Abstract
Altered gut microbiota is associated with autism spectrum disorders (ASD), a group of complex, fast growing but difficult-to-diagnose neurodevelopmental disorders worldwide. However, the role of the oral microbiota in ASD remains unexplored. Via high-throughput sequencing of 111 oral samples in 32 children with ASD and 27 healthy controls, we demonstrated that the salivary and dental microbiota of ASD patients were highly distinct from those of healthy individuals. Lower bacterial diversity was observed in ASD children compared to controls, especially in dental samples. Also, principal coordinate analysis revealed divergences between ASD patients and controls. Moreover, pathogens such as Haemophilus in saliva and Streptococcus in plaques showed significantly higher abundance in ASD patients, whereas commensals such as Prevotella, Selenomonas, Actinomyces, Porphyromonas, and Fusobacterium were reduced. Specifically, an overt depletion of Prevotellaceae co-occurrence network in ASD patients was obtained in dental plaques. The distinguishable bacteria were also correlated with clinical indices, reflecting disease severity and the oral health status (i.e. dental caries). Finally, diagnostic models based on key microbes were constructed, with 96.3% accuracy in saliva. Taken together, this study characterized the habitat-specific profile of the oral microbiota in ASD patients, which might help develop novel strategies for the diagnosis of ASD.
Similar content being viewed by others
Introduction
Autism spectrum disorders (ASD) are complex neurodevelopmental disorders unfolded in the first couple years of life, and characterized by impairments in language and social interactions, often with restricted interests and repetitive behaviors1. ASD affect 1-in-68 children in America, with the prevalence increasing dramatically over the past decades2. Currently, the fifth edition of Diagnostic and Statistical Manual of Mental Disorders (DSM-5) is considered the “gold standard” for ASD diagnosis. However, DSM-5 encompasses descriptive criteria merely based on empirical data, with no laboratory or other diagnostic tests3. As a result, diagnosis according to the relatively time-consuming DSM-5 guideline is not sufficiently scientific, and needs further validation4. Thus, developing precise and reliable diagnosis tools for ASD remains a challenge.
Genetic and environmental factors both play roles in the pathogenesis of ASD5. Available twin studies showed that environmental factors are more important than genetic predisposition6. Among such factors, microbial dysbiosis is of increasing interest, with accumulating reports in animal models and human epidemiologic studies linking disruptive alterations in the gut microbiota to ASD symptomology7,8,9,10,11,12. Although the mechanisms by which gut microorganisms might impact the central nervous system (CNS) are not fully understood, the human microbiota and related metabolites have been reported to affect a variety of complex behaviors, including social, emotional, and anxiety-like behaviors, also contributing to brain development and modulating cognition through imbalances in the microbiota-gut-CNS axis2,13,14. Consequently, microbial dysbiosis might play a causal role in the development of mental disorders, in a pathway mediated through the host’s metabolism.
While growing evidence suggests a key role for the gut microbiota in ASD, the neurologic effects of microorganisms inhabiting the oral cavity have been overlooked. Oral microbial communities can undergo dynamic fluctuations in response to various intrinsic and extrinsic factors and these may in turn affect brain function15. The oral cavity harbors approximately 700 predominant taxa, and serves as a reservoir for infection at distant body sites, including the neural system2,16. Current studies have documented the effects of the oral microbiota on neural function17,18,19,20,21. For example, oral microbial dysregulation is associated with Parkinson’s disease, Alzheimer’s disease, multiple sclerosis, and migraine17,18,19,20,21. However, the association of oral microbiota shifts with ASD remains unexplored.
Compared to the gut, the oral cavity is anatomically characterized as a unique and complex habitat because of the presence of both soft (i.e. mucosa) and hard (teeth) tissues. Microbiota colonize different oral habitats which are considered different physical niches; these organisms differ markedly in community structure and interact with each other continuously22. Additionally, oral samples, especially saliva specimens, are easier to collect compared to stool or intestinal mucosal biopsies. Therefore, it is anticipated that there would be great public interest in oral sampling-based research and clinical diagnosis, particularly for diseases for which sampling otherwise is challenging, such as ASD.
In this study, we collected samples from two distinct intraoral habitats, including saliva and dental plaques, in children with and without ASD. Microbial populations were assessed by 16S rRNA gene sequencing. Dental plaques retain diverse microorganisms and are known to reflect poor dental hygiene and oral health behavior, which were reported to be greatly inadequate in ASD children, consistent with our clinical observation and the current literature23,24. On the other hand, saliva, as a representative of the overall oral microbial communities, has been widely used to explore the associations of the oral microbiota with epidemiological characteristics and systemic diseases25,26. To our knowledge, this study provides the first evidence for oral microbial alterations in children with ASD, which would help expand the understanding of the relationship between oral microbiota and ASD, and develop more objective strategies for ASD diagnosis.
Methods
Ethics Statement
This study was approved by the Ethics Committee of the School and Hospital of Stomatology, Tongji University, and conducted in compliance with relevant guidelines and regulations. Written informed consent was obtained from the parents or guardians of all subjects, prior to the study.
Subject Recruitment
Children diagnosed with ASD by meeting the DMS-5 criteria were recruited locally from Shanghai Children’s Medical Center. Diagnosis was further confirmed at Shanghai Mental Health Center with the criteria for the International Classification of Diseases and Related Health Problems, Tenth Revision (ICD-10). The enrolled ASD subjects were between 7 and 14 years of age, with no previous medical treatment (except for rehabilitation training) or antibiotic/antifungal use within 3 months of sample collection. Gender- and age-matched healthy children, who were unrelated to the autistic individuals, were recruited from primary schools. The detailed eligibility criteria are presented in Supplementary Table S1. The parents or guardians of the enrolled subjects were asked to fill out the Aberrant Behavior Checklist (ABC) questionnaire to preliminarily evaluate the severity of ASD27.
Sample Collection and Oral Examination
The participants were instructed to refrain from eating and drinking, and to avoid oral hygiene practice, for at least 3 hours prior to sample collection. Before dental sampling, approximately 1 ml of non-stimulated, naturally outflowed saliva was consistently collected and transferred into 1.5 ml sterile Eppendorf tubes (Axygen, CA, USA). Then, the first permanent molars were isolated using cotton rolls and gentle air-drying. Supragingival plaques were obtained separately from caries-free molars in four quadrants (upper right, upper left, lower right and lower left) per subject using sterile Gracey curettes, and pooled in 1.5 ml sterile Eppendorf tubes containing sterile phosphate-buffered saline (PBS). All samples were immediately placed on ice, transported to the local laboratory within 2 hours, and stored at −80 °C until DNA extraction. In total, 111 specimens from 32 ASD children and 27 healthy children were obtained.
The 111 samples collected were divided into four groups according to study populations and sampling habitats: HS, salivary samples from healthy controls (n = 27); HP, dental samples from healthy controls (n = 26); AS, salivary samples from ASD patients (n = 32); AP, dental samples from ASD patients (n = 26).
Oral conditions in each subject, including dentition status, plaque index (PLI), bleeding on probing (BOP), gingival index (GI), sulcus bleeding index (BI), probing depth (PD), and decayed-missing-filled teeth/surfaces (DMFT/DMFS), were recorded after sampling (described in Supplementary Table S2). All examinations were performed by the same experienced dentist according to routine criteria.
Microbial DNA extraction and PCR amplification
DNA from dental and salivary samples was extracted with the OMEGA-soil DNA Kit (OmegaBio-Tek, USA) according to the manufacturer’s instructions. DNA purity was evaluated based on A260/A280 ratio using a NanoDrop2000 Spectrophotometer (Thermo Fisher Scientific, USA). DNA integrity and size were assessed by 1.0% agarose gel electrophoresis. A negative control containing the buffer only was included during DNA extraction and quantification. DNA samples were frozen at −20 °C until use. The V3-V4 region of the 16S ribosomal RNA gene was amplified using primers 338 F (5′-ACTCCTACGGGAGGCAGCA-3′) and 806 R (5′-GGACTACHVGGGTWTCTAAT-3′) via PCR (95 °C for 2 min, followed by 25 cycles at 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s, and a final extension at 72 °C for 5 min)28.
16S rRNA gene sequencing
PCR amplicons were separated by agarose gel electrophoresis, purified with the AxyPrep DNA Gel Extraction Kit (Axygen Biosciences, Union City, CA, USA) according to the manufacturer’s instructions, and quantified using QuantiFluor™ -ST (Promega, USA). The purified amplicons were pooled in equimolar and paired-end sequences (2 × 250) on a MiSeq platform (Illumina, USA), according to standard protocols. Raw reads were deposited in the NCBI SRA database (Accession number SRP097646).
Processing of sequencing data
Raw FASTQ files were demultiplexed and quality-filtered using QIIME (version 1.9.1), with the following criteria: (1) 300 bp reads were truncated at any site receiving an average quality score <20 over a 50 bp sliding window, and truncated reads shorter than 50 bp were discarded; (2) exact barcode matching, with 2 nucleotide mismatch in primer matching and reads containing ambiguous characters removed; (3) only sequences overlapping over more than 10 bp were assembled according to the overlapping sequence. Reads which could not be assembled were discarded. High-quality reads were clustered into operational taxonomic units (OTUs) with the UPARSE software29. The taxonomy of each 16S rRNA gene sequence was analyzed by RDP Classifier (http://rdp.cme.msu.edu/) against the Human Oral Microbiota Database, a curated dataset for oral taxa30.
Statistical and Data Analyses
Demographics and clinical parameters of subjects between the ASD and healthy groups were compared by the t-test (age, BMI), Chi-square test (gender) and Mann-Whitney U-test (DMFT/DMFS, PLI, BOP, GI, BI, PD). p < 0.05 was considered statistically significant. Rarefaction curves were generated with Mothur to assess the current sequencing depth. At a 97% level of nucleotide similarity cutoff, the OTUs were used for assessing α-diversity indices (ACE, Shannon and Shannoneven) and Good’s coverage. Phylogenetic α-diversity measures, such as principal coordinate analysis (PCoA), were conducted according to (un)weighted UniFrac distances based on the detected genera in each sample. Independent Student’s t-test and Permutational Multivariate Analysis of Variance (PERMANOVA) were applied to evaluate α and β diversity among different groups. Histograms based on different taxonomic levels were generated to depict the overall microbial composition and abundance. The data of RDP taxonomy were used in LEfSe (http://huttenhower.sph.harvard.edu/lefse/) to identify the taxa differentiating the microbial communities specific to different groups31. Microbiota features of ASD subjects were compared to those of healthy controls by the Wilcoxon rank sum test using p-values and FDR Q-values (modified version of FDR32) for non-normally distributed variables. Only OTUs with p < 0.05 and FDR Q < 0.05 were considered significant and used in the network analysis. Spearman’s rank correlation coefficients were determined between the OTUs in dental and salivary samples from healthy children and autistic patients. Then, co-occurrence networks were generated using Cytoscape (version 3.4.0). Associations of microbes with oral health indices were assessed by Spearman’s rank correlation, and visualized by heat maps. The Random Forest algorithm and 10-fold cross validation were used to select microbial markers and their combinations. The obtained combinations were assessed by Receiver Operator Characteristic (ROC) analysis for performance33. All analyses were performed in the R software package (version 3.3.1; The R Project for Statistical Computing, http://www.R-project.org).
Results
Overall structure of the oral microbiota
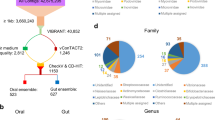
The demographic and clinical characteristics of all participants are depicted in Table 1 and Supplementary Table S2. To investigate the shifts in structure and composition of oral microbial communities in ASD, 111 samples were sequenced by the Illumina MiSeq technology. After processing, a total of 3,769,401 valid reads were obtained, with an average of 33,959 ± 4,253 reads per sample. All sequences (excluding singletons) were classified into 405 OTUs at a 97% similarity level, representing 12 phyla, 132 genera and 308 species. The predominant taxa were similar to those previously reported in non-ASD populations25,34, and are shown in Supplementary Fig. S2. The Good’s coverage was 99.86%, and rarefaction curves approached asymptotes for most samples with the current sequencing (Supplementary Fig. S1).
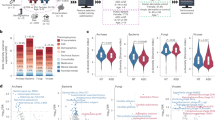
Alpha diversity (Fig. 1a–c and Supplementary Table S3), richness index (ACE), diversity index (Shannon), and evenness index (Shannoneven) were all significantly lower in ASD subjects compared to healthy controls (P < 0.05) in dental samples; however, in saliva samples, none of these indices showed significant differences (P > 0.05), although a downward trend was observed. Dental plaques displayed a higher level of bacterial diversity than saliva, with a lower level of bacterial richness (Fig. 1a,b). Moreover, principal coordinate analysis (PCoA) based on (un)weighted UniFrac metrics was performed to identify any differences in organismal structure of oral microbiota (Fig. 2a, Supplementary Fig. S3). The salivary and dental bacterial communities showed non-overlapping clustering, revealing that the oral microbiota mainly differed by habitats rather than the disease status.
Overall structural comparison of the oral microbiota. Sequences were randomly subsampled to obtain equal numbers of sequences (21,795) from each dataset. ACE index (a, representing community richness), Shannon index (b, representing diversity), and Shannoneven index (c, representing evenness) were calculated for dental and salivary samples. The bars depict mean ± SD of relative abundance rates. *p < 0.05; **p < 0.01; ***p < 0.001; NS, p > 0.05. HS (saliva samples collected from healthy controls, n = 27); HP (plaque samples from healthy controls, n = 26); AS (saliva samples from ASD patients, n = 32); AP (plaque samples collected from ASD patients, n = 26).
Comparison of β-diversity of the oral microbiota. A principal coordinate analysis (PCoA) score plot for all samples (principal coordinates 1 and 2) based on weighted UniFrac distance was generated to depict the divergence in distribution of the oral microbiota (a). Each point represents the oral microbiota composition of one sample. PERMANOVA shows statistically significant community differences between ASD and healthy controls within each habitat, as well as between saliva and dental plaques, regardless of the disease status (b).
We also observed divergences in bacterial communities between ASD patients and healthy subjects in both salivary and dental samples, with specimens from healthy subjects tending to cluster more closely (Fig. 2a). The PERMANOVA identified significant differences in PCoA plots between ASD and healthy children in both habitats (P < 0.01; Fig. 2b).
Taxonomy-based analysis of microbial changes in ASD patients
To identify the distinguishing taxa of saliva and dental plaque samples, LEfSe was performed. The cladogram based on LDA score >2 showed multiple discriminative features of saliva and the dental microbiota between healthy and ASD subjects at different taxonomic levels (Supplementary Fig. S4). More distinguishable taxa between ASD and control groups were observed in saliva compared to dental samples. Then, taxa with significant differences were further assessed by the Wilcoxon rank-sum test, with stringent criteria (P < 0.05 and FDR Q < 0.05). Despite high inter-individual variability, predominant phyla in all groups were Firmicutes, Proteobacteria, Actinobacteria, Bacteroidetes and Fusobacteria, constituting 98.06% and 94.86% of salivary and dental microbes, respectively. The phylum Proteobacteria was more abundant in ASD patients (both in salivary and dental samples) compared to controls (FDR Q = 0.017 and 0.014, respectively), while the other phyla were more abundant in healthy controls (Fig. 3a,b). Meanwhile, some genera differed significantly in abundance between ASD and healthy children (Fig. 3c,d). Specifically, among genera with a mean relative abundance >1%, ASD was associated with increased abundance rates of Streptococcus (FDR Q = 0.02 in plaques) and Haemophilus (FDR Q = 0.007 in saliva), and decreased rates of Prevotella (FDR Q = 0.009 in plaques), Selenomonas (FDR Q = 0.042 in plaques), Actinomyces (FDR Q = 0.002 in saliva), Porphyromonas (FDR Q = 0.03 in saliva), and Fusobacterium (FDR Q = 0.011 and 0.025 in plaques and saliva, respectively). Interestingly, in children with ASD, Rothia showed a significantly higher abundance in plaques (FDR Q = 0.023) but lower abundance in saliva (FDR Q = 0.033) compared to controls, indicating that this genus might play habitat-specific roles in ASD children.
Changes of oral microbiota at the phylum and genus levels between ASD and control subjects. The graphs depict Wilcoxon rank sum test results with p < 0.05 and FDR Q < 0.05 from comparisons of relative abundance between ASD and healthy children at the bacterial phylum level in saliva (a) and plaque (b) samples. Similar comparisons at the genus level in saliva and plaque samples are shown in (c) and (d), respectively (p < 0.05 and FDR Q < 0.05).
Co-occurrence networks of salivary and dental bacterial communities in patients with ASD and healthy controls
The differential OTUs identified by the Wilcoxon rank-sum test (P < 0.05, FDR Q < 0.05, Supplementary Tables S4, S5) in salivary and dental samples (53 and 43 OTUs, respectively) were used for network analysis. By assessing Spearman’s correlation coefficients for all OTU markers, a network plot displaying potential co-occurrence relationships was generated (Fig. 4, Spearman’s correlation coefficients |r| >0.5). OTUs were more highly interconnected in control groups than in ASD counterparts, especially in dental samples, indicating that ASD was not only associated with decreased richness of commensals, but also related to reduced mutual effects within these bacteria. Among these commensals, an interesting finding was the obvious reduction of Prevotellaceae co-occurrence network patterns in dental samples, including Prevotella and Alloprevotella clusters (Fig. 4b). In contrast, the OTUs in ASD groups formed smaller clusters and were less interconnected. Two ASD-enriched shared OTUs, including OTU139 (Rothia aeria) and OTU285 (Streptococcus mutans), and saliva-specific OTUs, i.e. OTU159, OTU189, and OTU361, which were all annotated to Haemophilus, were negatively correlated with control-enriched OTUs, suggesting unknown antagonistic or mutually exclusive relationships. Moreover, clusters containing 9 OTUs belonging to Actinomyces, Solobacterium, Alloprevotella, Leptotrichia, Peptostreptococcus and Peptostreptococcacea [XI][G-1], were all enriched in both salivary and dental samples from healthy controls but not in the ASD groups.
OTUs enriched in salivary and dental samples in ASD patients and healthy controls. A co-occurrence network was deduced from 53 OTUs in salivary samples (a) and 43OTUs in dental samples (b). Nodes depict OTUs with IDs and affiliated genus names displayed in the center. The size of the nodes indicates the abundance of each OTU. The color of a node indicates its taxonomic assignment. Connecting lines represent Spearman correlation coefficients above 0.5 (blue) or below −0.5 (red).
Next, co-variations were computed between the relative abundance rates of the most abundant 50 OTUs and clinical indices of ASD as well as oral health-related indices (Fig. 5a,b). A separation was found between control- and ASD-enriched OTUs based on ABC scores, which are used to preliminarily determine the severity of ASD27. Notably, most ASD-enriched phylotypes, such as Haemophilus sp. (OTU159, R = 0.445, p < 0.001) and Rothia aeria (OTU139, R = 0.381, P = 0.004), were positively correlated to the ABC score, to which control-enriched phylotypes were negatively correlated, both in salivary and dental samples. No saliva phylotypes were highly correlated with DMFT/S, which reflects the severity of dental caries. However, in dental plaques, 6 phylotypes belonging to the genera Streptococcus, Actinomyces and Capnocytophaga were positively correlated with DMFT/S. These findings indicated that dental caries had a greater impact on the microbial structure of dental plaques compared to saliva. Furthermore, Aggregatibacter segnis (OTU220) was positively correlated to BOP, GI, and PD, which reflects gingival inflammation.
Correlation between OTUs and various clinical indices. The abundance of OTUs (top 50) enriched in saliva (a) and dental plaques (b) of ASD children and controls were analyzed for covariation with clinical variables using Spearman’s rank correlation. The correlation effect is indicated by a color gradient from blue (negative correlation) to red (positive correlation). Correlation coefficients and p-values (*p < 0.05) are shown. OTUs and clinical variables in the heat maps were ordered using unsupervised hierarchical clustering. BMI, Body Mass Index; DMFT, Decayed-Missing-Filled Teeth; DMFS, Decayed-Missing-Filled Surface; PLI, Plaque Index; GI, Gingival Index; BI, Bleeding Index; PD, Probing Depth; BOP, Bleeding on Probing; ABC score, scale of ASD severity.
Diagnostic performance of salivary and dental microbiota
We subsequently performed the Random Forests (RF) analysis to examine whether ASD and healthy children could be discriminated based on microbiota composition. RF analysis assigned a variable importance score to each OTU, according to the extent of its contribution to the model. Based on the variable importance profile, we selected sixty OTUs with the highest variable importance sores for tenfold cross-validation, which was used to define the minimum number of top-ranking taxa according to decrease in classification error rate. Finally, 36 OTUs were identified in saliva with an error of 0.09 and 54 OTUs in dental plaques with an error of 0.17 (Fig. 6a,b). These findings indicated a more accurate discrimination value for salivary microbiota.
Diagnostic value assessment of salivary and dental samples. Cross-validation error of the random forest ASD classifier was performed based on OTUs in saliva (a) and dental plaques (b). The grey curve represents a trial of 10-fold cross-validation. The black dot indicates the final number of OTUs used for classification and related errors. (c) The AUCs of all combinations in saliva are depicted in the above panel. The following panel shows the magnified portion in the grey rectangle, indicating the AUC of the MIA_AS7HS13 combination reached the highest value. (d) The ROC curve of the above MIA in ASD diagnosis is depicted. The area under the ROC curve (AUC) was calculated and is shown in the center. The arrow shows the optimal cut-off point in the salivary microbiota. (e) The heat map indicates the ability of salivary potential biomarkers to discriminate between healthy and ASD subjects.
Then, we used the top OTUs to construct a prediction model for the diagnosis of ASD. To facilitate the clinical application of the identified markers, a more straightforward index, “microbial index of ASD (MIA)”, was developed for the discrimination of patients based on the relative abundance of the above OTUs:
After computing all combinations of ASD and control enriched bacteria in saliva and dental plaques, respectively, MIA was used to generate receiver operating characteristic (ROC) curves. The detailed AUC values for all combinations are shown in Supplementary Table S6. In total, 20 key OTUs (7 in ASD patients and 13 in healthy children, Fig. 6c) were obtained in saliva as well as 6 (all in healthy controls, Supplementary Fig. S5) in dental plaques, with areas under the ROC curve (AUCs) reaching the highest value. AUC, cut-off point, sensitivity, specificity and p-values are depicted in Fig. 6d and Supplementary Fig. S5. The area under the ROC curve of MIA in saliva reached 96.3%, and was higher than that of MIA in dental plaques, indicating that the salivary microbiota was more efficient for ASD diagnosis than the dental counterpart. In saliva, the cut-off point for MIA was determined at -482.5 (sensitivity of 0.889 and specificity of 0.906), indicating that individuals with MIA exceeding −482.5 might have a risk of ASD. Moreover, correlations between salivary biomarkers and ASD subjects were obtained in the heat map, indicating the ability of these biomarkers to distinguish healthy from autistic hosts (Fig. 6e).
Discussion
In this cross-sectional study, an initial comparative analysis of oral microbiota in ASD children with that of age, sex and BMI matched healthy controls was conducted via 16S rRNA gene sequencing. We identified alterations in the composition and taxonomy of the oral microbiota in ASD patients. Previous studies have reported strong correlations between ASD and altered gut microbiota using different methods7,8,9,10,11,12,35. Some of our findings broadly corroborated these reports evaluating intestinal microbiota. For example, several studies assessing the gut microbiota in stool or intestinal biopsies found no visible changes in diversity and richness between ASD subjects and controls (unrelated healthy controls or neurotypical siblings)7,9,36; nevertheless, Kang et al. reported significantly reduced diversity of gut microbiota in autistic children compared to neurotypical subjects12. In this research, our data on richness and diversity showed no differences in salivary samples; however, a minor but statistically significant decrease in dental samples was observed in ASD children, compared to non-ASD controls. Lower bacterial diversity may reduce the microbial integrity and fail to provide resistance to environmental stressors, such as the intake of pathogenic microbes37.
In addition, the increasing or decreasing trends of many taxa at different levels in this study were similar to those obtained for intestinal studies of ASD population. Several gut microbiota studies have revealed changes in relative abundance of phylotypes such as Proteobacteria, Bacteroidetes, and Actinobacteria7,12,38,39. Consistent with the latter reports, in this research, significantly higher abundance of Proteobacteria (in both salivary and dental samples) and lower abundance rates of Actinobacteria (in saliva) and Bacteroidetes (in dental plaques) were observed in ASD children. At the genus level, increased abundance of Streptococcus and Haemophilus, as well as markedly reduced abundance of Prevotella, Alloprevotella, Selenomonas, Actinomyces, Porphyromonas and Fusobacterium in ASD samples were observed, suggesting a characteristic dysbiotic signature. The family Prevotellaceae, including the genera Prevotella and Alloprevotella, also showed a relatively low abundance in children with ASD. This was evident in dental samples as demonstrated in the co-occurrence network (Fig. 4). In agreement with these findings, Finegold et al. recorded a depletion of Prevotella in the gut of autistic patients compared to sibling controls38. Meanwhile, Kang et al. reported that Prevotella, as the most significantly altered genus between autistic and neurotypical subjects, decreases dramatically in abundance in stool samples from autistic patients12. A low abundance of Prevotella was also detected in feces from patients with Parkinson’s disease and untreated Multiple Sclerosis, supporting the relevance of this bacterium in CNS disorders40,41. Prevotella is a commensal microorganism in multiple human habitats, including the intestine and oral cavity; it does not only interact with the immune system but also plays a key role in degrading a broad spectrum of saccharides42,43,44. Interestingly, it was reported that autistic children may have deficiencies in saccharide metabolism and impaired carbohydrate digestion36,45. Prevotella species also have essential genes for the biosynthesis of vitamins44, which were reported to mitigate ASD symptoms46,47. Future studies are warranted to further interrogate the role of Prevotellaceae, also evaluating its therapeutic potential for ASD.
In addition to a reduction in autochthonous bacteria, increased amounts of potential pathogens, including Haemophilus, Corynebacterium, Cardiobacterium, Kingella, Streptococcus and Rothia, were observed in ASD patients. These pathogens are quite different from intestinal bacterial pathogens, such as Clostridium, Sutterella, and Desulfovibrio12,38,48,49, which were not detected in the present study. In the genus Haemophilus, for example, all seven members were more abundant in salivary samples from ASD patients, with OTU159, OTU189 and OTU361 significantly discriminative. They were also positively correlated to ABC scores (Fig. 5a), indicating an association with ASD symptom severity. Their nearest neighbor, Haemophilus parainfluenzae, a Gram-negative and facultative anaerobic bacteria, is not only prevalent in oral diseases50, but also considered a potential pathogen causing diseases in remote target organs and systems, e.g. bacteremia and infectious endocarditis51,52. These evidences indicate that H. parainfluenzae and its metabolites could gain access to the bloodstream and trigger disease outbreaks. H. parainfluenzae was also reported to bypass the blood-brain barrier (BBB), enter the cerebrospinal fluid, and cause meningitis53,54. Thus, we hypothesized that H. parainfluenzae and its metabolites in the oral cavity might enter the blood circulation and even cross the BBB to affect brain development in ASD children.
Another putative pathogen showing overgrowth in dental samples was Streptococcus, a potent immunogenic trigger that was reported to affect the risk of CNS dysfunction such as Tourette Syndrome (another neurodevelopmental illness), Sydenham chorea and bacterial meningitis55,56. Indeed, Streptococcus was recorded to cause neurological damage by producing neurotoxins such as streptomycin, streptodornase, and streptokinase57. Studies of the intestinal microbiota indicated that Streptococcus might be responsible for bacterial infection in Parkinson’s disease and liver cirrhosis, probably originating from the mouth57,58,59. This finding indicated that specific oral bacteria could invade the gut and subsequently influence remote organs. Moreover, Rothia was highly abundant in ASD-associated dental samples, but interestingly, showed low abundance in saliva (Fig. 3c,d). This discrepancy could be explained at the species level. Among Rothia species, R. mucilaginosa (OTU279) abundance was significantly reduced in ASD salivary samples (Supplementary Table S4). However, R. aeria (OTU139) was significantly more abundant in ASD children in both habitats and positively correlated to ABC scores (Fig. 5a,b). Several studies have reported R. aeria infection in sepsis, arthritis and pneumonia, most of which were associated with dental pathology60,61,62. We conjecture that Rothia aeria might be involved in systemic inflammation and immune response in the pathogenesis of ASD.
Based on the putative pathogens identified in this study, we ventured to hypothesize the plausible mechanisms underlying the oral microbial disturbance in the pathogenesis of ASD. It was documented that oral bacteria could easily and frequently gain access to the bloodstream through routine dental procedures such as chewing, brushing and dental treatment, causing bacteremia subsequently63. A previous study demonstrated that the most likely pathway for the dissemination of oral microorganisms to the brain is hematogenous spread64. The pro-inflammatory responses provoked by oral bacteria might impair BBB integrity, allowing bacteria to reach the brain and quietly contribute to the pathogenesis of neurological diseases18. In addition, previous studies of Alzheimer’s disease described neuronal pathways18,65. The olfactory and trigeminal nerves may serve as direct passages for oral bacteria to the CNS or as transmitters of neural signals from the oral cavity to the brain. The orofacial region, which is enriched with vessels and nerves, is anatomically close to the brain, facilitating the cross-talk between the oral microbiota and the CNS. However, due to the exploratory nature of this study, the above attempts to explain the mechanisms underlying the link between ASD and the oral microbiota are highly speculative. Further research is needed for better understanding.
ASD diagnosis in clinical practice is currently guided by DSM-5 criteria, which are based on a consensus regarding clusters of clinical symptoms, with no trusted or objective laboratory measure3. Its precision and reliability were overstated in the past decades. Therefore, DSM-5 validity has been challenged by several groups, including the National Institute of Mental Health4. Although tools have been developed for autism diagnosis, the desire for clinically useful and reliable biomarkers remains strong66. In the present study, the MIA diagnosis model based on oral microbial biomarkers was proposed and achieved 96.3% accuracy, showing a great potential for improved diagnostic sensitivity and specificity. Compared to the gut microbiota usually collected from feces or biopsies, the oral microbiota is easier to sample and potentially mirrors the ASD-associated microbiota profile in situ. In addition to microbial markers, clinical indices could be combined to further improve the diagnosis of ASD. Among the demographical and clinical variables assessed in this study, age, BMI, PLI and PD showed no significant differences between ASD and control subjects. However, other parameters, including ABC scores (reflecting ASD severity roughly), DMFT/S (reflecting dental caries severity), BI, GI, and BOP (reflecting dental hygiene), differed significantly between the ASD and healthy cohorts, with higher severity in ASD individuals. ASD children were reported to have a relatively poor oral health condition due to inadequate tooth brushing, consistent with our clinical observation23,24. This might be related to lack of the necessary manual dexterity of autistic children as well as difficulties encountered by trainers and parents while brushing the children’s teeth. These oral health indicators might be used to evaluate the risk of ASD. Furthermore, inflammatory factors and metabolites in the oral cavity, such as interleukins, lipopolysaccharides and short-chain fatty acids, could also be included in the algorithm for ASD diagnosis to achieve a higher accuracy and more precise results. Future studies are still warranted to select the optimal combination of the indicators discussed above.
In conclusion, we preliminarily identified the oral microbial taxa associated with ASD, and delineated different features of salivary and dental microbiota between ASD and healthy individuals, providing a comprehensive cross-sectional description of oral microbiota alteration in ASD. This study deciphered dysbiosis in the oral microbial community of ASD patients, suggesting a potential role for microorganisms in the progression of this disease, although a causal relationship cannot be discerned from the association alone. We also suggest a diagnostic approach based on oral microbial features, providing new insights into the design of novel and readily accessible strategies for the diagnosis of ASD. Although this was more sensitive and objective compared to the empirical DSM method, verification of the current findings in large cohorts is required.
Accession codes
The data are available at the NCBI Sequence Read Archive (SRA) with the accession number SRP097646.
References
Johnson, C. P. & Myers, S. M. & American Academy of Pediatrics Council on Children With, D. Identification and evaluation of children with autism spectrum disorders. Pediatrics 120, 1183–1215 (2007).
Rosenfeld, C. S. Microbiome Disturbances and Autism Spectrum Disorders. Drug Metabolism and Disposition 43, 1557–1571 (2015).
Kulage, K. M., Smaldone, A. M. & Cohn, E. G. How will DSM-5 affect autism diagnosis? A systematic literature review and meta-analysis. Journal of autism and developmental disorders 44, 1918–1932 (2014).
Lane, C. The NIMH Withdraws Support for DSM-5. Psychology today (2013).
Sandin, S. et al. The familial risk of autism. JAMA 311, 1770–1777 (2014).
Hallmayer, J., Cleveland, S. & Torres, A. et al. Genetic heritability and shared environmental factors among twin pairs with autism. Archives of General Psychiatry 68, 1095–1102 (2011).
Hsiao, E. Y. et al. Microbiota modulate behavioral and physiological abnormalities associated with neurodevelopmental disorders. Cell 155, 1451–1463 (2013).
Finegold, S. M. et al. Pyrosequencing study of fecal microflora of autistic and control children. Anaerobe 16 (2010).
Son, J. S. et al. Comparison of Fecal Microbiota in Children with Autism Spectrum Disorders and Neurotypical Siblings in the Simons Simplex Collection. PloS one 10, e0137725 (2015).
Williams, B. L., Hornig, M., Parekh, T. & Lipkin, W. I. Application of Novel PCR-Based Methods for Detection, Quantitation, and Phylogenetic Characterization of Sutterella Species in Intestinal Biopsy Samples from Children with Autism and Gastrointestinal Disturbances. mBio 3, e00261–00211 (2012).
Adams, J. B. et al. Gastrointestinal flora and gastrointestinal status in children with autism – comparisons to typical children and correlation with autism severity. BMC Gastroenterology 11, 22 (2011).
Kang, D. W. et al. Reduced incidence of Prevotella and other fermenters in intestinal microflora of autistic children. PloS one 8, e68322 (2013).
Collins, S. M., Surette, M. & Bercik, P. The interplay between the intestinal microbiota and the brain. Nat Rev Micro 10, 735–742 (2012).
Cryan, J. F. & Dinan, T. G. Mind-altering microorganisms: the impact of the gut microbiota on brain and behaviour. Nat Rev Neurosci 13, 701–712 (2012).
Ding, T. & Schloss, P. D. Dynamics and associations of microbial community types across the human body. Nature 509, 357–360 (2014).
Krishnan, K., Chen, T. & Paster, B. J. A practical guide to the oral microbiome and its relation to health and disease. Oral diseases 23, 276 (2016).
Pereira, P. A. B. et al. Oral and nasal microbiota in Parkinson’s disease. Parkinsonism & related disorders 38, 61–67 (2017).
Shoemark, D. K. & Allen, S. J. The microbiome and disease: reviewing the links between the oral microbiome, aging, and Alzheimer’s disease. Journal of Alzheimer’s disease 43, 725–738 (2015).
Singhrao, S. K. et al. Oral inflammation, tooth loss, risk factors, and association with progression of Alzheimer’s disease. Journal of Alzheimers Disease Jad 42, 723 (2014).
Gonzalez, A. et al. Migraines Are Correlated with Higher Levels of Nitrate-, Nitrite-, and Nitric Oxide-Reducing Oral Microbes in the American Gut Project Cohort. mSystems 1 (2016).
Farrokhi, V. et al. Bacterial lipodipeptide, Lipid 654, is a microbiome-associated biomarker for multiple sclerosis. Clinical & translational immunology 2, e8 (2013).
Mager, D. L., Ximenez-Fyvie, L. A., Haffajee, A. D. & Socransky, S. S. Distribution of selected bacterial species on intraoral surfaces. Journal of Clinical Periodontology 30, 644–654 (2003).
Jaber, M. A. Dental caries experience, oral health status and treatment needs of dental patients with autism. Journal of Applied Oral Science 19, 212–217 (2011).
Lu, Y. Y., Wei, I. H. & Huang, C. C. Dental health - a challenging problem for a patient with autism spectrum disorder. General hospital psychiatry 35(214), e211–213 (2013).
Ling, Z. et al. Pyrosequencing Analysis of the Salivary Microbiota of Healthy Chinese Children and Adults. Microbial Ecology 65, 487–495 (2013).
Simón-Soro, Á. et al. Microbial Geography of the Oral Cavity. Journal of Dental Research 92, 616–621 (2013).
Aman, M. G., Singh, N. N., Stewart, A. W. & Field, C. J. Psychometric characteristics of the aberrant behavior checklist. American Journal of Mental Deficiency 89, 492 (1985).
Lu, H. et al. Deep sequencing reveals microbiota dysbiosis of tongue coat in patients with liver carcinoma. Sci Rep 6, 33142 (2016).
Edgar, R. C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nature methods 10, 996–998 (2013).
Dewhirst, F. E. et al. The human oral microbiome. Journal of bacteriology 192, 5002–5017 (2010).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biology 12, R60 (2011).
Storey, J. D. The positive false discovery rate: a Bayesian interpretation and the q-value. Annals of Statistics 31, 2013–2035 (2003).
Robin, X. et al. pROC: an open-source package for R and S + to analyze and compare ROC curves. BMC Bioinformatics 12, 77 (2011).
Xu, H. et al. Plaque Bacterial Microbiome Diversity in Children Younger than 30 Months with or without Caries Prior to Eruption of Second Primary Molars. PloS one 9, e89269 (2014).
Wang, L. et al. Gastrointestinal microbiota and metabolite biomarkers in children with autism spectrum disorders. Biomarkers in Medicine 8, 331–344 (2014).
Williams, B. L. et al. Impaired Carbohydrate Digestion and Transport and Mucosal Dysbiosis in the Intestines of Children with Autism and Gastrointestinal Disturbances. PloS one 6, e24585 (2011).
Stecher, B. et al. Salmonella enterica serovar typhimurium exploits inflammation to compete with the intestinal microbiota. PLoS biology 5, 2177–2189 (2007).
Finegold, S. M. et al. Pyrosequencing study of fecal microflora of autistic and control children. Anaerobe 16, 444–453 (2010).
De Angelis, M. et al. Fecal microbiota and metabolome of children with autism and pervasive developmental disorder not otherwise specified. PloS one 8, e76993 (2013).
Scheperjans, F. et al. Gut microbiota are related to Parkinson’s disease and clinical phenotype. Movement Disorders 30, 350–358 (2015).
Jangi, S. et al. Alterations of the human gut microbiome in multiple sclerosis. Nature communications 7, 12015 (2016).
Consortium, T. H. M. P. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Wu, G. D. et al. Linking Long-Term Dietary Patterns with Gut Microbial Enterotypes. Science 334, 105–108 (2011).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 506, 516–516 (2011).
Horvath, K. et al. Gastrointestinal abnormalities in children with autistic disorder. The Journal of Pediatrics 135, 559–563 (1999).
Berding, K. & Donovan, S. M. Microbiome and nutrition in autism spectrum disorder: current knowledge and research needs. Nutrition Reviews 74, 723–736 (2016).
Lonsdale, D., Shamberger, R. J. & Audhya, T. Treatment of autism spectrum children with thiamine tetrahydrofurfuryl disulfide: a pilot study. Neuro Endocrinology Letters 23, 303–308 (2002).
Finegold, S. M. Desulfovibrio species are potentially important in regressive autism. Med Hypotheses 77, 270–274 (2011).
Parracho, H. M., Bingham, M. O., Gibson, G. R., & McCartney, A. L. Differences between the gut microflora of children with autistic spectrum disorders and that of healthy children. Journal of Medical Microbiology 54 (2005).
Hu, X., Zhang, Q., Hua, H. & Chen, F. Changes in the salivary microbiota of oral leukoplakia and oral cancer. Oral oncology 56, e6–8 (2016).
Sen Yew, H. et al. Association between HACEK bacteraemia and endocarditis. Journal of Medical Microbiology 63, 892–895 (2014).
Faure, E. et al. Haemophilus parainfluenzae endocarditis in young adults. Medecine et maladies infectieuses (2016).
Coureuil, M., Lecuyer, H., Bourdoulous, S. & Nassif, X. A journey into the brain: insight into how bacterial pathogens cross blood-brain barriers. Nature reviews. Microbiology 15, 149–159 (2017).
Johansson Kostenniemi, U., Norman, D., Borgstrom, M. & Silfverdal, S. A. The clinical presentation of acute bacterial meningitis varies with age, sex and duration of illness. Acta paediatrica 104, 1117–1124 (2015).
Hornig, M. & Lipkin, W. I. Immune-mediated animal models of Tourette syndrome. Neuroscience & Biobehavioral Reviews 37, 1120–1138 (2013).
Libbey, J. E. & Fujinami, R. S. Role for antibodies in altering behavior and movement. Autism Research 3, 147 (2010).
Li, W. et al. Structural changes of gut microbiota in Parkinson’s disease and its correlation with clinical features. Science China Life Sciences (2017).
Qin, N. et al. Alterations of the human gut microbiome in liver cirrhosis. Nature 513, 59–64 (2014).
Riordan, S. M. & Williams, R. The intestinal flora and bacterial infection in cirrhosis. Journal of Hepatology 45, 744–757 (2006).
Verrall, A. J. et al. Rothia aeria as a cause of sepsis in a native joint. Journal of clinical microbiology 48, 2648–2650 (2010).
Monju, A. et al. First case report of sepsis due to Rothia aeria in a neonate. Journal of clinical microbiology 47, 1605 (2009).
de Steenhuijsen Piters, W. A. et al. Dysbiosis of upper respiratory tract microbiota in elderly pneumonia patients. The ISME journal 10, 97–108 (2016).
Olsen, I. Update on bacteraemia related to dental procedures. Transfusion & Apheresis Science 39, 173–178 (2008).
Moazzam, A. A. et al. Intracranial bacterial infections of oral origin. Journal of Clinical Neuroscience 22, 800 (2015).
Miklossy, J. Alzheimer’s disease - a neurospirochetosis. Analysis of the evidence following Koch’s and Hill’s criteria. Journal of Neuroinflammation 8, 1–16 (2011).
Zander, E., Sturm, H. & Bölte, S. The added value of the combined use of the Autism Diagnostic Interview–Revised and the Autism Diagnostic Observation Schedule: Diagnostic validity in a clinical Swedish sample of toddlers and young preschoolers. Autism 19, 187–199 (2014).
Acknowledgements
This study was supported by funds from the National Natural Science Foundation of China (81371129& 81670973). We would like to thank Zhengsheng Xue, Ping Wu and Chang Han for help in this project.
Author information
Authors and Affiliations
Contributions
F.C., M.W. and Y.Q. designed the study. Y.Q., M.W., Y.F., Z.Z. and L.C. collected the clinical samples. Y.Q. performed the experiments, analyzed the data and wrote the manuscript. F.C. and M.W. revised the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Qiao, Y., Wu, M., Feng, Y. et al. Alterations of oral microbiota distinguish children with autism spectrum disorders from healthy controls. Sci Rep 8, 1597 (2018). https://doi.org/10.1038/s41598-018-19982-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-19982-y
This article is cited by
-
Autism spectrum disorders and the gastrointestinal tract: insights into mechanisms and clinical relevance
Nature Reviews Gastroenterology & Hepatology (2024)
-
Autistic individuals have worse oral status than neurotypical controls: a systematic review and meta-analysis of observational studies
Clinical Oral Investigations (2024)
-
The Association Between Functional Variants in Long Non-coding RNAs and the Risk of Autism Spectrum Disorder Was Not Mediated by Gut Microbiota
Molecular Neurobiology (2024)
-
Autism Severity, Adverse Childhood Experiences, and Oral Health: A Comparative Study of Adolescents in the United States
Journal of Autism and Developmental Disorders (2024)
-
Altered oral microbiota and immune dysfunction in Chinese elderly patients with schizophrenia: a cross-sectional study
Translational Psychiatry (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.