, also present (albeit at lower and variable frequency) in all cell lines ( Fig. 1 and Supplementary Fig. 2 ). A high frequency of C > G mutations in the p53 gene was observed previously in both Hupki MEFs (with human TP53 sequences) and MEFs with the murine Trp53 gene17. Our WES results show that this phenomenon is global and may be due to specific mutagenic pressures inherent to the experimental conditions. Gene Ontology Biological Process analysis of genes affected by non-synonymous C > G mutations in the spontaneously immortalised cell lines identified 11 genes involved in regulation of apoptosis/programmed cell death (GO 0042981; GO 0043067) as high scoring categories (enrichment p-value < 0.05, Fisher's exact test, Supplementary Table 2 ), a finding consistent with the cultures overcoming senescence and with individual cells acquiring immortalised properties. This signature did not show any similarity to those reported by Alexandrov et al.2. Although C > G mutations have been associated with two signatures linked to APOBEC activity in human cancers2, they occur in a different sequence context of 5′-TCA-3′. A recent genome-wide analysis of gingivo-buccal oral SCC from Indian patients reported a high frequency of C > G mutations (although the sequence context was not reported) in three tobacco users carrying a high mutation load in their tumours38. These authors proposed that the unexpectedly high numbers of C > G mutations in their sample set may be caused by oxidative damage. The DNA lesion 8-oxoguanine caused by exposure to reactive oxygen species39,40 can lead to this transversion. The elevated numbers of C > G substitutions in spontaneously immortalising MEFs may be a cell culture artefact caused by high oxygen levels of standard incubation conditions and culturing the cells at physiological levels of oxygen can test this premise. Further investigation of the origin of these C > G substitutions in the MEF in vitro assay is warranted.
Although the number of cell lines analysed in the present study is limited, we identified several recurrently mutated genes among oncogenes and tumour suppressor genes classified as cancer drivers, or regulators of the epigenome, an emerging new class of potential driver genes. We note, however, that most of these mutations are likely to be passenger events occurring in the MEF immortalisation/transformation process, analogous to observations in human cancers. Interestingly, while mutations in all categories of genes but histone genes were observed in the carcinogen-exposed lines, the HxAID-Tg cell lines accumulated mutations mainly in histone genes ( Supplementary Datasets 1 and 2 ). A study to explore the reasons for this observation will require larger numbers of immortalised cell lines both with and without the AID transgene. The p53 status of emerging immortalised cells may also influence the subset of target genes subsequently mutated and selected for, but again, to address this speculation properly, an extensive set of cell line replicates will be needed.
Exome-wide analysis of MEFs thus joins epigenetic profiling and senescence bypass screens in the modern assembly of in vitro tools to elucidate cancer biology9,41. Analysis of more cell lines and detailed functional analyses of the specific mutations will be important in order to distinguish driver from passenger mutations in WES-MEF studies in a robust, statistically sound manner. Analysis of indels will be considered in future studies with more cell line replicates as the number of indels called in the current sample set was too small to derive meaningful interpretations of how indels might contribute to particular mutational signatures. The analyses of indels might also provide a more complete picture of mutations in tumour suppressor genes as this type of cancer gene is more often altered by indels. The present study is limited to a small number of cell lines and these have acquired typical human tumour TP53 mutations during immortalisation. The p53 gene mutation is the most common specific alteration known to drive senescence bypass and immortalisation of MEFs42,43. It will be interesting to investigate to what extent the p53 status influences global mutation patterns and the subset of mutated driver genes by comparing WES data from cell lines retaining the wild-type p53 gene sequence with cell lines that have acquired p53 mutations typical of human tumours.
In summary, the present study demonstrates the potential of the MEF immortalisation assay to reveal mutation signatures of human carcinogens. Although the use of mouse cells can be seen as a limitation because of differences in metabolism and DNA repair between humans and mice, in vitro cell models offer various strategies to accommodate or even exploit these distinctions, such as the addition of human liver microsomes to the culture, or breeding of mice with transgenic or knock-in strains expressing human genes to investigate various parameters relevant to a particular cancer risk factor. The ability of the MEF immortalisation model to recapitulate human carcinogen mutation signatures observed from whole-genome analysis of human tumours suggests that the model can provide important clues about the involvement of potential carcinogens in instances where aetiological and mechanistic evidence is deficient.
Methods
Hupki MEF cell lines
This study included immortalised MEF cell lines derived from primary cultures exposed to carcinogens that were reported previously ( Supplementary Table 1 and references therein). They were generated following a procedure referred to in the literature as the 3T3 protocol44, with minor adaptations. Briefly, fibroblasts from 13.5-day old Trp53tm/Holl mouse embryos harbouring a knock-in humanised version of the p53 gene (Hupki MEFs) were seeded into six-well plates, exposed to cancer agents or solvent during early passages and maintained in culture with occasional passaging until cultures emerged from senescence. Immortalised cultures were passaged at low density for several passages thereafter, prior to screening for the presence of a heterozygous or homo/hemizygous TP53 mutation and their designation as established cell lines. Cell lines chosen for the present analysis had acquired a dysfunctional TP53 mutation during immortalisation ( Supplementary Table 1 ). Acquisition of Trp53 gene mutations frequently occurs during senescence bypass and establishment in culture10,17,42,43 providing a convenient way to assess the identity and clonal origin of the immortalised cultures.
Cell lines with the AID transgene were established for the present study by crossing Hupki mice45,46 with AID transgenic mice47. From the interbred colony, we harvested MEFs from embryos homozygous for the (non-mutated) knock-in TP53 allele and either with or without the transgene (referred to as HxAID-Tg and MEFs respectively). T12.5 flasks (6 per MEF genotype) were seeded with 5 × 104 cells and cultured until the cells emerged from senescence, regained uniform morphology and could sustain repeated passaging at >1:10 dilution. Genomic DNAs from two immortalised cell lines per condition (independent biological duplicates) were prepared for WES analysis.
DNA preparation and WES
Genomic DNA (gDNA) was extracted from cells using DNeasy blood and tissue kit (QIAGEN) and checked for purity, concentration and integrity by OD260/280 ratio using NanoDrop Instruments (NanoDrop Technologies, Wilmington, DE, USA) and agarose gel electrophoresis. DNA was sheared by fragmentation by Covaris (Covaris, Inc., Woburn, MA USA) or Bioruptor (Diagenode, Inc., Denville, NJ, USA) and purified using Agencourt AMPure XP beads (Beckman Coulter, Fullerton, CA, USA). DNA samples were then tested for size distribution and concentration using an Agilent Bioanalyzer 2100 or Tapestation 2200 and by OD260/280 ratio. Fragment ends were repaired and Illumina libraries were generated using NEBNext reagents (New England Biolabs, Ipswich, MA, USA). Libraries were then subjected to exome enrichment using SureSelect XT Mouse All Exon Kit (Agilent Technologies, Wilmington, DE USA) following manufacturer's instructions. Enrichment was verified by qPCR and the quality, quantity and fragment size distribution of DNA determined by an Agilent Bioanalyzer or Tapestation. The libraries were sequenced in paired-end 100 nucleotide (nt) reads using the Illumina HiSeq2500 platform according to manufacturer's protocols.
MEF whole-exome data processing
All FASTQ files were analysed with FastQC to check sample homogeneity and quality. The FASTQ sequences were next aligned to the mm9 mouse reference genome with Burrows-Wheeler Aligner (BWA, version 0.7.5a) and the resulting SAM file was sorted and compressed in BAM format using Picard SortSam (version 1.98). Duplicate reads in the resulting BAM files were flagged with Picard MarkDuplicates. Local realignment around indels was performed in three steps: firstly, creation of a table of possible indels using GATK (version 2.7-2) RealignerTargetCreator, secondly, realignment of reads around those targets with GATK IndelRealigner and lastly a correction of mate pair information was done using Picard FixMateInformation. The base quality score recalibration required two steps: first to generate a recalibration table with GATK BaseRecalibrator, then to print reads based on the previous table with GATK PrintReads. An average of 58.8 million reads (100 bp) were sequenced per sample, of which 98% were mapped, 77% on target with a mean coverage of 61 (see Supplementary Table 3 for detailed metrics of sequencing quality and coverage). The recalibrated BAM files were used to call variants with MuTect software (version 1.1.4) using default parameters (including reads quality >20 and calls made only if the position has at least 14 reads in the tumour sample and at least 8 reads in the normal sample). As MuTect is tuned to perform normal/tumour comparison, primary cell cultures were used as “normal” samples and immortalised cell lines as “tumour” samples. Each immortalised cell line was compared to two primary MEF cultures and only the overlapping calls were taken into consideration to maximize the chance of robust variant calls and to exclude potential polymorphisms.
Human genome-wide sequencing datasets
Publicly available somatic mutation data obtained from whole-genome or whole-exome sequencing of human tumours were retrieved from the Catalogue Of Somatic Mutations In Cancer (COSMIC) database or original papers (selecting only SBS): a set of 14,715 substitutions reported in 19 UTUC samples from AA-exposed patients14; a set of 6,066 substitutions observed in seven primary skin squamous cell carcinomas (COSMIC v67); a set of 3,026 mutations observed in 10 primary lung adenocarcinomas from heavy smokers48; a set of 10,455 mutations observed in eight glioblastomas recurring in patients treated with temozolomide49; a set of 288 mutations observed in eight primary astrocytomas49; and a set of 378 mutations observed in four primary glioblastomas50.
Annotations of mutation data
For all datasets (MEF and human sets), the chromosome number, genomic coordinates, reference and mutated nucleotides were extracted for each variants. Variants were annotated with AnnoVar (version 2013aug23) using refGene, knownGene, ensGene, cytoBand, genomicSuperDups and dbSNP128 databases for the mm9 mouse genome build. The human sets were annotated using additional databases: gwasCatalog, 1000 Genomes Project, NHLBI GO Exome Sequencing Project (ESP), COSMIC, dbSNP137 (hg19 build) and PolyPhen and SIFT databases for predicting the functional impact of mutations. Gene strand orientation was retrieved from the UCSC Genome Browser database using a Perl script developed by Heng Li at the Sanger Institute. Mutations were included in the analyses only if they could be successfully annotated. Variants present in the dbSNP128 polymorphism database were excluded. The comprehensive lists of all SBS identified in all MEF conditions and SBS from human tumour datasets are available as Supplementary Dataset 3 .
Functional annotation analysis
A comprehensive list of established cancer driver genes (oncogenes and tumour suppressor genes) and candidate drivers coding for modifiers of DNA, histones and regulators of chromatin structure was assembled from literature and somatic mutation database mining1,25,26. Selected gene classes were annotated with functional domain information obtained from the UniProt and ENSEMBL databases. The comprehensive list of functional gene classes was matched against genes with mutations found in all MEF cell lines. Non-synonymously mutated genes were further selected, considering both exposure-specific alterations and any other mutation type. Human orthologues of the selected genes were examined in the COSMIC database for frequency of mutations in human tumours. For oncogenes, positions corresponding to non-synonymous mutations in MEFs were identified in human orthologues and investigated for mutation status in the COSMIC database. Gene Ontology Biological Process analyses were performed with the NIH DAVID web tool using default settings.
Statistical analyses
Statistical analyses were performed using the free R software (R Core Team, 2013) v3.0.2 or Excel. For the strand bias analyses, the statistical difference in the number of SBS between the non-transcribed and the transcribed strand was evaluated through the Pearson's χ2 test, using the prop.test function available in the stats R package. The test evaluated, for each experimental condition, whether the proportions of SBS in the non-transcribed strand differed from 0.5, which is the expected value by chance. As multiple conditions were assessed in parallel, a false discovery rate (FDR) correction was applied using the p.adjust function from the stats R package.
For analyses of mutation signatures, mutations were classified into 96 types determined by the six possible substitutions (A:T > C:G, A:T > G:C, A:T > T:A, C:G > A:T, C:G > G:C, C:G > T:A) and the 16 combinations of flanking (5′ and 3′) nucleotides. First, a PCA analysis was performed using as input the 96 variables. A 95% confidence interval was computed, including either only MEF samples or MEF/human data, to define the limits on the PCA plot for each experimental condition. Such analysis was performed based on the available functions in the FactoMiner package available in the Bioconductor repository (R package version 1.25. http://CRAN.R-project.org/package=FactoMineR). Second, the catalogue of experimental mutations defined by their 96 types was decomposed into signatures using the non-negative matrix factorisation algorithm of Brunet with the Kullback-Leibler divergence penalty24,51. The number of signatures was pre-set to six (the expected number of signatures based on the number of conditions) but the process was otherwise unsupervised: no information regarding exposures was used for the extraction of the signatures. To evaluate the similarity between the signatures from the cell lines and from human tumours by Alexandrov et al.2,24, each signature was represented as a vector in 96-dimensional space. The tangent of the angle between each pair of vectors was taken as the distance metric: the tangent transformation serving to expand the scale to compensate for the geometry of high-dimensional space. This distance was used to compute the grid of distance from each of the six MEF signatures to each of the 27 human signatures and converted for presentation to a similarity matrix by taking the negative log of the distance with negative values (angles >45°) suppressed.
Change history
16 January 2017
A correction has been published and is appended to both the HTML and PDF versions of this paper. The error has not been fixed in the paper.
References
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Hollstein, M., McCann, J., Angelosanto, F. A. & Nichols, W. W. Short-term tests for carcinogens and mutagens. Mutat. Res. 65, 133–226 (1979).
Miller, J. H. Carcinogens induce targeted mutations in Escherichia coli. Cell 31, 5–7 (1982).
Hahn, W. C. & Weinberg, R. A. Modelling the molecular circuitry of cancer. Nat. Rev. Cancer 2, 331–341 (2002).
Collado, M. & Serrano, M. Senescence in tumours: evidence from mice and humans. Nat. Rev. Cancer 10, 51–57 (2010).
Fridman, A. L. & Tainsky, M. A. Critical pathways in cellular senescence and immortalization revealed by gene expression profiling. Oncogene 27, 5975–5987 (2008).
Odell, A., Askham, J., Whibley, C. & Hollstein, M. How to become immortal: let MEFs count the ways. Aging (Albany. NY) 2, 160–165 (2010).
Liu, Z. et al. Human tumor p53 mutations are selected for in mouse embryonic fibroblasts harboring a humanized p53 gene. Proc. Natl. Acad. Sci. U. S. A 101, 2963–2968 (2004).
Liu, Z. et al. p53 mutations in benzo(a)pyrene-exposed human p53 knock-in murine fibroblasts correlate with p53 mutations in human lung tumors. Cancer Res. 65, 2583–2587 (2005).
vom Brocke, J., Schmeiser, H. H., Reinbold, M. & Hollstein, M. MEF immortalization to investigate the ins and outs of mutagenesis. Carcinogenesis 27, 2141–2147 (2006).
Schmitz, K. M. & Petersen-Mahrt, S. K. AIDing the immune system-DIAbolic in cancer. Semin. Immunol. 24, 241–245 (2012).
Hoang, M. L. et al. Mutational signature of aristolochic acid exposure as revealed by whole-exome sequencing. Sci. Transl. Med. 5, 197ra102 (2013).
Poon, S. L. et al. Genome-wide mutational signatures of aristolochic acid and its application as a screening tool. Sci. Transl. Med. 5, 197ra101 (2013).
Yoshikawa, K. et al. AID enzyme-induced hypermutation in an actively transcribed gene in fibroblasts. Science 296, 2033–2036 (2002).
Whibley, C. et al. Wild-type and Hupki (human p53 knock-in) murine embryonic fibroblasts: p53/ARF pathway disruption in spontaneous escape from senescence. J Biol. Chem. 285, 11326–11335 (2010).
Chen, C. H. et al. Aristolochic acid-associated urothelial cancer in Taiwan. Proc. Natl. Acad. Sci. U. S. A 109, 8241–8246 (2012).
Moriya, M.. et al. TP53 Mutational signature for aristolochic acid: an environmental carcinogen. Int. J. Cancer 129, 1532–1536 (2011).
Sidorenko, V. S. et al. Lack of recognition by global-genome nucleotide excision repair accounts for the high mutagenicity and persistence of aristolactam-DNA adducts. Nucleic Acids Res. 40, 2494–2505 (2012).
Bransteitter, R., Pham, P., Calabrese, P. & Goodman, M. F. Biochemical analysis of hypermutational targeting by wild type and mutant activation-induced cytidine deaminase. J. Biol. Chem. 279, 51612–51621 (2004).
Pham, P., Bransteitter, R., Petruska, J. & Goodman, M. F. Processive AID-catalysed cytosine deamination on single-stranded DNA simulates somatic hypermutation. Nature 424, 103–107 (2003).
Duncan, B. K. & Miller, J. H. Mutagenic deamination of cytosine residues in DNA. Nature 287, 560–561 (1980).
Alexandrov, L. B., Nik-Zainal, S., Wedge, D. C., Campbell, P. J. & Stratton, M. R. Deciphering signatures of mutational processes operative in human cancer. Cell Rep. 3, 246–259 (2013).
Plass, C. et al. Mutations in regulators of the epigenome and their connections to global chromatin patterns in cancer. Nat. Rev. Genet. 14, 765–780 (2013).
Timp, W. & Feinberg, A. P. Cancer as a dysregulated epigenome allowing cellular growth advantage at the expense of the host. Nat. Rev. Cancer 13, 497–510 (2013).
Chan, H. M., Narita, M., Lowe, S. W. & Livingston, D. M. The p400 E1A-associated protein is a novel component of the p53 -->p21 senescence pathway. Genes Dev. 19, 196–201 (2005).
Schmeiser, H. H. et al. Aristolochic acid activates ras genes in rat tumors at deoxyadenosine residues. Cancer Res. 50, 5464–5469 (1990).
Schmeiser, H. H., Scherf, H. R. & Wiessler, M. . Activating mutations at codon 61 of the c-Ha-ras gene in thin-tissue sections of tumors induced by aristolochic acid in rats and mice. Cancer Lett. 59, 139–143 (1991).
Wang, Y. et al. ACB-PCR measurement of H-ras codon 61 CAA-->CTA mutation provides an early indication of aristolochic acid I carcinogenic effect in tumor target tissues. Environ. Mol. Mutagen. 53, 495–504 (2012).
Lundberg, A. S., Hahn, W. C., Gupta, P. & Weinberg, R. A. Genes involved in senescence and immortalization. Curr. Opin. Cell Biol. 12, 705–709 (2000).
Hollstein, M., Moriya, M., Grollman, A. P. & Olivier, M. . Analysis of TP53 mutation spectra reveals the fingerprint of the potent environmental carcinogen, aristolochic acid. Mutat. Res. 753, 41–49 (2013).
Lindahl, T. Instability and decay of the primary structure of DNA. Nature 362, 709–715 (1993).
Burns, M. B. et al. APOBEC3B is an enzymatic source of mutation in breast cancer. Nature 494, 366–370 (2013).
Burns, M. B., Temiz, N. A. & Harris, R. S. Evidence for APOBEC3B mutagenesis in multiple human cancers. Nat. Genet. 45, 977–983 (2013).
Chiba, T., Marusawa, H. & Ushijima, T. Inflammation-associated cancer development in digestive organs: mechanisms and roles for genetic and epigenetic modulation. Gastroenterology 143, 550–563 (2012).
Shimizu, T., Marusawa, H., Endo, Y. & Chiba, T. Inflammation-mediated genomic instability: roles of activation-induced cytidine deaminase in carcinogenesis. Cancer Sci. 103, 1201–1206 (2012).
Maitra, A. et al. Mutational landscape of gingivo-buccal oral squamous cell carcinoma reveals new recurrently-mutated genes and molecular subgroups. Nat. Commun. 4, 2873 (2013).
Kino, K. & Sugiyama, H. GC-->CG transversion mutation might be caused by 8-oxoguanine oxidation product. Nucleic Acids Symp. Ser. 139–140 (2000).
Paz-Elizur, T. et al. DNA repair activity for oxidative damage and risk of lung cancer. J. Natl. Cancer Inst. 95, 1312–1319 (2003).
Tommasi, S. et al. Mammalian cells acquire epigenetic hallmarks of human cancer during immortalization. Nucleic Acids Res. 41, 182–195 (2013).
Harvey, D. M. & Levine, A. J. p53 alteration is a common event in the spontaneous immortalization of primary BALB/c murine embryo fibroblasts. Genes Dev. 5, 2375–2385 (1991).
Kamijo, T. et al. Functional and physical interactions of the ARF tumor suppressor with p53 and Mdm2. Proc. Natl. Acad. Sci. U. S. A 95, 8292–8297 (1998).
Todaro, G. J. & Green H. . Quantitative studies of the growth of mouse embryo cells in culture and their development into established lines. J. Cell Biol. 17, 299–313 (1963).
Luo, J. L. et al. Knock-in mice with a chimeric human/murine p53 gene develop normally and show wild-type p53 responses to DNA damaging agents: a new biomedical research tool. Oncogene 20, 320–328 (2001).
Reinbold, M. et al. Common tumour p53 mutations in immortalized cells from Hupki mice heterozygous at codon 72. Oncogene 27, 2788–2794 (2008).
Okazaki, I. M. et al. Constitutive expression of AID leads to tumorigenesis. J. Exp. Med. 197, 1173–1181 (2003).
Imielinski, M. et al. Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell 150, 1107–1120 (2012).
Johnson, B. E. et al. Mutational analysis reveals the origin and therapy-driven evolution of recurrent glioma. Science 343, 189–193 (2014).
Yost, S. E. et al. High-resolution mutational profiling suggests the genetic validity of glioblastoma patient-derived pre-clinical models. PLoS. One 8, e56185 (2013).
Brunet, J. P., Tamayo, P., Golub, T. R. & Mesirov, J. P. Metagenes and molecular pattern discovery using matrix factorization. Proc. Natl. Acad. Sci. U. S. A 101, 4164–4169 (2004).
Acknowledgements
We gratefully acknowledge core financial support for whole-exome sequencing from the Deutsches Krebsforschungszentrum (DKFZ). We thank the Centre Léon Bérard in Lyon, France, for providing computational capacity. We thank Dr Zdenko Herceg for comments on the manuscript scientific contents and Sylvie Nouveau for carefully reading the manuscript.
Author information
Authors and Affiliations
Contributions
M.O., M.H., J.Z. designed the study. A.W., T.N., K.-R.M., H.M., M.H. conducted cell culture and animal experiments. H.M. generated the AID-Tg and HxAID-Tg mice. M.V., J.McK., J.A., L.H. performed WES data processing and primary analyses. G.B. and X.C. performed the statistical analyses. M.O., M.A., X.C., H.H., H.M., G.B., M.H., J.Z. analysed and interpreted data. M.H. initiated the study. M.O., M.H., J.Z. wrote the manuscript. All authors approved the present submission of the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Supplementary Information
Supplementary_Dataset_1
Supplementary Information
Supplementary_Dataset_2
Supplementary Information
Supplementary_Dataset_3
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Olivier, M., Weninger, A., Ardin, M. et al. Modelling mutational landscapes of human cancers in vitro. Sci Rep 4, 4482 (2014). https://doi.org/10.1038/srep04482
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep04482
This article is cited by
-
DNA damage and somatic mutations in mammalian cells after irradiation with a nail polish dryer
Nature Communications (2023)
-
Contributions of replicative and translesion DNA polymerases to mutagenic bypass of canonical and atypical UV photoproducts
Nature Communications (2023)
-
Mutational signatures: experimental design and analytical framework
Genome Biology (2020)
-
Chemically induced mutations in a MutaMouse reporter gene inform mechanisms underlying human cancer mutational signatures
Communications Biology (2020)
-
Exome sequencing identifies novel mutation signatures of UV radiation and trichostatin A in primary human keratinocytes
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.