Abstract
Cytoplasmic dynein-1 transports intracellular cargo towards microtubule minus ends. Dynein is autoinhibited and undergoes conformational changes to form an active complex that consists of one or two dynein dimers, the dynactin complex, and activating adapter(s). The Lissencephaly 1 gene, LIS1, is genetically linked to the dynein pathway from fungi to mammals and is mutated in people with the neurodevelopmental disease lissencephaly. Lis1 is required for active dynein complexes to form, but how it enables this is unclear. Here, we present a structure of two yeast dynein motor domains with two Lis1 dimers wedged in-between. The contact sites between dynein and Lis1 in this structure, termed ‘Chi,’ are required for Lis1’s regulation of dynein in Saccharomyces cerevisiae in vivo and the formation of active human dynein–dynactin–activating adapter complexes in vitro. We propose that this structure represents an intermediate in dynein’s activation pathway, revealing how Lis1 relieves dynein’s autoinhibited state.
Similar content being viewed by others
Main
Cytoplasmic dynein-1 (dynein) is the main minus-end-directed microtubule motor responsible for transporting vesicles, protein complexes, RNAs, organelles, and viruses along microtubules. Dynein also positions nuclei and the mitotic spindle during mitosis and meiosis1. Dynein is conserved across eukaryotes, with the exception of flowering plants and some algae, and requires numerous binding partners and regulators to function1,2. Mutations in dynein or its regulators cause neurodevelopmental and neurodegenerative diseases in humans, while homozygous deletion of dynein is embryonically lethal in mice3,4. By contrast, in the yeast S. cerevisiae, dynein and its regulators are conserved but non-essential, providing an important model system to study dynein’s mechanism and function.
Dynein is a 1.4-MDa complex consisting of a dimer of two motor-containing heavy chains composed of a ring of six AAA+ (ATPase associated with various cellular activities) domains, two intermediate chains, two light intermediate chains, and two copies of three different light chains5. The dynein AAA+ ring is dynamic; ATP binding and hydrolysis in AAA1 regulates dynein’s binding to and movement on microtubules. Opening and closing of dynein’s ring is coupled to movements of dynein’s mechanical element, the ‘linker,’ and rearrangements in dynein’s stalk and buttress are responsible for controlling the affinity of the microtubule-binding domain for microtubules6 (Fig. 1a).
a, Cartoons of the domain organization of dynein and Lis1. Major elements mentioned in the text are labeled. MTBD, microtubule binding domain. b, Schematic of dynein activation. Activation of dynein requires the relief of its autoinhibited conformation (Phi) and the formation of an active complex containing the dynactin complex and an activating adapter. Although Lis1 is known to be involved in this process, how it is involved is unknown. c, Cryo-EM map of the Chi dynein–Lis1 complex (non-symmetry expanded), shown in two orientations. The different Lis1 β-propellers that bind to previously identified sites on dynein are labeled. d, Model of the Chi dynein–Lis1 complex, shown in the same two orientations as in the map in c.
Dynein is proposed to exist largely in an autoinhibited ‘Phi’ conformation in cells, which has been observed in vitro7,8,9 (Fig. 1b). In contrast to Phi dynein, which contains only the dynein subunits, active dynein complexes can be >4 MDa and consist of one or two dynein dimers, the 1.1-MDa dynactin complex, and an activating adapter that mediates the interaction between dynein and dynactin and links dynein to its cargos10,11 (Fig. 1b). Dynein must undergo major conformational changes to transition from the autoinhibited Phi form to these active complexes9,12,13. Some activated dynein complexes contain two dynein dimers, which move faster than complexes containing a single dynein dimer (Fig. 1b)13,14,15,16, and can include a second activating adapter12. Many putative activating adapters have been described, and about a dozen of these have been shown to activate dynein motility in vitro, including members of the Hook, Bicaudal D (BicD), and Ninein families10,11,17. Yeast dynein requires dynactin and the presumed activating adapter Num1 for function in vivo as well18,19,20.
Dynein function in vivo also requires Lis1, which is an essential positive regulator of dynein, as shown by genetic studies in many organisms21,22,23,24,25,26,27. Like dynein, Lis1 is conserved from yeast to mammals. Lis1 is a dimer, with each monomer consisting of an N-terminal small dimerization domain, an alpha helix, and a C-terminal β-propeller following a flexible linker28,29 (Fig. 1a). Lis1 is the only dynein regulator known to bind directly to dynein’s AAA+ motor domain, with two known binding sites: one on dynein’s motor domain between AAA3 and AAA4 (sitering), and the other on dynein’s stalk, a long anti-parallel coiled coil that leads to dynein’s microtubule-binding domain (sitestalk)30,31,32,33. In humans, mutations in LIS1 (PAFAH1B1; GeneID: 5048) or the dynein heavy chain gene (DYNC1H1; GeneID: 1778) cause the neurodevelopmental disease lissencephaly and other malformations of cortical development4,34,35. Recent studies have shown that Lis1 has a role in forming active dynein complexes14,15,31,36. Human dynein–dynactin–activating adapter complexes form more readily in vitro in the presence of Lis1 and move faster on microtubules owing to the recruitment of a second dynein dimer to the complex14,15. In S. cerevisiae and the filamentous fungus Aspergillus nidulans, mutations in dynein that block the formation of the autoinhibited Phi conformation can partially rescue Lis1 deletion or mutations, suggesting that Lis1 may activate dynein by relieving autoinhibition21,36,37. However, the inability of Phi-blocking mutations to fully rescue Lis1 loss-of-function phenotypes suggests that Lis1 may have additional roles in regulating dynein beyond relieving the autoinhibited Phi conformation21,31,36. Despite the data summarized here, how Lis1 relieves dynein autoinhibition remains unknown.
Results
Structure of a Lis1-mediated dynein dimer
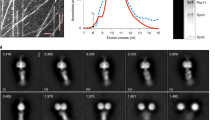
We have previously determined a high-resolution cryo-EM structure of the S. cerevisiae dynein motor domain (carrying an E2488Q substitution in the Walker B motif of its AAA3 domain) in the presence of ATP-vanadate and bound to two yeast Lis1 β-propellers, most likely coming from the same Lis1 dimer31. Although this data set appeared to consist primarily of particles containing one dynein motor domain bound to one Lis1 dimer, we revisited it to look for other structures. To our surprise, additional data processing revealed several two-dimensional (2D) class averages that contained two dynein motor domains, even though the dynein that we used for cryo-EM sample preparation was engineered to be monomeric. Using this subset of particles, we solved a structure of two dynein motor domains in complex with two Lis1 dimers to 4.1 Å. We called this structure ‘Chi’ because the dynein conformation resembles the Greek letter Chi, and because Chi follows Phi in the Greek alphabet (Fig. 1c,d, Supplementary Video 1, Extended Data Fig. 1, and Table 1). In our structure, two dynein motor domains, each bound by a Lis1 dimer, come together through dynein–Lis1 interactions which, to the best of our knowledge, have not been previously reported. This arrangement is consistent with previous studies that have reported the binding of two Lis1 dimers to dynein in solution14,38,39. In our Chi structure, the closed conformation of dynein’s ring and the characteristic bulge in the stalk near its contact site with the buttress indicate a weak microtubule-binding state40. The overall structure of each dynein–Lis1 complex is very similar to the structures that we have solved of individual dynein motor domains bound to a Lis1 dimer, also in a weak microtubule-binding state30,31. Importantly, the Chi conformation is not a consequence of the E2488Q substitution in dynein, or the use of ATP-vanadate in the sample: we also observed Chi in a data set collected from a sample containing wild-type dynein monomer, ATP, and Lis1 (Extended Data Fig. 2 and Table 1). The presence of only a single class average of Chi in that data set prevented us from obtaining a 3D reconstruction.
Dynein’s autoinhibited Phi conformation is characterized by interactions between its two motor domains that make them point in opposite directions (they are effectively ‘cross-legged’) (Figs. 1b and 2a), which prevents the motor from being able to bind to microtubules9. The Phi conformation is incompatible with the binding of Lis1 to dynein, as Lis1 bound to one dynein clashes with the second dynein in Phi (Extended Data Fig. 3a).
a, Interfaces in human dynein involved in stabilizing Phi (PDB: 5NUG)9. b, Interfaces in yeast dynein involved in stabilizing Chi. Note that all but one (stalk–stalk) of the interfaces in Chi also involve Lis1, which we do not show in this panel. The circled labels (3a, 3b, 3c) refer to panels in Figure 3, in which interfaces are shown in detail. c, Overlay of dynein in Phi (gray) and Chi (rainbow). As in panel (b), we omitted Lis1 in the Chi structure for clarity. The left panel shows the two structures superimposed. The right panel shows interatomic vectors linking equivalent alpha carbons in Phi and Chi. The length of the cylinders is proportional to the displacement of that atom between the two structures.
In Chi, the Lis1 proteins bound to the dynein motor domains act as wedges that keep the two dyneins apart, preventing most of the interactions that stabilize Phi (Fig. 2a,b and Extended Data Fig. 3b). The one exception is a contact between dynein’s stalk helices, which acts as a hinge where Phi opens to accommodate Lis1 and transition to Chi (Fig. 2a,b). The dynein–dynein Phi contacts that are disrupted in Chi are replaced by new contacts between Lis1 bound to sitering on one dynein and the AAA5 (Fig. 3a) and AAA6 (Fig. 3b) domains of the opposite dynein. The AAA5 and AAA6 Chi-specific interactions involve residues on Lis1’s β-propeller that are different from those involved in all previously characterized Lis1–dynein and Lis1–Lis1 contacts (Fig. 3d–g and Supplementary Video 1). The Lis1 bound to sitestalk does not interact with the opposite dynein molecule in Chi (Fig. 1c,d).
a–c, Chi is stabilized by three interfaces that, to the best of our knowledge, have not been previously reported: AAA5–Lis1ring (a); AAA6–Lis1ring (b); and a linker–linker interaction (c). In a, Chi is viewed from the dynein stalk and towards the motor domain. Figure 2b shows the location of these interactions in the context of full dynein. All three panels show the cryo-EM map colored by domain, with Lis1 in gray. The insets show the region of Chi that is highlighted in the main panel. d–g, Location of interaction interfaces in the Lis1ring–Lis1stalk dimer. d, Our model of Chi. Dynein is shown in surface representation (filtered to 8 Å), and Lis1 is shown as a ribbon diagram. The area within the square is enlarged in e. e, Residues involved in the four Lis1 interactions (binding of Lis1 to sitering and sitestalk, the Lis1–Lis1 dimer interface, and the interactions stabilizing Chi) are shown as sphered and colored by interface. Dynein is shown in white. f, The Lis1 dimer is viewed facing the dynein monomer, with which the Lis1–ring and Lis1–stalk interactions are formed. The dynein in front was removed for clarity. g, The Lis1 dimer viewed from the other side, facing the dynein monomer with which the Chi-stabilizing interactions are made. The dynein in front was removed for clarity.
The differences between Phi and Chi include changes in the conformation of the linker domain (Extended Data Fig. 4). In Phi dynein, the linker is bent and docked on AAA2 and AAA3 (Extended Data Fig. 4a), which is the same conformation seen in dynein alone in its ‘pre-power stroke’ state6. In contrast, in Chi, the linker is disengaged from AAA2 and AAA3 and shifted towards the opposing dynein molecule (Fig. 3c and Extended Data Fig. 4b,c); this accommodates the increased distance between the motor domains caused by the presence of Lis1 (Fig. 2c). This change in the linker conformation in turn causes a shift in the interface mediating the linker–linker interaction (Fig. 3c and Extended Data Fig. 4d–f); however, the resolution of the linker domain is too low to determine the exact nature of the residues involved.
The major features of the Chi structure—the disruption of most Phi-stabilizing interactions, a shift in the linker–linker interface, and the dependence of these on the presence of Lis1 and new interactions between Lis1 and dynein—suggest that Chi is an early intermediate in the activation of dynein that provides a mechanistic explanation for the role of Lis1 in this process. We set out to test this hypothesis and validate our structural model in vivo using S. cerevisiae and in vitro using recombinant human dynein–dynactin–activating adapter complexes.
Dynein and Lis1 Chi contacts are required in vivo in yeast
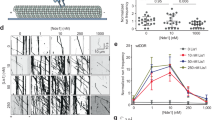
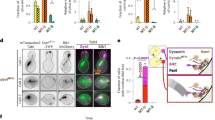
We first sought to determine whether the new contact sites between Lis1 and dynein at AAA5 (Fig. 4a) and AAA6 (Fig. 4b) that form Chi are important for the dynein pathway in vivo. For this, we turned to experiments in S. cerevisiae. In yeast, dynein functions to align the mitotic spindle, such that, upon the completion of mitosis, both the mother and daughter cell inherit a nucleus18,20,41,42,43. Deletion of the gene encoding dynein (DYN1; GeneID: 853928) and dynactin subunits, as well as Lis1, causes nuclear segregation defects that are readily quantifiable because they produce binucleate cells22. To disrupt the Chi-specific interactions (Fig. 4a,b), we mutated Asn213 in Lis1 to Ala (Lis1N213A), Trp288 to Asp (Lis1W288D), or both (Lis1N213A W288D) in the endogenous locus of yeast Lis1 (PAC1; GeneID: 854443). Western blots of strains containing FLAG-tagged versions of the mutants confirmed that Lis1N213A, Lis1W288D, and Lis1N213A W288D were expressed at wild-type levels (Extended Data Fig. 5a,b). We found that, among cells expressing any of these three mutants, the percentage of cells containing two nuclei was increased, similar to cells in which Lis1 was deleted (Fig. 4c). We obtained similar results when we disrupted Chi on dynein’s side. On the basis of our structure, we introduced F3446A in AAA5 and E3867A in AAA6 (Dyn1F3446A E3867A; Fig. 4a,b). Although we did not build the side chain for E3867 in our model owing to the lower resolution of that part of the map, this residue is a good candidate for mediating the interaction between dynein and Lis1-N213. These substitutions in dynein resulted in an increase in the percentage of binucleate cells similar to that observed following deletion of dynein (Fig. 4d). This suggests that the dynein–Lis1 interactions involved in forming Chi are required for Lis1’s regulation of dynein in vivo.
a, The yeast Chi AAA5–Lis1ring interface. Trp288 in Lis1 was mutated to Asp. b, The yeast Chi AAA6-Lis1ring interface. Asn213 was mutated to Ala. We did not build the side chain of E3867 beyond its β-carbon owing to the lower resolution of this part of the map. c, Quantitation (mean ± s.e.m.) of the percentage of cells displaying an aberrant binucleate phenotype for wild type (WT, dark gray), Lis1 deletion (light gray), Lis1N213A (orange), Lis1W288D (red) or Lis1N213A W288D (maroon). WT n = 12, ΔLis1 n = 11, Lis1N213A n = 10, Lis1W288D, and Lis1N213A, W288D n = 8 biological replicates from independent cultures, with at least 200 cells per condition in each replicate. Statistical analysis was done using a one-way ANOVA with Tukey’s multiple comparison test. WT and ΔLis1 ****P < 0.0001, WT and Lis1N213A ****P < 0.0001, WT and Lis1W288D **P = 0.0001, WT and Lis1N213A W288D ****P < 0.0001. Differences not noted are not statistically significant. d, Quantitation (mean ± s.e.m.) of the percentage of cells displaying an aberrant binucleate phenotype for wild type (blue), Dyn1 deletion (light blue), Dyn1F3446A, E3867A (green), and Dyn1F3446A, E3867A, D2868K (dark green). n = 3 biological replicates, with at least 200 cells per condition per replicate. Statistical analysis was done using a one-way ANOVA with Tukey’s multiple comparison test. WT and ΔDyn1 ***P = 0.0002, WT and Dyn1F3446A E3867A *P = 0.01, ΔDyn1 and Dyn1F3446A E3867A D2868K ***P = 0.0002, Dyn1F3446A E3867A and Dyn1F3446A E3867A D2868K *P = 0.015. e, Example images of endogenous dynein localization (DYN1-3XGFP) in dividing yeast cells expressing a fluorescently tagged SPB (SPC110-tdTomato) component and tubulin (TUB1-CFP). Scale bars are 3 μm. f–h, Quantification of dynein localization. Graphs show the number (mean ± s.e.m.) of dynein foci per cell localized to the cortex (f), SPB (g), and microtubule plus ends (h) in WT, Lis1Δ, Lis1N213A, Lis1W288D, and Lis1N213A W288D yeast strains, from three biological replicates. Statistical analysis was performed on the means of each biological replicate using a one-way ANOVA with Tukey’s multiple comparison test. f, WT and ΔLis1 **P = 0.0011, WT and Lis1N213A ***P = 0.0007, WT and Lis1W288D ***P = 0.0002, WT and Lis1N213A W288D ***P = 0.003. g,WT and ΔLis1 **P = 0.0023, WT and Lis1N213A *P = 0.0396, WT and Lis1W288D ***P = 0.0009, WT and Lis1N213A W288D ***P = 0.0003. h, WT and ΔLis1 ***P = 0.0006, WT and Lis1N213A *P = 0.02, WT and Lis1W288D *P = 0.049, WT and Lis1N213A W288D ***P = 0.0039. n.s. (not significant), P = 0.069 to >0.9981. n ≥ 40 cells per biological replicate; >120 cells total per condition. i, Quantitation (mean ± s.e.m.) of the percentage of cells displaying an aberrant binucleate phenotype for wild type (WT, dark gray), Lis1 deletion (light gray), Lis1N213A (orange), Lis1W288D (red), Lis1N213A W288D (maroon), Lis1 WT with the Phi-breaking dynein mutant dyneinD2868K (striped dark gray), Lis1 deletion with dyneinD2868K (striped light gray), Lis1N213A with dyneinD2868K (striped orange), Lis1W288D with dyneinD2868K (striped red), and Lis1N213A W288D with dyneinD2868K (striped maroon). The WT condition is the same as in d. n = 3 biological replicates, with at least 200 cells per condition per replicate. Statistical analysis was done using a one-way ANOVA with Tukey’s multiple comparison test. WT and ΔLis1 ****P < 0.0001, WT and Lis1N213A **P = 0.0035 WT and Lis1W288D *P = 0.045, WT and Lis1N213A W288D **P = 0.0014, ΔLis1 and ΔLis1 with dyneinD2868K **P = 0.0046, Lis1N213A W288D and Lis1N213A W288D with dyneinD2868K **P = 0.0036, Lis1N213A and Lis1N213A with dyneinD2868K *P = 0.0134, Lis1W288D and Lis1W288D with dyneinD2868K *P = 0.0244, Lis1N213A, W288D. Differences not noted are not statistically significant.
Dynein localization in yeast depends on dynein and Lis1 Chi contacts
Next, we wanted to determine how the Lis1–dynein interactions involved in forming Chi contribute to dynein’s localization in yeast cells (Fig. 4e–h). Yeast dynein assembles into fully active complexes with dynactin and Num1, the presumed yeast dynein activating adapter, at the cell cortex18,44. From this site, active dynein pulls on spindle pole body (SPB)-attached microtubules to align the mitotic spindle, which ultimately favors equal segregation of nuclei between mother and daughter cells. Dynein reaches the cortex by first localizing to microtubule plus ends, through either kinesin-dependent transport or cytosolic recruitment45,46,47. Dynein also localizes to the SPB, where microtubule minus ends are found46. Lis1 is required for dynein’s localization to all three sites20,22,48. To assess how the mutants that target the new interactions between dynein and Lis1 in Chi affect dynein localization, we introduced Lis1N213A, Lis1W288D and Lis1N213A W288D at the endogenous Lis1 (PAC1) locus. In these strains, dynein, α-tubulin, and a spindle pole body component were also tagged with a fluorescent protein (DYN1-3XGFP, TUB1-CFP, and SPC110-tdTomato, respectively) (Fig. 4e)22,30 and used to measure the number of dynein foci that were found at the cell cortex (Fig. 4f), SPBs (Fig. 4g), or microtubule plus ends (Fig. 4h) in each cell. All three mutants showed a striking reduction in the number of dynein foci at the cell cortex, similar to the strain with Lis1 deletion (Fig. 4f). Because the cortex is the site of dynein–dynactin–Num1 assembly, these data suggest that the Chi conformation is important for the assembly of active dynein complexes in vivo. All three mutants also significantly affected the ability of dynein to localize to microtubule plus ends. One mechanism dynein uses to reach microtubule plus ends is through kinesin transport in a complex that also contains Lis1 and Bik1/CLIP170 (refs. 20,49,50). Thus, it is possible that this transport complex uses the dynein–Lis1 interactions we identified in Chi, or potentially the Chi conformation itself. Together, our data show that the contacts we observe between dynein and Lis1 in the Chi structure are required for active dynein complex assembly and function in vivo.
Phi-disrupting mutations bypass the need for Chi in yeast
We hypothesize that Chi follows Phi as an intermediate in dynein’s activation pathway. This model predicts that mutations that prevent dynein from forming the autoinhibited Phi conformation should bypass the need for Chi to form. We tested this using genetic epistasis experiments. We generated yeast double mutant strains containing each of our Chi-breaking Lis1 substitutions (Lis1N213A, Lis1W288D, or Lis1N213A,W288D) combined with a Phi-breaking substitution in the endogenous dynein/DYN1 locus (dyneinD2868K)21, and performed nuclear segregation assays. In agreement with our model, introducing dynein with the Phi-breaking substitution into the strains carrying the Chi-breaking Lis1 mutants restored the percentage of cells with two nuclei to wild-type levels (Fig. 4i). We observed the same when dyneinD2868K was introduced into the Lis1-deletion strain (Fig. 4i). Introducing the Phi-breaking substitution into dynein already carrying Chi-disrupting alterations (dyneinF3446A E3867A D2868K) also fully rescued the increase in the percentage of binucleate cells (Fig. 4d).
Dynein and LIS1 Chi contacts are required for human dynein complex assembly
To assess the importance of the Chi conformation, or more generally of the new dynein–Lis1 interaction involved in stabilizing it, in the assembly of active human dynein–dynactin–activating adapter complexes at the molecular level, we next turned to reconstituting the complexes using human proteins. In vitro, human Lis1 protein (herein, LIS1) enhances the formation of dynein–dynactin–activating adapter complexes containing two dynein dimers, which move faster than the single dynein dimer complexes that form in the absence of LIS1 (refs. 14,15). To do this, we first had to identify Chi-disrupting substitutions in human LIS1 that were predicted to be equivalent to the ones we identified in yeast. Directly overlaying our recent structure of a human dynein–LIS1 complex51 onto each motor domain of yeast Chi did not work, as LIS1 bound to sitering is too far away from the opposite dynein to form the Chi-stabilizing interactions with AAA5 and AAA6 (Fig. 5a). This difference is due to human LIS1 having shorter peripheral loops and being slightly rotated relative to yeast Lis1 (ref. 51) when bound to dynein. To circumvent this problem, we modeled human Chi by aligning a copy of our structure of human dynein bound to two LIS1s to each motor domain in Chi dynein, and then manually moving the models so that they were within interacting distance while still maintaining the stalk–stalk interaction observed in both Chi and Phi dynein. Using this model, we made two sets of human LIS1 substitutions: p.N203A, p.D205A, and p.Y225A to disrupt both the LIS1–AAA5 and LIS1–AAA6 interfaces, and p.N203A, p.D205A, and p.D245A to disrupt the LIS1–AAA6 interface alone. Two of the sites we mutated in human LIS1, N203 and D245, are also conserved in yeast; Y225 corresponds to the W288 that we mutated in yeast Lis1 (Extended Data Fig. 6a).
a, A model of the human Chi AAA62 and AAA52–LIS11 interface. The arrow in the inset indicates the area highlighted in the main panel. b, Single-molecule velocity (median ± interquartile range) of TMR–dynein–dynactin–BICD2 complexes in the absence (white circles) or presence (black circles) of different human LIS1 constructs. The data points are represented as triangles, circles, squares, and hexagons corresponding to single measurements within each technical replicate. Four technical replicates were collected for each condition, and the number of data points (n) per each replicate is listed (no LIS1, n = 142, 91, 152, 131; LIS1, n = 147, 118, 119, 120; LIS1N203A D205A Y225A, n = 149, 91, 120, 103; LIS1N203A D205A D245A, n = 147 109 128 108). No LIS1 and LIS1 ***P = 0.0002, LIS1 and LIS1N203A D205A Y225A **P = 0.0051, LIS1 and LIS1N203A D205A D245A **P = 0.004. One-Way ANOVA with Tukey’s multiple comparison test. c, Processive runs (mean ± s.e.m.) of TMR–dynein–dynactin–BICD2 complexes in the absence (white circle) or presence (black circle) of different unlabeled human LIS1 constructs. The data points are represented as triangles, circles, squares, and hexagons corresponding to single measurements within each technical replicate. Four replicates per condition were collected, and the number of data points (n) per each replicate is listed (no LIS1, n = 14, 20, 20, 8; LIS1, n = 7, 16, 7, 5; Lis1N203A D205A Y225A, n = 15, 7, 16, 9; Lis1N203A, D205A D245A, n = 17, 9, 17, 9). No LIS1 and LIS1 *P = 0.023. One-Way ANOVA with Tukey’s multiple comparison test. d, Representative kymographs from single-molecule motility assays with purified TMR–dynein–dynactin–BICD2 in the absence (white circle) or presence (black circle) of different human LIS1 constructs. Scale bars, 10 μm (x) and 40 s (y). e, Schematic of the hypothesis for how Lis1 relieves dynein autoinhibition.
Next, we purified these mutant human LIS1 constructs (Extended Data Fig. 6b) and examined their ability to activate human dynein–dynactin complexes containing the BICD2 activating adapter in single-molecule motility assays. In these assays, activation of motility is read out by observing an increase in dynein complex velocity in the presence of LIS1, which results from the enhanced formation of dynein complexes containing two dynein dimers13,14,15. As we have shown previously, pre-incubation of dynein–dynactin–BICD2 with 300 nM wild-type human LIS1 increased the velocity and number of processive runs of these complexes (Fig. 5b–d)15. By contrast, there was no significant difference between dynein velocity and the number of processive runs when dynein–dynactin–BICD2 complexes were pre-incubated with the two human LIS1 mutants (LIS1N203A D205A Y225A and LIS1N203A D205A D245A), as compared with complexes that were formed in the absence of LIS1 (Fig. 5b–d). We also observed an increase in the number of diffusive events in the presence of LIS1 mutants (Extended Data Fig. 6c). These data indicate that the dynein–Lis1 contact sites found in the Chi structure are important for human LIS1’s role in forming the activated human dynein–dynactin–activating adapter complex, either by relieving the autoinhibited Phi conformation or by playing a role in a later step in the assembly of activated complexes, or a combination of both.
Discussion
On the basis of the data presented here, we propose that Chi is an intermediate state in the dynein activation pathway, providing the first structural and mechanistic explanation for how Lis1 relieves dynein autoinhibition. We propose that the wedging of Lis1 between the two dynein motor domains primes dynein for binding dynactin and activating adapter protein(s).
The Chi conformation has important implications for dynein activation. In dynein’s autoinhibited Phi state, the ‘tails,’ which precede the motor domain, make many contacts with each other until they reach their ‘neck’ region, where they move away from each other before rejoining again at the linker domain. Switching to the active dynactin-bound form requires the tail to undergo a large conformational change that turns the loose twofold symmetry present in Phi into the translational (‘parallel’) symmetry dynein adopts when bound to dynein and an activating adapter (Fig. 5e). This rearrangement involves breaking the linker–linker contacts, as well as many of the interactions between the tails and associated chains. We hypothesize that the Chi conformation primes dynein for binding dynactin and an activating adapter protein by stabilizing an intermediate conformation of the tail. In Chi, binding of Lis1 forces the linkers apart, which would cause some of these interactions to break. This would pull the dynein heavy chains apart from each other, disrupting the neck and potentially transmitting conformational changes all the way up the tail.
Methods
Electron microscopy sample preparation
Grids were prepared and imaged as previously described31. Briefly, monomeric yeast dyneinE2488Q was randomly biotinylated using water-soluble Sulfo ChromaLink biotin and then dialyzed into TEV buffer. Grid samples containing 150 nM dyneinE2488Q, 650 nM Lis1, 1.2 mM ATP, and 1.2 mM Na3VO4 were applied to streptavidin affinity grids and vitrified52,53.
For the imaging of wild-type yeast dynein monomer in the presence of ATP and Lis1, purified fresh dynein (not frozen) was incubated with apyrase for 30 min (apyrase, 0.1 U mL–1, NEB) prior to gel filtration on a Superose 6 Increase column pre-equilibrated in buffer containing 50 mM Tris, pH 8, 150 mM KCl, 2 mM EDTA, and 1 mM DTT. Eluted protein was concentrated to about 5 µM and diluted 1:1 with Lis1 or TEV buffer and incubated on ice for 10 min (final concentrations: 2.5 µM dynein, 2.5 µM Lis1). After an additional 30-s incubation on ice in the presence of ATP (final concentration: 1.25 mM ATP) samples were applied to plasma-cleaned (Solarus, Gatan) UltrAuFoil Holey Gold R 1.2/1. µM3 grids (Quantifoil). A vitrobot (FEI) was used to blot excess sample and plunge freeze the grids in liquid ethane. Grids were stored in liquid nitrogen until they were ready to be imaged.
Electron microscopy image collection and processing
Details on image collection and initial processing have been reported elsewhere31. Following particle picking in crYOLO5 (ref. 54), particles were extracted and binned to 3.93 Å pixel–1 in Relion 3.0 (ref. 55). Multiple rounds of 2D classification were carried out first in Relion 3.0 and subsequently in cryoSPARC56 to remove bad particles. Some class averages showed particles with more than one dynein molecule. Using particles from all good 2D classes, initial ab initio reconstructions failed to reconstruct a volume containing more than one dynein molecule. Particles belonging to 2D class averages that potentially contained more than one dynein molecule were manually selected and used in an ab initio reconstruction that resulted in an initial map for Chi dynein bound to Lis1. This volume was used together with our previously published map of monomeric dyneinE2488Q bound to Lis1 (EMDB-23829) to separate monomeric dyneinE2488Q from Chi dynein using heterogeneous refinement. We performed one additional round of heterogeneous refinement to further remove any monomeric dynein molecules. Using our final particles, we ran non-uniform refinement with C2 symmetry and optimized per-group CTF parameters enabled to give us a 4.1-Å map.
Wild-type yeast dynein monomer incubated in the presence of ATP and Lis1 was imaged using a Talos Arctica operated at 200 kV and equipped with a K2 Summit direct electron detector (Gatan). Automated data collection was performed using Leginon. A total of 4,525 movies was collected at ×36,000 magnification (1.16 Å pixel–1). The dose was ~4.63 e– Å–2 s–1 with a total exposure time of 11 s divided into 200 ms frames, for a total of 40 frames. The defocus range was set to 0.8–2.4 µm. All movies were aligned in CryoSPARC live57 using MotionCor2 (ref. 58) of the dose-weighted frames. CTF was estimated in dose-weighted images using CTFFIND4 (ref. 59). Images with CTF fits worse than 5 Å were excluded from further processing. Particles were initially selected using blob finder in CryoSPARC, and these peaks were used for Topaz60 model training and final particle picking using Topaz. Particles were extracted and binned to 4.64 Å pixel–1. Multiple rounds of 2D classification, with a varying number of online-EM interactions (30–40) and a batch size per class of between 200 and 400 dependent on the number of particles in each classification were carried out in cryoSPARC to remove bad particles.
Model building and refinement
Symmetry expansion followed by local refinement was used to improve the overall resolution of the motor domain to 3.6 Å. The yeast dyneinE2448Q–(Lis1)2 model (PDB 7MGM) was docked, and rigid body fit into the map using Phenix real space refine61. Discrepancies between the model and map were fixed manually in COOT6 (ref. 62) and then refined using a combination of Phenix real space refine and Rosetta Relax (v.13)63. Once the monomer model was finished, it was placed in the original map and refined using a similar strategy.
The resolution of the Chi map did not allow us to distinguish between ATP and ADP.Vi. Therefore, we chose to model ATP at AAA1, AAA2, and AAA3, to be consistent with our previously published structure of the yeast dynein motor domain bound to two Lis1s, which was obtained from the same data set31.
Cloning, plasmid construction, and mutagenesis
The pDyn1 plasmid (pACEBac1 expression vector containing insect cell codon-optimized dynein heavy chain (DYNC1H1) fused to a His-ZZ-TEV tag on the amino terminus and a carboxy terminal SNAPf tag (New England Biolabs) and the pDyn2 plasmid (the pIDC expression vector with codon-optimized DYNC1I2, DYNC1LI2, DYNLT1, DYNLL1, and DYNLRB1) were a gift from A. Carter (LMB-MRC). The pDyn1and pDyn2 plasmids were recombined in vitro with a Cre recombinase (New England Biolabs) to generate the pDyn3 plasmid. The presence of all six dynein chains was verified by PCR. The pFastBac plasmid with codon-optimized human full-length Lis1 (PAFAH1B1) fused to an N-terminal His-ZZ-TEV tag was a gift from A. Carter (LMB-MRC). The BICD2s construct (amino acids 25–398) fused to sfGFP on the N terminus and inserted into a pET28a expression vector was obtained as described previously17. Substitutions in LIS1 were made using a multi-site-directed mutagenesis kit (Agilent) and the following primers: 5′-TGTGAATGTCTTCACACAGGCGCCAGTTTGCACTTCCCAC-3′, 5′-GGTACGGCCAAATCAAGCTGGCACTCTGATAGCCAG-3′, and 5′-CAGTAGCCATCATGCCCGCTGGAGCTCATATAGTGTCTGCCTC-3′. The yeast gene encoding Lis1 (PAC1) previously cloned into a Topo 2.1 vector using the TOPO TA Cloning Kit (Thermo Fisher), was mutated using a site-directed mutagenesis kit (Agilent) with the primers: 5′-GGATGTTTTATTTACCAATTATACGGCCTCCAGCAAGAAGAACTATTTGGTG-3′ and 5′-cACCAAATAGTTCTTCTTGCTGGAGGCCGTATAATTGGTAAATAAAACATCC-3′ to insert the N213A substitution. PAC1 or pac11-N213A were mutated using the primers 5′-TTTGGGACTTCCACAATGGTGACTCGTTGAAAACATTTCAGCC-3′ and 5′-GGCTGAAATGTTTTCAACGAGTCACCATTGTGGAAGTCCCAAA-3′ to insert the W288D substitution. DYN1(3396–3921aa) was cloned into a pBlueHeron vector through Gibson assembly and was mutagenized with the primers 5′-GATTAGAAAATGCAATTAGAGCCGGAAGTGTAGTTATAATTC-3′ and 5′-GAATTATAACTACACTTCCGGCTCTAATTGCATTTTCTAATC-3′ to introduce the F3446A substitution. dyn1-F3446A was mutated using a site-directed mutagenesis kit (Agilent) with the primers 5′-GAGGAGACAAAGGCGGCAGAAGCACATGAGAAATTCAAAATGT-3′ and 5′-ACATTTTGAATTTCTCATGTGCTTCTGCCGCCTTTGTCTCCTC-3′ to introduce the E3867A substitution.
Yeast strains
The S. cerevisiae strains used in this study are listed in Supplementary Table 1. The endogenous genomic copy of PAC1 (encoding Lis1) was deleted using PCR-based methods as previously described64. In brief, K. lactis URA3 with homology arms complementary to regions upstream and downstream of the PAC1 or DYN1 genomic locus was generated using PCR. This fragment was transformed into a strain with the preferred genetic background using the lithium acetate method65 and screened by colony PCR. Point mutants were generated using QuikChange site-directed mutagenesis (Agilent) and verified by DNA sequencing. Mutated fragments were re-inserted into the klURA3 strains to reintroduce the mutated PAC1 or DYN1 gene. Positive clones lacking URA3 (klURA3-) were selected in the presence of 5-fluoroorotic acid, screened by colony PCR, and verified by DNA sequencing.
Nuclear segregation assay
Single colonies were picked and grown at 30 °C. Log-phase S. cerevisiae cells growing at 30 °C were transferred to 16 °C for 16 h. Cells were fixed with 75% ethanol for 1 h, sonicated for 5 s at 40% amplitude, and mounted in medium containing DAPI. Imaging was performed using an Apo TIRF 100 ×1.49 NA objective (Nikon, Plano Apo) on a Nikon Ti2 microscope with a Yokogawa-X1 spinning disk confocal system, MLC400B laser engine (Agilent), Prime 95B back-thinned sCMOS camera (Teledyne Photometrics), and a piezo Z-stage (Mad City Labs). The samples were blinded during imaging. The percentage of aberrant binucleate cells was calculated as the number of binucleate cells divided by the sum of wild-type and binucleate cells. Eight to 12 biological replicates from independent colonies were done for Figure 4c, and three were done for Figure 4d,i. At least 200 cells were counted for each biological replicate per condition.
Live-cell imaging
Single colonies were picked and grown at 30 °C. Log-phase live S. cerevisiae cells were mounted on a thin agarose pad made from SC medium pressed between two glass slides. Live cells genetically modified to express fluorescently labeled DYN1-3XGFP, CFP-TUB1, and SPC110-tdTomato were imaged using a Yokogawa W1 confocal scanhead mounted to a Nikon Ti2 microscope with an Apo TIRF 100 ×1.49 NA objective (Nikon, Plano Apo). The microscope was run with NIS Elements using the 488 nm 515 nm and 561 nm lines of a six-line (405 nm, 445 nm, 488 nm, 515 nm, 561 nm, and 640 nm) LUN-F-XL laser engine and Prime95B cameras (Photometrics). The DYN1-3×GFP foci localizing to the spindle pole body (SPB), microtubule plus end, and cell cortex were outlined as regions of interest in Fiji66, recorded, and analyzed for three replicates of at least 120 cells for each sample.
S. cerevisiae immunoprecipitations and western blots
Log-phase S. cerevisiae cells grown at 30 °C were pelleted at 4,000g, resuspended in water, and flash frozen in liquid nitrogen. Liquid-nitrogen-frozen yeast cell pellets were lysed by grinding in a chilled coffee grinder, resuspended in dynein-lysis buffer (30 mM HEPES pH 7.4, 50 mM potassium acetate, 2 mM magnesium acetate, 1 mM EGTA, 10% glycerol, 1 mM DTT) supplemented with 1 mM Pefabloc, 0.2% Triton X-100, cOmplete EDTA-free protease inhibitor cocktail tablet (Roche), and 1 mM Pepstatin A (Cayman Chemical Company), and spun at 50,000g for 1 h. The protein concentration of the clarified supernatants was quantified using a Bradford Protein Assay (Bio-Rad), and equal amounts of clarified lysates were incubated with anti-FLAG M2 Affinity Gel (Sigma) overnight at 4 °C. Beads were washed with dynein-lysis buffer, boiled in SDS sample buffer, and loaded onto a NuPAGE Bis-Tris gel (Invitrogen). Gels were transferred to a PVDF membrane that was blocked with PBS-T (PBS1X and 0.1% Tween-20) containing 5% milk and 1% BSA for 1 h at room temperature and blotted with a rabbit anti-FLAG antibody (1:3,000; Proteintech 20543-1-AP) overnight at 4 °C. Membranes were then incubated with a goat-anti-rabbit IRDye 6800RD secondary antibody (LI-COR) and were scanned in a ChemiDoc Imaging system (Bio-Rad).
Protein expression and purification
S. cerevisiae dynein
Protein purification steps were done at 4 °C unless otherwise indicated. S. cerevisiae dynein constructs were purified from S. cerevisiae using a ZZ tag as previously described67. Briefly, liquid-nitrogen-frozen yeast cell pellets were lysed by grinding in a chilled coffee grinder and resuspended in dynein-lysis buffer supplemented with 0.1 mM Mg-ATP, 0.5 mM Pefabloc, 0.05% Triton X-100, and cOmplete EDTA-free protease inhibitor cocktail tablet (Roche). The lysate was clarified by centrifuging at 264,900g for 1 h. The clarified supernatant was incubated with IgG Sepharose beads (GE Healthcare Life Sciences) for 1 h. The beads were transferred to a gravity flow column, washed with dynein-lysis buffer supplemented with 250 mM potassium chloride, 0.1 mM Mg-ATP, 0.5 mM Pefabloc, and 0.1% Triton X-100, and with TEV buffer (10 mM Tris-HCl pH 8.0, 150 mM potassium chloride, 10% glycerol, 1 mM DTT, and 0.1 mM Mg-ATP). Dynein was cleaved from IgG beads via incubation with 0.15 mg mL–1 TEV protease (purified in the Reck-Peterson lab) overnight at 4 °C. Cleaved dynein was concentrated using 100 kDa MWCO concentrator (EMD Millipore), filtered by centrifugation with Ultrafree-MC VV filter (EMD Millipore) in a tabletop centrifuge, and flash frozen in liquid nitrogen.
S. cerevisiae Lis1
S. cerevisiae Lis1 was purified from S. cerevisiae using 8×His and ZZ tags as previously described32. In brief, liquid-nitrogen-frozen pellets were ground in a pre-chilled coffee grinder, resuspended in buffer A (50 mM potassium phosphate pH 8.0, 150 mM potassium acetate, 150 mM sodium chloride, 2 mM magnesium acetate, 5 mM β-mercaptoethanol, 10% glycerol, 0.2% Triton X-100, 0.5 mM Pefabloc) supplemented with 10 mM imidazole (pH 8.0) and cOmplete EDTA-free protease inhibitor cocktail tablet, and spun at 118,300g for 1 h. The clarified supernatant was incubated with Ni-NTA agarose (QIAGEN) for 1 h. The Ni beads were transferred to a gravity column, washed with buffer A + 20 mM imidazole (pH 8.0), and eluted with buffer A + 250 mM imidazole (pH 8.0). The eluted protein was incubated with IgG Sepharose beads for 1 h. IgG beads were transferred to a gravity flow column and washed with buffer A + 20 mM imidazole (pH 8.0) and with modified TEV buffer (50 mM Tris-HCl pH 8.0, 150 mM potassium acetate, 2 mM magnesium acetate, 1 mM EGTA, 10% glycerol, 1 mM DTT). Lis1 was cleaved from the IgG beads by the addition of 0.15 mg mL–1 TEV protease (purified in the Reck-Peterson lab) for 1 h at 16 °C. Cleaved proteins were filtered by centrifuging with Ultrafree-MC VV filter (EMD Millipore) in a tabletop centrifuge and flash frozen in liquid nitrogen.
Human dynein
Full-length human SNAP-tagged dynein was expressed in Sf9 cells, as described previously11,15. Briefly, frozen Sf9 cell pellets from 2× 600 mL culture were resuspended in 80 mL of dynein-lysis buffer with 0.1 mM Mg-ATP, 0.5 mM Pefabloc, 0.05% Triton X-100, and cOmplete EDTA-free protease inhibitor cocktail tablet (Roche) and lysed using a Dounce homogenizer (10 strokes with a loose plunger and 15 strokes with a tight plunger). The lysate was clarified by centrifuging at 183,960g for 88 min in a Type 70 Ti rotor (Beckman). The clarified supernatant was incubated with 4 mL of IgG Sepharose 6 Fast Flow beads (GE Healthcare Life Sciences) for 3–4 h on a roller. The beads were transferred to a gravity flow column, washed with 200 mL of dynein-lysis buffer and 300 mL of TEV buffer (50 mM Tris-HCl pH 8.0, 250 mM potassium acetate, 2 mM magnesium acetate, 1 mM EGTA, 1 mM DTT, 0.1 mM Mg-ATP, 10% (vol/vol) glycerol). For fluorescent labeling of the C-terminal SNAPf tag, dynein-coated beads were labeled with 5 µM SNAP-Cell-TMR (New England Biolabs) in the column for 10 min at room temperature, and unbound dye was removed with a 300 mL wash with TEV buffer at 4 °C. The beads were then resuspended and incubated in 15 mL of TEV buffer supplemented with 0.5 mM Pefabloc and 0.2 mg mL–1 TEV protease (purified in the Reck-Peterson lab) overnight on a roller. The supernatant containing cleaved protein was concentrated using a 100 kDa MWCO concentrator (EMD Millipore) to 500 µL and purified via size-exclusion chromatography on a TSKgel G4000SWXL column (TOSOH Bioscience) with GF150 buffer (25 mM HEPES pH 7.4, 150 mM potassium chloride, 1 mM magnesium chloride, 5 mM DTT, 0.1 mM Mg-ATP) at 1 mL min–1. The peak fractions were collected, buffer-exchanged into a GF150 buffer supplemented with 10% glycerol, concentrated to 0.1–0.5 mg mL–1 using a 100 kDa MWCO concentrator (EMD Millipore). Purity was evaluated on SDS–PAGE gels, and protein aliquots were snap frozen in liquid N2 and stored at –80 °C.
Human dynactin
Dynactin (p62-HaloTag-3×FLAG) was purified from a stable 293T cell line, as previously described11,15,17. Briefly, frozen pellets from 293T cells (160 × 15-cm plates) were resuspended in dynein-lysis buffer supplemented with 0.1 mM Mg-ATP, 0.5 mM Pefabloc, 0.05% Triton, and cOmplete EDTA-free protease inhibitor cocktail tablet (Roche), and gently mixed at 4 °C for 15 min. The lysed cells were then centrifuged at 500,000g in a Ti70 rotor (Beckman) at 4 °C for 30 min. The clarified lysate was retrieved and added to 3 mL packed anti-FLAG M2 agarose resin (Sigma) and incubated with gentle mixing at 4 °C for 16 h. After incubation, the lysate-resin mixture was centrifuged at 1,000g for 2 min at 4 °C to pellet the resin, and the supernatant was decanted. The resin was transferred to a column at 4 °C, and the column was washed with 100 mL low-salt wash buffer (30 mM HEPES, pH 7.4; 50 mM potassium acetate; 2 mM magnesium acetate; 1 mM EGTA, pH 7.5; 10% glycerol; 1 mM DTT; 0.5 mM ATP; 0.5 mM Pefabloc; 0.02% Triton X-100), 100 mL high-salt wash buffer (30 mM HEPES, pH 7.4; 250 mM potassium acetate; 2 mM magnesium acetate; 1 mM EGTA, pH 7.5; 10% glycerol; 1 mM DTT; 0.5 mM ATP; 0.5 mM Pefabloc; 0.02% Triton X-100), and finally with 50 mL of low-salt wash buffer. The resin was resuspended in 800 µL of low-salt wash buffer containing 2 mg mL–1 3×-FLAG peptide (ApexBio) and incubated for 30 min at 4 °C. The mixture was retrieved and centrifuged through a small filter column to remove the resin. The eluate was then loaded onto a Mono Q 5/50 GL 1 mL column on an AKTA FPLC (GE Healthcare). The column was washed with 5 mL of Buffer A (50 mM Tris-HCl, pH 8.0; 2 mM magnesium acetate; 1 mM EGTA; 1 mM DTT) and then subjected to a 26 mL linear gradient from 35–100% Buffer B mixed with Buffer A (Buffer B = 50 mM Tris-HCl, pH 8.0; 1 M potassium acetate; 2 mM magnesium acetate; 1 mM EGTA; 1 mM DTT), followed by an additional 5 mL 100% Buffer B. Fractions containing pure dynactin (~75–80% Buffer B) were pooled and buffer-exchanged through iterative rounds of dilution and concentration on a 100 kDa MWCO centrifugal filter (Amicon Ultra, Millipore) using GF150 buffer with 10% glycerol. Purity was evaluated on SDS–PAGE gels and protein aliquots were snap frozen in liquid N2 and stored at –80 °C.
Human LIS1
LIS1 constructs were purified from frozen sf9 cell pellets from a 600 mL culture, as described previously68. Lysis and clarification steps were similar to full-length dynein purification except lysis was performed in LIS1-lysis buffer (30 mM HEPES pH 7.4, 50 mM potassium acetate, 2 mM magnesium acetate, 1 mM EGTA, 300 mM potassium chloride, 1 mM DTT, 0.5 mM Pefabloc, 10% (vol/vol) glycerol) supplemented with cOmplete EDTA-free protease inhibitor cocktail tablet (Roche) per 50 mL was used. The clarified supernatant was incubated with 0.5 mL of IgG Sepharose 6 Fast Flow beads (GE Healthcare Life Sciences) for 2-3 hours on a roller. The beads were transferred to a gravity flow column, washed with 20 mL of LIS1-lysis buffer, 100 mL of modified TEV buffer (10 mM Tris-HCl pH 8.0, 2 mM magnesium acetate, 150 mM potassium acetate, 1 mM EGTA, 1 mM DTT, 10% (vol/vol) glycerol) supplemented with 100 mM potassium acetate, and 50 mL of modified TEV buffer. LIS1 was cleaved from IgG beads via incubation with 0.2 mg mL–1 TEV protease overnight on a roller. The cleaved LIS1 was filtered by centrifuging with an Ultrafree-MC VV filter (EMD Millipore) in a tabletop centrifuge. Purity was evaluated on SDS–PAGE gels, and protein aliquots were snap frozen in liquid N2 and stored at –80 °C.
Human BICD2
BICD2 construct containing N-terminal sfGFP was expressed and purified as previously described17. In brief, BL-21[DE3] cells (New England Biolabs) were grown at optical density at 600 nm of 0.4–0.6, and protein expression was induced with 0.1 mM IPTG for 16 h at 18 °C. Frozen cell pellet from a 2 L culture was resuspended in 60 mL of lysis buffer (30 mM HEPES pH7.4, 50 mM potassium acetate, 2 mM magnesium acetate, 1 mM EGTA, 1 mM DTT and 0.5 mM Pefabloc, 10% (vol/vol) glycerol) supplemented with cOmplete EDTA-free protease inhibitor cocktail tablet (Roche) per 50 mL and 1 mg mL–1 lysozyme. The resuspension was incubated on ice for 30 min and lysed by sonication. The lysate was clarified by centrifuging at 500,000g for 30 min in Type 70 Ti rotor (Beckman). The clarified supernatant was incubated with 2 mL of IgG Sepharose 6 Fast Flow beads (GE Healthcare Life Sciences) for 2 h on a roller. The beads were transferred into a gravity flow column, washed with 100 mL of activating-adapter-lysis buffer supplemented with 150 mM potassium acetate and 50 mL of cleavage buffer (50 mM Tris-HCl pH 8.0, 150 mM potassium acetate, 2 mM magnesium acetate, 1 mM EGTA, 1 mM DTT, 0.5 mM Pefabloc, and 10% (vol/vol) glycerol). The beads were then resuspended and incubated in 15 mL of cleavage buffer supplemented with 0.2 mg mL–1 TEV protease overnight on a roller. The supernatant containing cleaved protein was concentrated using a 50 kDa MWCO concentrator (EMD Millipore) to 1 mL, filtered by centrifuging with Ultrafree-MC VV filter (EMD Millipore) in a tabletop centrifuge, diluted to 2 mL in buffer A (30 mM HEPES pH 7.4, 50 mM potassium acetate, 2 mM magnesium acetate, 1 mM EGTA, 10% (vol/vol) glycerol, and 1 mM DTT), and injected into a MonoQ 5/50 GL column (GE Healthcare and Life Sciences) at 1 mL min–1. The column was prewashed with 10 CV of buffer A, 10 CV of buffer B (30 mM HEPES pH7.4, 1 M potassium acetate, 2 mM magnesium acetate, 1 mM EGTA, 10% (vol/vol) glycerol, and 1 mM DTT) and again with 10 CV of buffer A at 1 mL min–1. For elution, a linear gradient was run over 26 CV from 0–100% buffer B. The peak fractions containing sfGFP-BICD2s were collected and concentrated using a 50 kDa MWCO concentrator (EMD Millipore) to 0.2 mL. Protein was then diluted to 0.5 mL in GF150 buffer and further purified using size-exclusion chromatography on a Superose 6 Increase 10/300GL column (GE Healthcare and Life Sciences) with GF150 buffer at 0.5 mL min–1. The peak fractions were collected, buffer-exchanged into a GF150 buffer supplemented with 10% glycerol, concentrated to 0.2–1 mg mL–1 using a 50 kDa MWCO concentrator (EMD Millipore), and flash frozen in liquid nitrogen.
TIRF microscopy
Imaging was performed with an inverted microscope (Nikon, Ti-E Eclipse) equipped with a 100×1.49 N.A. oil immersion objective (Nikon, Plano Apo). The xy position of the stage was controlled by ProScan linear motor stage controller (Prior). The microscope was equipped with an MLC400B laser launch (Agilent) equipped with 405 nm (30 mW), 488 nm (90 mW), 561 nm (90 mW), and 640 nm (170 mW) laser lines. The excitation and emission paths were filtered using appropriate single bandpass filter cubes (Chroma). The emitted signals were detected with an electron multiplying CCD camera (Andor Technology, iXon Ultra 888). Illumination and image acquisition was controlled by NIS Elements Advanced Research software (Nikon).
Single-molecule motility assays
Single-molecule motility assays were performed in flow chambers using the TIRF microscopy setup described above. To reduce non-specific binding biotinylated and PEGylated coverslips (Microsurfaces) with microtubules polymerized from tubulin prepared from bovine brain, as previously described17. Microtubules contained ~10% biotin-tubulin to allow for attachment to streptavidin-coated coverslip and ~10% Alexa Fluor 488 (Thermo Fisher Scientific) tubulin for visualization. Imaging was done in dynein-lysis buffer supplemented with 20 µM taxol, 1 mg mL–1 casein, 5 mM Mg-ATP, 71.5 mM β-mercaptoethanol, and an oxygen scavenger system containing 0.4% glucose, 45 μg mL–1 glucose catalase (Sigma-Aldrich), and 1.15 mg mL–1 glucose oxidase (Sigma-Aldrich). Images were recorded every 0.3 s for 3 min. Movies showing significant drift were not analyzed.
All movies were collected by measuring TMR–dynein signal with the following protein concentrations: 83.5 pM TMR-dynein, 665 pM unlabeled dynactin, 5 nM BICD2, and 300 nM LIS1. For conditions missing LIS1, the corresponding matching buffer was used. The dynein, dynactin, and BICD2 complexes were incubated on ice for 10 min prior to LIS1 or buffer addition. Each protein mixture was then incubated on ice for an additional 10 min prior to TIRF imaging.
TIRF motility data analysis
The velocity of moving particles was calculated from kymographs generated in Fiji as described previously47. Velocities were calculated only from molecules that moved processively (continuously moving along a microtubule track) for more than five frames. Non-motile or diffusive events were not considered in velocity calculations. Processive events were defined as events that move unidirectionally and do not exhibit directional changes greater than 600 nm. Diffusive events were defined as events that exhibit at least one bidirectional movement greater than 600 nm in each direction. Static events were defined as events that do not exhibit movement (less than 600 nm in each direction). Single-molecule movements that change apparent behavior (for example, shift from non-motile to processive) were considered multi-velocity events and counted as multiple events. Processive runs were calculated by counting the number of processive events for each microtubule in individual movies and dividing this number by the microtubule length. The percentage of diffusive events was calculated as diffusive events / (diffusive + stationary + processive) × 100.
Statistical analysis
Brightness and contrast were adjusted in Fiji for all images, videos, and kymographs. All statistical tests were generated using GraphPad Prism 9. The exact value of n, evaluation of statistical significance, P values, and specific statistical analysis are described in the corresponding figures and figure legends. All TIRF experiments were analyzed from four independent replicates, and individual analysis of each replicate showed similar results. For velocity analysis, frequency distributions were first calculated for each replicate, and data were fit to a Gaussian distribution to calculate mean values. Nuclear segregation assays in Figure 4c included 8–12 biological replicates from independent cultures per condition. Statistics were generated for all biological replicates (8–12) within each condition. Nuclear segregation assay analysis for Figure 4c,i included three biological replicates from independent cultures for each condition, and statistics were generated for the three biological replicates within each condition using a one-way ANOVA with Tukey’s multiple comparisons test of each mean. Live-cell imaging experiments to assess dynein localization include three biological replicates from three independent cultures. Each replicate included >40 mid-stage mitotic cells with an n of at least 120 cells per condition. Each cell was assessed for the number of dynein foci at the cortex, spindle pole body or microtubule tips. Statistical analysis was performed on the means of each biological replicate using a one-way ANOVA with Tukey’s multiple comparison test.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Cryo-EM maps and molecular models have been deposited in the EM Data Bank and Protein Data Bank, respectively. Accession codes are listed here and in Table 1. DyneinE2488Q bound to Lis1 in Chi conformation: EMD-27810 and PDB 8DZZ; dyneinE2488Q bound to Lis1 in Chi conformation, symmetry expanded: EMD-27811 and PDB 8E00. All other data will be made available upon request. Source data are provided with this paper.
References
Reck-Peterson, S. L., Redwine, W. B., Vale, R. D. & Carter, A. P. The cytoplasmic dynein transport machinery and its many cargoes. Nat. Rev. Mol. Cell Biol. 19, 382–398 (2018).
Wickstead, B. & Gull, K. Dyneins across eukaryotes: a comparative genomic analysis. Traffic 8, 1708–1721 (2007).
Harada, A. et al. Golgi vesiculation and lysosome dispersion in cells lacking cytoplasmic dynein. J. Cell Biol. 141, 51–59 (1998).
Lipka, J., Kuijpers, M., Jaworski, J. & Hoogenraad, C. C. Mutations in cytoplasmic dynein and its regulators cause malformations of cortical development and neurodegenerative diseases. Biochem. Soc. Trans. 41, 1605–1612 (2013).
Canty, J. T. & Yildiz, A. Activation and regulation of cytoplasmic dynein. Trends Biochem. Sci. 45, 440–453 (2020).
Cianfrocco, M. A., DeSantis, M. E., Leschziner, A. E. & Reck-Peterson, S. L. Mechanism and regulation of cytoplasmic dynein. Annu. Rev. Cell Dev. Biol. 31, 83–108 (2015).
Amos, L. A. Brain dynein crossbridges microtubules into bundles. J. Cell Sci. 93, 19–28 (1989).
Torisawa, T. et al. Autoinhibition and cooperative activation mechanisms of cytoplasmic dynein. Nat. Cell Biol. 16, 1118–1124 (2014).
Zhang, K. et al. Cryo-EM reveals how human cytoplasmic dynein is auto-inhibited and activated. Cell 169, 1303–1314 (2017).
McKenney, R. J., Huynh, W., Tanenbaum, M. E., Bhabha, G. & Vale, R. D. Activation of cytoplasmic dynein motility by dynactin-cargo adapter complexes. Science 345, 337 (2014).
Schlager, M. A., Hoang, H. T., Urnavicius, L., Bullock, S. L. & Carter, A. P. In vitro reconstitution of a highly processive recombinant human dynein complex. EMBO J. 33, 1855–1868 (2014).
Chaaban, S. & Carter, A. P. Structure of dynein–dynactin on microtubules shows tandem adaptor binding. Nature 610, 212–216 (2022).
Urnavicius, L. et al. Cryo-EM shows how dynactin recruits two dyneins for faster movement. Nature 554, 202–206 (2018).
Elshenawy, M. M. et al. Lis1 activates dynein motility by modulating its pairing with dynactin. Nat. Cell Biol. 22, 570–578 (2020).
Htet, Z. M. et al. LIS1 promotes the formation of activated cytoplasmic dynein-1 complexes. Nat. Cell Biol. 22, 518–525 (2020).
Grotjahn, D. A. et al. Cryo-electron tomography reveals that dynactin recruits a team of dyneins for processive motility. Nat. Struct. Mol. Biol. 25, 203–207 (2018).
Redwine, W. B. et al. The human cytoplasmic dynein interactome reveals novel activators of motility. eLife 6, e28257 (2017).
Heil-Chapdelaine, R. A., Oberle, J. R. & Cooper, J. A. The cortical protein Num1p is essential for dynein-dependent interactions of microtubules with the cortex. J. Cell Biol. 151, 1337–1344 (2000).
Kormanec, J., Schaaff-Gerstenschläger, I., Zimmermann, F. K., Perecko, D. & Küntzel, H. Nuclear migration in Saccharomyces cerevisiae is controlled by the highly repetitive 313 kDa NUM1 protein. Mol. Gen. Genet 230, 277–287 (1991).
Sheeman, B. et al. Determinants of S. cerevisiae dynein localization and activation: implications for the mechanism of spindle positioning. Curr. Biol. 13, 364–372 (2003).
Marzo, M. G., Griswold, J. M. & Markus, S. M. Pac1/LIS1 stabilizes an uninhibited conformation of dynein to coordinate its localization and activity. Nat. Cell Biol. 22, 559–569 (2020).
Lee, W.-L., Oberle, J. R. & Cooper, J. A. The role of the lissencephaly protein Pac1 during nuclear migration in budding yeast. J. Cell Biol. 160, 355–364 (2003).
Liu, Z., Xie, T. & Steward, R. Lis1, the Drosophila homolog of a human lissencephaly disease gene, is required for germline cell division and oocyte differentiation. Development 126, 4477–4488 (1999).
Markus, S. M., Punch, J. J. & Lee, W.-L. Motor- and tail-dependent targeting of dynein to microtubule plus ends and the cell cortex. Curr. Biol. 19, 196–205 (2009).
Willins, D. A., Liu, B., Xiang, X. & Morris, N. R. Mutations in the heavy chain of cytoplasmic dynein suppress the nudF nuclear migration mutation of Aspergillus nidulans. Mol. Gen. Genet 255, 194–200 (1997).
Xiang, X. & Qiu, R. Cargo-mediated activation of cytoplasmic dynein in vivo. Front. Cell Dev. Biol. 8, 598952 (2020).
Dix, C. I. et al. Lissencephaly-1 promotes the recruitment of dynein and dynactin to transported mRNAs. J. Cell Biol. 202, 479–494 (2013).
Tarricone, C. et al. Coupling PAF signaling to dynein regulation: structure of LIS1 in complex with PAF-acetylhydrolase. Neuron 44, 809–821 (2004).
Kim, M. H. et al. The structure of the N-terminal domain of the product of the lissencephaly gene Lis1 and its functional implications. Structure 12, 987–998 (2004).
DeSantis, M. E. et al. Lis1 has two opposing modes of regulating cytoplasmic dynein. Cell 170, 1197–1208 (2017).
Gillies, J. P. et al. Structural basis for cytoplasmic dynein-1 regulation by Lis1. eLife 11, e71229 (2022).
Huang, J., Roberts, A. J., Leschziner, A. E. & Reck-Peterson, S. L. Lis1 acts as a ‘clutch’ between the ATPase and microtubule-binding domains of the dynein motor. Cell 150, 975–986 (2012).
Toropova, K. et al. Lis1 regulates dynein by sterically blocking its mechanochemical cycle. eLife 3, e03372 (2014).
Parrini, E., Conti, V., Dobyns, W. B. & Guerrini, R. Genetic basis of brain malformations. Mol. Syndromol. 7, 220–233 (2016).
Reiner, O. et al. Isolation of a Miller–Dieker lissencephaly gene containing G protein beta-subunit-like repeats. Nature 364, 717–721 (1993).
Qiu, R., Zhang, J. & Xiang, X. LIS1 regulates cargo-adapter-mediated activation of dynein by overcoming its autoinhibition in vivo. J. Cell Biol. 218, 3630–3646 (2019).
Qiu, R., Zhang, J., Rotty, J. D. & Xiang, X. Dynein activation in vivo is regulated by the nucleotide states of its AAA3 domain. Curr. Biol. 31, 4486–4498.e6 (2021).
Gutierrez, P. A., Ackermann, B. E., Vershinin, M. & McKenney, R. J. Differential effects of the dynein-regulatory factor Lissencephaly-1 on processive dynein–dynactin motility. J. Biol. Chem. 292, 12245–12255 (2017).
Emre Kusakci, Z. M. H., John P. G., Reck-Peterson S. L., & Yildiz A. Lis1 binding regulates force-induced detachment of cytoplasmic dynein from microtubules. Preprint at bioRxiv https://doi.org/10.1101/2022.06.02.494578 (2022).
Schmidt, H., Zalyte, R., Urnavicius, L. & Carter, A. P. Structure of human cytoplasmic dynein-2 primed for its power stroke. Nature 518, 435–438 (2015).
Eshel, D. et al. Cytoplasmic dynein is required for normal nuclear segregation in yeast. Proc. Natl Acad. Sci. USA 90, 11172–11176 (1993).
Li, Y. Y., Yeh, E., Hays, T. & Bloom, K. Disruption of mitotic spindle orientation in a yeast dynein mutant. Proc. Natl Acad. Sci. USA 90, 10096–10100 (1993).
Muhua, L., Karpova, T. S. & Cooper, J. A. A yeast actin-related protein homologous to that in vertebrate dynactin complex is important for spindle orientation and nuclear migration. Cell 78, 669–679 (1994).
Lammers, L. G. & Markus, S. M. The dynein cortical anchor Num1 activates dynein motility by relieving Pac1/LIS1-mediated inhibition. J. Cell Biol. 211, 309–322 (2015).
Markus, S. M. & Lee, W.-L. Regulated offloading of cytoplasmic dynein from microtubule plus ends to the cortex. Dev. Cell 20, 639–651 (2011).
Moore, J. K., Stuchell-Brereton, M. D. & Cooper, J. A. Function of dynein in budding yeast: mitotic spindle positioning in a polarized cell. Cell Motil. Cytoskeleton 66, 546–555 (2009).
Roberts, A. J., Goodman, B. S. & Reck-Peterson, S. L. Reconstitution of dynein transport to the microtubule plus end by kinesin. eLife 3, e02641 (2014).
Markus, S. M. et al. Quantitative analysis of Pac1/LIS1-mediated dynein targeting: implications for regulation of dynein activity in budding yeast. Cytoskeleton 68, 157–174 (2011).
Carvalho, P., Gupta, M. L., Hoyt, M. A. & Pellman, D. Cell cycle control of kinesin-mediated transport of Bik1 (CLIP-170) regulates microtubule stability and dynein activation. Dev. Cell 6, 815–829 (2004).
Caudron, F., Andrieux, A., Job, D. & Boscheron, C. A new role for kinesin-directed transport of Bik1p (CLIP-170) in Saccharomyces cerevisiae. J. Cell Sci. 121, 1506–1513 (2008).
Reimer, J. M., DeSantis, M. E., Reck-Peterson, S. L. & Leschziner, A. E. Structures of human cytoplasmic dynein in complex with the lissencephaly 1 protein, LIS1. eLife 12, e84302 (2023).
Han, B.-G. et al. Long shelf-life streptavidin support-films suitable for electron microscopy of biological macromolecules. J. Struct. Biol. 195, 238–244 (2016).
Lahiri, I. et al. 3.1Å structure of yeast RNA polymerase II elongation complex stalled at a cyclobutane pyrimidine dimer lesion solved using streptavidin affinity grids. J. Struct. Biol. 207, 270–278 (2019).
Wagner, T. et al. SPHIRE-crYOLO is a fast and accurate fully automated particle picker for cryo-EM. Commun. Biol. 2, 218 (2019).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved single-particle cryo-electron microscopy. Nat. Methods. 4, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Bepler, T., Kelley, K., Noble, A. J. & Berger, B. Topaz-Denoise: general deep denoising models for cryoEM and cryoET. Nat. Commun. 11, 5208 (2020).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D Struct. Biol. 74, 531–544 (2018).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Struct. Biol. 66, 486–501 (2010).
Song, Y. et al. High-resolution comparative modeling with rosettaCM. Structure 21, 1735–1742 (2013).
Longtine, M. S. et al. Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 14, 953–961 (1998).
Gietz, R. D. & Schiestl, R. H. High-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2, 31–34 (2007).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Reck-Peterson, S. L. et al. Single-molecule analysis of dynein processivity and stepping behavior. Cell 126, 335–348 (2006).
Baumbach, J. et al. Lissencephaly-1 is a context-dependent regulator of the human dynein complex. eLife 6, e21768 (2017).
Acknowledgements
We thank the Nikon Imaging Center at UC San Diego and the Cryo-EM Facility at UC San Diego. We thank Z. Htet for early work preparing and collecting data on the Lis1 sample used for cryo-EM here. We also thank our funding sources: E.P.K. is funded by a Jane Coffin Childs Postdoctoral Fellowship; J.M.R. was a Merck Fellow of the Damon Runyon Cancer Research Foundation DRG-2370-19; A.A.K. was supported by American Cancer Society PF-18-190-01-CCG; A.L.’s lab by NIH R01 GM107214 and R35 GM145296; and S.R.P.’s lab by the Howard Hughes Medical Institute and NIH R35 GM141825.
Author information
Authors and Affiliations
Contributions
E.P.K. performed all the experiments in yeast; J.M.R. performed the cryo-EM data processing for the dyneinE2488Q construct, built the models of Chi, and analyzed the structures; A.A.K. performed the single-molecule experiments and collected and processed the data for wild-type dynein bound to Lis1; K.H.V.N. collected and processed data for full-length human dynein bound to LIS1; J.A.R. assisted with the yeast experiments; J.B.T. assisted with protein the protein biochemistry; I.L. prepared the grids with the dyneinE2488Q–Lis1 samples and collected the cryo-EM data; S.L.R.-P. and A.E.L. supervised the work. E.P.K., J.M.R., A.A.K., S.R.P.-L., and A.E.L. wrote and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Katarzyna Ciazynska, in collaboration with the Nature Structural & Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cryo-EM data processing workflow.
a, Processing workflow for Chi. Particles belonging to teal maps were carried into the next round of processing. b, Cryo-EM map of Chi (dyneinE2488Q-Lis1)2 colored by local resolution. c, Fourier shell correlation (FSC) plot for Chi. d, Symmetry expansion followed by local refinement map of dyneinE2488Q-(Lis1)2. e, Local refinement of symmetry expanded dyneinE2488Q-(Lis1)2 colored by local resolution. f, FSC plot for local refinement map.
Extended Data Fig. 2 The wild-type yeast dynein motor domain can form Chi in the presence of ATP.
a, Class average from a data set of wild-type yeast dynein motor domain incubated in the presence of Lis1 and ATP. b, Our model of Chi, rendered as a surface at 10 Å resolution, shown in the same orientation as the class average in (a), which corresponds to Chi being viewed from the stalks and towards the motor domains. c, The model in (b) is rotated to show a more canonical view of Chi. Note that Lis1-stalk2 appears to have partial density in the class average.
Extended Data Fig. 3 Binding of Lis1 to dynein is sterically incompatible with Phi.
a, Phi dynein (PDB 5NUG) was aligned to dynein1 in Chi, which is shown in gray in this panel. Phi’s dynein2, shown in color, clashes with the Lis1 bound at sitering in Chi’s dynein1. b, Close up of our model of Chi equivalent to the view in (a), showing Lis1 bound at sitering on dynein1. Part of dynein2, with which the Lis1 at sitering interacts in Chi, is also shown.
Extended Data Fig. 4 The conformation of the linker is different in Chi and Phi.
a, The linker docks onto AAA2 and AAA3 in Phi. b, In contrast, the linker is disengaged from the AAA ring in Chi. c, Overlay of Phi (a) and Chi (b) highlights the different positions of the linker. d, e, The N-termini of the linker adopt different paths in Phi (d) and Chi (e). f, Overlay of Phi (d) and Chi (e) shows how the linkers are pulled apart in Chi.
Extended Data Fig. 5 Western blot analysis of Lis1 mutant expression.
a, Cell lysates from wild type (WT), Lis1Δ or mutant Lis1 (Lis1N213A, Lis1W288D and Lis1N213A, W88D) S. cerevisiae strains containing Flag-tagged Lis1 (Pac1) were immunoprecipitated using anti-Flag agarose beads and immunoblotted for the FLAG peptide (α-FLAG). b, Quantitative analysis (mean ± s.e.m.) of Lis1 expression (α-FLAG band) normalized against the non-specific band observed at ~75 KDa for three independent immunoprecipitations. Statistics were generated using a One-way ANOVA with Tukey multiple comparisons of each mean test. **** WT and ΔLis1, ΔLis1 and Lis1N213A, ΔLis1 and Lis1W288D, ΔLis1 and Lis1N213A, W288D p < 0.0001. Differences not noted are not statistically significant. Source data are provided with the manuscript.
Extended Data Fig. 6 Chi mutations in human LIS1.
a, Sequence conservation between S. cerevisiae (S. cer) and human (H. sap) Lis1 in the vicinity of the dynein binding site in the Chi structure. Residues with 70% conservation or higher are shaded in light red. The residues mutated in this study: N203, D205, Y225, and D245 are highlighted by red boxes. b, Sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) of the human LIS1 constructs used for motility assays. c, Comparison of percent diffusive events (mean ± SEM) from single-molecule TMR-dynein-dynactin-BICD2 assays performed in the absence (white circle) or presence (black circle) of different LIS1 constructs. The data points are represented as triangles, circles, and squares, corresponding to the mean diffusive events measured within each technical replicate. Four replicates per condition were collected and the number of data points (n) per each replicate is listed (no LIS1, n = 14/20/20/8; LIS1, n = 7/16/7/5; LIS1N203A, D205A, Y225A, n = 15/7/16/9; LIS1N203A, D205A, D245A, n = 17/9/17/9). ** no LIS1 and LIS1N203A, D205A, D245A p = 0.0051, * no LIS1 and LIS1N203A, D205A, Y225A p = 0.017, * LIS1 and LIS1N203A, D205A, D245A ** p = 0.025 ns, not statistically significant, One-Way ANOVA with Tukey multiple comparisons of each mean. Source data are provided with the manuscript.
Supplementary information
Supplementary Information
Supplementary Table 1, yeast strains
Supplementary Video 1
The structure of S. cerevisiae Chi (dynein–Lis1).
Source data
Source Data Fig. 4
Statistical Source data and analyses
Source Data Fig. 5
Statistical Source data and analyses
Source Data Extended Data Fig. 5
Unprocessed western blot
Source Data Extended Data Fig. 5
Numerical values and statistical analyses
Source Data Extended Data Fig. 6
Unprocessed SDS–PAGE gel
Source Data Extended Data Fig. 6
Numerical values and statistical analyses
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Karasmanis, E.P., Reimer, J.M., Kendrick, A.A. et al. Lis1 relieves cytoplasmic dynein-1 autoinhibition by acting as a molecular wedge. Nat Struct Mol Biol 30, 1357–1364 (2023). https://doi.org/10.1038/s41594-023-01069-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-023-01069-6
This article is cited by
-
New pieces for the Lis1–dynein puzzle
Nature Structural & Molecular Biology (2023)
-
Nde1 promotes Lis1-mediated activation of dynein
Nature Communications (2023)