Abstract
At a proximal level, the physiological impacts of global climate change on ectothermic organisms are manifest as changes in body temperatures. Especially for plants and animals exposed to direct solar radiation, body temperatures can be substantially different from air temperatures. We deployed biomimetic sensors that approximate the thermal characteristics of intertidal mussels at 71 sites worldwide, from 1998-present. Loggers recorded temperatures at 10–30 min intervals nearly continuously at multiple intertidal elevations. Comparisons against direct measurements of mussel tissue temperature indicated errors of ~2.0–2.5 °C, during daily fluctuations that often exceeded 15°–20 °C. Geographic patterns in thermal stress based on biomimetic logger measurements were generally far more complex than anticipated based only on ‘habitat-level’ measurements of air or sea surface temperature. This unique data set provides an opportunity to link physiological measurements with spatially- and temporally-explicit field observations of body temperature.
Design Type(s) | observation design • time series design |
Measurement Type(s) | temperature of environmental material |
Technology Type(s) | biomimetic sensor |
Factor Type(s) | geographic location |
Sample Characteristic(s) | New South Wales • Queensland • State of Victoria • Province of British Columbia • Coquimbo • England • Scotland • County Clare • Galway • County Mayo • Baja California Peninsula • Auckland Region • Canterbury Region • Coromandel Peninsula • Northland Region • West Coast Region • State of California • Commonwealth of Massachusetts • State of Oregon • State of South Carolina • State of Washington • Eastern Cape Province • KwaZulu-Natal Province • Northern Cape Province • Western Cape Province • intertidal zone |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
Increasingly, researchers are emphasizing the need to consider physiological mechanisms when forecasting the effects of global climate change on organisms and ecosystems1–3. Specifically, studies have highlighted a need to understand how environmental conditions vary in space and time4 in addition to the details of how organisms respond to those variables5–8 as a means of evaluating inter- and intraspecific vulnerability (‘winners and losers’)9,10, the probability of invasion by non-native species11,12, changes in patterns of abundance and distribution13,14, and declines in biodiversity15 and ecosystem services16.
Notably, there is concern that simple correlations between environmental measurements (such as air, land surface and sea surface temperature) and species distributions may fail under the novel conditions presented by climate change17, highlighting the need to extrapolate from experiments conducted under controlled conditions to projections of future climate impacts3,18. There has also been an emphasis on considering the cumulative impacts of physiological stress14,19 on patterns of growth20 and reproduction21 rather than focusing solely on lethal extremes19.
However, making connections between the lab and field can be far more complex than is often assumed4. For example, a number of theoretical and empirical studies have explored the often over-riding importance of spatial and temporal variability in environmental parameters9,22, which is not captured when experiments are based only on monthly, yearly or decadal averages23,24. Moreover, while large-scale measurements of environmental conditions made by satellites, buoys, and weather stations provide critical insights into rates of environmental change on large scales25, at a proximal level these habitat-level measurements may not always serve as good indicators of physiological stress4,26. In fact, the only ‘environmental signals’ that matter to an organism are those that the organism actually experiences27. Making connections across scales that span from organismal to biogeographic is no easy matter, but is crucial if we are to effectively forecast ongoing responses to environmental change28,29.
One of the most obvious examples of the complex ways climate defines weather patterns, and weather then drives niche-level organismal responses30, is how climate change is ultimately reflected as changes in plant and animal body temperatures. The vast majority of organisms on Earth are ectothermic poikilotherms, so that their body temperatures and thus levels of physiological performance change with ambient environmental conditions. For terrestrial and intertidal ectotherms (and even some shallow-water corals31), body temperatures are driven by multiple environmental parameters, most notably solar radiation, air and water temperatures and wind speed32–34. The structure of an organism’s microhabitat, and especially its exposure to direct solar radiation, can have enormous implications for its body temperature, such that animal temperatures are only close to air temperature in fully shaded microhabitats26,35. While many animals can behaviourally select among these microhabitats as a means of thermoregulation36, others are functionally sessile and thus have body temperatures determined by very local topography. To further complicate matters, the size, morphology and colour of organisms, as well as their ability to form aggregations37,38 can affect heat exchange so that two organisms exposed to identical microclimatic conditions can have very different body temperatures39,40. To contend with these issues, multiple authors have developed heat budget models that factor-in the characteristics of the organism26,33,41 to predict body temperatures using weather data as inputs.
An alternative approach—and one that is required to validate biophysical (heat budget) models—is to use in situ sensors specifically tailored to record temperatures relevant to the organism being studied, either directly or through the use of biomimics42. Biomimetic sensors (biomimics) match the thermal characteristics (size, morphology, colour, material properties) of living organisms43,44, serving as an effective tool for recording organismal body temperature in their natural environment45,46. Here we report on a long-term data set of temperatures recorded by biomimetic loggers thermally matched to bivalves (mussels) in the intertidal zone, one of the most physiologically harsh habitats on Earth. Over the course of a 24-hr period, intertidal animals and algae are alternately exposed to water at high tide and to air, wind and solar radiation at low tide. Thus, their temperature not only depends on local weather conditions but also on the timing and duration of low tide47. We have previously shown, for example, that consistent differences in the timing of low tide relative to high levels of solar radiation create geographic mosaics in low tide temperature, where mussel body temperatures at higher latitude sites can be much higher than those at low latitude sites40,47,48. As ecosystem engineers49 mussels in particular have a large influence on the stability and biodiversity of the intertidal community and so quantifying their survival and physiological performance has significant ecosystem-level consequences50,51.
Methods
We used biomimetic loggers to estimate temperatures of the mussels Mytilus californianus (West coast of North America), M. edulis and Geukensia demissa (East coast of North America), M. chilensis (Chile), Perna perna (South Africa) and P. canaliculus (New Zealand). We also deployed unmodified commercial loggers directly on rock surfaces at multiple sites (Australia, Ireland, Mexico, Scotland, U.K., U.S.) that recorded temperatures relevant to barnacles, newly settled mussels and other organisms that are sufficiently small that their temperatures mirror those of the underlying rock52.
Each biomimetic sensor (‘Robomussel’; Fig. 1) consisted of either a commercially-available TidbiT logger (TB132-20+50 and UTB1-001; Onset Computer Corporation, Pocasset, MA) encased in black-coloured polyester resin (Evercoat Premium Marine Resin, Illinois Tool Works, Inc.), or a real mussel shell filled with silicone and encasing a Tidbit or a Thermochron iButton logger (DS1922L-F5; Maxim Integrated, San Jose, California). Both instruments are factory calibrated: Tidbit loggers have a reported accuracy of 0.21 °C and a stability (drift) of 0.1 °C per year (http://www.onsetcomp.com/products/data-loggers/utbi-001) and ibuttons have an accuracy of 0.5 °C (https://datasheets.maximintegrated.com/en/ds/DS1922L-DS1922T.pdf); the drift is reported by the manufacturer to be negligible, especially when compared to the ~2 °C accuracy of the biomimic loggers (see Technical Validation below). Because of loss due to waves, each logger was typically used for only 2–3 years. Details on logger designs and field tests are described in detail in previous publications44,45,53. In brief, logger thermal characteristics were calculated using empirical measurements of shell and tissue mass against length from adult Mytilus californianus collected on the west coast of North America. In addition to morphology (which determines convective heat flux) and colour (which affects solar heat load), the primary consideration is the maintenance of thermal inertia (the tendency of an object to resist temperature change as a function of external forcing). Mass/length relationships were combined with measurements of the specific heat capacity of shell and tissue to estimate total thermal inertia as a function of size45. This was then compared to the thermal mass of polyester resin mussels of different lengths. The point where the two curves intersect is~8 cm shell length; this was the size of the epoxy loggers. Silicone molds were cast from a representative 8 cm mussel, and were in turn used to pour two-part polyester resin (Evercoat) around the commercial TidbiT logger.
In some cases, iButton loggers were encased in ~8 cm mussel shells filled with silicone, which has a mass*specific heat similar to that of water. Comparisons of these instruments against adjacent mussels showed that silicone-filled shells recorded temperatures within ~1 °C of living animals54. However, these loggers were considerably less durable and required more frequent maintenance (~bimonthly) than epoxy mussels (every 6–10 months), and so were used only infrequently at most sites. At some sites where the targeted mussel species is smaller (e.g., M. edulis in the Gulf of Maine), we used 4 cm mussel shells. Loggers of differing size were never used at the same site, and are distinguished from one another in the database. Nevertheless, any direct comparison between data collected by loggers of different sizes should be made with caution, as size can affect mussel temperature by several degrees55.
Robomussels were deployed primarily on hard rock substrate, in growth position (posterior upward) in intact beds using Z-spar splash zone epoxy putty (Fig. 1). Care was taken to ensure that the logger was completely surrounded by other mussels, as tests showed that loggers deployed as solitary individuals tended to yield anomalously high readings. On the east coast of North America, loggers were also deployed at soft sediment (marsh) sites in mud substrate by attaching the loggers to dowel rod.
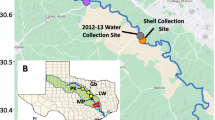
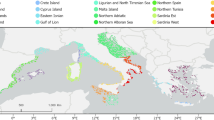
Deployment began in 1998 at the Hopkins Marine Station in Pacific Grove, California54, and was expanded to other sites beginning in 2000 (Table 1 (available online only), Fig. 2). Total deployment time varied by location, ranging from less than a year to almost 18 years (average deployment time of 4 years). The number of loggers deployed and lost due to wave dislodgement also varied at each site, but a standard protocol was to deploy at least 3 loggers in the middle of mussel beds on horizontal, unshaded surfaces. At most sites, loggers were deployed at the upper edge of the mussel bed (‘upper’), half way between the upper and mid levels (‘upper mid’), mid level (‘mid’), half way between the mid and lower edge of the bed (‘lower mid’) and at the bottom of the mussel bed (‘lower’).
Loggers were programmed to record at intervals of 10–30 min and left in the field for periods up to 9 months before they were removed for downloading, and replaced with another logger. Every effort was made to place this new logger in precisely the same position in the bed as the logger being retrieved. All logger clock times were set to GMT. In the U.S., the absolute tidal elevation (height above chart datum) was measured with a Trimble R8 GNSS GPS system capable of sub-cm resolution. Temperature records were also used to record wave swash by comparing sudden drops in temperature (an indication of first wave splash following exposure at low tide) against predicted tidal elevations. The measurements of ‘Effective Shore Level’ can subsequently be compared against buoy records of significant wave height in order to estimate wave splash as a function of nearshore wave height at each site56,57.
Code availability
Code written in R58 was used to trim data recorded by each logger before and after deployment. A separate software program (SiteParser) is also available on the Northeastern website to determine the incidence of wave splash56,57. This is accomplished by comparing rapid (user-defined) drops in temperature, indicative of the return of the tide, against predicted (Xtide software, www.flaterco.com/xtide) or measured (tidesandcurrents.noaa.gov) tide height for each site. By comparing these measurements against measured logger tidal elevations, it is possible to calculate the ‘effective shore level’ of a logger as a function of nearshore wave height56. This also provides a method of dividing logger temperatures into aerial and submerged records. Notably, the choice of temperature drop determines both the accuracy of the division between aerial and submerged records, as well as the total amount of data available. Specifically, the choice of a larger temperature drop tends to increase certainty as to temperature divisions, but can restrict the amount of data to days when such drops are observed. For this reason, the database provides data that have not been analyzed in this manner, but instead provides tools for the user to do so. A link to the open source SiteParser software program is provided on the Northeastern database website, along with links to all metadata including (when available) logger elevations.
Data Records
Data from all loggers are archived in two databases. The first is a searchable database maintained by Northeastern University (www.northeastern.edu/helmuthlab/Research/Database.html) and provides unrestricted access to data as well as to associated links such as the SiteParser software described above. Metadata for each microsite are included as a downloadable spreadsheet, which includes, for each site: Country, Region, Site name, and GPS coordinates (Table 1 (available online only)). The metadata file also includes information specific to each microsite, including: Biomimic logger type (unmodified ibutton, unmodified TidBit, epoxy [8 cm] mussel logger, shell (silicone-filled) mussel logger [4 or 8 cm length]), Substrate (rocky, muddy, tidepool), Tidal elevation zone (low, lower mid, mid, upper mid, or upper), Wave exposure (protected or exposed), and Start and end dates (Table 2). At the Northeastern website, data can be viewed and downloaded using a series of drop-down menus (Fig. 3). Given the range of selections, the database provides the range of dates over which data meeting those criteria are available (this information is also included in the metadata file). Data from each logger can be downloaded as raw data, as well as daily, monthly or annual maxima, minima and averages. Note that data include both aerial and submerged temperatures, but raw data can be parsed using the software provided. In instances where multiple microsites meet the selected criteria, the program takes the average at each time point from the maximum number of loggers available before calculating summary statistics. Data from all microsites can be downloaded as raw data to avoid this averaging procedure.
Raw data in text file format as well as associated metadata are also archived in a public repository (Data Citation 1). Files are organized in to a series of subfolders organized by Country, Region and Site (Table 1 (available online only)). Metadata identical to those available at the Northeastern site are also included as a downloadable file. Each data file contains information specific to the microsite in its header, and follows a 10 letter/6 number naming convention as follows: BM (indicating biomimetic logger database); Logger type (RM for mussel loggers [‘Robomussels’] or RB for unmodified loggers [‘Robobarnacles’]); 6 letter site code (Table 1 (available online only); Country, Region, Site); two-digit microsite ID and four digit Year.
Technical Validation
Comparisons of logger temperatures against tissue temperatures of adjacent live mussels made using thermocouples are presented in four publications44,45,54,59. The first compared temperatures recorded by a thermistor with the tip embedded in a silicone-filled shell against point measurements made from adjacent mussels in the field in Pacific Grove, California and found an average difference of ~0.75 °C (ref. 54). The second involved a more comprehensive set of tests of epoxy (polyester) loggers in both the field and in a wind tunnel fitted with a heat lamp45. In the laboratory experiments, the average difference between loggers and live mussels in artificial beds was ~2.2 °C (ref. 45). Notably, the average difference between live mussels and unmodified loggers (TidbiTs) in the same experiment was 14.6 °C. Field-tests yielded similar results, with an average error of 2.7 °C between robomussels and live mussels45. A follow-up study with additional laboratory tests over a wider range of temperatures (10–50 °C) reported a Root Mean Square Error (RMSE) of 3.84 °C with a correlation coefficient of 0.89 between loggers and live mussels, with a bias of 0.8 °C where loggers tended to overestimate temperatures slightly under extreme conditions44. Finally, iButton loggers placed in the middle of silicone-filled Geukensia demissa shells were tested in a wind tunnel in artificial beds under a range of wind speeds; results showed average differences of ~1.0–1.5 °C (ref. 59).
Usage Notes
Portions of the logger data presented here have been used in multiple field studies, and have provided context for laboratory studies. At small scales, biomimetic loggers (both loggers that we deployed as well as similar loggers made by other researchers) have been used to record differences in temperature among microhabitats (shaded and unshaded surfaces) and tidal elevations (Fig. 4) and the results compared to measurements of biochemical indicators of stress such as heat shock proteins54,60, gene expression61, reproductive condition62, and to the fine-scale distribution of native and non-native species63. At biogeographic scales, robomussels have been used to document thermal mosaics across large latitudinal gradients40,48 (Fig. 5) and the results related to patterns of mortality64, physiological stress65–67 and growth68,69, as well as interspecific differences in physiological stress39 and geographic distribution70. Measurements from mussel biomimetics have been used to test heat budget models that estimate animal temperature using data from weather stations and satellites71–73. Robomussels have also been used as part of controlled laboratory experiments that strive to replicate realistic field conditions37,74. Finally robomussel data can be used to estimate wave splash and water temperature56,57, although in this regard they do not present a major advantage over unmodified loggers.
Additional Information
How to cite this article: Helmuth, B. et al. Long-term, high frequency in situ measurements of intertidal mussel bed temperatures using biomimetic sensors. Sci. Data 3:160087 doi: 10.1038/sdata.2016.87 (2016).
References
References
Rapacciuolo, G. et al. Beyond a warming fingerprint: individualistic biogeographic responses to heterogeneous climate change in California. Global Change Biol. 20, 2841–2855 (2014).
Chown, S. L., Gaston, K. J. & Robinson, D. Macrophysiology: large-scale patterns in physiological traits and their ecological implications. Func. Ecol 18, 159–167 (2004).
Pörtner, H. O. & Farrell, A. P. Physiology and climate change. Nature 322, 690–692 (2008).
Kearney, M. Habitat, environment and niche: what are we modelling? Oikos 115, 186–191 (2006).
Jansen, J. M. et al. Geographic and seasonal patterns and limits on the adaptive response to temperature of European Mytilus spp. and Macoma balthica populations. Oecologia 154, 23–34 (2007).
Kroeker, K. J. et al. The role of temperature in determining species’ vulnerability to ocean acidification: A case study using Mytilus galloprovincialis. PLoS ONE 9, E100353 (2014).
Monaco, C. J. & Helmuth, B. Tipping points, thresholds, and the keystone role of physiology in marine climate change research. Adv. Mar. Biol. 60, 123–160 (2011).
Queirós, A. M. et al. Scaling up experimental ocean acidification and warming research: from individuals to the ecosystem. Global Change Biol. 21, 130–143 (2015).
Seebacher, F. & Franklin, C. E. Determining environmental causes of biological effects: the need for a mechanistic physiological dimension in conservation biology. Philosophical Transactions of the Royal Society B 367, 1607–1614 (2012).
Somero, G. N. The physiology of climate change: how potentials for acclimatization and genetic adaptation will determine ‘winners’ and ‘losers’. J. Exp. Biol. 213, 912–920 (2010).
Kelley, A. L. The role thermal physiology plays in species invasion. Conservation Physiology 2, cou045 (2014).
Lockwood, B. L. & Somero, G. N. Invasive and native blue mussels (genus Mytilus) on the California coast: The role of physiology in a biological invasion. J. Exp. Mar. Biol. Ecol. 400, 167–174 (2011).
Pörtner, H. O. Climate variations and the physiological basis of temperature dependent biogeography: systemic to molecular hierarchy of thermal tolerance in animals. Comparative Biochemistry and Physiology Part A 132, 739–761 (2002).
Woodin, S. A., Hilbish, T. J., Helmuth, B., Jones, S. J. & Wethey, D. S. Climate change, species distribution models, and physiological performance metrics: predicting when biogeographic models are likely to fail. Ecology and Evolution 3, 3334–3346 (2013).
Wernberg, T. et al. Impacts of climate change in a global hotspot for temperate marine biodiversity and ocean warming. J. Exp. Mar. Biol. Ecol 400, 7–16 (2011).
Mumby, P. J. et al. Revisiting climate thresholds and ecosystem collapse. Frontiers in Ecology and the Environment 9, 94–96 (2011).
Brown, C. J. et al. Quantitative approaches in climate change ecology. Global Change Biol 17, 3697–3713 (2011).
Somero, G. N. The physiology of global change: Linking patterns to mechanisms. Annual Review of Marine Science 4, 39–61 (2012).
Wethey, D. S. et al. Response of intertidal populations to climate: Effects of extreme events versus long term change. J. Exp. Mar. Biol. Ecol. 400, 132–144 (2011).
Thomas, Y. et al. Modelling spatio-temporal variability of Mytilus edulis (L.) by forcing a dynamic energy budget model with satellite-derived environmental data. J. Sea Res. 66, 308–317 (2011).
Sarà, G. et al. Growth and reproductive simulation of candidate shellfish species at fish cages in the Southern Mediterranean: Dynamic Energy Budget (DEB) modelling for integrated multi-trophic aquaculture. Aquaculture 324, 259–266 (2012).
Vasseur, D. A. et al. Increased temperature variation poses a greater risk to species than climate warming. Proceedings of the Royal Society B 281, 20132612 (2014).
Helmuth, B. et al. Beyond long-term averages: Making biological sense of a rapidly changing world. Climate Change Responses 1, 10–20 (2014).
Kingsolver, J. G. & Woods, H. A. Beyond thermal performance curves: Modeling time-dependent effects of thermal stress on ectotherm growth rates. Amer. Nat 187, 283–294 (2016).
Lima, F. P. & Wethey, D. S. Three decades of high-resolution coastal sea surface temperatures reveal more than warming. Nat. Commun 3, 704 (2012).
Kearney, M. R., Isaac, A. P. & Porter, W. P. microclim: Global estimates of hourly microclimate based on long-term monthly climate averages. Scientific Data 1, 140006 (2014).
Helmuth, B. et al. Organismal climatology: analyzing environmental variability at scales relevant to physiological stress. J. Exp. Biol. 213, 995–1003 (2010).
Pawar, S., Dell, A. I. & Savage, V. M. in Aquatic Functional Biodiversity: An ecological and evolutionary perspective (eds Belgrano, A., Woodward, G. & Jacob, U. ) 3–36 (2015).
Selkoe, K. A. et al. Principles for managing marine ecosystems prone to tipping points. Ecosystem Health and Sustainability 1, 17 (2015).
Kearney, M. & Porter, W. Mechanistic niche modelling: combining physiological and spatial data to predict species ranges. Ecol. Letters 12, 334–350 (2009).
Jimenez, I. M., Kühl, M., Larkum, A. W. D. & Ralph, P. J. Heat budget and thermal microenvironment of shallow-water corals: Do massive corals get warmer than branching corals? Limnol. Oceanogr. 53, 1548–1561 (2008).
Marshall, D. J., McQuaid, C. D. & Williams, G. A. Non-climatic thermal adaptation: implications for species' responses to climate warming. Biology Letters 6, 669–673 (2010).
Mislan, K. A. S. & Wethey, D. S. Gridded meteorological data as a resource for mechanistic macroecology in coastal environments. Ecol. Appl. 21, 2678–2690 (2011).
Mislan, K. A. S., Helmuth, B. & Wethey, D. S. Geographical variation in climatic sensitivity of intertidal mussel zonation. Global Ecology and Biogeography 23, 744–756 (2014).
Potter, K. A., Woods, H. A. & Pincebourde, S. Microclimatic challenges in global change biology. Global Change Biol. 19, 2932–2939 (2013).
Sunday, J. M. et al. Thermal-safety margins and the necessity of thermoregulatory behavior across latitude and elevation. Proc. Nat. Acad. Sci 111, 5610–5615 (2014).
Nicastro, K. R., Zardi, G. I., McQuaid, C. D., Pearson, G. A. & Serrão, E. A. Love thy neighbour: Group properties of gaping behaviour in mussel aggregations. PLoS ONE 7, e47382 (2012).
Miller, L. P. & Denny, M. W. Importance of behavior and morphological traits for controlling body temperature in littorinid snails. Biol. Bull. 220, 209–223 (2011).
Broitman, B. R., Szathmary, P. L., Mislan, K. A. S., Blanchette, C. A. & Helmuth, B. Predator-prey interactions under climate change: the importance of habitat vs body temperature. Oikos 118, 219–224 (2009).
Helmuth, B. S. et al. Climate change and latitudinal patterns of intertidal thermal stress. Science 298, 1015–1017 (2002).
Kearney, M., Simpson, S. J., Raubenheimer, D. & Helmuth, B. Modelling the ecological niche from functional traits. Philosophical Transactions of the Royal Society B 365, 3469–3483 (2010).
Dzialowski, E. M. Use of operative temperature and standard operative temperature models in thermal biology. J. Thermal Biol. 30, 317–334 (2005).
Lathlean, J. A., Ayre, D. J., Coleman, R. A. & Minchinton, T. E. Using biomimetic loggers to measure interspecific and microhabitat variation in body temperatures of rocky intertidal invertebrates. Mar. Freshwater Res. 66, 86–94 (2014).
Lima, F. P. et al. in Advances in Biomimetics, 499–522 (INTECH publishing, 2011).
Fitzhenry, T., Halpin, P. M. & Helmuth, B. Testing the effects of wave exposure, site, and behavior on intertidal mussel body temperatures: Applications and limits of temperature logger design. Mar. Biol. 145, 339–349 (2004).
Seabra, R., Wethey, D. S., Santos, A. M. & Lima, F. P. Understanding complex biogeographic responses to climate change. Scientific Reports 5, 12930 (2015).
Mislan, K. A. S., Wethey, D. S. & Helmuth, B. When to worry about the weather: role of tidal cycle in determining patterns of risk in intertidal ecosystems. Global Change Biol. 15, 3056–3065 (2009).
Helmuth, B. et al. Mosaic patterns of thermal stress in the rocky intertidal zone: implications for climate change. Ecol. Monogr. 76, 461–479 (2006).
Gutiérrez, J. L., Jones, C. G., Strayer, D. L. & Iribarne, O. O. Mollusks as ecosystem engineers: the role of shell production in aquatic habitats. Oikos 101, 79–90 (2003).
Smith, J. R., Fong, P. & Ambrose, R. F. Dramatic declines in mussel bed community diversity: response to climate change? Ecology 87, 1153–1161 (2006).
Paine, R. T. Intertidal community structure: Experimental studies on the relationship between a dominant competitor and its principal predator. Oecologia 15, 93–120 (1974).
Wethey, D. S. Biogeography, competition, and microclimate: the barnacle Chthamalus fragilis in New England. Int. Comp. Biol 42, 872–880 (2002).
Harley, C. D. G. & Helmuth, B. S. T. Spatial variation in invertebrate upper limits, thermal stress, and effective tidal height. Amer. Zool. 41, 1466 (2001).
Helmuth, B. S. T. & Hofmann, G. E. Microhabitats, thermal heterogeneity, and patterns of physiological stress in the rocky intertidal zone. Biol. Bull. 201, 374–384 (2001).
Helmuth, B. S. T. Intertidal mussel microclimates: Predicting the body temperature of a sessile invertebrate. Ecol. Monogr. 68, 51–74 (1998).
Gilman, S. E. et al. Evaluation of effective shore level as a method of characterizing intertidal wave exposure regimes. Limnology and Oceanography: Methods 4, 448–457 (2006).
Harley, C. D. G. & Helmuth, B. S. T. Local and regional scale effects of wave exposure, thermal stress, and absolute vs. effective shore level on patterns of intertidal zonation. Limnol. Oceanogr. 48, 1498–1508 (2003).
R Core Development Team . A language and environment for statistical computing. R Foundation for Statistical Computing, (2016).
Jost, J. & Helmuth, B. Morphological and ecological determinants of body temperature of Geukensia demissa, the Atlantic ribbed Mussel, and their effects on mussel mortality. Biol. Bull. 213, 141–151 (2007).
Petes, L. E., Mouchka, M. E., Milston-Clements, R. H., Momoda, T. S. & Menge, B. A. Effects of environmental stress on intertidal mussels and their sea star predators. Oecologia 156, 671–680 (2008).
Gracey, A. Y. et al. Rhythms of gene expression in a fluctuating intertidal environment. Current Biology 18, 1–7 (2008).
Petes, L. E., Menge, B. A. & Harris, A. L. Intertidal mussels exhibit energetic trade-offs between reproduction and stress resistance. Ecol. Monogr. 78, 387–402 (2008).
Schneider, K. R. & Helmuth, B. Spatial variability in habitat temperature may drive patterns of selection between an invasive and native mussel species. Mar. Ecol. Prog. Ser. 339, 157–167 (2007).
Zardi, G., Nicastro, K., McQuaid, C. D., Hancke, L. & Helmuth, B. The combination of selection and dispersal helps explain genetic structure in intertidal mussels. Oecologia 165, 947–958 (2011).
Place, S. P., O'Donnell, M. J. & Hofmann, G. E. Gene expression in the intertidal mussel Mytilus californianus: physiological response to environmental factors on a biogeographic scale. Mar. Ecol. Prog. Ser. 356, 1–14 (2008).
Logan, C. A., Kost, L. E. & Somero, G. N. Latitudinal differences in Mytilus californianus thermal physiology. Mar. Ecol. Prog. Ser. 450, 93–105 (2012).
Tagliarolo, M. & McQuaid, C. D. Field measurements indicate unexpected, serious underestimation of mussel heart rates and thermal tolerance by laboratory studies. PLoS ONE 11, e0146341 (2016).
Kroeker, K. J. et al. Interacting environmental mosaics drive geographic variation in mussel performance and species interactions. Ecol. Letters 19, 771–779 (2016).
Blanchette, C. A., Helmuth, B. & Gaines, S. D. Spatial patterns of growth in the mussel, Mytilus californianus, across a major oceanographic and biogeographic boundary at Point Conception, California, USA. J. Exp. Mar. Biol. Ecol 340, 126–148 (2007).
Tagliarolo, M. & McQuaid, C. D. Sub-lethal and sub-specific temperature effects are better predictors of mussel distribution than thermal tolerance. Mar. Ecol. Prog. Ser. 535, 145–159 (2015).
Gilman, S. E., Wethey, D. S. & Helmuth, B. Variation in the sensitivity of organismal body temperature to climate change over local and geographic scales. Proc. Nat. Acad. Sci 103, 9560–9565 (2006).
Helmuth, B. et al. Hidden signals of climate change in intertidal ecosystems: what (not) to expect when you are expecting. J. Exp. Mar. Biol. Ecol 400, 191–199 (2011).
Wethey, D. S., Brin, L. D., Helmuth, B. & Mislan, K. A. S. Predicting intertidal organism temperatures with modified land surface models. Ecological Modelling 222, 3568–3576 (2011).
Schneider, K. R. Heat stress in the intertidal: comparing survival and growth of an invasive and native mussel under a variety of thermal conditions. Biol. Bull. 215, 253–264 (2008).
Data Citations
Helmuth, B. Dryad https://doi.org/10.5061/dryad.6n8kf (2016)
Acknowledgements
The collection of robomussel data has primarily been supported by grants from NSF (IBN-9985878, OCE-0323364, OCE-0926581 and IBN- 1557868); NASA (NNG04GE43G, NNX07AF20G, NNX11AP77G), NOAA (NA04NOS4780264), the National Geographic Society to B.H., N.S.F. IBN-1557868 to B.H. and M.Z., and the David and Lucile Packard Foundation to PISCO, the Partnership for Interdisciplinary Studies of Coastal Oceans (B.M., G.E.H., C.B.) and by the Chilean Ministry of Economics (MINECON NC120086) to B.B. This work includes research supported by the South African Research Chairs Initiative of the Department of Science and Technology and the National Research Foundation (C.D.M.). This is publication number 468 of the Partnership for Interdisciplinary Studies of Coastal Oceans (PISCO) and 343 of the Marine Science Center, Northeastern University. Numerous people helped to collect loggers over the last 18 years, and we are very grateful for their help: Tameka Breland, Susan Bolte, Nick Burnett, Cari Cardoni, Ryan Craig, Sean Craig, Colette Dryden, Tara Fitzhenry, Kristi Gardner, Shawn Gerrity, Chris Haas, Steven Hawkins, Maxine Henry, Lindsay Hunter, Scott Johnson, Nicole Kish, Ruth Milston-Clements, Gayle Murphy, Tom O’Keefe, Susanne Pender, Cathy Pfister, Camryn Pennington, Brittany Poirson, Sylvain Pincebourde, Eric Sanford and Morgan Timmerman-Helmuth. The findings and conclusions in this article are those of the authors and do not necessarily represent the views of any of the authors' institutions or agencies.
Author information
Authors and Affiliations
Contributions
Helmuth led the study for the entire duration of the record. Logistics of deployment, retrieval and data management were, at various times, overseen by Choi, Matzelle, Szathmary, Gilman, Mislan, Yamane, Tockstein and Strickland. All other authors managed instrument deployment and retrieval at field sites: Massachusetts (Choi, Torossian, Morello); Scotland (Burrows); Ireland (Power, Gosling); U.K. (Hilbish, Mieszkowska); British Columbia and Washington (Nishizaki, Carrington, Harley); Oregon (Menge, Petes, Foley, Johnson, Poole, Noble, Richmond, Robart, Robinson, Sapp); California (Denny, Mach, Miller, O′Donnell, Sones, Hilbish, Harley, Hofmann, Zippay, Blanchette, Macfarlan); Baja California (Carpizo-Ituarte, Ruttenberg, Peña Mejía); Chile (Broitman); New Zealand (Mislan, Petes, Ross, Menge) and South Africa (McQuaid, Lathlean, Monaco, Nicastro, Zardi). Helmuth led the writing of the manuscript and all authors contributed to editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
ISA-Tab metadata
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0 Metadata associated with this Data Descriptor is available at http://www.nature.com/sdata/ and is released under the CC0 waiver to maximize reuse.
About this article
Cite this article
Helmuth, B., Choi, F., Matzelle, A. et al. Long-term, high frequency in situ measurements of intertidal mussel bed temperatures using biomimetic sensors. Sci Data 3, 160087 (2016). https://doi.org/10.1038/sdata.2016.87
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2016.87
This article is cited by
-
Transcriptome wide analyses reveal intraspecific diversity in thermal stress responses of a dominant habitat‐forming species
Scientific Reports (2023)
-
Loss of transcriptional plasticity but sustained adaptive capacity after adaptation to global change conditions in a marine copepod
Nature Communications (2022)
-
Coastal upwelling generates cryptic temperature refugia
Scientific Reports (2022)
-
High resolution spatiotemporal patterns of seawater temperatures across the Belize Mesoamerican Barrier Reef
Scientific Data (2020)
-
Cycles of heat and aerial-exposure induce changes in the transcriptome related to cell regulation and metabolism in Mytilus californianus
Marine Biology (2020)