Abstract
Rice is a staple crop for over half of the global population. However, blast disease caused by Magnaporthe orzae can result in more than a 30% loss in rice yield in epidemic years. Although some major resistance genes bolstering blast resistance have been identified in rice, their stacking in elite cultivars usually leads to yield penalties. Here we report that OsUBC45, a ubiquitin-conjugating enzyme functioning in the endoplasmic reticulum-associated protein degradation system, promotes broad-spectrum disease resistance and yield in rice. OsUBC45 is induced upon infection by M. oryzae, and its overexpression enhances resistance to blast disease and bacterial leaf blight by elevating pathogen-associated molecular pattern-triggered immunity (PTI) while nullifying the gene-attenuated PTI. The OsUBC45 overexpression also increases grain yield by over 10%. Further, OsUBC45 enhances the degradation of glycogen synthase kinase 3 OsGSK3 and aquaporin OsPIP2;1, which negatively regulate the grain size and PTI, respectively. The OsUBC45 reported in our study has the potential for improving yield and disease resistance for sustainable rice production.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Gene sequence information of rice from this study can be found at https://www.ricedata.cn/gene/, under the following accession numbers: OsUBC45, LOC_Os03g19500; OsPIP2;1, LOC_Os07g26690; OsGSK3, LOC_Os02g14130; OsBiP5, LOC_Os08g09770; OsCNX, LOC_Os04g32950; OsPDIL 2-1, LOC_Os05g06430; OsDOA10B, LOC_Os08g01040. Source data are provided with this paper.
References
Dean, R. et al. The top 10 fungal pathogens in molecular plant pathology. Mol. Plant Pathol. 13, 414–430 (2012).
Savary, S. et al. The global burden of pathogens and pests on major food crops. Nat. Ecol. Evol. 3, 430–439 (2019).
Li, W., Chern, M., Yin, J., Wang, J. & Chen, X. Recent advances in broad-spectrum resistance to the rice blast disease. Curr. Opin. Plant Biol. 50, 114–120 (2019).
Jones, J. D. & Dangl, J. L. The plant immune system. Nature 444, 323–329 (2006).
Mundt, C. C. Use of multiline cultivars and cultivar mixtures for disease management. Annu. Rev. Phytopathol. 40, 381–410 (2002).
Xiao, W. et al. Improvement of rice blast resistance by developing monogenic lines, two-gene pyramids and three-gene pyramid through MAS. Rice 12, 78 (2019).
Smalle, J. & Vierstra, R. D. The ubiquitin 26S proteasome proteolytic pathway. Annu. Rev. Plant Biol. 55, 555–590 (2004).
Romisch, K. Endoplasmic reticulum-associated degradation. Annu. Rev. Cell Dev. Biol. 21, 435–456 (2005).
Park, J. H. et al. EMR, a cytosolic-abundant ring finger E3 ligase, mediates ER-associated protein degradation in Arabidopsis. New Phytol. 220, 163–177 (2018).
Zhang, R. et al. Medicago falcata MfSTMIR, an E3 ligase of endoplasmic reticulum-associated degradation, is involved in salt stress response. Plant J. 98, 680–696 (2019).
Chen, Q. et al. ERAD-related E2 and E3 enzymes modulate the drought response by regulating the stability of PIP2 aquaporins. Plant Cell 33, 2883–2898 (2021).
Zhang, L. et al. Regulation of the stability and ABA import activity of NRT1.2/NPF4.6 by CEPR2-mediated phosphorylation in Arabidopsis. Mol. Plant 14, 633–646 (2021).
Bae, H. & Kim, W. T. Classification and interaction modes of 40 rice E2 ubiquitin-conjugating enzymes with 17 rice ARM-U-box E3 ubiquitin ligases. Biochem. Biophys. Res. Commun. 444, 575–580 (2014).
Wang, R. et al. An ORFeome of rice E3 ubiquitin ligases for global analysis of the ubiquitination interactome. Genome Biol. 23, 154 (2022).
Cui, F. et al. Arabidopsis ubiquitin conjugase UBC32 is an ERAD component that functions in brassinosteroid-mediated salt stress tolerance. Plant Cell 24, 233–244 (2012).
Muller, J. et al. Conserved ERAD-Like quality control of a plant polytopic membrane protein. Plant Cell 17, 149–163 (2005).
Gao, X. et al. Rice qGL3/OsPPKL1 functions with the GSK3/SHAGGY-Like Kinase OsGSK3 to modulate brassinosteroid signaling. Plant Cell 31, 1077–1093 (2019).
Li, J. et al. An ERAD-related E2-E3 enzyme pair controls grain size and weight through the brassinosteroid signaling pathway in rice. Plant Cell 35, 1076–1091 (2022).
Ai, G. et al. A Phytophthora sojae CRN effector mediates phosphorylation and degradation of plant aquaporin proteins to suppress host immune signaling. PLoS Pathog. 17, e1009388 (2021).
Rodrigues, O. et al. Aquaporins facilitate hydrogen peroxide entry into guard cells to mediate ABA- and pathogen-triggered stomatal closure. Proc. Natl Acad. Sci. USA 114, 9200–9205 (2017).
Tian, S. et al. Plant aquaporin AtPIP1;4 links apoplastic H2O2 induction to disease immunity pathways. Plant Physiol. 171, 1635–1650 (2016).
Chen, X. et al. Functional modulation of an aquaporin to intensify photosynthesis and abrogate bacterial virulence in rice. Plant J. 108, 330–346 (2021).
Wang, X. et al. The aquaporin TaPIP2;10 confers resistance to two fungal diseases in wheat. Phytopathology 111, 2317–2331 (2021).
Zhang, H. et al. A Gγ protein regulates alkaline sensitivity in crops. Science 379, eade8416 (2023).
Wang, X. et al. The plasma membrane NADPH oxidase OsRbohA plays a crucial role in developmental regulation and drought-stress response in rice. Physiol. Plant. 156, 421–443 (2016).
Ning, Y., Liu, W. & Wang, G. L. Balancing immunity and yield in crop plants. Trends Plant Sci. 22, 1069–1079 (2017).
Nelson, R., Wiesner-Hanks, T., Wisser, R. & Balint-Kurti, P. Navigating complexity to breed disease-resistant crops. Nat. Rev. Genet. 19, 21–33 (2018).
Wang, J. et al. A single transcription factor promotes both yield and immunity in rice. Science 361, 1026–1028 (2018).
Liu, M. M. et al. Inducible overexpression of Ideal Plant Architecture1 improves both yield and disease resistance in rice. Nat. Plants 5, 389–400 (2019).
Wang, L. et al. Arabidopsis UBC13 differentially regulates two programmed cell death pathways in responses to pathogen and low-temperature stress. New Phytol. 221, 919–934 (2019).
Liu, X. et al. Rice ubiquitin-conjugating enzyme OsUBC26 is essential for immunity to the blast fungus Magnaporthe oryzae. Mol. Plant Pathol. 22, 1613–1623 (2021).
Hao, Y. et al. Apollon ubiquitinates SMAC and caspase-9, and has an essential cytoprotection function. Nat. Cell Biol. 6, 849–860 (2004).
Liu, T. Y. et al. PHO2-dependent degradation of PHO1 modulates phosphate homeostasis in Arabidopsis. Plant Cell 24, 2168–2183 (2012).
Pan, W. et al. The UBC27-AIRP3 ubiquitination complex modulates ABA signaling by promoting the degradation of ABI1 in Arabidopsis. Proc. Natl Acad. Sci. USA 117, 27694–27702 (2020).
Chen, Q., Liu, R. J., Wang, Q. & Xie, Q. ERAD tuning of the HRD1 complex component AtOS9 is modulated by an ER-bound E2, UBC32. Mol. Plant 10, 891–894 (2017).
Hachez, C., Besserer, A., Chevalier, A. S. & Chaumont, F. Insights into plant plasma membrane aquaporin trafficking. Trends Plant Sci. 18, 344–352 (2013).
Lu, K. et al. Phosphorylation of a wheat aquaporin at two sites enhances both plant growth and defense. Mol. Plant 15, 1772–1789 (2022).
Chen, S. et al. Ubiquitin ligase OsRINGzf1 regulates drought resistance by controlling the turnover of OsPIP2;1. Plant Biotechnol. J. 20, 1743–1755 (2022).
Yang, J. et al. A novel protein Com1 is required for normal conidium morphology and full virulence in Magnaporthe oryzae. Mol. Plant Microbe Interact. 23, 112–123 (2010).
Liu, Y. et al. A designer rice NLR immune receptor confers resistance to the rice blast fungus carrying noncorresponding avirulence effectors. Proc. Natl Acad. Sci. USA 118, e2110751118 (2021).
Sun, X. et al. Xa26, a gene conferring resistance to Xanthomonas oryzae pv. oryzae in rice, encodes an LRR receptor kinase-like protein. Plant J. 37, 517–527 (2004).
He, F., Chen, S., Ning, Y. & Wang, G. L. Rice (Oryza sativa) protoplast isolation and its application for transient expression analysis. Curr. Protoc. Plant Biol. 1, 373–383 (2016).
Zhang, Y. et al. A highly efficient rice green tissue protoplast system for transient gene expression and studying light/chloroplast-related processes. Plant Methods 7, 30 (2011).
Yoo, S. D., Cho, Y. H. & Sheen, J. Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat. Protoc. 2, 1565–1572 (2007).
Thaminy, S., Miller, J. & Stagljar, I. The split-ubiquitin membrane-based yeast two-hybrid system. Methods Mol. Biol. 261, 297–312 (2004).
Chen, H. et al. Firefly luciferase complementation imaging assay for protein-protein interactions in plants. Plant Physiol. 146, 368–376 (2008).
Sang, Y. & Macho, A. P. Analysis of PAMP-triggered ROS burst in plant immunity. Methods Mol. Biol. 1578, 143–153 (2017).
Acknowledgements
We thank L. Luo from Shanghai Agrobiological Gene Center, China, for providing the CRISPR/Cas9-knockout lines and overexpression lines of OsPIP2;1. We also thank J. Huang from Nanjing Agricultural University for providing the OsGSK3 antibody. This work is supported by grants from the National Natural Science Foundation of China (32293244 to Y.-L.P.; 32072368 to Q.C.), the National Rice Industry Program from the Ministry of Agriculture and Rural Affairs (CARS-01-36 to Y.-L.P.), the 111 Project (grant no. B13006 to Y.-L.P.) from the Ministry of Education, the Staple Crop Disease Resistance Breeding Program from China Agricultural University (Y.-L.P.) and Pinduoduo-China Agricultural University Research Fund (Y.-L.P.).
Author information
Authors and Affiliations
Contributions
Q.C. and Y.-L.P. designed the research. Y.W. performed most of the experiments. J. Yue, N.Y., Y.Z., C.Z., X.W., J. Yang and W.Z. contributed to the assays of rice protoplasts, western blots and plant inoculation. Q.C., Y.-L.P., V.B., Y.W. and Q.X. wrote the manuscript. Y.N., L.L. and H.Z. participated in the discussion of the results and modification of manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Food thanks Tsutomu Kawasaki and the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 UPR is induced by chitin treatment.
The expression of UPR genes was induced by chitin treatment. Ten-day-old WT seedlings (ZH11) were treated with ddH2O or 10 μg ml−1 chitin for 6 h. The expression of the UPR genes was determined by qPCR. Data are presented as mean values +/− SEM, n = 3 replications. Asterisks indicate significant differences evaluated by two-tailed Student’s t test analysis, *p < 0.05, **p < 0.01. For exact p values, refer to Source Data.
Extended Data Fig. 2 Analysis of the evolutionary tree and tissue-specific expression of OsUBC45.
a, Schematic of OsUBC45. A UBCc domain (yellow) and a transmembrane domain (blue) are predicted in OsUBC45. The red bar is a ER membrane retention signal IEGK as predicted by the psort II program. b, The neighbor-joining phylogenetic tree of OsUBC45 and its orthologues from human and Arabidopsis. Bootstrap values from 1000 replicates are indicated at each node; the scale represents branch length. c, The expression level of OsUBC45 in different tissues was determined by qPCR. The individual tissues came from the rice in the filling stage. Data are presented as mean values +/− SEM, n = 3 replications. For exact p values, refer to Source Data. d, OsUBC45 interacted with OsDOA10B (LOC_Os08g01040) in a yeast two-hybrid assay. OsUBC45 was inserted into pPR3-N, while OsDOA10B was inserted into pBT3-STE. The transformants were grown on SD-Leu-Trp (SD-LW) and SD-Leu-Trp-His-Ade (SD-LWHA) plates containing 1 mM 3-AT. e, Ubiquitin-conjugating enzyme activity of His-MBP-OsUBC45. The OsUBC45-ubiquitin adduct (indicated by arrow) were detected using both anti-MBP and anti-ubiquitin antibodies.
Extended Data Fig. 3 Rice yield and grain size were reduced in the osubc45 mutants.
a, Morphology of the WT and osubc45 mutant plants at the mature stage. Bar = 5 cm. b, Plant height of the WT and osubc45 mutant plants. Data are presented as mean values +/− SEM, n = 5 independent plants. Asterisks indicate significant differences evaluated by two-tailed Student’s t-test analysis, **p < 0.01. c, Number of panicles per WT and osubc45 mutant plants. Data are presented as mean values +/− SEM, n = 5 independent hills. Asterisks indicate significant differences evaluated by two-tailed Student’s t-test analysis, *p < 0.05, **p < 0.01. d, Panicle length of WT and osubc45-9 and osubc45-23. Data are presented as mean values +/− SEM, n = 10 independent panicles. Asterisks indicate significant differences evaluated by two-tailed Student’s t-test, *p < 0.05 and **p < 0.01. e-f, Primary branches per panicle (e) and secondary branches per panicle (f) of WT and osubc45 mutants. Data are presented as mean values +/− SEM, n = 10 independent panicles. Asterisks indicate significant differences evaluated by two-tailed Student’s t-test, **p < 0.01. g, Grain number per panicle of WT, osubc45-9 and osubc45-23. Data are presented as mean values +/− SEM, n = 10 independent panicles. Asterisks indicate significant differences evaluated by two-tailed Student’s t-test, **p < 0.01. h-i, Grain length (h) and Grain width (i) of WT and osubc45 mutant plants. Data are presented as mean values +/− SEM, n = 10 grains, 3 replications were performed. Asterisks indicate significant differences evaluated by two-tailed Student’s t-test (**p < 0.01). For exact p values, refer to Source Data.
Extended Data Fig. 4 The osubc45 mutants were sensitive to rice blast by spray inoculation.
a, The osubc45 mutants increased the susceptibility to rice blast by spray inoculation. The WT and osubc45 mutant plants were spray-inoculated with the virulent isolate RB22. Leaves were photographed at 7 dpi. b, Lesion number of the leaves was measured at 7 dpi. Data are presented as mean values +/− SEM, n = 3 replications. Asterisks indicate significant differences evaluated by two-tailed Student’s t-test, **p < 0.01. For exact p values, refer to Source Data.
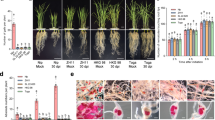
Extended Data Fig. 5 Effects of overexpression of OsUBC45 on rice growth and yield.
a, Morphology of WT and OsUBC45 overexpression lines at maturity. Bar = 5 cm. b, Plant height of the WT and OsUBC45 overexpression plants. Data are presented as mean values +/− SEM, n = 5 independent plants. c, Panicle number per plant of (WT and OsUBC45 transgenic lines). Data are presented as mean values +/− SEM, n = 5 independent plants. d, Primary branches per panicle of WT and OsUBC45 transgenic lines. e, Secondary branches per panicle of WT and OsUBC45 transgenic plants. (d)-(e) Data are presented as mean values +/− SEM, n = 10 independent panicles (*p < 0.05, **p < 0.01). For exact p values, refer to Source Data.
Extended Data Fig. 6 OsUBC45 enhanced resistance to rice blast by spray inoculation.
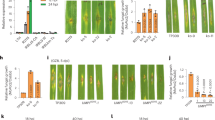
a, OsUBC45 transgenic plants increased resistance to rice blast by spray inoculation. WT and OsUBC45 transgenic plants were sprayed with virulent isolate RB22. Leaves were photographed at 7 dpi. b, Lesion number of the inoculated leaves was measured at 7 dpi. Data are presented as mean values +/− SEM, n = 3 replications. Asterisks indicate significant differences evaluated by two-tailed Student’s t-test, **p < 0.01. c, Lesion length of the drop inoculation in Fig. 4d. Data are presented as mean values +/− SEM, n = 6 independent lesions. Asterisks indicate significant differences evaluated by two-tailed Student’s t-test, **p < 0.01. For exact p values, refer to Source Data.
Extended Data Fig. 7 DGS1 interacted with OsGSK3 and promoted its degradation.
a, Proteins that were identified to interact with OsUBC45 by yeast two-hybrid screening. b, OsUBC45 interacts with OsGSK3 by LCI assay. c, The expression of OsGSK3 in WT, osubc45 mutants and OsUBC45 overexpression lines were determined by qPCR. Data are presented as mean values +/− SEM (n = 3 replications). d, Chitin treatment did not affect the degradation of OsGSK3. Protein levels of OsGSK3 in WT, osubc45 mutants and OsUBC45-OE plants were detected using anti-OsGSK3 antibody with/without chitin treatment. e, DGS1 interacted with OsGSK3 in yeast. DGS1 was inserted into pBT3-STE, while OsGSK3 was inserted into pPR3-N. The transformants were grown on SD-LW and SD-LWHA plates containing 2 mM 3-AT. f, DGS1 interacted with OsGSK3 by LCI assay. g, DGS1 promoted the degradation of OsGSK3-HA-Nluc in rice protoplasts. Different combinations of plasmids were transformed into rice protoplasts of WT. Proteins from the rice protoplasts were used for protein gel blot analysis.
Extended Data Fig. 8 DGS1 interacted with OsPIP2;1 and promoted its degradation.
a, OsUBC45 interacted with OsPIP2;1 according to a yeast two-hybrid assay. OsUBC45 was inserted into pBT3-STE, while OsPIP2;1 was inserted into pPR3-N. The transforms were grown on SD-LW and SD-LWHA plates containing 3 mM 3-AT. b, The expression of OsPIP2;1 in WT, osubc45 mutants and OsUBC45 overexpression lines were determined by qPCR. Data are presented as mean values +/− SEM, n = 3 replications. c, DGS1 interacted with OsPIP2;1 in yeast two-hybrid assays. DGS1 was inserted into pBT3-STE, while OsPIP2;1 was inserted into pPR3-N. The transformants were grown on SD-LW and SD-LWHA plates containing 2 mM 3-AT. d, DGS1 associated with OsPIP2;1 in the LCI assay. e, DGS1 negatively regulated the stability of OsPIP2;1-Myc in rice protoplasts. Different combinations of plasmids were used to transform rice protoplasts of WT. Proteins isolated from the rice protoplasts were used for western blot analysis.
Extended Data Fig. 9 OsPIP2;1 increased susceptibility to bacterial blight.
a, The expression of OsPIP2;1 is inhibited by M. oryzae treatment. The WT plants were inoculated with the M. oryzae virulent isolate RB22 for the corresponding times. Data are presented as mean values +/− SEM, n = 3 replications. Asterisks indicate significant differences evaluated by two-sided Student’s t-tests, **p < 0.01. b, OsPIP2;1 negatively regulated bacterial blight resistance. Leaves were photographed at 14 dpi. c, Lesion length of bacterial blight in leaves (b). Data are presented as mean values +/− SEM, n = 3 independent lesions. Asterisks indicate significant differences evaluated by two-tailed Student’s t-test (*p < 0.05, **p < 0.01). For exact p values, refer to Source Data.
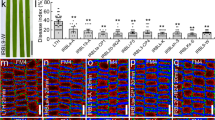
Extended Data Fig. 10 OsUBC45 positively regulates disease resistance by accumulating more ROS under M. oryzae treatment.
a, H2O2 contents of WT, osubc45-9 and OsUBC45-OE2 seedlings after inoculation with M. oryzae for 24 h. Seedlings treated with 0.02% Tween-20 served as mock controls. Data are presented as mean values +/− SEM, n = 3 replications. Asterisks indicate significant differences evaluated by two-sided Student’s t-tests, **p < 0.01. FW, fresh weight. b, DAB staining of different rice leaves infected with M. oryzae for 48 h. H2O2 accumulation was shown as dark-brown spots. c, Relative DAB staining intensity was measured by Image J. Data are presented as mean values +/− SEM, n = 3 independent leaves. Asterisks indicate significant differences evaluated by two-tailed Student’s t-test, **p < 0.01. d, AUR, AR and H2DCF-DA probing of H2O2 in leaf sheath cells of WT, osubc45-9 and OsUBC45-OE2 plants infected with M. oryzae for 48 h. Bars = 50 μm. e–g, Statistical analysis of H2O2 accumulation in (d). Fluorescence intensity of AUR (e). Fluorescence intensity of AR (f). Fluorescence intensity of H2DCF-DA (g). Data are presented as mean values +/− SEM, n = 5 independent views. Asterisks indicate significant differences evaluated by two-tailed Student’s t-test, **p < 0.01. For exact p values, refer to Source Data.
Supplementary information
Supplementary Table
List of primer sequences (5′→3′) used in this study.
Source data
Statistical Source Data
All statistical source data.
Source Data of gels or blots
All unprocessed western blots and/or gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, Y., Yue, J., Yang, N. et al. An ERAD-related ubiquitin-conjugating enzyme boosts broad-spectrum disease resistance and yield in rice. Nat Food 4, 774–787 (2023). https://doi.org/10.1038/s43016-023-00820-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43016-023-00820-y
This article is cited by
-
Early molecular events in the interaction between Magnaporthe oryzae and rice
Phytopathology Research (2024)
-
Genome‑wide identification and expression analysis of the UBC gene family in wheat (Triticum aestivum L.)
BMC Plant Biology (2024)
-
DGS1 improves rice disease resistance by elevating pathogen-associated molecular pattern-triggered immunity
aBIOTECH (2024)