Abstract
Two unique housefly strains, MSS and N-MRS, were selected and used to clarify mechanisms of sex-associated malathion resistance in the housefly, Musca domestica. Compared with the lab-susceptible CSS strain, susceptible females and resistant males were observed in the malathion-susceptible MSS strain, while the malathion-resistant near-isogenic line, N-MRS, achieved similar resistance level between genders. Significant synergistic effect of the esterase-inhibitor DEF on resistant houseflies pointed to the important involvement of esterase in this specific malathion resistance. Examination of the carboxylesterase gene MdαE7 in malathion resistant housefly populations found seven, non-synonymous SNP mutations (Ser250-Thr, Trp251-Ser, Met303-Ile, Leu354-Phe, Ser357-Leu, Trp378-Arg and Ser383-Thr), not found in susceptible houseflies, revealing a strong correlation between these mutations and the development of malathion resistance. Further genetic analysis conducted with bioassays by topical application and nucleotide polymorphism detection provided a first line of molecular evidence for a linkage between a male-determining factor and MdαE7 gene in the MSS and N-MRS males. This linkage results in a much higher level of malathion resistance for males than females in the MSS strain. Lastly, quantitative real-time PCR showed that MdαE7 was over expressed in the resistant strain due to the increased transcription level of mRNA rather than gene duplication.
Similar content being viewed by others
Introduction
The housefly, Musca domestica L. is a serious threat to human and animal health by carrying more than 100 pathogens1. Insecticides have been utilized for decades as the main tool to control houseflies. Malathion, an organophosphate (OP) insecticide, is widely used against houseflies owing to its toxicity profile2. Unfortunately, resistance to malathion in housefly was detected since the 1960s3.
Esterase-mediated resistance has been identified as one of the main mechanisms in OP resistance among insects4. In several dipteran and hemipteran insects, it is generally accepted that OP resistance is mainly mediated by enhanced metabolic detoxification or sequestration through quantitative changes in activity of general esterases5. Quantitative changes, such as gene duplication or up-regulation, or a combination of both, lead to the overproduction of enzymes6. The most comprehensive research on insecticide detoxification by gene amplification involved the overexpression of a specific carboxylesterase in Myzus persicae. This study demonstrated that the duplication of the genes E4, or its paralog FE4, were responsible for enhanced degradation and sequestration of insecticides7. Studies on other species, for example Culex pipiens (Diptera: Culicidae)8 and Nilaparvata lugens (Hemiptera: Delphacidae)9, provided more evidence for this resistance mechanism. Additionally, the up-regulation and elevated expression of esterase-encoding genes were observed in many species, such as B-biotype of Bemisia tabaci (Hemiptera: Aleyrodidae)10 and Aphis gossypii (Glover) (Hemiptera: Aphididae)11.
Structural changes in carboxylesterase, which confer an enhanced ability to metabolize the insecticide, are considered to be another important contributor in esterase-based OP resistance. In dipteran insects, a malathion-specific carboxylesterase (MCE), which hydrolyzes the insecticide to the less toxic and more easily excreted monoacids, has been implicated to be involved in resistance to malathion12. Research by van Asperen and Oppenoorth found that resistance to OP in housefly was always accompanied by reduced activity of ali-esterase in several resistant strains13. This relationship was explained by the so-called ‘mutant ali-esterase theory’, which proposed that the reduced ability to hydrolyze carboxylesterase substrates associated with an acquired ability to hydrolyze organophosphate substrates was caused by structural mutations in carboxylesterase14,15. Furthermore, mutational studies on OP resistance provided powerful, molecular evidence for this theory. LcαE7 and MdαE7, a pair of homologous genes encoding carboxylesterase E3 in Lucilia cuprina and Musca domestica, respectively, were identified to be associated with both the loss of E3′s carboxylesterase activity and the acquisition of a novel OP hydrolysis activity. The same mutation Gly137Asp in both genes was proposed to be responsible for diazinon resistance, while another substitution Trp251Leu in LcαE7 and Trp251Ser in MdαE7 was involved in malathion resistance2,16,17. Since these seminal studies, mutation of esterase genes mediating OP resistance has been identified in many insect species18.
In houseflies of the so-called “standard” strains where females carry XX and males carry XY sex chromosomes sex is determined by a male-determining factor (M factor) located on the Y chromosome that blocks the female-determining factor (F factor)19. However, in many populations, the heterozygous or homozygous presence of the M factor has been discovered on one or multiple autosomes, or even rarely on the X chromosome20,21. Due to the presence of multiple M factors in the same individual, the F factor has evolved a dominant, constitutive mutation, F D, to trigger female development19. The first case of sex-associated insecticide resistance in housefly, reported in 1990, showed females were more resistant against permethrin than males22. Later studies focusing on malathion as well as spinosad resistance indicated that the correlation of insecticide resistance to sex in housefly was not coincidental23,24. Interestingly, unlike the other two insecticides, the higher level of malathion resistance was observed in males rather than females. Kerr reported a segregation linkage between an autosomal M factor and a DDT resistance gene through research into the genetic basis of DDT resistance25. Moreover, similar conclusions were found in both pyrethroids and OP resistance22,23. Due to the finding that M factors can be present in different linkage groups in diverse housefly populations, the likelihood of an association between the M factor and a transposable genetic element was proposed26. Subsequent genetic analysis showing the M factor moved to chromosome 2 from chromosome 3 as a consequence of selection with malathion supported this suggestion23. Recently, the M factor has been identified to be originated from a duplication of a spliceosomal factor gene27. However, the molecular basis of this linkage is still unclear despite two decades of research.
Near-isogenic lines (NILs) refer to inbred populations of organisms with an approximately identical genetic background to the recurrent parent, except for loci chromosomally close to the target genotype. Although the susceptible and resistant strains used for insecticide resistance research are typically from two different populations, NILs can be used for avoiding the phenotypic interference from nonresistant factors found in more widely differing genetic backgrounds. Several insect NILs have been utilized for genetic analyses of insecticide resistance28,29,30,31, resistance gene mapping and cloning32, and fitness tests33.
In the housefly genome, a total of 92 esterase genes were predicted34. Given that the carboxylesterase gene MdαE7 was suggested to be responsible for malathion resistance of houseflies2,17, in this manuscript we report the establishment of a NIL of malathion-resistant (N-MRS) housefly to minimize the differences in genetic backgrounds, theoretically enabling clarification of the involvement of multiple mutations and MdαE7 overexpression in sex-specific malathion resistance. This allowed us to provide molecular evidence for a possible role of an M factor in insecticide resistance of the housefly.
Results
Toxicity analysis and biochemical assays
As the result of selection, compared to the susceptible CSS females, a 11-fold decrease of LD50 to malathion was observed in the malathion-susceptible MSS females. Meanwhile, surprisingly, the MSS males obtained 10-fold malathion resistance to the CSS males, with a significantly higher LD50 than the MSS females. After the process of repeated backcrossing and self-breeding under malathion selection, the near-isogenic resistant to malathion N-MRS females and males developed 56- and 54-fold resistance relative to the CSS strain, respectively (Table 1).
The effects of the esterase inhibitor DEF as a synergist of malathion toxicity in females and males are shown in Table 1. The synergistic ratios for the MSS males, N-MRS females, and N-MRS males pretreated with DEF were 176, 294, and 127, respectively, while the corresponding value for the MSS females was 1.86.
Identification of mutations related to malathion resistance in MdαE7
The total coding region of the carboxylesterase MdαE7 gene was successfully cloned and sequenced. Alignment of nucleotide and amino acid sequences of the cDNA fragment revealed a total of seven, non-synonymous mutations that consistently occurred in the resistant houseflies, but not the susceptible MSS females (Table 2): Ser250-Thr (S250T), Trp251-Ser (W251S), Met303-Ile (M303I), Leu354-Phe (L354F), Ser357-Leu (S357L), Trp378-Arg (W378R) and Ser383-Thr (S383T). The former six mutations were caused by single nucleotide substitutions, while in the last mutation two nucleotides (G1148 and T1149) were replaced at the Ser383 codon. However, a polymorphic allele at C1148 that can produce a substitution codon encoding threonine (Thr383) renders irrelevant the replacement of the third nucleotide in this codon because the amino acid switch only requires the G1148C substitution.
Based on the three-dimensional structure of LcαE7 (76% identical with MdαE7), which has been solved by X-ray crystallography, MdαE7 gene was successfully modeled after energy minimization. Similar with LcαE7 35, the structure of active sites in MdαE7 are comprised of a canonical, catalytic triad of the α/β-hydrolase fold (Ser218, His471 and Glu351), an oxyanion hole (Ala219, Gly136 and Gly137) and an extremely asymmetrical cavity for substrate binding. This model of the substrate binding pocket includes a small pocket (Leu354, Tyr457, Met460 and Ala472) and a large pocket (Trp251, Met308, Phe309, Phe355, Tyr420 and Phe421) (Fig. 1), revealing that two of the detected mutation sites, both Trp251 and Leu354 are in the active site region. Moreover, five of the substitutions are predicted to be close to the substrate binding cavity except for Trp378 and Ser383, which were, respectively, 21.0 and 28.0 Å away from Ser218, the catalytic center of the molecule.
The structure of the active cavity of MdαE7. The catalytic triad and oxyanion hole are colored magenta and blue, respectively. The substrate binding pocket is divided into a small pocket and a large pocket, colored green and yellow, respectively. Mutations are displayed as sticks and colored dark red, except Trp251 and Leu354, which are located in the binding pocket.
Correlation of polymorphic allele frequencies with malathion resistance in housefly
We conducted a point mutation screen of resistant houseflies to search for polymorphisms in 80 adult individuals from each gender of the MSS and N-MRS strains to explore their correlation with malathion resistance. We found significant differences in the occurrence of polymorphisms among different populations (Table 3). Only homozygous susceptible (wild-type) alleles of each of the seven mutation sites were carried by the MSS individuals, while homozygous mutant alleles were only detected in the N-MRS strain. Notably, homozygous susceptible alleles mainly occurred in the MSS females (96.25% to 100%) while the majority of MSS males (80% to 98.75%) were heterozygous for these alleles, showing extremely unbalanced expression of polymorphisms between genders. In contrast, consistently high frequencies of homozygous mutant alleles were examined in both the N-MRS females (86.25% to 100%) and males (93.75% to 100%).
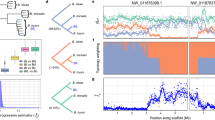
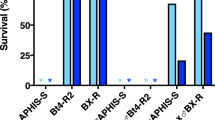
We separately examined the prevalence of each mutation in the F2 females and males in order to further evaluate the relationship between the polymorphic allele frequencies and increased levels of resistance to malathion. Four-day-old houseflies were treated with different concentrations of malathion and assembled into four groups (A to D), for each gender, based on their similar levels of malathion resistance (low to high, respectively) (Table 4). The results of SNP detection revealed that the frequencies of polymorphic expression at each codon site were strongly correlated with the levels of resistance in housefly (Fig. 2). High expression of homozygous susceptible alleles and low expression of heterozygous alleles in the F2 females were observed in groups A1 and B1, with relatively low malathion resistance. However, a significantly decreased frequency of homozygous susceptible alleles and increased frequency of heterozygous alleles was detected in group C1 and D1, accompanied by an increase in malathion resistance. Among the F2 males, heterozygotes were only found in the former three groups (A2, B2, and C2), with significantly higher occurrence in groups A2 and B2 than group C2. In confirmation of our predictions, our results showed an increasing trend of malathion-tolerant individuals carrying homozygous mutant alleles, for example, comprising the majority of group D2.
SNP mutation allele frequency distribution for each F2 generation housefly group, distinguished by gender and level of malathion tolerance. The frequency of allele expression shown along the Y axis is the percentage of houseflies carrying the corresponding homozygous or heterozygous allele(s). Housefly groups are shown along the X axis; A1, B1, C1 and D1 represent the groups in F2 females that were dead under LD10, between LD10 and LD50, between LD50 and LD90 and alive above LD90 dosage treatment, respectively; and A2, B2, C2 and D2 represent the groups in F2 males that were dead under LD10, between LD10 and LD50, between LD50 and LD90 and alive above LD90 dosage treatment, respectively. Error bars represent standard errors of the means (n = 4 independent replicates of treatment).
Linkage analysis
At the seven mutations, we observed that the majority of MSS females carried homozygous susceptible alleles, while the MSS males were primarily heterozygous for these same alleles. In view of current knowledge of the M factor, one possible explanation is that the mutant allele and M factor are carried on the same chromosome in heterozygous MSS males, so only male offspring can inherit the mutant form. Additionally, rare heterozygotes of certain codon sites still present in the MSS females may be caused by recombination36. Therefore, a series of genetic crosses was carried out to identify whether the mutant MdαE7 gene is also linked to Mfactor in the N-MRS males.
Offspring from the F1, F2, and BC generation were separately tested for malathion resistance (Table 5). The F1 hybrids showed a similar level of resistance between females and males, with a LD50 of 359.80 and 466.05 μg/fly, respectively. However, in the F2 generation, a high degree of resistance to malathion was maintained in males, while a nearly 14-fold decrease was observed in females compared with their parental houseflies. The BC females and males also expressed remarkably different levels of resistance with a resistance ratio of 0.18- and 36-fold compared with the CSS strain, respectively.
To validate our hypothesis of a linkage between the MdαE7 gene and Mfactor in the N-MRS males at a molecular level, we examined the frequency of polymorphic expression at the seven codon sites in the F2 and BC generation (Table 3). Among the seven mutation sites found in F2 females, 56.72% to 59.55% of individuals had the homozygous susceptible alleles and 40.45% to 43.28% of individuals had heterozygous alleles. However, only heterozygous and homozygous mutant alleles were expressed in the F2 males, with a frequency of 50.99% to 51.72% and 48.28% to 49.01%, respectively. As for the BC offspring, no nucleotide substitutions at the seven codon sites were detected in any females, and individuals carrying heterozygous alleles were, for the most part of males, notably comprising the entire group with mutations at codon sites 378 and 383.
Since the inheritance mode of malathion resistance in housefly has been demonstrated as single, autosomal and incompletely dominant3,37, according to our hypothesis mentioned above, the F1 offspring should be all heterozygotes in both genders, regardless if the linkage exists or not (Fig. 3A). Following the law of linkage, a 1:1 segregation ratio of homozygous susceptible to heterozygous alleles is expected in F2 females with an intermediate level of malathion tolerance relative to MSS and F1 females, which carried homozygous susceptible and heterozygous alleles, respectively. As for F2 males, heterozygous individuals were expected to segregate in equal number to homozygous mutants, resulting in their intermediate resistance between the MSS males and the N-MRS strain. Meanwhile, BC females and males are expected to carry only homozygous susceptible and heterozygous alleles, respectively (Fig. 3B), resulting in a BC population of susceptible females and males with similar resistance to that of F1 males. As described above, the results of both toxicity analysis and SNP detection are relatively consistent with our assumption. However, the higher-than-expected segregation ratio (nearly 6:4) in the F2 females is possibly attributable to the use of only 43 individuals from the F2♀- > LD90 group for SNP detection. The unexpected expression of homozygous susceptible alleles in the BC males may be owing to rare recombination36 or rare heterozygotes in the N-MRS males.
Analysis of the MdαE7 gene expression
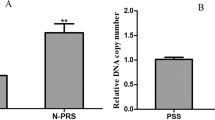
Comparisons of MdαE7 gene expression were made from two aspects: expression between genders of the same strain and expression between the MSS and N-MRS strains of the same gender (Table 6). The N-MRS strain showed significantly higher mRNA expression than the MSS strain in each gender, with a p-value of 0.004 and 0.035 for females and males, respectively. However, the DNA expression did not differ between strains with both genders. In addition, no significant differences of DNA and mRNA expressions were observed between genders in both strains.
Discussion
Near-isogenic lines are important material for the study of insecticide resistance. With the aid of a rotating set of genetic methods and concurrent malathion selection, we introduced purified malathion-resistant gene(s) into the genetic background of the susceptible MSS strain without loss, to generate the resistant N-MRS strain for further study. Our data from synergistic bioassays confirmed the involvement of esterase in sex-specific resistance to malathion.
Our investigation unveiled seven, non-synonymous SNP mutations in the MdαE7 gene in the malathion resistant populations. Several studies have reported the existence and crucial contribution of the W251S non-synonymous mutation in malathion resistant houseflies. It has been proposed that the replacement of W251 with smaller residues creates more space to accommodate substrates with bulkier acid moieties, and reduces the steric obstruction to the inversion that must occur around phosphorus during hydrolysis of OP38,39,40,41. Additionally, another four mutations, S250T, M303I, S357L and S383T were detected in certain housefly strains with high malathion resistance2. Notably, two substitutions in our study, L354F and W378R, were the first discovery of these mutations in malathion resistant houseflies. In this case, L354F likely coincides with the carboxylesterase binding, which is the structural equivalent of the anionic site in cholinesterase, providing the π electrons to accommodate the quaternary nitrogen of acetylcholine42. Kinetic assays have shown that substitution by insertion of a small aliphatic side chain resulted in a greater increase of esterase activity than that with a larger aromatic side chain in the residue F354 of LcαE7 41, which was surprising, given our detection of the L354F mutation in MdαE7. Since all seven mutations have also been documented in houseflies resistant to pyrethroid insecticides43, it has been suggested that carboxylesterase evolution via qualitative mechanisms may cause broad insecticide resistance in insects.
A strategy of comparing polymorphic codon usage bias in individuals between different populations and groups allowed us to supply proof of the potential association between each mutation and malathion resistance in housefly. Among all the mutations, a clear shift of polymorphic distribution was detected from those with primarily homozygous susceptible alleles, through those with mostly heterozygous alleles, to those with all, or nearly all, homozygous mutant alleles, correlating with increasing resistance to malathion, and ultimately supported our predictions. Future studies will examine the effects of individual or combinations of mutations on the degradation or sequestration of malathion and other insecticides.
Genetic crosses conducted for this study, in combination with bioassays and SNP detection, identified the coexistence of an M factor and mutant MdαE7 gene on the same chromosome in both MSS and N-MRS males. Notably, this linkage in MSS males also offered a compelling explanation for their surprisingly higher level of resistance than the susceptible MSS females. To further previous research that focused on the linkage between an autosomal M factor and malathion resistance factors23, our study is the first report on the molecular characterization of this linkage in both the susceptible strain and the resistant near-isogenic line, thus providing valuable insights into the important role the M factor may play in sex-associated insecticide resistance of housefly.
In our work, we found that overproduction of MdαE7 gene transcripts in the N-MRS strain indicated that the increased expression of the carboxylesterase gene was associated with malathion resistance in housefly. Similar to a resistance study in other insect species44, the combination of quantitative and qualitative changes in carboxylesterase may further contribute to the malathion resistance of insects. Moreover, our analysis suggested that up-regulation of the MdαE7 gene was mainly responsible for the overabundance of the gene product. Many types of regulatory mutations can lead to changes in gene expression, and can occur either in cis (disruption or deletion of an upstream regulatory element in the promoter region of the gene) or in trans (disruption of a gene coding for a transcription factor that binds the above-mentioned cis-elements)45. Although we have obtained evidence for both cis- and trans-regulation of metabolic resistance involving glutathione S-transferases’ and cytochrome P450 monooxygenases’ activity, the regulatory mechanisms of esterase-mediated resistance are still poorly understood10. In our estimation, it is noteworthy that MdαE7 was 2.1-fold higher expressed in males than females in the N-MRS strain, in spite of no significant difference between them. Given the linkage between M factor and MdαE7 gene in the N-MRS males, it is reasonable to consider the possible involvement of an M factor in the cis-regulation of esterase genes, suggesting a novel direction for further investigation into the regulatory factors of gene overexpression.
Methods
Insects
Three housefly strains were used in this study.
The CSS strain, a susceptible strain derived from National Taiwan University in 1987, has been reared in laboratory without exposure to any insecticides. Malathion toxicity in this strain was included to evaluate resistance levels in each population tested in our study.
The MSS strain was selected from the CSS strain. Unmated houseflies less than 3 h old from the CSS strain were collected in this selection. Single pairs of houseflies were reared and crossed individually. Based on a previous bioassay of CSS females, partial progeny of each pair were treated with an appropriate dosage of malathion, and the most susceptible pair was selected according to the highest mortality after 24 h. Progeny without malathion treatment from the selected pair were chosen for further breeding. The MSS strain was produced after two generations with this selection.
The near-isogenic line to malathion resistance (N-MRS) strain was established using the protocols of Shan31, with some modifications. A wild housefly strain (CFD), originally collected from a garbage dump in Beijing, China, in 1998, and maintained in the laboratory without exposure to insecticides, showed high resistance to malathion. The CFD males and unmated MSS females were collected 3 h after emergence and crossed to produce a hybrid F1 population. An F2 population was derived from the self-breeding of F1 flies. The bioassay was conducted on four-day-old F2 females, while the rest of F2 population (collected within 24 h of emergence) was treated with a proper dosage of malathion to kill all the susceptible ones and most of the heterozygotes. The survivors produced an F3 generation. F3 males were backcrossed to virgin MSS females, both collected within 3 h of emergence, to produce the BC1 population. Subsequently, the BC1F1 population was derived from the self-breeding of BC1, and the same treatment for F2 was repeated on BC1F1. The N-MRS strain was thus constructed after 7 repeated processes of backcrossing and self-breeding with malathion selection.
Bioassay and analysis for synergism
Topical applications were performed by delivery of a 1.1-μL drop of insecticide (in acetone) onto the thoracic notum of four-day-old adult houseflies46. Malathion was dissolved in acetone to produce 5–7 concentrations that gave a mortality rate greater than 0 and less than 100%. Control groups were treated with a 1.1-μL drop of acetone alone. Each of the three replicates consisted of 15 to 20 flies per dose. Treated houseflies were placed in 240 ml plastic jars with a piece of sponge saturated in sugar water. Mortality was assessed after 24 h. Flies that did not move normally were scored as dead. All tests were performed at 25 ± 1 °C.
Synergistic assays were applied to study the biochemical mechanism of resistance. Both females and males were separately tested by using S, S, S-tributylphosphorotrithioate (DEF), an esterase inhibitor47. DEF was dissolved in acetone and applied by topical application at a dose of 0.98 μg/fly 1 h before the malathion treatment. The dosage of DEF was selected experimentally as the highest dose with no control mortality.
Sequence analysis and protein modeling of MdαE7
Total RNA was extracted from a pool of adult houseflies of each gender, from the MSS and N-MRS strains, with a TRIzol kit (Invitrogen, Carlsbad, CA), following the manufacturer’s instructions. The first-strand cDNA was synthesized using PrimeScriptTM RT Reagent kit with gDNA Eraser (Takara Biotechnology, Dalian, China). To screen for putative mutations related to resistance, the full length of MdαE7 cDNA was amplified by using primers AE7.30 (5′ATGAATTTCAAAGTTAGTCAA3′) and AE7.11 (5′AAACAATTCCTTCTTTTTATCGA3′)17. Amplification started with an initial denaturation step of 94 °C for 5 min, followed by 35 cycles of PCR reaction (94 °C for 30 sec, 48 °C for 30 sec, and 72 °C for 2 min) and a final extension of 72 °C for 10 min. Purified PCR products were cloned into the pMD 18-T vector and sequenced by Beijing Genomics Institute (BGI). Sequence analyses of MdαE7 cDNA fragments were repeated three times for each housefly sample, with different preparations of mRNA, and three TA clones sequenced from each replicate. Alignment was performed using DNAman (Lynnon Biosof, USA).
The deduced amino acid sequence of the MdαE7 gene was modeled against the 3D structure of the LcαE7 gene (PDB accession no.5ikx)48 using the Swiss-model homology modeling server49,50,51, refined using Dokholyan Lab’s Chiron Server52, checked and validated using Structural Analysis and Verification Server from UCLA (http://services.mbi.ucla.edu/SAVES/) and viewed using Pymol Viewer.
Genetic cross and malathion treatment
To identify the potential linkage between the M factor and malathion resistance in Musca domestica, MSS females and N-MRS males within 12 h of emergence were put into a cage for mass mating to produce the hybrid F1. A backcross by F1 males to virgin MSS females, as well as a self-cross of the F1 generation, were subsequently conducted, and the offspring were referred to as BC and F2 progeny, respectively. Bioassays were performed on both genders from each generation.
Based on the bioassay results of the F2 generation, a corresponding dose range of LD10, LD50 and LD90 were obtained and utilized to generate 4 groups divided by different levels of malathion resistance in each gender. According to the previous method53, four-day-old F2 females and males were separately treated with malathion at their respective dose of LD10. Eight hours after treatment, the dead houseflies of each gender were collected and designated as group A (i.e., F2♀- < LD10, or F2♂- < LD10). The survivors of the LD10 dose were then exposed to malathion at the dose of LD50. Eight hours later, the dead houseflies of both genders were collected and designated group B (i.e., F2♀-LD10−50, or F2♂-LD10−50), and live flies were continually exposed to a malathion LD90 dose. Eight hours after treatment, the dead and surviving houseflies were separately collected and designated as group C (i.e., F2♀-LD50−90, or F2♂-LD50−90) and group D (i.e., F2♀- > LD90, or F2♂- > LD90), respectively. Each treatment was repeated 4 times, and a total of 1858 females and 1748 males were treated in this experiment.
Nucleotide polymorphism (SNP) detection in MdαE7
Houseflies collected from the previous cross and malathion treatment were used in SNP determination. Four replications were conducted and a total of 80 individual houseflies from each population were utilized, with 20 for each replication. A total of 60, 31 and 43 houseflies were collected in the groups of F2♂- < LD10, F2♂- > LD90 and F2♀- > LD90, respectively, and all houseflies from the three groups were used for SNP detection. Extraction of total RNA and synthesis of the first strand cDNAs from each individual housefly were described above. Primer pair 1 (forward: 5′CGGCGAAGCAAATCGTAACT3′; reverse: 5′AAGCGATGCATGGGGAAGAG3′) was designed to amplify the specific fragment from each of the individual houseflies on which the polymorphisms reside. Amplification began with an initial step of 5-min denaturation at 94 °C. The subsequent cycling conditions consisted of denaturation at 94 °C for 30 sec, annealing at 55 °C for 30 sec, and extension at 72 °C for 1 min for 35 cycles. The final step included a 10-min extension at 72 °C. PCR products were directly sequenced and the allelic polymorphisms of each mutation were analyzed using the chromatogram viewer Chromas (Technelysium Pty Ltd). The frequencies of polymorphic expression were recorded for each mutation in all tested individuals.
Real-time PCR
The relative transcription level and copy number of the MdαE7 gene in females and males from the MSS and N-MRS strains were separately tested by quantitative real-time PCR. Since abdomen was reported to be the principal site of carboxylesterase activity in adult houseflies54,55, both genomic DNA and total RNA were extracted from abdomens of four-day-old houseflies. Modified from the method of Cao11, the standard curves of MdαE7 and the internal reference gene GAPDH were generated. Briefly, primer pair 2 (forward: 5′CGCTTCCTACAATACGCTTC3′; reverse: 5′CATCGGCATGGCTTACACC3′) was designed for MdαE7 and primer pair 3 (forward: 5′GGTCATCATCTCCGCTCCATC3′; reverse: 5′CAGTGGTGGCATGGACAGTGG3′) was for GAPDH to detect the relative expression levels of mRNA and DNA. To construct the standard plasmid, the respective fragment derived from a PCR reaction with primer pair 2 or 3 was inserted into the pMD 18-T vector. The plasmid was extracted and quantified by spectrophotometry. The plasmid mass was converted into copy concentration using the formula copies/μL = C × 10−6 × 6.02 × 1023/(660 × L), where C is the plasmid concentration (μg/μL) and L is the plasmid length (bp). The standard curve was generated using the threshold cycle of a serial 10-fold dilution of plasmid templates. PCR of 20 μL reactions were conducted using 1 μL of cDNA (equivalent to 1 μg of total RNA), or 4 μL of DNA (equivalent 0.8 μg of DNA), 10 μL of 2× SYBR Premix Ex Taq (Takara), 0.4 μL of each primer (10 mM), 0.4 μL of Rox (Takara), and 7.8 μL (for RNA) or 4.8 μL (for DNA) ddH2O. The reactions were performed on ABI PRISM 7500 HT Sequence Detection Systems, the PCR program for which consisted of an initial step at 95 °C for 2 min, followed by 40 cycles of 95 °C for 15 sec and 60 °C for 30 sec. Specificity of the amplification product was assessed by a melting curve generated by a final dissociation stage at 95 °C for 15 sec, 60 °C for 15 sec and 95 °C for 15 sec. All samples including the non-template control were run in three replicates and the experiment was independently conducted three times with different RNA and DNA preparations. The accurate initial copy number of MdαE7 and GAPDH gene were calculated by their corresponding standard curves, respectively. The relative DNA and mRNA expressions of MdαE7 was normalized by GAPDH (amount of MdαE7 gene/ amount of GAPDH gene).
Data analysis
For the bioassays, data were pooled and probit analysis was made by POLO-Plus 2.0 software (LeOra Software Lnc., Berkeley, CA)56. Statistical analysis of LD50 was based on non-overlap of 95% confidence intervals57. The resistance ratio (RR), the LD50 of the resistant population divided by the LD50 of the reference susceptible population, presents the level of resistance. Synergism ratios (SR) were calculated by dividing the LD50 of insecticide alone by the LD50 of insecticide with DEF.
For all two-sample comparisons, the statistically significant differences of gene expression were determined using a Student’s t-test (SPSS v20.0 software), with a value of p < 0.05 considered statistically significant.
References
Pavela, R. Insecticidal properties of several essential oils on the house fly (Musca domestica L.). Phytotherapy Research 22, 274–278, https://doi.org/10.1002/ptr.2300 (2008).
Taskin, V. & Kence, M. The genetic basis of malathion resistance in housefly (Musca domestica L.) strains from Turkey. Genetika 40, 1475–1482, https://doi.org/10.1023/B:RUGE.0000048663.17417.97 (2004).
Nguy, V. D. & Busvine, J. R. Studies on the genetics of resistance to parathion and malathion in the housefly. Bulletin of the World Health Organization 22, 531–542 (1960).
Hemingway, J. The molecular basis of two contrasting metabolic mechanisms of insecticide resistance. Insect Biochemistry and Molecular Biology 30, 1009–1015, https://doi.org/10.1016/S0965-1748(00)00079-5 (2000).
Baker, J. E., Fabrick, J. A. & Zhu, K. Y. Characterization of esterases in malathion-resistant and susceptible strains of the pteromalid parasitoid Anisopteromalus calandrae. Insect Biochemistry and Molecular Biology 28, 1039–1050, https://doi.org/10.1016/S0965-1748(98)00095-2 (1998).
Panini, M., Manicardi, G. C., Moores, G. & Mazzoni, E. An overview of the main pathways of metabolic resistance in insects. Invertebrate Survival Journal 13, 326–335 (2016).
Bass, C. et al. The evolution of insecticide resistance in the peach potato aphid, Myzus persicae. Insect Biochemistry and Molecular Biology 51, 41–51, https://doi.org/10.1016/j.ibmb.2014.05.003 (2014).
Hemingway, J., Hawkes, N. J., McCarroll, L. & Ranson, H. The molecular basis of insecticide resistance in mosquitoes. Insect Biochemistry and Molecular Biology 34, 653–665, https://doi.org/10.1016/j.ibmb.2004.03.018 (2004).
Small, G. J. & Hemingway, J. Molecular characterization of the amplified carboxylesterase gene associated with organophosphorus insecticide resistance in the brown planthopper, Nilaparvata lugens. Insect Molecular Biology 9, 647–653, https://doi.org/10.1046/j.1365-2583.2000.00229.x (2001).
Alon, M., Alon, F., Nauen, R. & Morin, S. Organophosphates’ resistance in the B-biotype of Bemisia tabaci (Hemiptera: Aleyrodidae) is associated with a point mutation in an ace1-type acetylcholinesterase and overexpression of carboxylesterase. Insect Biochemistry and Molecular Biology 38, 940–949, https://doi.org/10.1016/j.ibmb.2008.07.007 (2008).
Cao, C. W., Zhang, J., Cao, X. W., Liang, P. & Guo, H. L. Overexpression of carboxylesterase gene associated with organophosphorous insecticide resistance in cotton aphids, Aphis gossypii (Glover). Pesticide Biochemistry and Physiology 90, 175–180, https://doi.org/10.1016/j.pestbp.2007.11.004 (2008).
Krueger, H. R. & O’Brien, R. D. Relationship between metabolism and differential toxicity of malathion in insects and mice. Journal of Economic Entomology 52, 1063–1067, https://doi.org/10.1093/jee/52.6.1063 (1960).
van Asperen, K. & Oppenoorth, F. J. Organophosphate resistance and esterase activity in houseflies. Entomologia Experimentalis et Applicata 2, 48–57, https://doi.org/10.1007/BF00344520 (1959).
Oppenoorth, F. J. & van Asperen, K. Allelic genes in the housefly producing modified enzymes that cause organophosphate resistance. Science 132, 298–299, https://doi.org/10.1126/science.132.3422.298 (1960).
van Asperen, K. Biochemistry and genetics of esterases in houseflies (Musca domestica) with special reference to the development of resistance to organophosphorus compounds. Entomologia Experimentalis et Applicata 7, 205–214, https://doi.org/10.1007/BF00366837 (1964).
Campbell, P. M., Newcomb, R. D., Russell, R. J. & Oakeshott, J. G. Two different amino acid substitutions in the ali-esterase, E3, confer alternative types of organophosphorus insecticide resistance in the sheep blowfly, Lucilia cuprina. Insect Biochemistry and Molecular Biology 28, 139–150, https://doi.org/10.1016/S0965-1748(97)00109-4 (1998).
Claudianos, C., Russell, R. J. & Oakeshott, J. G. The same amino acid substitution in orthologous esterases confers organophosphate resistance on the housefly and a blowfly. Insect Biochemistry and Molecular Biology 29, 675–686, https://doi.org/10.1016/S0965-1748(99)00035-1 (1999).
Cui, F. et al. Two single mutations commonly cause qualitative change of nonspecific carboxylesterases in insects. Insect Biochemistry and Molecular Biology 41, 1–8, https://doi.org/10.1016/j.ibmb.2010.09.004 (2011).
Dübendorfer, A., Hediger, M., Burghardt, G. & Bopp, D. Musca domestica, a window on the evolution of sex-determining mechanisms in insects. International Journal of Developmental Biology 46, 75–79, https://doi.org/10.5167/uzh-509 (2002).
Hamm, R. L. & Scott, J. G. A high frequency of male determining factors in male Musca domestica (Diptera: Muscidae) from Ipswich, Australia. Journal of Medical Entomology 46, 169–172, https://doi.org/10.1603/033.046.0121 (2009).
Denholm, I., Franco, M. G., Rubini, P. G. & Vecchi, M. Identification of a male determinant on the X chromosome of housefly (Musca domestica L.) populations in South-East England. Genetical Research, Cambridge 42, 311–322, https://doi.org/10.1017/S0016672300021790 (1983).
Shnon, T. & Scott, J. G. Autosomal sex-associated pyrethroid resistance in a strain of house fly (Diptera: Muscidae) with a male-determining factor on chromosome three. Journal of Economic Entomology 83, 686–689, https://doi.org/10.1093/jee/83.3.686 (1990).
Kence, M. & Kence, A. Genetic consequences of linkage between malathion resistance and an autosomal male-determining factor in house fly (Diptera: Muscidae). Journal of Economic Entomology 85, 1566–1570, https://doi.org/10.1093/jee/85.5.1566 (1992).
Højland, D. H., Scott, J. G., Vagn Jensen, K. M. & Kristensen, M. Autosomal male determination in a spinosad-resistant housefly strain from Denmark. Pest Management Science 70, 1114–1117, https://doi.org/10.1002/ps.3655 (2014).
Kerr, R. W. Inheritance of DDT resistance in a laboratory colony of the house fly, Musca domestica. Australian Journal of Biological Sciences 23, 377–400, https://doi.org/10.1071/BI9700377 (1970).
Green, M. M. Transposable elements in Drosophila and other diptera. Annual Review of Genetics 14, 109–120, https://doi.org/10.1146/annurev.ge.14.120180.000545 (1980).
Sharma, A. et al. Male sex in houseflies is determined by Mdmd, a paralog of the generic splice factor gene CWC22. Science 356, 642–645, https://doi.org/10.1126/science.aam5498 (2017).
Shanahan, G. J. Genetics of diazinon resistance in larva of Lucilia cuprine (Wiedemann) (Diptera: Calliphoridae). Bulletin of Entomological Research 69, 225–228, https://doi.org/10.1017/S0007485300017685 (1979).
Roush, R. T. & Wolfenbarger, D. A. Inheritance of methomyl resistance in the tobacco budworm (Lepidoptera: Noctuidae). Journal of Economic Entomology 78, 1020–1022, https://doi.org/10.1093/jee/78.5.1020 (1985).
White, N. D. G. & Bell, R. J. Inheritance of malathion resistance in a strain of Tribolium castaneum (Coleoptera: Tenebrionidae) and effects of resistance genotypes on fecundity and larval survival in malathion-treated wheat. Journal of Economic Entomology 81, 381–386, https://doi.org/10.1093/jee/81.1.381 (1988).
Shan, C., Zhang, Y., Ma, Z. & Gao, X. Inheritance of propoxur resistance in a near-isogenic line of Musca domestica (Diptera: Muscidae). Journal of Economic Entomology 109, 873–878, https://doi.org/10.1093/jee/tow001 (2016).
Dong, K. & Scott, J. G. Linkage of kdr-type resistance and the para-homologous sodium channel gene in German cockroaches (Blattella germanica). Insect Biochemistry and Molecular Biology 24, 647–654, https://doi.org/10.1016/0965-1748(94)90051-5 (1994).
Mckenzie, J. A., Whitten, M. J. & Adena, M. A. The effect of genetic background on the fitness of the diazinon resistance genotypes of the Australian sheep blowfly, Lucilia cuprina. Heredity 49, 1–9, https://doi.org/10.1038/hdy.1982.60 (1982).
Scott, J. G. et al. Genome of the house fly, Musca domestica L., a global vector of diseases with adaptations to a septic environment. Genome Biology 15, 466, https://doi.org/10.1186/s13059-014-0466-3 (2014).
Jackson, C. J. et al. Structure and function of an insect α-carboxylesterase (αEsterase7) associated with insecticide resistance. PNAS 110, 10177–10182, https://doi.org/10.1073/pnas.1304097110 (2013).
Feldmeyer, B., Pen, I. & Beukeboom, L. W. A microsatellite marker linkage map of the housefly, Musca domestica: evidence for male recombination. Insect Molecular Biology 19, 575–581, https://doi.org/10.1111/j.1365-2583.2010.01016.x (2010).
Harris, R. L., Wearden, S. & Roan, C. C. Preliminary study of the genetics of house fly (Musca domestica) resistance to malathion. Journal of Economic Entomology 54, 40–45, https://doi.org/10.1093/jee/54.1.40 (1961).
Claudianos, C., Crone, E., Coppin, C., Russell, R. J. & Oakeshott, J. G. A genomics perspective on mutant aliesterases and metabolic resistance to organophosphates. Acs Symposium Series 808, 90–101, https://doi.org/10.1021/bk-2002-0808.ch005 (2002).
Scott, J. G. & Zhang, L. The house fly aliesterase gene (MdαE7) is not associated with insecticide resistance or P450 expression in three strains of house fly. Insect Biochemistry and Molecular Biology 33, 139–144, https://doi.org/10.1016/S0965-1748(02)00238-2 (2003).
Devonshire, A. L. et al. Kinetic efficiency of mutant carboxylesterases implicated in organophosphate insecticide resistance. Pesticide Biochemistry and Physiology 76, 1–13, https://doi.org/10.1016/S0048-3575(03)00054-3 (2003).
Heidari, R. et al. Hydrolysis of organophosphorus insecticides by in vitro modified carboxylesterase E3 from Lucilia cuprina. Insect Biochemistry and Molecular Biology 34, 353–363, https://doi.org/10.1016/j.ibmb.2004.01.001 (2004).
Järv, J. Stereochemical aspects of cholinesterase catalysis. Bioorganic Chemistry 12, 259–278, https://doi.org/10.1016/0045-2068(84)90010-5 (1984).
Zhang, L. et al. Quantitative and qualitative changes of the carboxylesterase associated with beta-cypermethrin resistance in the housefly, Musca domestica (Diptera: Muscidae). Comparative Biochemistry Physiology Part B: Biochemistry and Molecular Biology 156, 6–11, https://doi.org/10.1016/j.cbpb.2010.01.011 (2010).
Pan, Y., Guo, H. & Gao, X. Carboxylesterase activity, cDNA sequence, and gene expression in malathion susceptible and resistant strains of the cotton aphid, Aphis gossypii. Comparative Biochemistry Physiology Part B: Biochemistry and Molecular Biology 152, 266–270, https://doi.org/10.1016/j.cbpb.2008.12.002 (2009).
Feyereisen, R. Molecular biology of insecticide resistance. Toxicology Letters 82–83, 83–90, https://doi.org/10.1016/0378-4274(95)03470-6 (1995).
Scott, J. G. & Georghiou, G. P. Rapid development of high-level permethrin resistance in a field-collected strain of house fly (Diptera: Muscidae) under laboratory selection. Journal of Economic Entomology 78, 316–319, https://doi.org/10.1093/jee/78.2.316 (1985).
Li, J., Wang, Q. M., Zhang, L. & Gao, X. W. Characterization of imidacloprid resistance in the housefly Musca domestica (Diptera: Muscidae). Pesticide Biochemistry and Physiology 102, 109–114, https://doi.org/10.1016/j.pestbp.2011.10.012 (2012).
Fraser, N. J. et al. Evolution of protein quaternary structure in response to selective pressure for increased thermostability. Journal of Molecular Biology 428, 2359–2371, https://doi.org/10.1016/j.jmb.2016.03.014 (2016).
Biasini, M. et al. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Research 42, W252–W258, https://doi.org/10.1093/nar/gku340 (2014).
Arnold, K., Bordoli, L., Kopp, J. & Schwede, T. The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22, 195–201, https://doi.org/10.1093/bioinformatics/bti770 (2006).
Kiefer, F., Arnold, K., Künzli, M., Bordoli, L. & Schwede, T. The SWISS-MODEL Repository and associated resources. Nucleic Acids Research 37, D387–D392, https://doi.org/10.1093/nar/gkn750 (2009).
Ramachandran, S., Kota, P., Ding, F. & Dokholyan, N. V. Automated minimization of steric clashes in protein structures. Proteins Structure Function & Bioinformatics 79, 261–270, https://doi.org/10.1002/prot.22879 (2011).
Li, T. et al. Multiple mutations and mutation combinations in the sodium channel of permethrin resistant mosquitoes, Culex quinquefasciatus. Scientific Reports 2, 781, https://doi.org/10.1038/srep00781 (2012).
Ahmad, S. Localization of aliesterase and acetylcholinesterase enzymes in various tissues of susceptible and organophosphate resistant Musca domestica L. Comparative & General Pharmacology 1, 273–279, https://doi.org/10.1016/0010-4035(70)90020-0 (1970).
Ahmad, S. Larval and adult housefly carboxylesterase: Isozymic composition and tissue pattern. Insect Biochemistry 6, 541–547, https://doi.org/10.1016/0020-1790(76)90082-2 (1976).
Russell, R. M., Robertson, J. L. & Savin, N. E. POLO: A new computer program for probit analysis. Bulletin of the Esa 23, 209–213, https://doi.org/10.1093/besa/23.3.209 (1977).
Liu, N. & Scott, J. G. Genetics of resistance to pyrethroid insecticides in the house fly, Musca domestica. Pesticide Biochemistry and Physiology 52, 116–124, https://doi.org/10.1006/pest.1995.1036 (1995).
Acknowledgements
This research was supported by the National Basic Research Programme of China (Contract No. 2012CB114103).
Author information
Authors and Affiliations
Contributions
X.G. contributed to the conception and design of this research. Y.Z., J.L., Z.M. and C.S. conducted experiments and analyzed the data. Y.Z. wrote the main manuscript text. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, Y., Li, J., Ma, Z. et al. Multiple mutations and overexpression of the MdaE7 carboxylesterase gene associated with male-linked malathion resistance in housefly, Musca domestica (Diptera: Muscidae). Sci Rep 8, 224 (2018). https://doi.org/10.1038/s41598-017-17325-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-17325-x
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.