Abstract
Although the human olfactory system is capable of discriminating a vast number of odors, we do not currently understand what chemical features are encoded by olfactory receptors. In large part this is due to a paucity of data in a search space covering the interactions of hundreds of receptors with billions of odorous molecules. Of the approximately 400 intact human odorant receptors, only 10% have a published ligand. Here we used a heterologous luciferase assay to screen 73 odorants against a clone library of 511 human olfactory receptors. This dataset will allow other researchers to interrogate the combinatorial nature of olfactory coding.
Design Type(s) | cellular process design |
Measurement Type(s) | receptor activity |
Technology Type(s) | Dual-Luciferase Reporter Assay System |
Factor Type(s) | gene list |
Sample Characteristic(s) | Homo sapiens • Mus musculus |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
Previous functional analysis of olfactory receptors (ORs) in olfactory neurons and in heterologous cells found that different odorants are recognized by unique, but overlapping ensembles of ORs1–4. These findings suggest that specific patterns of ORs activated by an odorant code for the odorant’s identity, but there are few, if any, explicit predictions relating OR activity patterns to olfactory perception.
Matching mammalian ORs to ligands has seen limited success, and the picture is even worse when considering human ORs; ligands have been published for only 49 of the approximately 400 intact human ORs5–21. This lack of data is a critical bottleneck in the field; matching ligands to ORs is critical for understanding the olfactory system at all levels and is essential for building viable models of olfaction. The characterization of OR responses to ligands in the empty neuron system of Drosophila melanogaster22 has allowed researchers in the field to choose rationally diverse odorant sets23 and specifically manipulate subpopulations of ORs to dissect olfactory coding24,25. Extending this idea by matching odorants to human ORs has the added advantage that humans can directly communicate their perception of odor intensity, pleasantness, and quality.
In addition, understanding the role of a single OR in olfactory perception allows us to look at evolutionary changes in OR genes in a new light. For example, the knowledge that Tas1r2 is a pseudogene in seven of twelve species in the order Carnivora26 is difficult to interpret in isolation. The knowledge that Tas1r2 is the primary mediator of sweet taste in mice, however, suggests that carnivores do not need to taste sweet and therefore there is no selective pressure on the gene. As several genome sequencing projects are examining both genetic variation within humans27 and across species28, understanding the role of OR genes in olfactory perception becomes crucial to the interpretation of how and why genetic changes occur over the course of evolution.
In a recent manuscript we conducted a high-throughput screen of 511 human odorant receptors against 73 odorants15. The resulting screen identified agonists for 27 odorant receptors, including 18 that were previously orphan receptors. We went on to characterize how genetic variation in these receptors alters both in vitro responses and influences olfactory perception. In this manuscript we present the full screening data to permit wider reuse and reanalysis.
In summary, this dataset addresses a major bottleneck in the field, namely how the physical stimulus in olfaction is transduced into receptor responses. In addition, the G-protein coupled receptor class accounts for approximately 50% of therapeutic drug targets29. The ORs, being GPCRs, offer the opportunity to examine the strategies employed by this receptor class to recognize a wide variety of ligand structural features and thus will provide insight into fundamental principles of ligand recognition by GPCRs. Matching odorants to ORs will provide a valuable resource to the field and allow more specific explorations of links between odor, behavior and ecology.
Methods
These methods are expanded from descriptions in our previous work15.
Cloning
OR open reading frames were amplified from genomic DNA using Phusion polymerase and subcloned into pCI expression vectors (Promega) containing the first 20 residues of human rhodopsin (Rho tag). Human ORs were amplified from the pooled genomic DNA of 20 participants from the International Hapmap Consortium, while mouse ORs were amplified from the genomic DNA of C57/BL6 mice. The sequences of the cloned receptors were verified by sequencing (3100 Genetic Analyzer, Applied Biosystems). Clones that were present in the 1000 Genomes Project, but not cloned from our pooled genomic DNA sample, were created using an overlap extension polymerase chain reaction protocol30.
Luciferase assay
The Dual-Glo Luciferase Assay System (Promega) was used to measure receptor responses as previously described31. Hana3A cells were transfected with 5 ng/well of RTP1S32, 5 ng/well of pRL-SV40, 10 ng/well of CRE-luciferase, 2.5 ng/well of M3 (ref. 33), and 5 ng/well of odorant receptor. 1 M odorant stocks are diluted in DMSO. 24 hours following transfection, transfection media was removed and replaced with the appropriate concentration of odor diluted from the 1 M stocks in CD293 (Gibco). Four hours following odor stimulation luminescence was measured using a Polarstar Optima plate reader (BMG). All luminescence values were divided by the Renilla Luciferase activity to control for transfection efficiency in a given well. Data were analyzed with Microsoft Excel, GraphPad Prism 4, and MATLAB.
Primary screen design
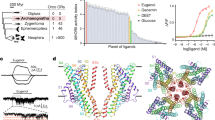
Our screen design is outlined in Figure 1. In the primary screen we stimulated 511 human ORs with 73 odorants used in previous psychophysical testing12,34. We applied the majority of odorants at a concentration of 100 μM. All plates in the primary screen included 85 test wells, five broadly-tuned odorant receptors (Olfr1079, OR2W1, Olfr1377, Olfr73, Olfr1341), and six wells transfected with Oflr544 which served as a standard. Of the six wells transfected with Olfr544, three were challenged with the diluent (CD293) and three were challenged with 10 μM of a known ligand for Olfr544 (nonanedioic acid). Each screening run consisted of twelve plates where each of the 96-wells were transfected with the same set of receptors. One plate had no odor in all test wells and served as a baseline. The other eleven plates were each challenged with a different test odor.
This figure was reprinted from our previous publication15, where it was included as Supplementary Figure 1.
Secondary screen design
To rank hits from the Primary Screen we standardized each plate, setting the mean Olfr544 response to nonanedioic acid minus the mean Olfr544 response to the no-odor control to a value of 1. We then subtracted the baseline response for each receptor from the no-odor plate from the response to the odor challenge and ranked the resulting values. We selected the top 5% of odorant/receptor pairs from the primary screen, although not more than the top ten ligands for a given receptor. We then performed a secondary screen in which each odorant receptor was tested against a no-odor control as well as 1, 10 and 100 μM of odor. Each comparison was performed in triplicate, where each measure was collected from separate wells, but each well contained cells from the same parent plate of cells. Note that we began the secondary screen before completion of the entire primary screen, so some odor/receptor combinations outside of the overall top 5% were tested.
Dose-response design
We then constructed dose-response curves using concentrations ranging from 10 nM to 10 mM for the odor/receptor pairs that were significantly different from baseline in the Secondary Screen. Each odorant receptor-odorant dose was tested in triplicate, where each measure was collected from separate wells, but each well contains cells from the same parent plate of cells. A vector-only control was included for each odorant. We fit the data to a sigmoidal curve. We counted an odorant as an agonist if the 95% confidence intervals of the top and bottom parameters did not overlap, the standard deviation of the fitted log EC50 was less than 1 log unit, and the extra sums-of-squares test confirmed that the odorant activated the receptor significantly more than the control, which was transfected with an empty vector. This data identified 25 odorant receptors with a significant response to at least one agonist15 (Figures 2 and 3).
The responses of cells transfected with either a plasmid encoding the indicated odorant receptor or an empty vector to the indicated odorants. Responses have been normalized such that each receptor has a minimum response of zero and a maximum response of one. Error bars, s.e.m. over three replicates. Abbreviations for the odorants are as follows: +CAR are shown, (+)-carvone; LIN, linalool; GA, geranyl acetate; COUM, coumarin; OTHI, octanethiol; C3HEX, cis-3-hexen-1-ol; EUG, eugenol; EUGME, eugenol methyl ether; ANIS, anisaldehyde; ANDI, 4,16-androstadien-3-one; AND, 5α-androst-16-en-3-one; DMHDMF, caramel furanone; +MEN, (+)-menthol; 3PPP, 3-phenyl propyl propionate; VAN, vanillin; LYR, lyral; 2EF, 2-ethyl fenchol; IVA, isovaleric acid; APA, allyl phenyl acetate. This figure was modified from our previous publication15, where it was included as Figure 1.
The responses of cells transfected with either a plasmid encoding the indicated odorant receptor or an empty vector to the indicated odorants. Error bars, s.e.m. over three replicates. Abbreviations for the odorants are as follows: +CAR are shown, (+)-carvone; LIN, linalool; GA, geranyl acetate; COUM, coumarin; OTHI, octanethiol; C3HEX, cis-3-hexen-1-ol; EUG, eugenol; EUGME, eugenol methyl ether; ANIS, anisaldehyde; ANDI, 4,16-androstadien-3-one; AND, 5α-androst-16-en-3-one; DMHDMF, caramel furanone; +MEN, (+)-menthol; 3PPP, 3-phenyl propyl propionate; VAN, vanillin; LYR, lyral; 2EF, 2-ethyl fenchol; IVA, isovaleric acid; APA, allyl phenyl acetate. This figure was modified from our previous publication15, where it was included as Figure 1.
Data Records
The data for this manuscript have been deposited in figshare (Data Citation 1). A summary of the clones tested in each phase of the screen is presented in Supplementary Table 1.
Data record 1—primary screen
The raw screening results are presented in a tab-separated values file (Data Citation 1). Each row represents an experiment from a single well.
Plate. A unique ID for a 96-well plate on a given date.
Well. A number assigned to each well of the 96-well plate. The wells are sequentially numbered with the upper-leftmost well assigned as 1 and the lower-leftmost well assigned as 85 (see Figure 4).
Concentration. The concentration of the odorant applied in uM. A ‘9999’ indicates no odor was applied (DMSO was diluted 1:10,000 in CD293).
Luc. The number of photons counted by the plate reader when the well was treated with the luciferase substrate. This is the cAMP reporter, and therefore correlates with receptor responses to odorants.
RL. The number of photons counted by the plate reader when the well was treated with the Renilla luciferase substrate. This is the constitutively active reporter, which serves as a control for cell death and transfection efficiency.
OR. A unique ID for each olfactory receptor clone.
Odor. A unique ID for the odorant applied to the well.
Date. The date the experiment was run in MM/DD/YY format.
Data record 2—secondary screen
The raw screening results are presented in a tab-separated values file (Data Citation 1). Each row represents an experiment from a single well.
Date. The date the experiment was run in MM/DD/YY format.
OR. A unique ID for each olfactory receptor clone.
Odor. A unique ID for the odorant applied to the well.
Concentration. The concentration of the odorant applied in uM. A ‘0’ indicates no odor was applied (CD293 only). Note that rows for the ‘no odor’ condition will contain an ‘Odor’ label to facilitate pairing controls with the matched experiments at other concentrations.
NormalizedLuc. The Luc/RL ratio from each well.
Data record 3—dose-response
The Luc/RL ratios are presented in a tab-separated values file (Data Citation 1). Each row represents an experiment from a single well. The EC50 for each odor/receptor pair that passed this phase of screening is listed in Table 1 (available online only).
Concentration. The molarity applied to the well. Note that the no-odor condition was coded as −12 in this column.
NormLuc. The Luc/RL ratio from each well.
OR. A unique ID for each olfactory receptor clone.
Odor. A unique ID for the odorant applied to the well.
Date. The date the experiment was run in MM/DD/YY format.
Data record 4—receptors
The receptor information is presented in a tab-separated values file (Data Citation 1). Each row represents a single olfactory receptor.
OR. A unique ID for the olfactory receptor, used in Data Records 1–3.
Gene. The gene name of the olfactory receptor encoded in a given plasmid, followed by the amino acid changes from the hg19 reference sequence for the gene. For example, ‘OR6Y1 V252I’ encodes the gene OR6Y1, but while the hg19 reference sequence has a ‘V’ as the 252nd amino acid, this clone encodes an ‘I’ at position 252. Note that some plasmids were cloned from older builds of the genome, which may start at a different methionine than the current model. These differences from h19 reference may not appear here. Please consult the nucleotide sequence for a more thorough description of differences from the reference sequence.
NucleotideSeq. The nucleotide sequence for the olfactory receptor encoded in a given plasmid. Note that the rhodopsin tag is not included in this field.
Data record 5—odors
The odor information is presented in a tab-separated values file (Data Citation 1). Each row represents a single odor. Further synonyms can be found in a file which correlates all of the CIDs in PubChem with submitted synonyms (ftp://ftp.ncbi.nlm.nih.gov/pubchem/Compound/Extras/CID-Synonym-filtered.gz).
Odor. A unique ID for the odorant, used in Data Records 1–3.
CASRegistryNum. The Chemical Abstracts Service number for the odorant, when available.
OdorName. Common name for the odorant applied to the well.
CID. PubChem Compound Identification number, a non-zero integer PubChem accession identifier for a unique chemical structure, when available.
SMILES. Simplified Molecular-Input Line-Entry System string, an ASCII string identifier for a unique chemical structure.
Technical Validation
The screen included two types of negative controls. Cells transfected with each receptor clone were challenged with a no-odor stimulation (CD293 alone) to control for baseline receptor activity. Cells transfected with an empty vector were challenged with all of the tested odorants to control for nonspecific activation. All plates in the primary screen included five broadly-tuned odorant receptors and six wells transfected with Olfr544 (also known as MOR42-3 or S6) which served as a standard. Of the six wells transfected with Olfr544, three were challenged with the diluent (CD293), and three were challenged with 10 μM of nonanedioic acid. Rankings from the primary screen consistently predicted results from later screens (Figure 5). The ultimate validation of this assay is prediction of behaviour, and previous results from similar in vitro assays have been shown to predict human olfactory perception12,15,35,36.
Usage Notes
We have included R37 scripts to facilitate analysis of the data. The included R scripts, Supplementary Files 1 and 2, have been implemented as a hosted Shiny38 application (http://www.monell.org/supplemental_files/jmainland/jm0714) to facilitate browsing the data39–41. The R markdown file, Supplementary File 3, includes code to carry out routine normalization of the primary screen, fit an ANOVA to data from the secondary screen, fit a sigmoid to the dose-response data, and create Figures 2, 3 and 542,43.
Additional information
Table 1 is only available in the online version of this paper.
How to cite this article: Mainland, J. D. et al. Human olfactory receptor responses to odorants. Sci. Data 2:150002 doi: 10.1038/sdata.2015.2 (2015).
References
References
Araneda, R. C., Kini, A. D. & Firestein, S. The molecular receptive range of an odorant receptor. Nat. Neurosci. 3, 1248–1255 (2000).
Katada, S., Hirokawa, T., Oka, Y., Suwa, M. & Touhara, K. Structural basis for a broad but selective ligand spectrum of a mouse olfactory receptor: mapping the odorant-binding site. J. Neurosci. 25, 1806–1815 (2005).
Malnic, B., Hirono, J., Sato, T. & Buck, L. B. Combinatorial receptor codes for odors. Cell 96, 713–723 (1999).
Saito, H., Kubota, M., Roberts, R., Chi, Q. & Matsunami, H. RTP family members induce functional expression of mammalian odorant receptors. Cell 119, 679–691 (2004).
Wetzel, C. H. et al. Specificity and sensitivity of a human olfactory receptor functionally expressed in human embryonic kidney 293 cells and Xenopus Laevis oocytes. J. Neurosci. 19, 7426–7433 (1999).
Spehr, M. et al. Identification of a testicular odorant receptor mediating human sperm chemotaxis. Science 299, 2054–2058 (2003).
Sanz, G., Schlegel, C., Pernollet, J.-C. & Briand, L. Comparison of odorant specificity of two human olfactory receptors from different phylogenetic classes and evidence for antagonism. Chem. Senses 30, 69–80 (2005).
Matarazzo, V. et al. Functional characterization of two human olfactory receptors expressed in the baculovirus Sf9 insect cell system. Chem. Senses 30, 195–207 (2005).
Jacquier, V., Pick, H. & Vogel, H. Characterization of an extended receptive ligand repertoire of the human olfactory receptor OR17-40 comprising structurally related compounds. J. Neurochem. 97, 537–544 (2006).
Neuhaus, E. M., Mashukova, A., Zhang, W., Barbour, J. & Hatt, H. A specific heat shock protein enhances the expression of mammalian olfactory receptor proteins. Chem. Senses 31, 445–452 (2006).
Schmiedeberg, K. et al. Structural determinants of odorant recognition by the human olfactory receptors OR1A1 and OR1A2. J. Struct. Biol. 159, 400–412 (2007).
Keller, A., Zhuang, H., Chi, Q., Vosshall, L. B. & Matsunami, H. Genetic variation in a human odorant receptor alters odour perception. Nature 449, 468–472 (2007).
Menashe, I. et al. Genetic elucidation of human hyperosmia to isovaleric acid. PLoS Biol. 5, e284 (2007).
Saito, H., Chi, Q., Zhuang, H., Matsunami, H. & Mainland, J. D. Odor coding by a Mammalian receptor repertoire. Sci. Signal. 2, ra9 (2009).
Mainland, J. D. et al. The missense of smell: functional variability in the human odorant receptor repertoire. Nat. Neurosci. 17, 114–120 (2014).
Adipietro, K. A., Mainland, J. D. & Matsunami, H. Functional evolution of mammalian odorant receptors. PLoS Genet. 8, e1002821 (2012).
Braun, T., Voland, P., Kunz, L., Prinz, C. & Gratzl, M. Enterochromaffin Cells of the Human Gut: Sensors for Spices and Odorants. Gastroenterology 132, 1890–1901 (2007).
Fujita, Y. et al. Deorphanization of Dresden G protein-coupled receptor for an odorant receptor. J. Recept. Signal Transduct. Res. 27, 323–334 (2007).
Mashukova, A., Spehr, M., Hatt, H. & Neuhaus, E. M. Beta-arrestin2-mediated internalization of mammalian odorant receptors. J. Neurosci. 26, 9902–9912 (2006).
Jaeger, S. R. et al. A mendelian trait for olfactory sensitivity affects odor experience and food selection. Curr. Biol. 23, 1601–1605 (2013).
Busse, D. et al. A Synthetic Sandalwood Odorant Induces Wound Healing Processes in Human Keratinocytes via the Olfactory Receptor OR2AT4. J. Invest. Dermatol. 134, 2823–2832 (2014).
Hallem, E. A & Carlson, J. R. Coding of odors by a receptor repertoire. Cell 125, 143–160 (2006).
Haddad, R. et al. A metric for odorant comparison. Nat. Methods 5, 425–429 (2008).
Olsen, S. R. & Wilson, R. I. Lateral presynaptic inhibition mediates gain control in an olfactory circuit. Nature 452, 956–960 (2008).
Bhandawat, V., Olsen, S. R., Gouwens, N. W., Schlief, M. L. & Wilson, R. I. Sensory processing in the Drosophila antennal lobe increases reliability and separability of ensemble odor representations. Nat. Neurosci. 10, 1474–1482 (2007).
Jiang, P. et al. Major taste loss in carnivorous mammals. Proc. Natl Acad. Sci. USA 109, 4956–4961 (2012).
The 1000 Genomes Project Consortium et al. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Genome 10K Consortium. Genome 10K: a proposal to obtain whole-genome sequence for 10,000 vertebrate species. J. Hered. 100, 659–674 (2009).
Klabunde, T. & Hessler, G. Drug design strategies for targeting G-protein-coupled receptors. ChemBioChem 3, 928–944 (2002).
Heckman, K. L. & Pease, L. R. Gene splicing and mutagenesis by PCR-driven overlap extension. Nat. Protoc. 2, 924–932 (2007).
Zhuang, H. & Matsunami, H. Evaluating cell-surface expression and measuring activation of mammalian odorant receptors in heterologous cells. Nat. Protoc. 3, 1402–1413 (2008).
Zhuang, H. & Matsunami, H. Synergism of accessory factors in functional expression of mammalian odorant receptors. J. Biol. Chem. 282, 15284–15293 (2007).
Li, Y. R. & Matsunami, H. Activation state of the M3 muscarinic acetylcholine receptor modulates mammalian odorant receptor signaling. Sci . Signal. 4, ra1 (2011).
Keller, A., Hempstead, M., Gomez, I. A., Gilbert, A. N. & Vosshall, L. B. An olfactory demography of a diverse metropolitan population. BMC Neurosci. 13, 122 (2012).
McRae, J. F. et al. Genetic variation in the odorant receptor OR2J3 is associated with the ability to detect the “grassy” smelling odor, cis-3-hexen-1-ol. Chem. Senses 37, 585–593 (2012).
McRae, J. F. et al. Identification of regions associated with variation in sensitivity to food-related odors in the human genome. Curr. Biol. 23, 1596–1600 (2013).
R Development Core Team. R: A Language and Environment for Statistical Computing http://www.r-project.org (2009).
RStudio and Inc. Shiny: Web Application Framework for R http://cran.r-project.org/package=shiny (2014).
Wickham, H . ggplot2: elegant graphics for data analysis. Media 35 (Springer, 2009).
Wickham, H. The Split-Apply-Combine strategy for data. J. Stat. Softw. 40, 1–29 (2011).
Ritz, C., Streibig, J. C., Ritz, C. & Streibig, J. C. Bioassay Analysis using R. J. Stat. Softw. 12, 1–22 (2005).
Robin, X. et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinformatics 12, 77 (2011).
Wickham, H. Reshaping Data with the reshape Package. J. Stat. Softw. 21, 1–20 (2007).
Data Citations
Mainland, J. D., Yun, R. L., Zhou, T., Liu, W. L. L., & Matsunami, H. Figshare (2014) http://dx.doi.org/10.6084/m9.figshare.979135
Acknowledgements
This work was supported by R01 DC005782, R01 DC012095, R03 DC011373, R01 DC013339, the Defense Advanced Research Project Agency RealNose Project, and an NRSA postdoctoral fellowship F32 DC008932 to J.D.M.
Author information
Authors and Affiliations
Contributions
J.D.M. conceived and designed the project, supervised the screen, carried out the analysis and wrote the paper. Y.R.L. carried out portions of the screen and analysis. T.Z. carried out portions of the screen. W.L.L.L. carried out portions of the screen. H.M. conceived, designed, and supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interest.
ISA-Tab metadata
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0 Metadata associated with this Data Descriptor is available at http://www.nature.com/sdata/ and is released under the CC0 waiver to maximize reuse.
About this article
Cite this article
Mainland, J., Li, Y., Zhou, T. et al. Human olfactory receptor responses to odorants. Sci Data 2, 150002 (2015). https://doi.org/10.1038/sdata.2015.2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2015.2
This article is cited by
-
Characterization of human oxidoreductases involved in aldehyde odorant metabolism
Scientific Reports (2023)
-
Comparative Evaluation of Sensitivity to Hexanal Between Human and Canine Olfactory Receptors
Biotechnology and Bioprocess Engineering (2019)
-
Computational genome-wide survey of odorant receptors from two solitary bees Dufourea novaeangliae (Hymenoptera: Halictidae) and Habropoda laboriosa (Hymenoptera: Apidae)
Scientific Reports (2017)
-
Genetic variants affecting equivalent protein family positions reflect human diversity
Scientific Reports (2017)
-
Olfactory receptor 10J5 responding to α-cedrene regulates hepatic steatosis via the cAMP–PKA pathway
Scientific Reports (2017)