Abstract
To examine the interrelationship of aging extension and modification of lipid metabolism under chronic caloric restriction (CCR; reduced concentration of the green algae Tetraselmis suecica) in the monogonont rotifer Brachionus koreanus, we assessed life cycle parameters, fatty acid composition, and expression of sirtuin and genes related to lipid metabolism. B. koreanus in the 5% T. suecica group showed an increased life span but decreased reproduction. Based on this finding, we chose 5% T. suecica for further experiments and compared the data with those for 100% T. suecica. Upregulation of sirtuin gene expression was observed under CCR. In addition, despite the reduction in the amount of total fatty acid (FA) and the area of triacylglycerol, increases in the ratios of saturated fatty acid and monounsaturated fatty acid (MUFA) to total FA in 5%-exposed B. koreanus were observed. Furthermore, mRNA expression analysis confirmed that CCR promoted the synthesis of MUFA through Δ9 desaturase. Moreover, expression of the docosahexaenoic acid (DHA) synthesizing gene Δ4 desaturase was also upregulated, together with DHA content. These data suggest that CCR modified protein acetylation and lipid metabolism, leading to a decrease in reproduction and consequently resulting in life span extension.
Similar content being viewed by others
Introduction
The energy trade-off between life span and reproduction is commonly observed in most animal taxa1. This phenomenon can be affected by various abiotic and biotic factors, such as temperature2,3, pH4, and metabolic changes5. Among them, chronic caloric restriction (CCR) is a well-known regulator leading to an energy trade-off in various organisms such as mice6,7,8, the grasshopper Romalea microptera9, the nematode Caenorhabditis elegans10, and the rotifer Brachionus manjavacas11. Based on previous findings, a simple hypothesis to explain this phenomenon is that this is one of the strategies organisms use to overcome unfavorable changes in their surrounding environment.
Many studies on the mechanisms of prolonging life span have focused on protein deacetylase. From yeast to human, life span is regulated by protein modification12,13. In particular, the gene encoding sirtuin, one of the protein deacetylases, is a well-known marker of life span regulation14. Sirtuin genes were initially identified as transcription-silencing protein deacetylases in yeast. Calorie restriction was shown to extend life span through upregulation of yeast sirtuin genes15,16. In addition, upregulation of sirtuin2 was reported to extend life span in the fruit fly Drosophila melanogaster17, although there is still some controversy regarding this finding18. In mammals, seven sirtuin genes have been identified, and sirtuin1 was found to have a similar function to sirtuin2 in yeast in terms of longevity19. Therefore, study of the function of sirtuin genes is important in relation to life span.
In addition to protein deacetylase mechanisms, recent studies have revealed that CCR affects lipid metabolism. In mice, increasing the ratio of dietary saturated fatty acid (SFA) to monounsaturated fatty acid (MUFA) increases the life span by lowering the ratio of dietary polyunsaturated fatty acid (PUFA) composition7. Other researchers have found that CCR extends life span and affects overall lipid metabolism (e.g., circulating lipid, hepatic cholesterol level, triacylglycerol)6. In addition, an increase in life span was positively correlated with an increase in MUFA synthesis in H3K4me3 (trimethylates histone H3 on lysine 4) methyltransferase-deficient C. elegans20. However, since lipid metabolism plays a multifunctional role in various mechanisms such as long-term energy storage, intercellular and intracellular signaling, and membrane homeostasis, confirmation of the direct correlation between lipid metabolism and CCR remains a challenging issue.
As microzooplankton, rotifers (phylum Rotifera) are widely distributed throughout aquatic ecosystems and function as a bridge between producers and higher-level consumers in aquatic food chains21. Because of their small size (100–200 μm), easy maintenance, slow locomotion, and short reproductive cycle (~24 h), rotifers have established as an excellent experimental model organism in aquatic research22,23. Rotifers have also been used as a model species for aging experiments in response to temperature24, antioxidants25, and CCR11,26,27. In addition, a recent study used rotifers to screen drugs for follow-up in a vertebrate model28. Therefore, B. koreanus is a suitable organism to confirm the energy trade-off between lifespan and reproduction. Also, B. koreanus has the additional advantage over other aquatic species of the ability to reveal mechanisms as we have successfully identified its whole RNA sequence29.
The purpose of this investigation is to examine the interrelationship of aging extension and modification of lipid metabolism under CCR in B. koreanus. To assess the effect of CCR, we initially observed the life cycle parameters (life span, cumulative offspring, and offspring production per day) under eight different dietary conditions from ad libitum feeding with T. suecica (100%) to starvation (0%). Based on our results, we chose 5% T. suecica, which showed an increase in life span and decrease in fecundity, for further analysis of modulation of the expression of sirtuin and genes related to lipid metabolism and fatty acid composition under CCR in the rotifer B. koreanus. This study provides new insight into the interplay between CCR and the relationships between protein deacetylase genes, lipid metabolism, and life cycle parameters.
Results
Effects of CCR on Life Cycle Parameters
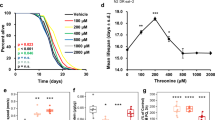
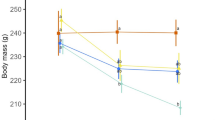
Life cycle parameters were observed after exposure to different food concentrations (100, 75, 50, 25, 10, 5, 1, and 0%). In life span assessment (Fig. 1A and Suppl. Table 3), a significant (P < 0.05) extension of life span (about 48.6%) was observed only in the 5%-exposed group compared with the 100%-exposed group. Maximum and minimum life spans were 15 and 8 days in the 5%-exposed group and 10 and 6 days in the 100%-exposed group, respectively. In terms of cumulative offspring (Fig. 1B), 1%- and 5%-exposed groups showed a delay, and the 0% group did not produce any offspring. However, no changes in the mean total offspring were observed between the 100%- and 5%-exposed groups (Fig. 1B; Suppl. Table 3). Regarding the average number of offspring per day (Fig. 1C), CCR treatment resulted in a reduction in daily reproduction throughout the experiment, but the number of reproductive days was increased, especially in the 5%-exposed group. Based on in-vivo data, further experiments were performed using 100% being the control and 5% as test group.
Effects of chronic caloric restriction on life cycle parameters in B. koreanus exposed to different concentrations of T. suecica (100 [Control], 75, 50, 25, 10, 5, 1, and 0%). (A) Life span, (B) cumulative offspring and (C) number of offspring per day. Asterisk (*) indicates significant difference between test and control groups (P < 0.05, Bonferroni’s correction).
Modulation of Sirtuin Genes under CCR
The mRNA expression of sirtuin genes of B. koreanus was measured under 5% T. suecica CCR conditions in a time-dependent manner (24, 48, and 96 h) (Fig. 2; Suppl. Table 2). With the exception of sirtuin 4, all of the sirtuin genes were significantly induced (P < 0.05) under CCR (5%-exposed group) compared with the control.
The Area of Lipid Droplets in Response to CCR
The area of LDs was measured after exposure to 100% and 5% T. suecica for 24, 48, and 96 h (Fig. 3). As expected, the area of the 5% group was significantly decreased (P < 0.05) compared with the control (100%) for all exposure times.
Lipid accumulation in B. koreanus exposed to 100% and 5% T. suecica for 24, 48, and 96 h. (A) Confocal microscopy analysis of Nile red-stained B. koreanus: (a) bright field, (b) fluorescent, and (c) merged. Red fluorescence indicates accumulated neutral lipids. (B) The area of triacylglycerol in B. koreanus is represented as a percentage of the relative area using the LAS image analysis tool (n = 30). Significant differences compared with the control value are indicated by an asterisk (*) (P < 0.05, Student’s t-test).
Effect of CCR on Fatty Acid Composition
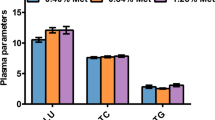
Fatty acid composition was measured after exposure to 100% and 5% T. suecica for 24, 48, and 96 h. Similar to the results for LDs area, the amount of total fatty acid was significantly decreased (P < 0.05) in the 5%-exposed group at all time points (Suppl. Table 4). Analysis of fatty acid composition in the 100% and 5% group revealed a decrease in most of the fatty acids (e.g., C18:1n-9, C20:3n-3, and C20:5n-3) in the 5%-exposed group. However, results for single fatty acids (Fig. 4) showed an increasing trend in the relative amounts of SFAs and MUFAs (P < 0.05). Regardless of the reduction in the relative amount of PUFA, a significant increase (P < 0.05) in the relative amount of DHA (C22:6n-3) was observed.
Modulation of Genes Related to Lipid Metabolism under CCR
The mRNA expression of lipid metabolism-related genes was measured in the 5%-exposed group compared with the control (100%) at 24, 48, and 96 h (Fig. 5; Suppl. Table 2). Genes involved in de novo lipogenesis (DNL: acetyl-CoA carboxylase [ACC], ATP-citrate lyase [ACLY], and β-keto-acyl-[acyl-carrier-protein] synthase [KAS]) were significantly upregulated (P < 0.05) under CCR. Desaturases and elongases were also significantly modulated. Although food supply was lower in the 5%-exposed group than in the 100%-exposed group (control), Δ9 desaturase (at 24 h), Δ4 desaturase (at 96 h), and elongase 6 (at all time points) mRNAs were significantly upregulated (P < 0.05) compared with the control (100%).
Transcription profile of lipid metabolism-related genes after exposure to 100% or 5% T. suecica at 24, 48, and 96 h represented by a heat map. (A) De novo lipogenesis genes (ATP-citrate lyase [ACLY], acetyl-CoA carboxylase [ACC], and β-keto-acyl-[acyl-carrier-protein] synthase [KAS]), (B) fatty acid structure modification genes (desaturase [DES] and elongase [ELO]), and (C) triacylglycerol formation genes (monoacylglycerol acyltransferase [MGAT], diacylglycerol acyltransferase [DGAT], and Lipin 1 and 2).
Discussion
Life span extension and reproductive retardation under CCR are common phenomena in most animals. However, studies on the interrelationship of the life cycle parameter outcomes and lipid metabolism under CCR over the entire lifetime are still insufficient. Therefore, in this study, we assessed life cycle parameters (e.g., life span and fecundity), measured the expression of protein deacetylase sirtuin genes and lipid metabolism–related genes, and examined fatty acid composition to investigate the aging extension and modification of lipid metabolism in the monogonont rotifer B. koreanus under CCR.
Among the various food concentrations tested (from 0% to 100% of T. suecica), only the 5%-exposed group demonstrated a delay in cumulative offspring and an increase in reproductive days with significant life span extension in B. koreanus (Fig. 1; Suppl. Table 3), indicating that life span extension and fecundity (cumulative offspring) under CCR showed an inverse relationship. This phenomenon, known as the disposable soma hypothesis, is also referred to as the “cost of reproduction” in rotifers30,31. Rotifers have been widely used to investigate the aging system28,32. For example, extension of the life span was demonstrated in a less than 25% treatment group compared to the 100% treatment group in B. manjavacas11. Also, in mass culture of the rotifer B. plicatilis, fasting after hatching (days 1–4) reduced the reproductive success rate but increased life span33. This phenomenon has been observed not only in the rotifer Brachionus species11,33, but also in mice6,7,8, grasshopper9, and nematode10. Taken together, these results are considered to be one of the strategy to reduce reproduction (cumulative offspring) and prolong the life span under conditions of insufficient food source.
We identified sirtuin genes, which are well-known protein deacetylases, in B. koreanus using RNA-seq.29 and a whole-genome database (GenBank no. PJRA00000000) produced in our laboratory and analyzed their mRNA expression under CCR. Based on the results of the life cycle parameters, the 5%-exposed group was suitable candidates for analysis of mRNA expression of sirtuin genes compared to the 100%-exposed group as the 5%-exposed group demonstrated an increase in mean life span. In B. koreanus, all sirtuin genes except sirtuin 4 were upregulated under CCR over 96 h (Fig. 2; Suppl. Table 2). In general, sirtuins are involved in cellular functions such as DNA repair, inflammatory response, cell cycle, and apoptosis14. In addition, recent studies have focused on the strong relationship between sirtuin expression and aging processes. For example, sirtuin 2 was first identified in yeast, and a correlation between sirtuin activity and longevity was revealed16,34. Furthermore, in mice, the expression of sirtuin genes increased life span35,36, whereas sirtuin knockout decreased life span37,38, indicating that expression of sirtuin genes is a key element in prolonging the life span. Furthermore, calorie restriction has been reported to extend life span through upregulation of sirtuins in yeast and mice8,15,16. To summarize, upregulation of sirtuin mRNA expression under CCR, which is similarly conserved from yeast to mammals, as well as in B. koreanus, implies the functional conservation of sirtuins as life span regulators.
Analysis of LDs and fatty acid composition showed that CCR modulated lipid metabolism in B. koreanus (Fig. 3; Suppl. Table 4). In our CCR model in the rotifer B. koreanus, the food exposure was only 5% of that of the control; therefore, as expected, the area of LDs and the absolute values of total and single fatty acids were decreased. Indeed, the changes in the LDs are likely commonly observed indication of the stressful condition in aquatic organisms. For example, under high temperature shift, trend between the survival of each generation and lipid contents was negatively correlated in the estuarine copepod Eurytemora affinis39. Furthermore, our results are concordant with findings in mice exposed to CCR, which showed a reduction in total body weight and decrease in TAG and cholesterol7,8. Despite the decrease in total fatty acid and the area of LDs, the ratio of SFA to MUFA among total fatty acids in the 5%-exposed group showed a relative increase in B. koreanus (Fig. 4). This result is consistent with the findings from mice and nematode. In mice7, the composition of dietary lipids has an effect on the life span. In particular, administration of an energy source containing a low ratio of PUFA and high ratio of MUFA and SFA resulted in life span extension. In addition, a knockout mutant of H3K4me3 methyltransferase, a life span prolonging gene, in the nematode C. elegans12 exhibited differences in the modulation of lipid metabolism in intestine and germline compared with the wild type20. For example, in the intestine, an increase in the synthesis of NL and MUFA and their synthesizing genes was observed, but levels of the germline target genes were reduced. Taken together, these data show that CCR reduces total fatty acid, the area of LDs, and reproduction, while promoting MUFA synthesis, which can further implicate to the extension of life span at the cost of the physiological changes.
The correlation between the expression of lipid metabolism-related genes and fatty acid composition was investigated in the rotifer B. koreanus under CCR (Fig. 5; Suppl. Table 2). DNL genes (ACC, ACLY, and KAS) are involved in the synthesis of fatty acids from glucose, the final metabolite of which is palmitic acid (PA, C16:0)40. In the present study, the amount of PA was initially decreased at 24 h but was increased significantly compared to the 100%-exposed group at 96 h, suggesting that increased expression of DNL genes led to the synthesis of PA from glucose. Also, the increase in PA might influence the proportion of MUFA in B. koreanus. The synthesis of MUFA is affected by an increase in Δ9 desaturase, which is the key gene involved in the production of MUFA (palmitoleic acid [16:1n-9] and oleic acid [C18:1n-9]) from saturated fatty acid (PA and stearic acid [C18:0])41. These results are consistent with findings in the H3K4me3 methyltransferase mutant of C. elegans, which showed life span extension and upregulation of Δ9 desaturases, leading to an increase in the MUFA content12,20.
An increase in Δ4 desaturase in the rotifer B. koreanus might have been involved in the synthesis of docosahexaenoic acid (DHA, C22:6n-3), which was the only increased PUFA, under CCR. In general, DHA has anti-inflammatory activity and is metabolized into inflammation-resolving mediators (e.g., resolvins, protectins, and maresins) through cyclooxygenase and lipoxygenase pathways42,43. There are two DHA synthetic pathways: a pathway created by β-oxidation after the action of elongase and Δ6 desaturase on docosapentaenoic acid (DPA, C22:5n-3) or direct synthesis of DHA from DPA using Δ4 desaturase42. We measured the composition of FA of T. suecica in normal condition, and interestingly, DHA was not detected (Suppl. Table 5). Other studies also reported very low DHA content in T. suecica44. Therefore, DHA in B. koreanus was only synthesized from other FA utilizing enzymes. Therefore, our results indicate that an increase in Δ4 desaturase expression modulates DHA synthesis. In terms of TAG synthesis in the rotifer B. koreanus, expression of MGAT and Lipin 2 genes was increased, which is consistent with the results of the H3K4me3 methyltransferase mutant C. elegans20. However, unlike the C. elegans experiment, the energy source administered to experimental organisms was limited to 5% of the control, thus the area of TAG appeared to decrease by approximately 60% compared with the control (100%).
To summarize the relationship between fatty acid composition and expression of lipid metabolism-related genes (Figs 4, 5; Suppl. Table 2), the mRNA expression level of lipid metabolism genes was analyzed. Based on our results, we suggest that the rotifer B. koreanus utilizes its energy to synthesize SFA and MUFA using the DNL pathway and Δ9 desaturase, respectively. Also, Δ4 desaturase is specifically involved in the synthesis of DHA, which is the only constituent of PUFA that is increased under CCR.
In conclusion, the phenomenon called “cost of reproduction” was confirmed in the monogonont rotifer B. koreanus, via utilizing of aging extension mechanism under CCR through increase in mRNA expression (sirtuin and MUFA synthesis genes) and the proportional increase in MUFA (Fig. 6). From this investigation, it can be suggested that one of the survival strategies to endure food shortages is to modulate the mechanism (protein deacetylation and lipid metabolism) in B. koreanus, thereby lowering reproduction and increasing life span. This study will provide a valuable insight into the mechanism of aging extension and will contribute a step toward establishment of B. koreanus as a model species for in-depth mechanistical studies.
Methods
Culture and Assessment of Life Cycle Parameters
The species identification was confirmed by morphological analysis45,46 and sequencing of the mitochondrial DNA gene CO147. In this study, one mL ASW with various concentrations of T. suecica (100 [6 × 105 cells/ml], 75, 50, 25, 10, 5, 1, and 0%) were renewed every 24 h. We determined the number of T. suecica cells based on the reports on life-span extension by caloric restriction in the rotifer B. manjavacas11,26 and set 6 × 105 cells/ml as 100% for this experiment. The detailed information on the procedure for the assessment of life cycle parameters is incorporated in the Supplementary file. All experiments were performed in biological triplicate.
Measurement of Triacylglycerol in Response to CCR
To examine the effects of chronic caloric restriction on neutral lipid accumulation in vivo, Nile red staining was performed. Two groups of B. koreanus were fed 100% or 5% T. suecica for 24, 48, and 96 h. The 100% group was considered the control. A detailed Nile red staining method is provided in the Supplementary file.
Analysis of Fatty Acid Composition under CCR
To analyze variations in fatty acid composition in response to chronic caloric restriction in B. koreanus (100% and 5% T. suecica at 24, 48, and 96 h), we followed the protocol provided by Hama and Handa48 with minor modifications, and detailed information is provided in the Supplementary section. All experiments were performed in triplicate.
Expression of Sirtuin and Genes Related to Lipid Metabolism
To examine the expression of sirtuin and lipid metabolism-related genes, in silico analysis of B. koreanus RNA-seq information was performed29. The identification of sirtuin and lipid-metabolism-related genes were performed using BLAST analysis. To investigate the CCR-induced modulation of sirtuin and lipid metabolism-related genes, we measured mRNA expression levels over 96 h (24, 48, and 96 h) in response to 5% T. suecica exposure. All experiments were performed in technical triplicate. The information regarding identification of genes and the protocols for the mRNA expression are further provided in the Supplementary file.
Statistical Analysis
For statistical analysis, SPSS ver. 18.0 (SPSS Inc., Chicago, IL, USA) was used and the data are presented as mean ± S.D. Student’s paired t-test and one-way ANOVA were used to analyze the significant differences between the control and test groups, followed by Tukey’s test. Using R statistical software (version 3.2.1 of the R Foundation for Statistical Computing Platform©, 2015), Kaplan–Meier survival curves for significance were calculated. Overall, the differences with P < 0.05 were considered significant.
References
Kooijman, S. A. & Troost, T. A. Quantitative steps in the evolution of metabolic organisation as specified by the dynamic energy budget theory. Biol. Rev. Camb. Philos. Soc. 82, 113–142 (2007).
Kauler, P. & Enesco, H. E. The effect of temperature on life history parameters and cost of reproduction in the rotifer Brachionus calyciflorus. J. Freshwater Ecol. 26, 399–408 (2011).
Xi, Y.-L., Ge, Y.-L., Chen, F., Wen, X.-L. & Dong, L.-L. Life history characteristics of three strains of Brachionus calyciflorus (Rotifera) at different temperatures. J. Freshwater Ecol. 20, 707–713 (2005).
Yin, X. W. & Niu, C. J. Testing the cost of reproduction with the rotifer Brachionus patulus at different pH levels. Limnology 12, 145–149 (2011).
Speakman, J. R. et al. Uncoupled and surviving: individual mice with high metabolism have greater mitochondrial uncoupling and live longer. Aging Cell 3, 87–95 (2004).
Kuhla, A. et al. Lifelong caloric restriction reprograms hepatic fat metabolism in mice. J. Gerontol. A Biol. Sci. Med. Sci. 69, 915–922 (2014).
López-Domínguez, J. A. et al. The influence of dietary fat source on life span in calorie restricted mice. J. Gerontol. A Biol. Sci. Med. Sci. 70, 1181–1188 (2015).
Miller, K. N. et al. Aging and caloric restriction impact adipose tissue, adiponectin, and circulating lipids. Aging Cell 16, 497–507 (2017).
Heck, M. J., Pehilvanovic, M., Purcell, J. U., Hahn, D. A. & Hatle, J. D. Life-extending dietary restriction reduces oxidative damage of proteins in grasshoppers but does not alter allocation of ingested nitrogen to somatic tissues. J. Gerontol. A Biol. Sci. Med. Sci. 72, 616–623 (2017).
Smith, E. D. et al. Age- and calorie-independent life span extension from dietary restriction by bacterial deprivation in Caenorhabditis elegans. BMC Dev. Biol. 5, 49 (2008).
Gribble, K. E. & Mark Welch, D. B. Life-span extension by caloric restriction is determined by type and level of food reduction and by reproductive mode in Brachionus manjavacas (Rotifera). J. Gerontol. A Biol. Sci. Med. Sci. 68, 349–358 (2012).
Han, S. & Brunet, A. Histone methylation makes its mark on longevity. Trends Cell. Biol. 22, 42–49 (2012).
Sen, P., Shah, P. P., Nativio, R. & Berger, S. L. Epigenetic mechanisms of longevity and aging. Cell 166, 822–839 (2016).
Wątroba, M. et al. Sirtuins, epigenetics and longevity. Ageing Res. Rev. 5, 11–19 (2017).
Kennedy, B. K., Austriaco, N. R. Jr, Zhang, J. & Guarente, L. Mutation in the silencing gene SIR4 can delay aging in S. cerevisiae. Cell. 80, 485–496 (1995).
Guarente, L. Sir2 links chromatin silencing, metabolism, and aging. Genes Dev. 14, 1021–1026 (2000).
Whitaker, R. et al. Increased expression of Drosophila Sir2 extends life span in a dose-dependent manner. Aging 5, 682–691 (2013).
Burnett, C. et al. Absence of effects of Sir2 overexpression on lifespan in C. elegans and Drosophila. Nature 477, 482–485 (2011).
Chang, H.-C. & Guarente, L. SIRT1 and other sirtuins in metabolism. Trends Endocrinol. Metab. 25, 138–145 (2014).
Han, S. et al. Mono-unsaturated fatty acids link H3K4me3 modifiers to C. elegans lifespan. Nature 544, 185–190 (2017).
Hutchinson, G. E. A Treatise on Limnology (Wiley, 1957).
Dahms, H.-U., Hagiwara, A. & Lee, J.-S. Ecotoxicology, ecophysiology, and mechanistic studies with rotifers. Aquat. Toxicol. 101, 1–12 (2011).
Lubzens, E. Raising rotifers for use in aquaculture. Hydrobiologia 147, 245–255 (1987).
Johnston, R. K. & Snell, T. W. Moderately lower temperatures greatly extend the lifespan of Brachionus manjavacas (Rotifera): Thermodynamics or gene regulation? Exp. Gerontol. 78, 12–22 (2016).
Snell, T. W., Fields, A. M. & Johnson, R. K. Antioxidants can extend lifespan of Brachionus manjavacas (Rotifera), but only in a few combinations. Biogerontology 13, 261–275 (2012).
Gribble, K. E., Jarvis, G., Bock, M. & Mark Welch, D. B. Maternal caloric restriction partially rescues the deleterious effects of advanced maternal age on offspring. Aging Cell 13, 623–630 (2014).
Snell, T. W. & Johnston, R. K. Glycerol extends lifespan of Brachionus manjavacas (Rotifera) and protects against stressors. Exp. Gerontol. 57, 47–56 (2014).
Snell, T. W. et al. Repurposing FDA-approved drugs for anti-aging therapies. Biogerontology 17, 907–920 (2016).
Lee, B.-Y. et al. Whole transcriptome analysis of the monogonont rotifer Brachionus koreanus provides molecular resources for developing biomarkers of carbohydrate metabolism. Comp. Biochem. Physiol. D 14, 33–41 (2015).
Sarma, S. S. S., Nandini, S. & Gulati, R. D. Cost of reproduction in selected species of zooplankton (rotifers and cladocerans). Hydrobiologia 481, 89–99 (2002).
Stelzer, C. P. Evolution of rotifer life histories. Hydrobiologia 546, 335–346 (2005).
Snell, T. W. Rotifers as models for the biology of aging. Int. Rev. Hydrobiol. 99, 84–95 (2014).
Yoshinaga, T., Hagiwara, A. & Tsukamoto, K. Life history response and age-specific tolerance to starvation in Brachionus plicatilis O.F. Müller (Rotifera). J. Exp. Mar. Biol. Ecol. 287, 261–271 (2003).
Sinclair, D. A. & Guarente, L. Extrachromosomal rDNA circles-a cause of aging in yeast. Cell 91, 1033–1042 (1997).
Bellizzi, D. et al. A novel VNTR enhancer within the SIRT3 gene, a human homologue of SIR2, is associated with survival at oldest ages. Genomics 85, 258–263 (2005).
Kanfi, Y. et al. The sirtuin SIRT6 regulates lifespan in male mice. Nature 483, 218–221 (2012).
Kawahara, T. L. et al. SIRT6 links histone H3 lysine 9 deacetylation to NF-kappa B-dependent gene expression and organismal life span. Cell 136, 62–74 (2009).
Vazquez, B. N. et al. SIRT7 promotes genome integrity and modulates non-homologous end joining DNA repair. EMBO J. 35, 1488–1503 (2016).
Souissi, A., Souissi, S. & Hansen, B. W. Physiological improvement in the copepod Eurytemora affinis through thermal and multigenerational selection. Aquacult. Res. 47, 2227–2242 (2014).
Castro, L. F., Tocher, D. R. & Monroig, O. Long-chain polyunsaturated fatty acid biosynthesis in chordates: Insights into the evolution of Fads and Elovl gene repertoire. Prog. Lipid Res. 62, 25–40 (2016).
Sassa, T. & Kihara, A. Metabolism of very long-chain Fatty acids: genes and pathophysiology. Biomol. Ther. 22, 83–92 (2014).
Calder, P. C. Marine omega-3 fatty acids and inflammatory processes: effects, mechanisms and clinical relevance. Biochim. Biophys. Acta. 1851, 469–484 (2015).
Serhan, C. N., Dalli, J., Colas, R. A., Winkler, J. W. & Chiang, N. Protectins and maresins: New pro-resolving families of mediators in acute inflammation and resolution bioactive metabolome. Biochim. Biophys. Acta. 1851, 397–413 (2015).
Patil, V., Källqvist, T., Olsen, E., Vogt, G. & Gislerød, H. R. Fatty acid composition of 12 microalgae for possible use in aquaculture feed. Aquacult. Int. 15, 1–9 (2007).
Hwang, D.-S., Dahms, H.-U., Park, H. G. & Lee, J.-S. A new intertidal Brachionus and intrageneric phylogenetic relationships among Brachionus as revealed by allometry and CO1-ITS1 gene analysis. Zool. Stud. 52, 13 (2013).
Mills, S. et al. Fifteen species in one: deciphering the Brachionus plicatilis species complex (Rotifera, Monogononta) through DNA taxonomy. Hydrobiologia 796, 39–58 (2017).
Hwang, D.-S. et al. Complete mitochondrial genome of the monogonont rotifer, Brachionus koreanus (Rotifera, Brachionidae). Mito. DNA 25, 29–30 (2014).
Hama, T. & Handa, N. Pattern of organic matter production by natural phytoplankton population in a eutrophic lake. I. Intracellular products. Arch. Hydrobiol. 109, 107–120 (1987).
Acknowledgements
This work was supported by a grant from the Development of Techniques for Assessment and Management of Hazardous Chemicals in the Marine Environment (20150342) of the Ministry of Oceans and Fisheries, Korea funded to Jae-Seong Lee and was also supported by a grant from the Development of practical technique to establish fisheries forensic center of the Ministry of Oceans and Fisheries, Korea funded to Kyung-Hoon Shin.
Author information
Authors and Affiliations
Contributions
M.-C.L. and J.-S.L. designed the experiments, analyzed the data, and wrote the manuscript. M.-C.L., J.C.P., D.-S.Y., J.H., S.K., and S.K. performed the experiments. M.-C.L., J.C.P., A.-S.O., K.-H.S., A.H., and J.-S.L. discussed the experiments and worked through potential problems during their execution.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, MC., Park, J.C., Yoon, DS. et al. Aging extension and modifications of lipid metabolism in the monogonont rotifer Brachionus koreanus under chronic caloric restriction. Sci Rep 8, 1741 (2018). https://doi.org/10.1038/s41598-018-20108-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-20108-7
This article is cited by
-
A transcriptomic examination of encased rotifer embryos reveals the developmental trajectory leading to long-term dormancy; are they “animal seeds”?
BMC Genomics (2024)
-
Effects of microplastics on reproductive characteristics and mechanisms of the marine rotifer Brachionus plicatilis
Scientific Reports (2024)
-
Adequate nutrient intake mitigate the toxic effects of bromate on the rotifer Brachionus calyciflorus
Environmental Science and Pollution Research (2024)
-
Impacts of nanoplastics on life-history traits of marine rotifer (Brachionus plicatilis) are recovered after being transferred to clean seawater
Environmental Science and Pollution Research (2022)
-
A collective analysis of lifespan-extending compounds in diverse model organisms, and of species whose lifespan can be extended the most by the application of compounds
Biogerontology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.