Abstract
Receptor activator of nuclear factor-kappa B (RANK) ligand (RANKL) binds RANK on the surface of osteoclast precursors to trigger osteoclastogenesis. Recent studies have indicated that osteocytic RANKL has an important role in osteoclastogenesis during bone remodelling; however, the role of osteoblastic RANKL remains unclear. Here we show that vesicular RANK, which is secreted from the maturing osteoclasts, binds osteoblastic RANKL and promotes bone formation by triggering RANKL reverse signalling, which activates Runt-related transcription factor 2 (Runx2). The proline-rich motif in the RANKL cytoplasmic tail is required for reverse signalling, and a RANKL(Pro29Ala) point mutation reduces activation of the reverse signalling pathway. The coupling of bone resorption and formation is disrupted in RANKL(Pro29Ala) mutant mice, indicating that osteoblastic RANKL functions as a coupling signal acceptor that recognizes vesicular RANK. RANKL reverse signalling is therefore a potential pharmacological target for avoiding the reduced bone formation associated with inhibition of osteoclastogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Sims, N. A. & Martin, T. J. Coupling the activities of bone formation and resorption: a multitude of signals within the basic multicellular unit. Bonekey Rep. 3, 481 (2014).
Florencio-Silva, R., Sasso, G. R., Sasso-Cerri, E., Simoes, M. J. & Cerri, P. S. Biology of bone tissue: structure, function, and factors that influence bone cells. BioMed Res. Int. 2015, 421746 (2015).
Kong, Y. Y. et al. OPGL is a key regulator of osteoclastogenesis, lymphocyte development and lymph-node organogenesis. Nature 397, 315–323 (1999).
Suda, T. et al. Modulation of osteoclast differentiation and function by the new members of the tumor necrosis factor receptor and ligand families. Endocr. Rev. 20, 345–357 (1999).
Nakashima, T. et al. Evidence for osteocyte regulation of bone homeostasis through RANKL expression. Nat. Med. 17, 1231–1234 (2011).
Xiong, J. et al. Matrix-embedded cells control osteoclast formation. Nat. Med. 17, 1235–1241 (2011).
Kisiswa, L. et al. TNFα reverse signaling promotes sympathetic axon growth and target innervation. Nat. Neurosci. 16, 865–873 (2013).
Sugamori, Y. et al. Peptide drugs accelerate BMP-2-induced calvarial bone regeneration and stimulate osteoblast differentiation through mTORC1 signaling. BioEssays 38, 717–725 (2016).
Boyce, B. F. & Xing, L. The RANKL/RANK/OPG pathway. Curr. Osteoporos. Rep. 5, 98–104 (2007).
Martin, T. J. & Sims, N. A. RANKL/OPG; Critical role in bone physiology. Rev. Endocr. Metab. Disord. 16, 131–139 (2015).
Fei, Q. et al. Osteogenic growth peptide enhances the proliferation of bone marrow mesenchymal stem cells from osteoprotegerin-deficient mice by CDK2/cyclin A. Acta Biochim. Biophys. Sin. (Shanghai) 42, 801–806 (2010).
Andersen, T. L. et al. Understanding coupling between bone resorption and formation: are reversal cells the missing link? Am. J. Pathol. 183, 235–246 (2013).
Negishi-Koga, T. et al. Suppression of bone formation by osteoclastic expression of semaphorin 4D. Nat. Med. 17, 1473–1480 (2011).
Huynh, N. et al. Characterization of regulatory extracellular vesicles from osteoclasts. J. Dent. Res. 95, 673–679 (2016).
Komori, T. Regulation of osteoblast differentiation by transcription factors. J. Cell. Biochem. 99, 1233–1239 (2006).
Kadono, Y. et al. Strength of TRAF6 signalling determines osteoclastogenesis. EMBO Rep. 6, 171–176 (2005).
Singha, U. K. et al. Rapamycin inhibits osteoblast proliferation and differentiation in MC3T3–E1 cells and primary mouse bone marrow stromal cells. J. Cell. Biochem. 103, 434–446 (2008).
Trajkovic, K. et al. Ceramide triggers budding of exosome vesicles into multivesicular endosomes. Science 319, 1244–1247 (2008).
Tomimori, Y. et al. Evaluation of pharmaceuticals with a novel 50-hour animal model of bone loss. J. Bone Miner. Res. 24, 1194–1205 (2009).
Kariya, Y., Honma, M., Aoki, S., Chiba, A. & Suzuki, H. Vps33a mediates RANKL storage in secretory lysosomes in osteoblastic cells. J. Bone Miner. Res. 24, 1741–1752 (2009).
Saksela, K., Cheng, G. & Baltimore, D. Proline-rich (PxxP) motifs in HIV-1 Nef bind to SH3 domains of a subset of Src kinases and are required for the enhanced growth of Nef+ viruses but not for down-regulation of CD4. EMBO J. 14, 484–491 (1995).
Cuevas, B. D. et al. Tyrosine phosphorylation of p85 relieves its inhibitory activity on phosphatidylinositol 3-kinase. J. Biol. Chem. 276, 27455–27461 (2001).
Takeshita, S. et al. Osteoclast-secreted CTHRC1 in the coupling of bone resorption to formation. J. Clin. Invest. 123, 3914–3924 (2013).
Pederson, L., Ruan, M., Westendorf, J. J., Khosla, S. & Oursler, M. J. Regulation of bone formation by osteoclasts involves Wnt/BMP signaling and the chemokine sphingosine-1-phosphate. Proc. Natl Acad. Sci. USA 105, 20764–20769 (2008).
Sato, N. et al. MyD88 but not TRIF is essential for osteoclastogenesis induced by lipopolysaccharide, diacyl lipopeptide, and IL-1α. J. Exp. Med. 200, 601–611 (2004).
George, J., Kuboki, Y. & Miyata, T. Differentiation of mesenchymal stem cells into osteoblasts on honeycomb collagen scaffolds. Biotechnol. Bioeng. 95, 404–411 (2006).
Maruyama, Z. et al. Runx2 determines bone maturity and turnover rate in postnatal bone development and is involved in bone loss in estrogen deficiency. Dev. Dyn. 236, 1876–1890 (2007).
Alles, N. et al. Suppression of NF-κB increases bone formation and ameliorates osteopenia in ovariectomized mice. Endocrinology 151, 4626–4634 (2010).
Dempster, D. W. et al. Standardized nomenclature, symbols, and units for bone histomorphometry: a 2012 update of the report of the ASBMR Histomorphometry Nomenclature Committee. J. Bone Miner. Res. 28, 2–17 (2013).
Pini, A. et al. Design and use of a phage display library. Human antibodies with subnanomolar affinity against a marker of angiogenesis eluted from a two-dimensional gel. J. Biol. Chem. 273, 21769–21776 (1998).
Watzl, C. Production and use of trimeric isoleucine zipper fusion proteins to study surface receptor ligand interactions. Curr. Protoc. Protein Sci. 19, 19.11.1–19.11.13 (2006).
Sakai, A. et al. Bone marrow cell development and trabecular bone dynamics after ovariectomy in ddy mice. Bone 23, 443–451 (1998).
Van Deun, J. et al. EV-TRACK: transparent reporting and centralizing knowledge in extracellular vesicle research. Nat. Methods 14, 228–232 (2017).
Vizcaino, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44, D447–D456 (2016).
Acknowledgements
We thank K. Toritani, M. Maeda, and M. A. Al Mamun for their technical assistance. This work was supported in part by JSPS KAKENHI grant numbers JP16H05445, JP24390349, JP26713045, JP24890048, JP25670632; and by MEXT KAKENHI grant number JP22136015.
Reviewer information
Nature thanks S. Takeda. M. Zaidi and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Y.I. and S.A. performed in vitro and in vivo experiments, analysed data, and wrote the manuscript. M.Ho. designed the study, performed in vitro experiments, analysed data, and wrote and revised the manuscript. Y.S., M.K., G.K. and Y.T. designed and/or performed experiments using the calvarial defect model. M.Ha. performed construction and evaluation of biologics. Y.K. performed RANK suppression experiments. N.U. and J.M.P. designed and/or performed experiments using RANKL-deficient cells. K.A. and H.S. supervised the study and revised the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Incorporation of RANK into mOC-SEVs.
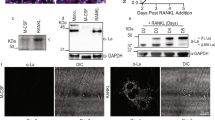
a, RANK content over time (top) in osteoclasts (lines) and their secretion products that were fractionated by serial centrifugation (columns) (n = 3); BLQ, below the limit of quantification. Bottom, osteoclast maturation over time with images of TRAP-stained cells and TRAP activity (n = 4). b, Transmission electron micrograph of precipitates from culture supernatants of mOCs by ultracentrifugation at 100,000g. Scale bar, 100 nm. c, RANK content in SEVs collected from osteoclast culture supernatants over time using ExoQuick-TC SEV-precipitating reagent (n = 3). d, Co-precipitation analyses of mOC-SEVs. Top, RANK content in co-precipitates (n = 3). Bottom, co-precipitates analysed for SEV markers (CD9, CD63, and CD81) and an endoplasmic reticulum protein (calnexin) by immunoblotting. GST pull-down (GPD) assays were used to precipitate RANK, and immuno-precipitation (IP) was used to precipitate CD9, CD63, and CD81, respectively. e, Confocal laser scanning micrograph of osteoclasts labelled for RANK (green), CD9 (red), and nuclei (blue). Scale bar, 20 μm. In a, c, d, data are mean, with individual points representing biologically independent samples.
Extended Data Fig. 2 Osteogenic potency of mOC-SEVs under physiological conditions.
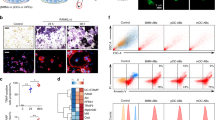
a, Images of von Kossa-stained ST2 cells stimulated with SEVs derived from mOCs, pOCs, or unstimulated RAW 264.7 cells. b, Changes in mRNA expression levels of osteoblast markers in ST2 cells after 5 days of stimulation with SEVs derived from mOCs, pOCs, or unstimulated RAW 264.7 cells (n = 3). c, Expression of osteoblast markers in mouse primary osteoblasts stimulated with mOC-SEVs, mOC-SEVs plus sRANKL, or vehicle (n = 3). d, Von Kossa-stained primary osteoblasts stimulated with mOC-SEVs, mOC-SEVs plus sRANKL, or vehicle. e, Von Kossa-stained ST2 cells stimulated with serially-centrifuged conditioned medium from mOCs, pOCs, or unstimulated RAW 264.7 cells. f, Comparison of osteoblast-stimulating effect of conditioned medium from mOCs with that of purified mOC-SEVs. Von Kossa-stained ST2 cells stimulated with serially-centrifuged conditioned medium from mOCs, or with 2.5, 5.0, and 10.0 μg protein per ml purified mOC-SEVs. g, Dual-energy X-ray absorptiometry measurements of bone mineral density (BMD) and bone mineral content (BMC). Measurements were performed on the injured skull region of the mouse calvarial defect model after treatment with mOC-SEVs, mOC-SEVs plus sRANKL, BMP-2 (positive control), or vehicle (n = 4). In b, c, g, data are mean, with individual scatter points representing biologically independent samples or mice. One-way ANOVA followed by Student–Newman–Keuls test; *P < 0.05, **P < 0.01.
Extended Data Fig. 3 Influence of RANKL-binding molecules on the osteogenic effects of mOC-SEVs.
a, The inhibitory effect of OPG on the activation of PI3K–Akt–mTORC1 pathway in ST2 cells stimulated with mOC-SEVs. Akt Thr308 phosphorylation (p-Akt), PRAS40 Thr246 phosphorylation (p-PRAS40) and S6K1 Thr389 phosphorylation (p-S6K1) indicate PI3K, Akt and mTORC1 activation, respectively. b, The inhibitory effect of OPG on osteoblast-marker expression levels in ST2 cells after 5 days of stimulation with mOC-SEVs (n = 3). c, Von Kossa-stained ST2 cells stimulated with mOC-SEVs or vehicle in the presence or absence of OPG. d, Inhibitory effect of the WP9QY peptide, αR-tri, or OPG on RANK-Fc binding to GST–sRANKL immobilized on ELISA plate (n = 3 biologically independent samples). e, Von Kossa-stained primary osteoblasts derived from RANKL-deficient mice or wild-type littermates stimulated with mOC-SEVs, WP9QY peptide, mOC-SEVs plus WP9QY peptide, or vehicle. f, mRNA expression levels of osteoblast markers in primary osteoblasts derived from RANKL-deficient mice or wild-type littermates after 5 days of stimulation with mOC-SEVs, WP9QY peptide, mOC-SEVs plus WP9QY peptide, or vehicle (n = 3). In b, f, data are mean, with individual points representing biologically independent samples. One-way ANOVA followed by Student–Newman–Keuls test; *P < 0.05, **P < 0.01.
Extended Data Fig. 4 Activation of PI3K–Akt–mTORC1–Runx2 axis by vesicular RANK.
a, The effect of rapamycin on nuclear translocation of Runx2 in ST2 cells stimulated with mOC-SEVs. Histone deacetylase 2 (Hdac2) was used as reference nuclear protein. b, c, Influence of PI3K inhibitor (LY294002) (b) or mTORC1 inhibitor (rapamycin) (c) on activation of the PI3K–Akt–mTORC1 pathway in ST2 cells stimulated with mOC-SEVs.
Extended Data Fig. 5 Osteoblast-stimulating effects of RANKL-crosslinking agents.
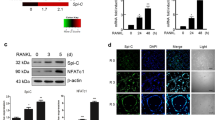
a, Influence of mTORC1 inhibitor (rapamycin) on nuclear Runx2 in ST2 cells stimulated with αR-tri or αB-tri. b, Time course of nuclear Runx2 in mouse primary osteoblasts stimulated with αR-tri or αB-tri. c, Time course of osteoblastic marker expression in mouse primary osteoblasts stimulated with αR-tri, αB-tri, or vehicle (n = 3). d, Activation of PI3K–Akt–mTORC1 pathway in ST2 cells stimulated with RANKL-crosslinking agents, RANK-Fc plus beads, OPG-Fc plus beads, RANK-Fc plus IgM, and OPG-Fc plus IgM. Vehicle, RANK-Fc, OPG-Fc, beads, and IgM were used as respective controls. 10 μg ml−1 mOC-SEVs (containing about 1 ng ml−1 RANK) and 1 μg ml−1 RANK-Fc or OPG-Fc was used. e. Expression of osteoblast markers in ST2 cells after 5 days of stimulation with RANKL-crosslinking agents. (n = 3). c, e, Data are mean, with individual points representing biologically independent samples. One-way ANOVA followed by Student–Newman–Keuls test; *P < 0.05, **P < 0.01.
Extended Data Fig. 6 Confirmation of biphasic property of RANKL reverse signalling.
a, Alizarin red staining of ST2 cells stimulated with mOC-SEVs, αR-tri, αB-tri, or vehicle during the early or late stage of differentiation. b, Alizarin red-stained ST2 cells infected with DN-Runx2 adenovirus before stimulation with αR-tri or αB-tri during the early or late stage of differentiation. c, d, Time courses of expression of early osteoblastic markers (Col1a1 mRNA and P1NP value reflecting the expression of type I collagen protein) and late osteoblastic markers (osteocalcin mRNA and Gla-osteocalcin protein) in mouse primary osteoblasts stimulated during the early stage (c) or late stage (d) of differentiation with mOC-SEVs, αR-tri, αB-tri, or vehicle (n = 3). e, f, Von Kossa-stained (e) or alizarin red-stained (f) mouse primary osteoblasts stimulated with mOC-SEVs, αR-tri, αB-tri, or vehicle during the early stage or late stage of differentiation. In c and d, data are mean, with individual points representing biologically independent samples.
Extended Data Fig. 7 The effect of neutralizing mOC-SEVs.
a, Influence of suppressing mRNA expression of Rab family members in osteoclasts on the secretion of vesicular RANK from mOCs (n = 3). b, Influence of suppressing mRNA expression of Rab family members in osteoclasts on formation of multinuclear cells (MNCs) and their mineral-resorbing activity (n = 3). c, Influence of GW4869 on the secretion of vesicular RANK from mOCs (n = 3). d, Influence of GW4869 on formation of MNCs and their mineral-resorbing activity (n = 3). e, Binding affinity of αI-tri1 or αI-tri2 to the IGSF8 ectodomain (n = 3 biologically independent samples). f, Influence of αI-tri1/2 on formation of MNCs and their mineral-resorbing activity (n = 3). g, Influence of pre-incubating mOC-SEVs with αI-tri1/2, αB-tri, GST–sRANKL or vehicle on the stimulatory activity of mOC-SEVs on ST2 cells; activation of the PI3K–Akt–mTORC1 pathway in ST2 cells is shown. In a–d, f, data are mean, with individual points representing biologically independent samples. Statistical analyses, one-way ANOVA followed by Student–Newman–Keuls test; *P < 0.05, **P < 0.01.
Extended Data Fig. 8 Effect of introducing RANKL Pro29Ala mutation.
a, Src family kinase inhibitors (PP2 or dasatinib) inhibit the activation of the PI3K–Akt–mTORC1 pathway in ST2 cells stimulated with mOC-SEVs. b, Overexpression of kinase-dead Src mutant (Δkinase) inhibits PI3K activation in ST2 cells stimulated with mOC-SEVs. c, Overexpression of RANKL mutants (Pro29Ala or Pro39Ala) inhibits PI3K activation in ST2 cells stimulated with mOC-SEVs. d, Co-precipitation of Src and Fyn with RANKL in ST2 cells stimulated with RANK beads. RANKL Pro29Ala or Pro39Ala mutations decrease co-precipitation. e, Schematic diagram depicting the introduction of RANKL Pro29Ala point mutation. DTA, diphtheria toxin subunit A; Neo, neomycin-resistance gene. f, Expression and cell-surface presentation of RANKL in primary osteoblasts derived from wild-type and RANKLP29A mutant mice. g, Von Kossa-stained primary osteoblasts derived from RANKLP29A mutant mice and wild-type littermates stimulated with mOC-SEVs.
Extended Data Fig. 9 Phenotypes of RANKLP29A mutant mice.
a, Bone morphometric parameters in 17-week-old male RANKLP29A mice (n = 8) and wild-type littermates (n = 4) under physiological conditions. b, Bone morphometric parameters in OVX or sham operated 16-week-old female RANKLP29A mice (n = 7 (sham), 7 (OVX)) and wild-type littermates (n = 5 (sham), 5 (OVX)). c, Bone morphometric parameters in 8-week-old male RANKLP29A mice and wild-type littermates treated with daily subcutaneous injection of 20 or 100 μg kg−1 day−1 parathyroid hormone (PTH) over the course of 2 weeks (n = 3 in each group). d, Expression levels over time of RANKL and OPG mRNA in primary osteocytes derived from RANKLP29A mice and wild-type littermates stimulated with αR-tri or αB-tri (n = 3). e, TRAP activity in bone marrow macrophages (BMMs) and primary osteocyte co-culture derived from RANKLP29A mice and wild-type littermates (n = 3). f, Time-courses of the expression levels of sclerostin mRNA in primary osteocytes derived from RANKLP29A mutant mice and wild-type littermates stimulated with αR-tri or αB-tri (n = 3). In a–f, data are mean, with individual points representing biologically independent samples or mice. Unpaired two-sided Student’s t-test (a) or one-way ANOVA followed by Student–Newman–Keuls test (b–f.); *P < 0.05, **P < 0.01.
Supplementary Information
Supplementary Information
This file contains a Supplementary Note, Supplementary Methods, Supplementary References and Supplementary Figure 1 (the unprocessed images presented in the main and Extended Data Figures).
Supplementary Table 1
This file contains a list of proteins incorporated in mOC-SEVs.
Source Data
Rights and permissions
About this article
Cite this article
Ikebuchi, Y., Aoki, S., Honma, M. et al. Coupling of bone resorption and formation by RANKL reverse signalling. Nature 561, 195–200 (2018). https://doi.org/10.1038/s41586-018-0482-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0482-7
Keywords
This article is cited by
-
Pan-histone deacetylase inhibitor vorinostat suppresses osteoclastic bone resorption through modulation of RANKL-evoked signaling and ameliorates ovariectomy-induced bone loss
Cell Communication and Signaling (2024)
-
The role and applications of extracellular vesicles in osteoporosis
Bone Research (2024)
-
Mechanistic insights into bone remodelling dysregulation by human viral pathogens
Nature Microbiology (2024)
-
Development of an optogenetics tool, Opto-RANK, for control of osteoclast differentiation using blue light
Scientific Reports (2024)
-
Unraveling the intricacies of osteoclast differentiation and maturation: insight into novel therapeutic strategies for bone-destructive diseases
Experimental & Molecular Medicine (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.