Abstract
DNA mismatch repair (MMR) is thought to contribute to the onset and progression of Huntington disease (HD) by promoting somatic expansion of the pathogenic CAG nucleotide repeat in the huntingtin gene (HTT). Here we have studied constitutional HTT CAG repeat size in two cohorts of individuals with Lynch syndrome (LS) carrying heterozygous loss-of-function variants in the MMR genes MLH1 (n = 12/60; Lund cohort/Bochum cohort, respectively), MSH2 (n = 15/88), MSH6 (n = 21/23), and controls (n = 19/559). The sum of CAG repeats for both HTT alleles in each individual was calculated due to unknown segregation with the LS allele. In the larger Bochum cohort, the sum of CAG repeats was lower in the MLH1 subgroup compared to controls (MLH1 35.40 CAG repeats ± 3.6 vs. controls 36.89 CAG repeats ± 4.5; p = 0.014). All LS genetic subgroups in the Bochum cohort displayed lower frequencies of unstable HTT intermediate alleles and lower HTT somatic CAG repeat expansion index values compared to controls. Collectively, our results indicate that MMR gene haploinsufficiency could have a restraining impact on constitutional HTT CAG repeat size and support the notion that the MMR pathway is a driver of nucleotide repeat expansion diseases.
Similar content being viewed by others
Introduction
Huntington disease (HD) is one of at least nine Mendelian CAG/polyglutamine diseases, in which CAG nucleotide repeat expansions encode elongated stretches of glutamines in the respective disease-associated protein1. In HD, the underlying pathogenic CAG repeat is located in exon 1 in the huntingtin gene (HTT)2. By not fully understood mechanisms during meiosis, pathogenic HTT CAG repeat expansions (≥ 36 CAG repeats) can arise from the transition of harmless but unstable intermediate alleles (27–35 CAG repeats) to HD incomplete-penetrance alleles (36–39 CAG repeats) or HD full-penetrance alleles (≥ 40 CAG repeats) upon inheritance2. The CAG repeat expansion in HD triggers a protracted cascade of events, leading particularly to the degeneration of medium-spiny neurons in the striatum but also neuronal loss in other brain regions, causing movement, cognitive and psychiatric disorders with a wide spectrum of signs and symptoms3,4. HD full-penetrance allele repeat size inversely correlates with the age of disease onset (AO)5 but does not fully explain AO variability. In addition to a variable CAA interruption of the HTT CAG repeat that modulates AO6,7,8, genome-wide association studies (GWAS) have also identified several DNA maintenance genes as modifiers of AO including genes encoding components of the DNA mismatch repair (MMR) pathway9,10,11,12,13 (for review, see14). The MMR pathway is involved in the correction of misaligned DNA strands, the event which frequently occurs in genomic regions with mono- di-, or trinucleotide repeats (also termed short tandem repeats; STRs or microsatellites) in both replicating and non-replicating cells15. There is now a large body of experimental evidence showing that MMR contributes to the AO and disease progression in HD by promoting somatic expansion of the pathogenic CAG repeat, highlighting the MMR pathway as a potential target for therapeutic interventions14. Individuals heterozygous for germline loss-of-function (LoF) pathogenic variants in either of the MMR genes MLH1, MSH2, MSH6 or PMS2 have Lynch syndrome (LS), an autosomal dominant predisposition mainly to colorectal cancer and endometrial cancer (for a recent review of LS, see16). The biochemical hallmark of LS cancer is deficient MMR (dMMR) due to somatic inactivation of the wild type allele of the affected MMR gene, and as a consequence LS cancer cells show immunohistochemical loss of MMR protein expression and accumulate nucleotide repeat aberrations known as microsatellite instability (MSI). In gametes of individuals with LS, the state of haploidy at meiosis implies that oocytes and sperm cells carrying MMR LoF alleles are subject to dMMR. In the present study we therefore hypothesized that MMR gene haploinsufficiency could affect constitutional HTT CAG repeat size. To test this hypothesis, we determined HTT CAG repeat size in individuals with LS.
Results
Characterization and analysis of the Lund cohort
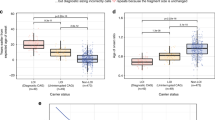
Initially, a cohort of individuals with LS (n = 65) was investigated (Lund cohort; Fig. 1 a). A total of 48 individuals with LoF variants in either of MLH1 (n = 12), MSH2 (n = 15) and MSH6 (n = 21) were subjected to constitutional HTT CAG repeat size estimation and compared with controls (n = 19) (Fig. 1a). The sum of CAG repeats did not differ significantly between individuals with LoF variants in MLH1 (37.75 CAG ± 6.3; mean ± SD), MSH2 (37.93 ± 5.8), MSH6 (36.95 ± 4.0) and controls (36.84 ± 4.4) (Fig. 2; Table 1). Three individuals had one HTT allele with CAG repeats in the intermediate allele interval (MLH1, n = 1, MSH2, n = 1, controls, n = 1; Table 1). The remaining alleles were in the normal allele interval (≤ 26 CAG repeats).
Characterization and analysis of the Bochum cohort
Subsequently, a larger cohort of individuals with LS (n = 207) was investigated (Bochum cohort; Fig. 1b). A total of 171 individuals (mean age 47.1 ± 12.9 years) with LoF variants in either of MLH1 (n = 60; mean age 47.6 ± 13.1 years), MSH2 (n = 88; mean age 46.6 ± 13.1 years) and MSH6 (n = 23; mean age 48 ± 11.5 years) were subjected to constitutional HTT CAG repeat size estimation and compared with controls (n = 559; mean age 27.3 ± 9.4 years) (Fig. 1b). The sum of CAG repeats in individuals with LoF variants in MLH1 was significantly smaller than in controls (35.40 CAG ± 3.6 vs. 36.89 CAG ± 4.5 in controls, Student’s t-test with Bonferroni correction, p = 0.014, CI − 2.766 to − 0.310) (Fig. 3; Table 1) and remained significantly smaller also after removal of individuals with HTT intermediate alleles in an additional analysis (35.19 CAG ± 3.2 vs. 36.34 CAG ± 4.0 in controls; Student’s t-test, p = 0.031, CI − 2.202 to − 0.104) (Table 1). The sum of CAG repeats in individuals with LoF variants in MSH2 (36.41 CAG ± 5.2) and MSH6 (36.17 CAG ± 4.2) did not differ significantly from controls (Fig. 3; Table 1). Thirty-four individuals had one HTT allele with CAG repeats in the intermediate allele interval (MLH1 n = 1, MSH2 n = 3, MSH6 n = 1, controls n = 29; Table 1). The remaining alleles were in the normal allele interval. The fraction of individuals with an intermediate allele among individuals with LS did not differ significantly from that in controls, but the fraction was consistently lower in all LS genetic subgroups (Table 1). The mean somatic HTT CAG expansion index (EI) value, which typically is increased in tissues from individuals with HD17, did not differ significantly between individuals with LoF variants in MLH1 (EI = 0.099), MSH2 (EI = 0.122), MSH6 (EI = 0.100) and controls (EI = 0.131) (Table 1). However, notably all LS genetic subgroups showed a lower mean EI value compared to controls (Table 1).
Discussion
Investigation of HTT CAG repeat size in lymphocyte DNA from 217 individuals from two different LS cohorts, showed a small but statistically significant CAG repeat size reduction in a subgroup of 60 MLH1 LoF heterozygotes from the larger cohort from Bochum. The frequencies of HTT intermediate alleles and somatic EI values were consistently lower in all LS genetic groups compared to controls in the Bochum cohort, but the observed differences were individually not statistically significant. The CAG repeat size in the Bochum MLH1 LS subgroup remained significantly smaller compared to controls also after removal of individuals with HTT intermediate alleles. Nucleotide repeat instability and an increased mutational burden is a known phenomenon in dMMR cancers in LS patients following somatic “second hit” of the remaining wild-type MMR allele16, and in all tissues in individuals with constitutional biallelic MMR deficiency18. However, to the best of our knowledge, MMR gene haploinsufficiency in humans has to date not been reported to affect constitutional nucleotide repeat size. A recent whole genome sequencing (WGS) study of non-neoplastic tissue samples from individuals with LS failed to detect any changes in the repertoire of mutational processes or mutation rates19. Yet, subtle nucleotide repeat variations could have escaped detection using WGS technology due to limited methodological accuracy in regions with STRs compared to PCR fragment-based analyses20. Like previous population-based observations21, we found a large CAG repeat size variation between HTT alleles, both intra- and inter-individually which, together with the lack of parental HTT repeat size data prevent us from a more detailed data interpretation. Clearly, the deciphering of which HTT allele has cosegregated with the LS allele in each individual, e.g., by LS family trio analyses, would have enhanced our data interpretation considerably, allowing us to identify individuals, or certain repeats size intervals including variable CAA interruptions that may account for the observed repeat variation in the Bochum LS cohort. Although HTT intermediate alleles appear under-represented in the Bochum LS cohort, especially in individuals with MLH1-associated LS (1.7%), compared to the controls used (5.2%) and to reported population-based frequencies (6.8%;21), interpretation of data should be made with caution due to the limited size of the cohort subgroups and the absence of LS family trio data. Possibly, the observed frequency of HTT intermediate alleles in LS in our study could reflect intergenerational CAG repeat contractions of such alleles into the normal repeat-size interval. However, HTT intermediate alleles alone are not responsible for the observed CAG repeat-size reduction in the Bochum MLH1 LS subgroup as the removal of this category of alleles from our calculations had little impact. Somatic EI values did not differ significantly between the LS subgroups and controls, but notably the values were consistently lower in all LS genetic subcategories. As EI values normally are positively age-dependent22 and since the mean age in the Bochum LS group was higher than in the control group, the EI value gap between the two groups could potentially be an underestimate. Clearly, the use of age-matched controls would have sharpened interpretation of EI values. Experimentally, in mouse models of HD, there is long-standing evidence that reduced expression of the MMR proteins Msh2, Msh3, Mlh1 or Mlh3 counteracts somatic CAG repeat expansion23,24 and de-escalates the HD experimental pathogenic process25. More recently, reduced expression of the endo- and exonuclease Fan1 was shown to promote somatic CAG repeat expansion in an Mlh1-dependent manner, i.e., suppression of Mlh1 blocked Fan1-induced repeat expansion26,27. There is now mounting evidence that the MMR pathway contributes to the expansion of unstable pathogenic nucleotide repeats in HD and other human hereditary neurodegenerative diseases14. Given the present results and current knowledge in this field of research, it could be speculated that individuals with LS could be less prone to HTT CAG repeat expansion. In summary, this study indicates that MMR gene haploinsufficiency, in particular for MLH1, could be associated with a propensity for reduced constitutional HTT CAG repeat size. Further investigations, e.g., with larger LS case samples and LS family trio WGS analyses are required to confirm our results. Additional studies should also be encouraged to explore the possible impact of MMR gene haploinsufficiency on other nucleotide repeat regions in the human genome.
Methods
Cohort information
Lymphocyte DNA was retrieved from two different cohorts of index individuals diagnosed with LS from Sweden and Germany (Lund cohort and Bochum cohort, respectively) carrying germline class 4 (likely pathogenic) or class 5 (pathogenic) variants in MLH1, MSH2, MSH6 and PMS2 according to variant classification criteria by The American College of Medical Genetics and Genomics (ACMG)28 or The International Society of Gastrointestinal Hereditary Tumours variant database29, and from controls (Fig. 1). A subgroup of the Lund cohort was previously presented in a pre-publication (Dalene Skarping et al. 2022, MedRxiv, https://doi.org/10.1101/2022.05.28.22275723). Controls in the present study were individuals diagnosed with immunohistochemically MMR proficient colorectal cancers during 1999–2011 from whom tumor tissue DNA had also been archived (Lund cohort) or self-reported healthy university students (Bochum cohort). Controls from Bochum were excluded if they or any of their close relatives suffered from neurological and/or mental illnesses, as assessed by a self-report questionnaire. Individuals with LS-associated missense variants predicted to cause single amino acid substitutions were excluded to avoid variants with partial LoF, and variants with unclear pathogenic mechanism. Other types of LS-associated variants, i.e., nonsense variants, variants altering the reading frame or splicing, deletions or duplications of exon(s) were considered complete LoF alleles. Individuals with variants in the MMR gene PMS2 were excluded due to the limited number of such individuals in both cohorts (Fig. 1).
HTT CAG repeat size estimation and somatic expansion ratio calculation
HTT germline CAG repeat size estimation was performed using standard protocols for PCR amplification and capillary electrophoresis fragment analysis with a validated accuracy of ± 1 CAG repeat for alleles with < 45 repetitions and ± 3 CAG repetitions for alleles with 45 or more repeats using PCR primers (Lund cohort) HD1: 5′ ATGAAGGCCTTCGAGTCCCTCAAGTCCTTC 3′ and HD3: 5′ Hex-GGCGGTGGCGGCTGTTGCTGCTGCTGC 3′ as described30, or (Bochum cohort) Hu4: (F) 6-FAM-5′-ATGGCGACCCTGGAAAAGCTGATGAA) and Hu5: (R) (5′-GGCGGTGGCGGCTGTTGCTGCTGCTGCTGC) as described31,32. A canonical glutamine-encoding repeat sequence in HTT was assumed. PCR products were resolved using the ABI 3500XL Genetic Analyzer (Applied Biosystems) using GeneMapper v6 software and GeneScan 500-ROX as internal size standard (Lund cohort), or ABI 3500XL Genetic Analyzer (Applied Biosystems), GeneMapper v4.1 software and GeneScan 500-ROX as internal size standard (Bochum cohort). Somatic CAG repeat EI values were derived from indices from GeneMapper peak height data and calculated as described17, considering only expansion peaks to the right of the highest (modal allele) peak, using 250 consecutively selected individuals from the Bochum control group as controls.
Statistical analyses
CAG repeat size was converted to integers according to clinical genetic laboratory diagnostic routines30. The methodological estimation error ± 1 repeat was excluded from statistical calculations. Since the methods used in this study do not unmask which HTT allele has co-segregated with the LS-associated variant, the sum of HTT CAG repeats in each individual was calculated and used in all analyses except for somatic EI calculations. Mean values for sum of CAG repeats and standard deviation (SD) with 95% confidence interval (CI) were calculated for each MMR gene. Student’s t-test was used. P-values < 0.05 were considered significant. Bonferroni correction was applied to adjust for multiple comparisons, i.e., MLH1, MSH2 and MSH6 vs. controls, respectively, following which P-values < 0.017 were considered significant. Calculations were performed using SPSS Statistics for Windows (SPSS Inc., Chicago, Ill., USA).
Ethics approval
This study was approved by The Regional Ethical Review Board in Lund, Sweden (application no. 2013/468 and application no. 2015/211), approved, or waived following anonymization procedures by the Swedish Ethical Review Agency (application no. 2019-02312 and application no. 2021-06254-02, respectively), and approved by the Ethics Review Board of the Ruhr University in Bochum, Germany, (application no. 18-6563-BR). Informed written consent was required and obtained from all individuals (Bochum cohort) or waived (Lund cohort) following anonymization of DNA samples prior to HTT CAG repeat size analysis (application no. 2021-06254-02). No individual-level data are published in this study. All methods were performed in accordance with the relevant local guidelines and regulations.
Data availability
This manuscript contains all relevant data to the study. The raw case data sets generated and/or analyzed during the current study are not openly available due to ethical considerations related to patient privacy. Access to raw data and detailed protocols may be considered on request to the corresponding authors SGM or HPN.
References
Paulson, H. Repeat expansion diseases. Handb. Clin. Neurol. 147, 105–123. https://doi.org/10.1016/B978-0-444-63233-3.00009-9 (2018).
The Huntington’s Disease Collaborative Research Group. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington’s disease chromosomes. Cell 72, 971–983. https://doi.org/10.1016/0092-8674(93)90585-e (1993).
Tabrizi, S. J., Flower, M. D., Ross, C. A. & Wild, E. J. Huntington disease: New insights into molecular pathogenesis and therapeutic opportunities. Nat. Rev. Neurol. 16, 529–546. https://doi.org/10.1038/s41582-020-0389-4 (2020).
Vonsattel, J. P. et al. Neuropathological classification of Huntington’s disease. J. Neuropathol. Exp. Neurol. 44, 559–577. https://doi.org/10.1097/00005072-198511000-00003 (1985).
Langbehn, D. R. et al. A new model for prediction of the age of onset and penetrance for Huntington’s disease based on CAG length. Clin. Genet. 65, 267–277. https://doi.org/10.1111/j.1399-0004.2004.00241.x (2004).
Findlay Black, H. et al. Frequency of the loss of CAA interruption in the HTT CAG tract and implications for Huntington disease in the reduced penetrance range. Genet. Med. 22, 2108–2113. https://doi.org/10.1038/s41436-020-0917-z (2020).
Genetic Modifiers of Huntington’s Disease (GeM-HD) Consortium. CAG Repeat Not Polyglutamine Length Determines Timing of Huntington's Disease Onset. Cell 178, 887–900 e814. https://doi.org/10.1016/j.cell.2019.06.036 (2019).
Wright, G. E. B. et al. Length of uninterrupted CAG, independent of polyglutamine size, results in increased somatic instability, hastening onset of Huntington disease. Am. J. Hum. Genet. 104, 1116–1126. https://doi.org/10.1016/j.ajhg.2019.04.007 (2019).
Bettencourt, C. et al. DNA repair pathways underlie a common genetic mechanism modulating onset in polyglutamine diseases. Ann. Neurol. 79, 983–990. https://doi.org/10.1002/ana.24656 (2016).
Genetic Modifiers of Huntington’s Disease (GeM-HD) Consortium. Identification of Genetic Factors that Modify Clinical Onset of Huntington's Disease. Cell 162, 516–526. https://doi.org/10.1016/j.cell.2015.07.003 (2015).
Goold, R. et al. FAN1 modifies Huntington’s disease progression by stabilizing the expanded HTT CAG repeat. Hum. Mol. Genet. 28, 650–661. https://doi.org/10.1093/hmg/ddy375 (2019).
Lee, J. M. et al. A modifier of Huntington’s disease onset at the MLH1 locus. Hum. Mol. Genet. 26, 3859–3867. https://doi.org/10.1093/hmg/ddx286 (2017).
Moss, D. J. H. et al. Identification of genetic variants associated with Huntington’s disease progression: A genome-wide association study. Lancet Neurol. 16, 701–711. https://doi.org/10.1016/S1474-4422(17)30161-8 (2017).
Iyer, R. R. & Pluciennik, A. DNA mismatch repair and its role in Huntington’s disease. J. Huntingtons Dis. 10, 75–94. https://doi.org/10.3233/JHD-200438 (2021).
Levinson, G. & Gutman, G. A. Slipped-strand mispairing: A major mechanism for DNA sequence evolution. Mol. Biol. Evol. 4, 203–221. https://doi.org/10.1093/oxfordjournals.molbev.a040442 (1987).
Curtius, K., Gupta, S. & Boland, C. R. Review article: Lynch Syndrome-a mechanistic and clinical management update. Aliment Pharmacol. Ther. 55, 960–977. https://doi.org/10.1111/apt.16826 (2022).
Mouro Pinto, R. et al. Patterns of CAG repeat instability in the central nervous system and periphery in Huntington's disease and in spinocerebellar ataxia type 1. Hum. Mol. Genet. 29, 2551–2567. https://doi.org/10.1093/hmg/ddaa139 (2020).
Aronson, M. et al. Diagnostic criteria for constitutional mismatch repair deficiency (CMMRD): Recommendations from the international consensus working group. J. Med. Genet. 59, 318–327. https://doi.org/10.1136/jmedgenet-2020-107627 (2022).
Lee, B. C. H. et al. Mutational landscape of normal epithelial cells in Lynch Syndrome patients. Nat. Commun. 13, 2710. https://doi.org/10.1038/s41467-022-29920-2 (2022).
Ibanez, K. et al. Whole genome sequencing for the diagnosis of neurological repeat expansion disorders in the UK: A retrospective diagnostic accuracy and prospective clinical validation study. Lancet Neurol. 21, 234–245. https://doi.org/10.1016/S1474-4422(21)00462-2 (2022).
Sundblom, J. et al. High frequency of intermediary alleles in the HTT gene in Northern Sweden—The Swedish Huntingtin Alleles and Phenotype (SHAPE) study. Sci. Rep. 10, 9853. https://doi.org/10.1038/s41598-020-66643-0 (2020).
Monckton, D. G. The contribution of somatic expansion of the CAG repeat to symptomatic development in Huntington’s Disease: A historical perspective. J. Huntingtons Dis. 10, 7–33. https://doi.org/10.3233/JHD-200429 (2021).
Dragileva, E. et al. Intergenerational and striatal CAG repeat instability in Huntington’s disease knock-in mice involve different DNA repair genes. Neurobiol. Dis. 33, 37–47. https://doi.org/10.1016/j.nbd.2008.09.014 (2009).
Manley, K., Shirley, T. L., Flaherty, L. & Messer, A. Msh2 deficiency prevents in vivo somatic instability of the CAG repeat in Huntington disease transgenic mice. Nat. Genet. 23, 471–473. https://doi.org/10.1038/70598 (1999).
Pinto, R. M. et al. Mismatch repair genes Mlh1 and Mlh3 modify CAG instability in Huntington’s disease mice: genome-wide and candidate approaches. PLoS Genet. 9, e1003930. https://doi.org/10.1371/journal.pgen.1003930 (2013).
Loupe, J. M. et al. Promotion of somatic CAG repeat expansion by Fan1 knock-out in Huntington’s disease knock-in mice is blocked by Mlh1 knock-out. Hum. Mol. Genet. 29, 3044–3053. https://doi.org/10.1093/hmg/ddaa196 (2020).
Porro, A. et al. FAN1-MLH1 interaction affects repair of DNA interstrand cross-links and slipped-CAG/CTG repeats. Sci. Adv. 7. https://doi.org/10.1126/sciadv.abf7906 (2021).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424. https://doi.org/10.1038/gim.2015.30 (2015).
The International Society for Gastrointestinal Hereditary Tumours Data Base, http://insight-database.org/.
Losekoot, M. et al. EMQN/CMGS best practice guidelines for the molecular genetic testing of Huntington disease. Eur. J. Hum. Genet. 21, 480–486. https://doi.org/10.1038/ejhg.2012.200 (2013).
Riess, O. et al. Precise mapping of the brain alpha 2-adrenergic receptor gene within chromosome 4p16. Genomics 19, 298–302. https://doi.org/10.1006/geno.1994.1061 (1994).
Rubinsztein, D. C., Barton, D. E., Davison, B. C. & Ferguson-Smith, M. A. Analysis of the huntingtin gene reveals a trinucleotide-length polymorphism in the region of the gene that contains two CCG-rich stretches and a correlation between decreased age of onset of Huntington’s disease and CAG repeat number. Hum. Mol. Genet. 2, 1713–1715. https://doi.org/10.1093/hmg/2.10.1713 (1993).
Acknowledgements
Acknowledgements to the Department of Clinical Genetics and Pathology, Office for Medical Service in Lund for support with personnel, equipment, and materials, and to Gabriele Schlüter for excellent technical assistance. This study was supported by grants from the Southern health care region in Sweden (SGM), the Swedish state under the agreement between the Swedish government and the county councils, the ALF-agreement for clinical research (KDS, ÅP), Knut and Alice Wallenberg Foundation (grant number 2019.0467; ÅP), the Swedish Research Council (grant number 2022-01092; ÅP), and by the E-Rare Research Programme ERA-NET E-Rare 3 “TreatPolyQ” awarded to HPN (BMBF grant number 01GM1804).
Funding
Open access funding provided by Lund University.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Data collection and analysis were performed by KDS and LA. The first draft of the manuscript was written by KDS, LA and SGM. All authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dalene Skarping, K., Arning, L., Petersén, Å. et al. Attenuated huntingtin gene CAG nucleotide repeat size in individuals with Lynch syndrome. Sci Rep 14, 4300 (2024). https://doi.org/10.1038/s41598-024-54277-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-54277-5
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.