Abstract
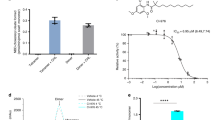
Cholesterol is an essential component of mammalian cell membranes, constituting up to 50% of plasma membrane lipids. By contrast, it accounts for only 5% of lipids in the endoplasmic reticulum (ER)1. The ER enzyme sterol O-acyltransferase 1 (also named acyl-coenzyme A:cholesterol acyltransferase, ACAT1) transfers a long-chain fatty acid to cholesterol to form cholesteryl esters that coalesce into cytosolic lipid droplets. Under conditions of cholesterol overload, ACAT1 maintains the low cholesterol concentration of the ER and thereby has an essential role in cholesterol homeostasis2,3. ACAT1 has also been implicated in Alzheimer’s disease4, atherosclerosis5 and cancers6. Here we report a cryo-electron microscopy structure of human ACAT1 in complex with nevanimibe7, an inhibitor that is in clinical trials for the treatment of congenital adrenal hyperplasia. The ACAT1 holoenzyme is a tetramer that consists of two homodimers. Each monomer contains nine transmembrane helices (TMs), six of which (TM4–TM9) form a cavity that accommodates nevanimibe and an endogenous acyl-coenzyme A. This cavity also contains a histidine that has previously been identified as essential for catalytic activity8. Our structural data and biochemical analyses provide a physical model to explain the process of cholesterol esterification, as well as details of the interaction between nevanimibe and ACAT1, which may help to accelerate the development of ACAT1 inhibitors to treat related diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The 3D cryo-EM density map has been deposited in the Electron Microscopy Data Bank under the accession number EMD-21390. Atomic coordinates for the atomic model have been deposited in the Protein Data Bank under the accession number 6VUM. Source Data for Figs. 1, 3, 4 and Extended Data Fig. 9 are provided with the paper. All other data are available from the corresponding authors upon reasonable request.
References
Radhakrishnan, A., Goldstein, J. L., McDonald, J. G. & Brown, M. S. Switch-like control of SREBP-2 transport triggered by small changes in ER cholesterol: a delicate balance. Cell Metab. 8, 512–521 (2008).
Rogers, M. A. et al. Acyl-CoA:cholesterol acyltransferases (ACATs/SOATs): enzymes with multiple sterols as substrates and as activators. J. Steroid Biochem. Mol. Biol. 151, 102–107 (2015).
Goldstein, J. L. & Brown, M. S. The LDL receptor. Arterioscler. Thromb. Vasc. Biol. 29, 431–438 (2009).
Hutter-Paier, B. et al. The ACAT inhibitor CP-113,818 markedly reduces amyloid pathology in a mouse model of Alzheimer’s disease. Neuron 44, 227–238 (2004).
Yagyu, H. et al. Absence of ACAT-1 attenuates atherosclerosis but causes dry eye and cutaneous xanthomatosis in mice with congenital hyperlipidemia. J. Biol. Chem. 275, 21324–21330 (2000).
Yang, W. et al. Potentiating the antitumour response of CD8+ T cells by modulating cholesterol metabolism. Nature 531, 651–655 (2016).
LaPensee, C. R. et al. ATR-101, a selective and potent inhibitor of acyl-CoA acyltransferase 1, induces apoptosis in H295R adrenocortical cells and in the adrenal cortex of dogs. Endocrinology 157, 1775–1788 (2016).
Guo, Z. Y., Lin, S., Heinen, J. A., Chang, C. C. & Chang, T. Y. The active site His-460 of human acyl-coenzyme A:cholesterol acyltransferase 1 resides in a hitherto undisclosed transmembrane domain. J. Biol. Chem. 280, 37814–37826 (2005).
Brown, M. S., Radhakrishnan, A. & Goldstein, J. L. Retrospective on cholesterol homeostasis: the central role of Scap. Annu. Rev. Biochem. 87, 783–807 (2018).
Porter, F. D. & Herman, G. E. Malformation syndromes caused by disorders of cholesterol synthesis. J. Lipid Res. 52, 6–34 (2011).
Xu, Y., Du, X., Turner, N., Brown, A. J. & Yang, H. Enhanced acyl-CoA:cholesterol acyltransferase activity increases cholesterol levels on the lipid droplet surface and impairs adipocyte function. J. Biol. Chem. 294, 19306–19321 (2019).
Masumoto, N. et al. Membrane bound O-acyltransferases and their inhibitors. Biochem. Soc. Trans. 43, 246–252 (2015).
Hofmann, K. A superfamily of membrane-bound O-acyltransferases with implications for Wnt signaling. Trends Biochem. Sci. 25, 111–112 (2000).
Ma, D. et al. Crystal structure of a membrane-bound O-acyltransferase. Nature 562, 286–290 (2018).
Chang, C. C., Huh, H. Y., Cadigan, K. M. & Chang, T. Y. Molecular cloning and functional expression of human acyl-coenzyme A:cholesterol acyltransferase cDNA in mutant Chinese hamster ovary cells. J. Biol. Chem. 268, 20747–20755 (1993).
Anderson, R. A. et al. Identification of a form of acyl-CoA:cholesterol acyltransferase specific to liver and intestine in nonhuman primates. J. Biol. Chem. 273, 26747–26754 (1998).
Cases, S. et al. ACAT-2, a second mammalian acyl-CoA:cholesterol acyltransferase. Its cloning, expression, and characterization. J. Biol. Chem. 273, 26755–26764 (1998).
Oelkers, P., Behari, A., Cromley, D., Billheimer, J. T. & Sturley, S. L. Characterization of two human genes encoding acyl coenzyme A:cholesterol acyltransferase-related enzymes. J. Biol. Chem. 273, 26765–26771 (1998).
Yu, C. et al. Role of the N-terminal hydrophilic domain of acyl-coenzyme A:cholesterol acyltransferase 1 on the enzyme’s quaternary structure and catalytic efficiency. Biochemistry 41, 3762–3769 (2002).
Nissen, S. E. et al. Effect of ACAT inhibition on the progression of coronary atherosclerosis. N. Engl. J. Med. 354, 1253–1263 (2006).
Cao, J. et al. Targeting Acyl-CoA:diacylglycerol acyltransferase 1 (DGAT1) with small molecule inhibitors for the treatment of metabolic diseases. J. Biol. Chem. 286, 41838–41851 (2011).
Long, T. et al. Structural basis for human sterol isomerase in cholesterol biosynthesis and multidrug recognition. Nat. Commun. 10, 2452 (2019).
Guo, Z. Y., Chang, C. C. & Chang, T. Y. Functionality of the seventh and eighth transmembrane domains of acyl-coenzyme A:cholesterol acyltransferase 1. Biochemistry 46, 10063–10071 (2007).
Das, A., Davis, M. A. & Rudel, L. L. Identification of putative active site residues of ACAT enzymes. J. Lipid Res. 49, 1770–1781 (2008).
Schneidman-Duhovny, D., Inbar, Y., Nussinov, R. & Wolfson, H. J. PatchDock and SymmDock: servers for rigid and symmetric docking. Nucleic Acids Res. 33, W363–W367 (2005).
Li, X., Roberti, R. & Blobel, G. Structure of an integral membrane sterol reductase from Methylomicrobium alcaliphilum. Nature 517, 104–107 (2015).
Liu, J., Chang, C. C., Westover, E. J., Covey, D. F. & Chang, T. Y. Investigating the allosterism of acyl-CoA:cholesterol acyltransferase (ACAT) by using various sterols: in vitro and intact cell studies. Biochem. J. 391, 389–397 (2005).
Li, X., Saha, P., Li, J., Blobel, G. & Pfeffer, S. R. Clues to the mechanism of cholesterol transfer from the structure of NPC1 middle lumenal domain bound to NPC2. Proc. Natl Acad. Sci. USA 113, 10079–10084 (2016).
Pfeffer, S. R. NPC intracellular cholesterol transporter 1 (NPC1)-mediated cholesterol export from lysosomes. J. Biol. Chem. 294, 1706–1709 (2019).
Long, T. et al. Structural basis for itraconazole-mediated NPC1 inhibition. Nat. Commun. 11, 152 (2020).
Chang, C. C. et al. Recombinant acyl-CoA:cholesterol acyltransferase-1 (ACAT-1) purified to essential homogeneity utilizes cholesterol in mixed micelles or in vesicles in a highly cooperative manner. J. Biol. Chem. 273, 35132–35141 (1998).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017)
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Terwilliger, T. et al. A fully automatic method yielding initial models form high-resolution cryo-electron microscopy maps. Nat. Methods 15, 905-908 (2018).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Acknowledgements
We thank M. Brown and J. Goldstein for support throughout the project. Data were collected at the UT Southwestern Medical Center Cryo-EM Facility (funded in part by the CPRIT Core Facility Support Award RP170644). We thank D. Stoddard for assistance with data collection; R. DeBose-Boyd, L. Esparza and L. Friedberg for technical support; E. Coutavas and E. Debler for discussion; and A. Lemoff at the UT Southwestern Proteomics Core for mass spectrometry identification. This work was supported by National Institutes of Health grants P01 HL020948, R01 GM134700 and R01 GM135343; the Endowed Scholars Program in Medical Science of UT Southwestern Medical Center; O’Donnell Junior Faculty Funds; and Welch Foundation grant I-1957 to X.L. X.Q. is the recipient of a DDBrown Fellow of the Life Sciences Research Foundation. X.L. is a Damon Runyon-Rachleff Innovator supported by the Damon Runyon Cancer Research Foundation (DRR-53-19) and a Rita C. and William P. Clements Jr. Scholar in Biomedical Research at UT Southwestern Medical Center.
Author information
Authors and Affiliations
Contributions
X.L. conceived the project and designed the research with T.L. T.L. and A.H. performed the biochemical study. T.L., Y.S. and X.Q. carried out cryo-EM work, built the model and refined the structure. T.L., Y.S., X.Q. and X.L. analysed the data, and all authors contributed to the preparation of the manuscript. X.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks David Drew, Savvas N. Savvides and Alan A. Tall for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Sequence alignment of human ACAT1 and ACAT2, hamster ACAT1 and ACAT2, chicken ACAT1 and zebrafish ACAT1.
The transmembrane helices and the residue numbers of human ACAT1 are indicated above the protein sequence. The specific residues necessary for ligand binding are labelled with an asterisk. Hs, Homo sapiens (human); cg, Cricetulus griseus (Chinese hamster); gg, Gallus gallus (chicken); dr, Danio rerio (zebrafish).
Extended Data Fig. 2 Expression and purification of human ACAT1 with nevanimibe.
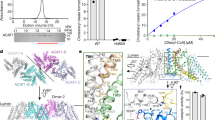
Representative gel-filtration chromatogram of ACAT1 (Superose 6 Increase 10/30 column). The SDS–PAGE gel (with molecular weight markers) is shown on the right, and ACAT1 is indicated with an arrow. The experiment was reproduced three times on separate occasions with similar results.
Extended Data Fig. 3 Processing of data acquired from 200 kV cryo-TEM.
a, Representative gel-filtration chromatogram (Superose 6 Increase 10/30 column) of cross-linked ACAT1. The SDS–PAGE gel (with molecular weight markers) is shown on the right, and ACAT1 is indicated with an arrow. The experiment was reproduced twice on separate occasions with similar results. b, The data processing workflow in RELION-3. The 3D classes and refinement results from the cryo-EM data are shown.
Extended Data Fig. 4 Processing of data acquired from 300 kV cryo-TEM.
The data processing workflow in RELION-3. The cryo-EM 3D classes as well as the masks used for the refinement are shown. After RELION-3 refinement, the final cryo-EM map was sharpened using post-process with a B-factor value of −150 Å2.
Extended Data Fig. 5 FSC curve and estimation of the local resolution.
a, FSC curve as a function of resolution using RELION-3 output. b, Density maps of the structure of ACAT1, coloured by local resolution estimation using ResMap.
Extended Data Fig. 6 Cryo-EM map of the structural elements of ACAT1.
a, The FSC curves calculated between the refined structure model and the half map used for refinement (orange), the other half map (green) and the full map (blue). b, The major helices of ACAT1-A and ACAT1-B. The density map and model of the complex are shown as mesh and cartoons, respectively. Cryo-EM maps are contoured at the 5σ level.
Extended Data Fig. 7 Structural comparison of ACAT1 and DltB.
a, Structural comparison from parallel to the membrane view (left) and from the lumen view (right). b, Structural comparison of the catalytic sites of ACAT1 and DltB. The catalytic histidine residues are shown as sticks.
Extended Data Fig. 8 Purification of human ACAT1 variants for activity assays.
Representative gel-filtration chromatograms (Superose 6 Increase 10/30 column) of ACAT1 variants in buffer containing 20 mM HEPES, pH 7.5, 150 mM NaCl and 1% CHAPS. The experiment was reproduced twice on separate occasions with similar results.
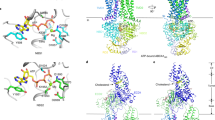
Extended Data Fig. 9 The putative allosteric sites of ACAT1.
a, b, Putative allosteric sites 1 (a) and 2 (b). Cholesterol (yellow) may form hydrophobic contacts that result in the stabilization of TM4 and TM6, respectively. c, Putative allosteric site 2 in the ACAT1 dimer. The cryo-EM maps of cholesterol are contoured at the 5σ level. d, Functional validation of the putative allosteric sites. Data are mean ± s.d. (n = 3 technically independent experiments). Each experiment was reproduced at least twice on separate occasions with similar results.
Supplementary information
Rights and permissions
About this article
Cite this article
Long, T., Sun, Y., Hassan, A. et al. Structure of nevanimibe-bound tetrameric human ACAT1. Nature 581, 339–343 (2020). https://doi.org/10.1038/s41586-020-2295-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2295-8
This article is cited by
-
SOAT1 regulates cholesterol metabolism to induce EMT in hepatocellular carcinoma
Cell Death & Disease (2024)
-
Structural insights into the transporting and catalyzing mechanism of DltB in LTA D-alanylation
Nature Communications (2024)
-
Metabolic reprogramming in colorectal cancer: regulatory networks and therapy
Cell & Bioscience (2023)
-
Hepatic steatosis is associated with dysregulated cholesterol metabolism and altered protein acetylation dynamics in chickens
Journal of Animal Science and Biotechnology (2023)
-
Mechanism of d-alanine transfer to teichoic acids shows how bacteria acylate cell envelope polymers
Nature Microbiology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.