Abstract
Ketone bodies are generated in the liver and allow for the maintenance of systemic caloric and energy homeostasis during fasting and caloric restriction. It has previously been demonstrated that neonatal ketogenesis is activated independently of starvation. However, the role of ketogenesis during the perinatal period remains unclear. Here, we show that neonatal ketogenesis plays a protective role in mitochondrial function. We generated a mouse model of insufficient ketogenesis by disrupting the rate-limiting hydroxymethylglutaryl-CoA synthase 2 enzyme gene (Hmgcs2). Hmgcs2 knockout (KO) neonates develop microvesicular steatosis within a few days of birth. Electron microscopic analysis and metabolite profiling indicate a restricted energy production capacity and accumulation of acetyl-CoA in Hmgcs2 KO mice. Furthermore, acetylome analysis of Hmgcs2 KO cells revealed enhanced acetylation of mitochondrial proteins. These findings suggest that neonatal ketogenesis protects the energy-producing capacity of mitochondria by preventing the hyperacetylation of mitochondrial proteins.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
We submitted RNA sequencing data to the Gene Expression Omnibus under accession GSE18853. The dataset of GC–MS analysis and iMAPQ analysis and other data that support the findings of this study are available from the corresponding author upon request. The dataset of acetylome analysis is available as source data. Source data are provided with this paper.
References
Krebs, H. A. The regulation of the release of ketone bodies by the liver. Adv. Enzyme Regul. 4, 339–354 (1966).
Arima, Y. et al. Myocardial ischemia suppresses ketone body utilization. J. Am. Coll. Cardiol. 73, 246–247 (2019).
McGarry, J. D. & Foster, D. W. Regulation of hepatic fatty acid oxidation and ketone body production. Annu. Rev. Biochem. 49, 395–420 (1980).
Rojas-Morales, P., Tapia, E. & Pedraza-Chaverri, J. Beta-hydroxybutyrate: a signaling metabolite in starvation response? Cell. Signal. 28, 917–992 (2016).
Shimazu, T. et al. SIRT3 deacetylates mitochondrial 3-hydroxy-3-methylglutaryl CoA synthase 2 and regulates ketone body production. Cell Metab. 12, 654–661 (2010).
Puchalska, P. & Crawford, P. A. Multi-dimensional roles of ketone bodies in fuel metabolism, signaling and therapeutics. Cell Metab. 25, 262–284 (2017).
Chapman, M. J., Miller, L. R. & Ontko, J. A. Localization of the enzymes of ketogenesis in rat liver mitochondria. J. Cell Biol. 58, 284–306 (1973).
Dashti, N. & Ontko, J. A. Rate-limiting function of 3-hydroxy-3-methylglutaryl coenzyme A synthase in ketogenesis. Biochem. Med. 22, 365–374 (1979).
Hegardt, F. G. Mitochondrial 3-hydroxy-3-methylglutaryl-CoA synthase: a control enzyme in ketogenesis. Biochem. J. 338, 569–582 (1999).
d’Avignon, D. A. et al. Hepatic ketogenic insufficiency reprograms hepatic glycogen metabolism and the lipidome. JCI Insight 3, e99762 (2018).
Cotter, D. G. et al. Ketogenesis prevents diet-induced fatty liver injury and hyperglycemia. J. Clin. Invest. 124, 5175–5190 (2014).
Cotter, D. G., Ercal, B., d’Avignon, D. A., Dietzen, D. J. & Crawford, P. A. Impact of peripheral ketolytic deficiency on hepatic ketogenesis and gluconeogenesis during the transition to birth. J. Biol. Chem. 288, 19739–19749 (2013).
Decaux, J. F., Robin, D., Robin, P., Ferre, P. & Girard, J. Intramitochondrial factors controlling hepatic fatty acid oxidation at weaning in the rat. FEBS Lett. 232, 156–158 (1988).
Ward Platt, M. & Deshpande, S. Metabolic adaptation at birth. Semin. Fetal Neonatal Med. 10, 341–350 (2005).
Baeza, J., Smallegan, M. J. & Denu, J. M. Mechanisms and dynamics of protein acetylation in mitochondria. Trends Biochem. Sci. 41, 231–244 (2016).
Mews, P. et al. Acetyl-CoA synthetase regulates histone acetylation and hippocampal memory. Nature 546, 381–386 (2017).
Sivanand, S., Viney, I. & Wellen, K. E. Spatiotemporal control of Acetyl-CoA metabolism in chromatin regulation. Trends Biochem. Sci. 43, 61–74 (2018).
Fisher-Wellman, K. H. et al. Respiratory phenomics across multiple models of protein hyperacylation in cardiac mitochondria reveals a marginal impact on bioenergetics. Cell Rep. 26, 1557–1572 (2019).
Cheng, A. et al. SIRT3 haploinsufficiency aggravates loss of GABAergic interneurons and neuronal network hyperexcitability in an Alzheimer’s disease model. J. Neurosci. 40, 694–709 (2020).
Xie, Z. et al. Metabolic regulation of gene expression by histone lysine beta-hydroxybutyrylation. Mol. Cell 62, 194–206 (2016).
Finley, L. W. et al. Succinate dehydrogenase is a direct target of sirtuin 3 deacetylase activity. PLoS ONE 6, e23295 (2011).
Hebert, A. S. et al. Calorie restriction and SIRT3 trigger global reprogramming of the mitochondrial protein acetylome. Mol. Cell 49, 186–199 (2013).
Someya, S. et al. Sirt3 mediates reduction of oxidative damage and prevention of age-related hearing loss under caloric restriction. Cell 143, 802–812 (2010).
Zhang, Y. et al. Protein–protein interactions and metabolite channelling in the plant tricarboxylic acid cycle. Nat. Commun. 8, 15212 (2017).
Hirschey, M. D. et al. SIRT3 deficiency and mitochondrial protein hyperacetylation accelerate the development of the metabolic syndrome. Mol. Cell 44, 177–190 (2011).
Brumbaugh, D. E. & Friedman, J. E. Developmental origins of nonalcoholic fatty liver disease. Pediatr. Res. 75, 140–147 (2014).
Soderborg, T. K. et al. The gut microbiota in infants of obese mothers increases inflammation and susceptibility to NAFLD. Nat. Commun. 9, 4462 (2018).
Ma, J. et al. High-fat maternal diet during pregnancy persistently alters the offspring microbiome in a primate model. Nat. Commun. 5, 3889 (2014).
Narita, T., Weinert, B. T. & Choudhary, C. Functions and mechanisms of non-histone protein acetylation. Nat. Rev. Mol. Cell Biol. 20, 156–174 (2019).
Choudhary, C., Weinert, B. T., Nishida, Y., Verdin, E. & Mann, M. The growing landscape of lysine acetylation links metabolism and cell signalling. Nat. Rev. Mol. Cell Biol. 15, 536–550 (2014).
Schwer, B. & Verdin, E. Conserved metabolic regulatory functions of sirtuins. Cell Metab. 7, 104–112 (2008).
Newman, J. C., He, W. & Verdin, E. Mitochondrial protein acylation and intermediary metabolism: regulation by sirtuins and implications for metabolic disease. J. Biol. Chem. 287, 42436–42443 (2012).
Parodi-Rullan, R. M., Chapa-Dubocq, X. R. & Javadov, S. Acetylation of mitochondrial proteins in the heart: the role of SIRT3. Front. Physiol. 9, 1094 (2018).
Fan, J. et al. Tyr phosphorylation of PDP1 toggles recruitment between ACAT1 and SIRT3 to regulate the pyruvate dehydrogenase complex. Mol. Cell 53, 534–548 (2014).
Wang, L. et al. GCN5L1 modulates cross-talk between mitochondria and cell signaling to regulate FoxO1 stability and gluconeogenesis. Nat. Commun. 8, 523 (2017).
Paik, W. K., Pearson, D., Lee, H. W. & Kim, S. Nonenzymatic acetylation of histones with acetyl-CoA. Biochim. Biophys. Acta 213, 513–522 (1970).
Wagner, G. R. & Hirschey, M. D. Nonenzymatic protein acylation as a carbon stress regulated by sirtuin deacylases. Mol. Cell 54, 5–16 (2014).
Casey, J. R., Grinstein, S. & Orlowski, J. Sensors and regulators of intracellular pH. Nat. Rev. Mol. Cell Biol. 11, 50–61 (2010).
Davies, M. N. et al. The acetyl group buffering action of carnitine acetyltransferase offsets macronutrient-induced lysine acetylation of mitochondrial proteins. Cell Rep. 14, 243–254 (2016).
Wagner, G. R. & Payne, R. M. Widespread and enzyme-independent Nε-acetylation and Nε-succinylation of proteins in the chemical conditions of the mitochondrial matrix. J. Biol. Chem. 288, 29036–29045 (2013).
Shimazu, T. et al. Suppression of oxidative stress by beta-hydroxybutyrate, an endogenous histone deacetylase inhibitor. Science 339, 211–214 (2013).
Lowe, D. M. & Tubbs, P. K. 3-hydroxy-3-methylglutaryl-coenzyme A synthase from ox liver. Purification, molecular and catalytic properties. Biochem. J. 227, 591–599 (1985).
Nakade, S. et al. Microhomology-mediated end-joining-dependent integration of donor DNA in cells and animals using TALENs and CRISPR/Cas9. Nat. Commun. 5, 5560 (2014).
Nakagawa, Y. et al. Culture time of vitrified/warmed zygotes before microinjection affects the production efficiency of CRISPR–Cas9-mediated knock-in mice. Biol. Open 6, 706–713 (2017).
Sakuma, T., Nakade, S., Sakane, Y., Suzuki, K. T. & Yamamoto, T. MMEJ-assisted gene knock-in using TALENs and CRISPR–Cas9 with the PITCh systems. Nat. Protoc. 11, 118–133 (2016).
Takihara, Y. et al. High mitochondrial mass is associated with reconstitution capacity and quiescence of hematopoietic stem cells. Blood Adv. 3, 2323–2327 (2019).
Inoue, T. & Matsumoto, K. Techniques for Illustrated Animal Experiments, Vol. 24 (Kyoritsu Shuppan, 1981).
Hino, S. et al. FAD-dependent lysine-specific demethylase-1 regulates cellular energy expenditure. Nat. Commun. 3, 758 (2012).
Umemoto, T., Hashimoto, M., Matsumura, T., Nakamura-Ishizu, A. & Suda, T. Ca2+–mitochondria axis drives cell division in hematopoietic stem cells. J. Exp. Med. 215, 2097–2113 (2018).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Anan, K. et al. LSD1 mediates metabolic reprogramming by glucocorticoids during myogenic differentiation. Nucleic Acids Res. 46, 5441–5454 (2018).
Acknowledgements
The authors thank Y. Kimura, M. Nagahiro and S. Tokunaga for their excellent technical assistance throughout the experiments. The Kumamoto University School of Medicine Core Laboratory for Medical Research and Education provided support for ddPCR, GC–MS and microscopic analysis. Y. Tanoue of the International Research Center for Medical Sciences performed the section electron microscopic examination. T. Keida (Kumamoto Mouse Clinic, Kumamoto University) performed a blood panel analysis. T. Motoyoshi (K.I. Stainer) provided excellent sections for histological analysis. Support for the iMPAQT analysis was provided by Kyushu Pro Search. M. P. Stokes and K. Abell (Cell Signaling Technology) performed the acetylome analysis. This study was supported by a grant-in-aid for Scientific Research (19K08520, 17K16014 and 18K08110) from the Ministry of Education, Culture, Sports, Science and Technology of Japan, the grant for Basic Research of the Japanese Circulation Society (2018), a grant from the Sumitomo Foundation (2018), a grant from the Takeda Foundation (2019), a grant from the Kanae Foundation (2019), a grant from the Japan Foundation for Applied Enzymology (2019), a grant from the Ono Medical Research Foundation (2019), a Kumamoto University Challenging Research Projects (2019) and a grant for Research on Development of New Drugs (16769865) from the Japan Agency for Medical Research and Development.
Author information
Authors and Affiliations
Contributions
Y.A., Y.N., T.T. and K. Tsujita conceived and designed the experiments. Y.A., Y.N., T.T., T.I., T. Yamada, S. Hanada, S. Hino and T.U. performed the experiments. Y.A., Y.N., T.T., T.I., T. Yamada, S.H., M.N., T.U., T. Suda, T.W., K. Nagaoka, Y.T., T.N., S.A., E.Y., Y.I., K.S., K.K., K.M., K. Nishiyama and K. Tsujita analysed the data. Y.A., Y.N., T.T., S. Hino, M.N., S. Hanada, T. Sakuma., T.Y., Y.K.K., K. Tonami, H.K., Y.S., K.Y., K.M., K. Nishiyama and N.N. contributed materials/analysis tools. Y.A., Y.N., T.T., T. Suda., T.N., S.A., K.M. and K. Tsujita wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Metabolism thanks Iain Scott, Heng Zhu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: George Caputa.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 (Related to Fig. 2): The sequencing analyses of Hmgcs2-targeted mice.
a, Sanger sequencing results of two lines (#9 and #10) of Hmgcs2 KO heterozygote mice. b, Alignments with the reference sequence of Hmgcs2. Green box represents the first ATG (start codon). Blue box represents the second ATG.
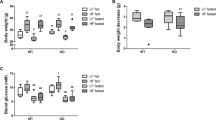
Extended Data Fig. 2 (Related to Fig. 3): Phenotype of Hmgcs2 KO mice.
a-c: Kaplan-Meier analysis of Hmgcs2 WT and KO of all (a), male (b), and female (c) mice. d,e: Concentration of β-hydroxybutyrate (d), Body weight (e), food intake (f), and feces (g) of 15 week-old Hmgcs2 WT and KO mice. The body weight of WT and Hmgcs2 KO mice does not differ significantly. However, when fed a normal diet, the food intake and feces volume increase significantly in Hmgcs2 KO mice (WT n = 7, KO n = 7). Two-way ANOVA analysis and Sidak’s multiple comparison test were applied (d). Welch’s two-sided t-tests were performed (e-f). h: Reactive oxygen species (ROS) level in postnatal day 3 (P3) isolated hepatocytes. Wt; Hmgcs2 WT, KO; Hmgcs2 KO, GeoMFI; geometrical mean fluorescence intensity. Welch’s two-sided t-test was performed. i: Oxygen consumption rate with/without palmitate. BSA; bovine serum albumin, FCCP; Carbonyl cyanide-p-trifluoromethoxyphenylhydrazone. Two-way ANOVA analysis and Sidak’s multiple comparison test were applied. All results are expressed as means ± standard deviation (SD).
Extended Data Fig. 3 (Related to Fig. 3): Generation of floxed mice at the Hmgcs2 locus.
a, Schematic illustration to generate a floxed allele at the Hmgcs2 locus, mediated by the CRIS-PITCh (v2) system. Two gene-specific gRNAs were designed upstream and downstream of exon 2. A PITCh donor plasmid was designed to carry two loxP sites flanking exon 2 and 50-bp homology arms. Arrows indicate the primer sets for PCR. b, Sequence data (sense strand) of region ‘B’. Gray area is the loxP sequence. c, Sequence data (antisense strand) of region ‘C’. Gray area is the loxP sequence. d,e, Genotyping of Hmgcs2+/+, Hmgcs2flox/flox, and Alb-cre;Hmgcs2 flox/+ mice.
Extended Data Fig. 4 (Related to Fig. 4): Phenotype in the liver of Hmgcs2 KO.
a, Section electron microscopy of postnatal day 3 (P3) hepatocytes. Red arrows represent peroxisomes. Scale bar = 0.5 µm. b, Protein content is represented as weight percent (%). WT; Hmgcs2 WT, n = 7, 13.1 ± 0.82%, KO; Hmgcs2 KO, n = 9, 9.10 ± 0.59%. c, Gas chromatography-mass spectrometry (GC-MS) based concentrations of citrate, alpha-ketoglutarate, fumarate, and malate, adjusted by each sample weight. Hmgcs2 WT (n = 6), Hmgcs2 KO (n = 8). d, GC-MS based concentrations of citrate, alpha-ketoglutarate, fumarate, and malate, adjusted by each protein content. Hmgcs2 WT (n = 6), Hmgcs2 KO (n = 8). e, Basal respiration, ATP production, and maximal respiration of primary hepatocytes from P3 Hmgcs2 WT and KO neonates (Related to Fig. 4g). f, Gating strategy of flow cytometry (Related to Fig. 4h). All results are expressed as means ± standard deviation (SD). Welch’s two-sided t-tests were performed (c-e).
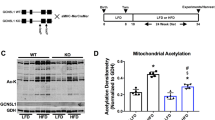
Extended Data Fig. 5 (Related to Fig. 5): Mitochondrial protein acetylation in Hmgcs2 KO livers.
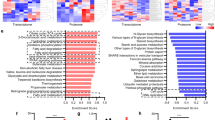
a, Heatmap for iMPAQT analysis (WT = 3, KO = 2). Green characters represent mitochondrial proteins. b, Protein fold changes between Hmgcs2 KO and WT mice (Related to Fig. 5a). c, Immunoprecipitation assays on P7 livers: IP, anti-acetylated lysine (AcK); IB, anti-LCAD. Red arrowheads indicate LCAD protein. LCAD, long-chain acyl-CoA dehydrogenase. d, Immunoprecipitation assays on postnatal day 7 (P7) livers: IP, anti-LCAD; IB, anti-acetylated lysine (left), anti-LCAD (right). Red arrowheads indicate LCAD protein. 800 µg purified mitochondrial proteins were used for immunoprecipitation assays. e, Serial changes in protein acetylation in the liver. Whole protein (10 µg) from WT livers (E18.5, P3, P7, P21, and Adult) was blotted using the anti-acetylated lysine antibody. (AcK; acetylated lysine). f, Western blot analysis for GCN5L1, UQCRFS1, and HMGCS2 between P7 Hmgcs2 wild type and knockout livers (n = 4 each). Welch’s two-sided t-test was performed. g, Expressions of Acly, Acss2, and Slc25a1 mRNA, adjusted by Rplp1 in P7 neonatal livers. (Hmgcs2 WT, n = 4, KO, n = 4). Welch’s two-sided t-test was performed. Results are expressed as means ± standard deviation (SD). h, Western blot analysis for AcK and total protein (TP) between RosaCAG-Sirt3 and wild-type adult heart (n = 4 each). Welch’s two-sided t-test was performed.
Extended Data Fig. 6 (Related to Fig. 6): Characteristics of histone protein acetylation in Hmgcs2 KO mice.
a, Bar plots of fold changes in histone protein acetylation. b, Expression of Hdac1, Hdac2, Hdac3, and Hdac8 transcripts, adjusted by Rplp1 in P7 neonatal livers. (Hmgcs2 WT, n = 4, KO, n = 4). Welch’s t-test was performed. c, Western blot analysis for HDAC1, HMGCS2, and βACTIN between P7 Hmgcs2 wild type and knockout livers (n = 4 each). Welch’s t-test was performed. d, Histone deacetylases (HDACs) fluorometric assay with β-hydroxybutyrate (βOHB). Relative fluorescent intensity with excitation at 360 nm and emission at 450 nm of 50 µg nuclear extracts from postnatal day 7 mice livers were measured. Before the measurements, nuclear extracts were incubated with 5 mM 3OHB (n = 4), 50 mM 3OHB(n = 4), 500 mM 3OHB(n = 4), or 1 µM trichostatin A (n = 4) for 20 minutes at 37 °C. RFU, relative fluorescent intensity. All lines represent the mean with standard error. After 8 minutes, significant differences were observed in all treatment groups in both WT and KO compared with control (p < 0.0001). Repeated measures two-way ANOVA and Dunnett’s multiple comparisons tests were performed. e, Chip-qPCR analysis of promoter and distal regions in CPT1A and ACADL genes in P7 liver tissue of Hmgcs2 WT and Hmgcs2 KO mice (WT = 5, KO = 5). Two-way ANOVA results are presented. f, Expressions of Per1, Ppargc1b, and Cpt1a mRNA, adjusted by Rplp1 in P7 neonatal livers. (Hmgcs2 WT, n = 4, KO, n = 4). Welch’s two-sided t-test was performed. All results are expressed as means ± standard deviation (SD).
Extended Data Fig. 7 (Related to Fig. 7): In vitro assay for acetylation of mitochondrial proteins.
a, Western blot analysis of primary hepatocytes for Hmgcs2, treated with siCtrl or siHmgcs2 (n = 3 each). Welch’s two-sided t-test was performed. b. Immunofluorescent analysis of primary hepatocytes for Hmgcs2, treated with siCtrl or siHmgcs2. c, Western blot analysis of primary hepatocytes for acetylated lysine, treated with siCtrl or siHmgcs2 (n = 3 each). Welch’s two-sided t-test was performed. AcK; acetylated lysine. d, Time-course curve of 13C-labeled fumarate (n = 3 at each time point). Results are expressed as means ± standard deviation (SD). e, Western blot analysis of HMGCS2 in isolated mitochondria from P7 Hmgcs2 WT and Hmgcs2 KO livers. f, 13C-labeled fumarate is significantly reduced in Hmgcs2 KO mitochondria. (0 min; n = 3 each; 120 min; n = 5 each). Two-way ANOVA and Sidak’s multiple comparison test were performed. g, Co-incubation with acetyl-CoA enhances mitochondrial protein acetylation. Western blot analysis of acetylated lysine in isolated mitochondria with/without acetyl-CoA (n = 4, each). *Welch’s two-sided t-test was performed.
Supplementary information
Supplementary Information
Supplementary Tables 1–6
Source data
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Unprocessed gels and western blots.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 6
Source data of acetylome analysis.
Source Data Fig. 7
Unprocessed western blots.
Source Data Fig. 7
Statistical source data.
Source Data Table 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed gels.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Unprocessed western blots.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
About this article
Cite this article
Arima, Y., Nakagawa, Y., Takeo, T. et al. Murine neonatal ketogenesis preserves mitochondrial energetics by preventing protein hyperacetylation. Nat Metab 3, 196–210 (2021). https://doi.org/10.1038/s42255-021-00342-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-021-00342-6
This article is cited by
-
Downregulation of HMGCS2 mediated AECIIs lipid metabolic alteration promotes pulmonary fibrosis by activating fibroblasts
Respiratory Research (2024)
-
HMGCS2 serves as a potential biomarker for inhibition of renal clear cell carcinoma growth
Scientific Reports (2023)
-
Metabolic Messengers: ketone bodies
Nature Metabolism (2023)
-
Pharmacological inhibition of Lin28 promotes ketogenesis and restores lipid homeostasis in models of non-alcoholic fatty liver disease
Nature Communications (2022)
-
Modulation of cellular processes by histone and non-histone protein acetylation
Nature Reviews Molecular Cell Biology (2022)