Abstract
The central role of MRI in neuro-oncology is undisputed. The technique is used, both in clinical practice and in clinical trials, to diagnose and monitor disease activity, support treatment decision-making, guide the use of focused treatments and determine response to treatment. Despite recent substantial advances in imaging technology and image analysis techniques, clinical MRI is still primarily used for the qualitative subjective interpretation of macrostructural features, as opposed to quantitative analyses that take into consideration multiple pathophysiological features. However, the field of quantitative imaging and imaging biomarker development is maturing. The European Imaging Biomarkers Alliance (EIBALL) and Quantitative Imaging Biomarkers Alliance (QIBA) are setting standards for biomarker development, validation and implementation, as well as promoting the use of quantitative imaging and imaging biomarkers by demonstrating their clinical value. In parallel, advanced imaging techniques are reaching the clinical arena, providing quantitative, commonly physiological imaging parameters that are driving the discovery, validation and implementation of quantitative imaging and imaging biomarkers in the clinical routine. Additionally, computational analysis techniques are increasingly being used in the research setting to convert medical images into objective high-dimensional data and define radiomic signatures of disease states. Here, I review the definition and current state of MRI biomarkers in neuro-oncology, and discuss the clinical potential of quantitative image analysis techniques.
Key points
-
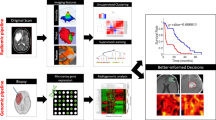
Imaging biomarkers offer the opportunity to move precision diagnostics forward, enabling better informed medical decision-making and tracking of biological changes before, during and after brain tumour treatment.
-
Guidelines and standards for data acquisition, image processing and validation processes for the development and eventual implementation of imaging biomarkers are provided by the European Society of Radiology and the Radiological Society of North America.
-
Radiomics is a rapidly emerging field of imaging research delivering an almost limitless supply of potential imaging biomarkers for improved patient and disease characterization.
-
The currently available evidence on imaging biomarkers and radiomics is still mostly at the discovery level; rigorous technical, biological and clinical validation are needed for clinical application.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Eckel-Passow, J. E. et al. Glioma groups based on 1p/19q, IDH, and TERT promoter mutations in tumors. N. Engl. J. Med. 372, 2499–2508 (2015).
Louis, D. N. et al. WHO classification of tumours of the central nervous system. Acta Neuropathol. 131, 803–820 (2016).
Suzuki, H. et al. Mutational landscape and clonal architecture in grade II and III gliomas. Nat. Genet. 47, 458–468 (2015).
Wijnenga, M. M. J. et al. The impact of surgery in molecularly defined low-grade glioma: an integrated clinical, radiological, and molecular analysis. Neuro Oncol. 20, 103–112 (2018).
Albert, N. L. et al. Response Assessment in Neuro-Oncology working group and European Association for Neuro-Oncology recommendations for the clinical use of PET imaging in gliomas. Neuro Oncol. 18, 1199–1208 (2016).
Galldiks, N. et al. PET imaging in patients with meningioma–report of the RANO/PET Group. Neuro Oncol. 19, 1576–1587 (2017).
Galldiks, N. et al. PET imaging in patients with brain metastasis–report of the RANO/PET group. Neuro Oncol. 21, 585–595 (2019).
Kessler, L. G. et al. The emerging science of quantitative imaging biomarkers terminology and definitions for scientific studies and regulatory submissions. Stat. Methods Med. Res. 24, 9–26 (2015). Introduction to imaging biomarkers and associated terminology.
FDA-NIH Biomarker Working Group. BEST (Biomarkers, EndpointS, and other Tools) Resource (FDA, 2021).
O’Connor, J. P. et al. Imaging biomarker roadmap for cancer studies. Nat. Rev. Clin. Oncol. 14, 169–186 (2017). Outline and recommendations for development and introduction of imaging biomarkers for cancer.
Law, I. et al. Joint EANM/EANO/RANO practice guidelines/SNMMI procedure standards for imaging of gliomas using PET with radiolabelled amino acids and [18F]FDG: version 1.0. Eur. J. Nucl. Med. Mol. Imaging 46, 540–557 (2019).
Sullivan, D. C. et al. Metrology standards for quantitative imaging biomarkers. Radiology 277, 813–825 (2015). Introduction and outline of metrology in the context of imaging biomarkers.
deSouza, N. M. et al. Validated imaging biomarkers as decision-making tools in clinical trials and routine practice: current status and recommendations from the EIBALL* subcommittee of the European Society of Radiology (ESR). Insights Imaging 10, 87 (2019).
Fournier, L. et al. Incorporating radiomics into clinical trials: expert consensus on considerations for data-driven compared to biologically driven quantitative biomarkers. Eur. Radiol. https://doi.org/10.1007/s00330-020-07598-8 (2021).
European Society of Radiology. ESR statement on the validation of imaging biomarkers. Insights Imaging 11, 76 (2020).
European Medicines Agency. Guideline on bioanalytical method validation (EMA, 2011).
Alberich-Bayarri, A., Neri, E. & Martí-Bonmatí, L. in Artificial Intelligence in Medical Imaging: Opportunities, Applications and Risks Ch. 10 (eds Ranschaert, E. R., Morozov, S. & Algra, P. R.) 119–128 (Springer, 2019).
Obuchowski, N. A. et al. Statistical issues in testing conformance with the Quantitative Imaging Biomarker Alliance (QIBA) profile claims. Acad. Radiol. 23, 496–506 (2016).
Huang, E. P. et al. Meta-analysis of the technical performance of an imaging procedure: guidelines and statistical methodology. Stat. Methods Med. Res. 24, 141–174 (2015).
Yankeelov, T. E. et al. Quantitative imaging in cancer clinical trials. Clin. Cancer Res. 22, 284–290 (2016).
Manfrini, E. et al. From research to clinical practice: a European neuroradiological survey on quantitative advanced MRI implementation. Eur. Radiol. https://doi.org/10.1007/s00330-020-07582-2 (2021).
Thust, S. C. et al. Glioma imaging in Europe: a survey of 220 centres and recommendations for best clinical practice. Eur. Radiol. 28, 3306–3317 (2018).
Erker, C. et al. Response assessment in paediatric high-grade glioma: recommendations from the Response Assessment in Pediatric Neuro-Oncology (RAPNO) working group. Lancet Oncol. 21, e317–e329 (2020).
Cha, S. Update on brain tumor imaging: from anatomy to physiology. AJNR 27, 475–487 (2006).
Sugahara, T. et al. Usefulness of diffusion-weighted MRI with echo-planar technique in the evaluation of cellularity in gliomas. J. Magn. Reson. Imaging 9, 53–60 (1999).
Higano, S. et al. Malignant astrocytic tumors: clinical importance of apparent diffusion coefficient in prediction of grade and prognosis. Radiology 241, 839–846 (2006).
Quantitative Imaging Biomarkers Alliance. QIBA Profile: Diffusion-weighted magnetic resonance imaging (DWI). Profile consensus (QIBA, 2019).
Bonekamp, D. et al. Diffusion tensor imaging in children and adolescents: reproducibility, hemispheric, and age-related differences. Neuroimage 34, 733–742 (2007).
Paldino, M. J., Barboriak, D., Desjardins, A., Friedman, H. S. & Vredenburgh, J. J. Repeatability of quantitative parameters derived from diffusion tensor imaging in patients with glioblastoma multiforme. J. Magn. Reson. Imaging 29, 1199–1205 (2009).
Pfefferbaum, A., Adalsteinsson, E. & Sullivan, E. V. Replicability of diffusion tensor imaging measurements of fractional anisotropy and trace in brain. J. Magn. Reson. Imaging 18, 427–433 (2003).
Sanvito, F., Castellano, A. & Falini, A. Advancements in neuroimaging to unravel biological and molecular features of brain tumors. Cancers 13, 424 (2021).
Aliotta, E. et al. Automated apparent diffusion coefficient analysis for genotype prediction in lower grade glioma: association with the T2-FLAIR mismatch sign. J. Neurooncol. 149, 325–335 (2020).
Zhang, L. et al. The utility of diffusion MRI with quantitative ADC measurements for differentiating high-grade from low-grade cerebral gliomas: evidence from a meta-analysis. J. Neurol. Sci. 373, 9–15 (2017).
Wang, Q. P., Lei, D. Q., Yuan, Y. & Xiong, N. X. Accuracy of ADC derived from DWI for differentiating high-grade from low-grade gliomas: systematic review and meta-analysis. Medicine 99, e19254 (2020).
Hales, P. W. et al. Arterial spin labelling and diffusion-weighted imaging in paediatric brain tumours. Neuroimage Clin. 22, 101696 (2019).
Suh, C. H., Kim, H. S., Jung, S. C. & Kim, S. J. Diffusion-weighted imaging and diffusion tensor imaging for differentiating high-grade glioma from solitary brain metastasis: a systematic review and meta-analysis. AJNR 39, 1208–1214 (2018).
Yu, Y. et al. Meta-analysis of the diagnostic performance of diffusion magnetic resonance imaging with apparent diffusion coefficient measurements for differentiating glioma recurrence from pseudoprogression. Medicine 99, e20270 (2020).
van Dijken, B. R. J., van Laar, P. J., Holtman, G. A. & van der Hoorn, A. Diagnostic accuracy of magnetic resonance imaging techniques for treatment response evaluation in patients with high-grade glioma, a systematic review and meta-analysis. Eur. Radiol. 27, 4129–4144 (2017).
Zulfiqar, M., Yousem, D. M. & Lai, H. ADC values and prognosis of malignant astrocytomas: does lower ADC predict a worse prognosis independent of grade of tumor?–a meta-analysis. Am. J. Roentgenol. 200, 624–629 (2013).
Suh, C. H., Kim, H. S., Jung, S. C., Choi, C. G. & Kim, S. J. Imaging prediction of isocitrate dehydrogenase (IDH) mutation in patients with glioma: a systemic review and meta-analysis. Eur. Radiol. 29, 745–758 (2019).
Aboian, M. S. et al. Imaging characteristics of pediatric diffuse midline gliomas with histone H3 K27M mutation. AJNR 38, 795–800 (2017).
Poussaint, T. Y. et al. Apparent diffusion coefficient histogram metrics correlate with survival in diffuse intrinsic pontine glioma: a report from the Pediatric Brain Tumor Consortium. Neuro Oncol. 18, 725–734 (2016).
Lober, R. M. et al. Diffusion-weighted MRI derived apparent diffusion coefficient identifies prognostically distinct subgroups of pediatric diffuse intrinsic pontine glioma. J. Neurooncol. 117, 175–182 (2014).
Tan, W. L. et al. Can diffusion tensor imaging noninvasively detect IDH1 gene mutations in astrogliomas? A retrospective study of 112 cases. AJNR 35, 920–927 (2014).
Aliotta, E. et al. Molecular subtype classification in lower-grade glioma with accelerated DTI. AJNR 40, 1458–1463 (2019).
Miloushev, V. Z., Chow, D. S. & Filippi, C. G. Meta-analysis of diffusion metrics for the prediction of tumor grade in gliomas. AJNR 36, 302–308 (2015).
Jiang, R. et al. The value of diffusion tensor imaging in differentiating high-grade gliomas from brain metastases: a systematic review and meta-analysis. PLoS ONE 9, e112550 (2014).
Abdalla, G. et al. The diagnostic role of diffusional kurtosis imaging in glioma grading and differentiation of gliomas from other intra-axial brain tumours: a systematic review with critical appraisal and meta-analysis. Neuroradiology 62, 791–802 (2020).
Zhang, H., Schneider, T., Wheeler-Kingshott, C. A. & Alexander, D. C. NODDI: practical in vivo neurite orientation dispersion and density imaging of the human brain. Neuroimage 61, 1000–1016 (2012).
Panagiotaki, E. et al. Noninvasive quantification of solid tumor microstructure using VERDICT MRI. Cancer Res. 74, 1902–1912 (2014).
Zaccagna, F. et al. Non-invasive assessment of glioma microstructure using VERDICT MRI: correlation with histology. Eur. Radiol. 29, 5559–5566 (2019).
Le Bihan, D. et al. MR imaging of intravoxel incoherent motions: application to diffusion and perfusion in neurologic disorders. Radiology 161, 401–407 (1986).
Falk Delgado, A., Nilsson, M., van Westen, D. & Falk Delgado, A. Glioma grade discrimination with MR diffusion kurtosis imaging: a meta-analysis of diagnostic accuracy. Radiology 287, 119–127 (2018).
Li, W. F. et al. An evidence-based approach to assess the accuracy of intravoxel incoherent motion imaging for the grading of brain tumors. Medicine 97, e13217 (2018).
Kadota, Y. et al. Differentiation between glioblastoma and solitary brain metastasis using neurite orientation dispersion and density imaging. J. Neuroradiol. 47, 197–202 (2020).
Lefranc, M. et al. Perfusion MRI as a neurosurgical tool for improved targeting in stereotactic tumor biopsies. Stereotact. Funct. Neurosurg. 90, 240–247 (2012).
Chakhoyan, A. et al. Validation of vessel size imaging (VSI) in high-grade human gliomas using magnetic resonance imaging, image-guided biopsies, and quantitative immunohistochemistry. Sci. Rep. 9, 2846 (2019).
Smits, M. et al. Repeatability and reproducibility of relative cerebral blood volume measurement of recurrent glioma in a multicentre trial setting. Eur. J. Cancer 114, 89–96 (2019).
Shin, W. et al. Quantitative cerebral perfusion using dynamic susceptibility contrast MRI: evaluation of reproducibility and age- and gender-dependence with fully automatic image postprocessing algorithm. Magn. Reson. Med. 58, 1232–1241 (2007).
Caseiras, G. B. et al. Relative cerebral blood volume measurements of low-grade gliomas predict patient outcome in a multi-institution setting. Eur. J. Radiol. 73, 215–220 (2010).
Jafari-Khouzani, K. et al. Repeatability of cerebral perfusion using dynamic susceptibility contrast MRI in glioblastoma patients. Transl. Oncol. 8, 137–146 (2015).
Bell, L. C. et al. Evaluating multisite rCBV consistency from DSC-MRI imaging protocols and postprocessing software across the NCI quantitative imaging network sites using a digital reference object (DRO). Tomography 5, 110–117 (2019).
Milchenko, M. V. et al. Comparison of perfusion- and diffusion-weighted imaging parameters in brain tumor studies processed using different software platforms. Acad. Radiol. 21, 1294–1303 (2014).
Orsingher, L., Piccinini, S. & Crisi, G. Differences in dynamic susceptibility contrast MR perfusion maps generated by different methods implemented in commercial software. J. Comput. Assist. Tomogr. 38, 647–654 (2014).
Quantitative Imaging Biomarkers Alliance. QIBA Profile: Dynamic susceptibility contrast MRI (DSC-MRI). Stage 2: Consensus profile (QIBA, 2020).
Delgado, A. F. & Delgado, A. F. Discrimination between glioma grades II and III using dynamic susceptibility perfusion MRI: a meta-analysis. AJNR 38, 1348–1355 (2017).
Abrigo, J. M. et al. Magnetic resonance perfusion for differentiating low-grade from high-grade gliomas at first presentation. Cochrane Database Syst. Rev. 1, CD011551 (2018).
Xing, Z. et al. Noninvasive assessment of IDH mutational status in World Health Organization grade II and III astrocytomas using DWI and DSC-PWI combined with conventional MR imaging. AJNR 38, 1138–1144 (2017).
Chuang, M. T., Liu, Y. S., Tsai, Y. S., Chen, Y. C. & Wang, C. K. Differentiating radiation-induced necrosis from recurrent brain tumor using MR perfusion and spectroscopy: a meta-analysis. PLoS ONE 11, e0141438 (2016).
Wang, L. et al. Evaluation of perfusion MRI value for tumor progression assessment after glioma radiotherapy: a systematic review and meta-analysis. Medicine 99, e23766 (2020).
Patel, P. et al. MR perfusion-weighted imaging in the evaluation of high-grade gliomas after treatment: a systematic review and meta-analysis. Neuro Oncol. 19, 118–127 (2017).
Smits, M. Imaging of oligodendroglioma. Br. J. Radiol. 89, 1060 (2016).
Suh, C. H., Kim, H. S., Jung, S. C., Choi, C. G. & Kim, S. J. Perfusion MRI as a diagnostic biomarker for differentiating glioma from brain metastasis: a systematic review and meta-analysis. Eur. Radiol. 28, 3819–3831 (2018).
Xu, W., Wang, Q., Shao, A., Xu, B. & Zhang, J. The performance of MR perfusion-weighted imaging for the differentiation of high-grade glioma from primary central nervous system lymphoma: a systematic review and meta-analysis. PLoS ONE 12, e0173430 (2017).
Choi, S. H. et al. Perfusion MRI as the predictive/prognostic and pharmacodynamic biomarkers in recurrent malignant glioma treated with bevacizumab: a systematic review and a time-to-event meta-analysis. J. Neurooncol. 128, 185–194 (2016).
Hoque, M. M. et al. The cerebral microvasculature: basic and clinical perspectives on stroke and glioma. Microcirculation 28, e12671 (2021).
Quantitative Imaging Biomarkers Alliance. QIBA Profile: DCE-MRI Quantification (DCEMRI-Q). Stage 1: public comment (QIBA, 2017).
Shukla-Dave, A. et al. Quantitative Imaging Biomarkers Alliance (QIBA) recommendations for improved precision of DWI and DCE-MRI derived biomarkers in multicenter oncology trials. J. Magn. Reson. Imaging 49, e101–e121 (2019).
Okuchi, S. et al. Diagnostic accuracy of dynamic contrast-enhanced perfusion MRI in stratifying gliomas: a systematic review and meta-analysis. Cancer Med. 8, 5564–5573 (2019).
Alsop, D. C. et al. Recommended implementation of arterial spin-labeled perfusion MRI for clinical applications: a consensus of the ISMRM perfusion study group and the European consortium for ASL in dementia. Magnetic Reson. Med. 73, 102–116 (2015).
Quantitative Imaging Biomarkers Alliance. EIBIR/QIBA ASL Biomarker Committee (QIBA, 2019).
Kong, L., Chen, H., Yang, Y. & Chen, L. A meta-analysis of arterial spin labelling perfusion values for the prediction of glioma grade. Clin. Radiol. 72, 255–261 (2017).
Brendle, C. et al. Glioma grading and determination of IDH mutation status and ATRX loss by DCE and ASL perfusion. Clin. Neuroradiol. 28, 421–428 (2018).
Yoo, R. E. et al. Arterial spin labeling perfusion-weighted imaging aids in prediction of molecular biomarkers and survival in glioblastomas. Eur. Radiol. 30, 1202–1211 (2020).
Bertholdo, D., Watcharakorn, A. & Castillo, M. Brain proton magnetic resonance spectroscopy: introduction and overview. Neuroimaging Clin. N. Am. 23, 359–380 (2013).
Wilson, M. et al. Methodological consensus on clinical proton MRS of the brain: review and recommendations. Magn. Reson. Med. 82, 527–550 (2019).
Howe, F. A. et al. Metabolic profiles of human brain tumors using quantitative in vivo 1H magnetic resonance spectroscopy. Magn. Reson. Med. 49, 223–232 (2003).
Choi, C. et al. 2-hydroxyglutarate detection by magnetic resonance spectroscopy in IDH-mutated patients with gliomas. Nat. Med. 18, 624–629 (2012).
Ishimaru, H. et al. Differentiation between high-grade glioma and metastatic brain tumor using single-voxel proton MR spectroscopy. Eur. Radiol. 11, 1784–1791 (2001).
Caivano, R. et al. 3 Tesla magnetic resonance spectroscopy: cerebral gliomas vs. metastatic brain tumors. Our experience and review of the literature. Int. J. Neurosci. 123, 537–543 (2013).
Esteve, F., Rubin, C., Grand, S., Kolodie, H. & Le Bas, J. F. Transient metabolic changes observed with proton MR spectroscopy in normal human brain after radiation therapy. Int. J. Radiat. Oncol. Biol. Phys. 40, 279–286 (1998).
Suh, C. H., Kim, H. S., Jung, S. C., Choi, C. G. & Kim, S. J. 2-Hydroxyglutarate MR spectroscopy for prediction of isocitrate dehydrogenase mutant glioma: a systemic review and meta-analysis using individual patient data. Neuro Oncol. 20, 1573–1583 (2018).
Andronesi, O. C. et al. Treatment response assessment in IDH-mutant glioma patients by noninvasive 3D functional spectroscopic mapping of 2-hydroxyglutarate. Clin. Cancer Res. 22, 1632–1641 (2016).
de la Fuente, M. I. et al. Integration of 2-hydroxyglutarate-proton magnetic resonance spectroscopy into clinical practice for disease monitoring in isocitrate dehydrogenase-mutant glioma. Neuro Oncol. 18, 283–290 (2016).
Choi, C. et al. Prospective longitudinal analysis of 2-hydroxyglutarate magnetic resonance spectroscopy identifies broad clinical utility for the management of patients with IDH-mutant glioma. J. Clin. Oncol. 34, 4030–4039 (2016).
Suh, C. H. et al. False-positive measurement at 2-hydroxyglutarate MR spectroscopy in isocitrate dehydrogenase wild-type glioblastoma: a multifactorial analysis. Radiology 291, 752–762 (2019).
Branzoli, F. et al. Cystathionine as a marker for 1p/19q codeleted gliomas by in vivo magnetic resonance spectroscopy. Neuro Oncol. 21, 765–774 (2019).
Gillies, R. J., Kinahan, P. E. & Hricak, H. Radiomics: images are more than pictures, they are data. Radiology 278, 563–577 (2016). Introduction to radiomics.
Lambin, P. et al. Radiomics: the bridge between medical imaging and personalized medicine. Nat. Rev. Clin. Oncol. 14, 749–762 (2017). Quality standards for radiomics studies.
Diehn, M. et al. Identification of noninvasive imaging surrogates for brain tumor gene-expression modules. Proc. Natl Acad. Sci. USA 105, 5213–5218 (2008).
TCGA Glioma Phenotpye Research Group. VASARI research project. Cancer Imaging Archive https://wiki.cancerimagingarchive.net/display/Public/VASARI+Research+Project (2020)
Gutman, D. A. et al. MR imaging predictors of molecular profile and survival: multi-institutional study of the TCGA glioblastoma data set. Radiology 267, 560–569 (2013).
Gevaert, O. et al. Glioblastoma multiforme: exploratory radiogenomic analysis by using quantitative image features. Radiology 273, 168–174 (2014).
Lohmann, P. et al. Radiomics in neuro-oncology: basics, workflow, and applications. Methods 188, 112–121 (2020). Comprehensive overview and explanation of radiomics.
Rudie, J. D., Rauschecker, A. M., Bryan, R. N., Davatzikos, C. & Mohan, S. Emerging applications of artificial intelligence in neuro-oncology. Radiology 290, 607–618 (2019). Overview of applications of radiomics and other computational methods in neuro-oncological imaging.
Starmans, M. P. A. et al. in Handbook of Medical Image Computing and Computer Assisted Intervention (eds Zhou, S. K., Rueckert, D. & Fichtinger, G.) 429–456 (Academic, 2020).
Wang, Q., Lei, D., Yuan, Y. & Zhao, H. Accuracy of magnetic resonance imaging texture analysis in differentiating low-grade from high-grade gliomas: systematic review and meta-analysis. BMJ Open 9, e027144 (2019).
Yang, Y. et al. Glioma grading on conventional MR images: a deep learning study with transfer learning. Front. Neurosci. 12, 804 (2018).
Kim, J. Y. et al. Incorporating diffusion- and perfusion-weighted MRI into a radiomics model improves diagnostic performance for pseudoprogression in glioblastoma patients. Neuro Oncol. 21, 404–414 (2019).
Hu, X., Wong, K. K., Young, G. S., Guo, L. & Wong, S. T. Support vector machine multiparametric MRI identification of pseudoprogression from tumor recurrence in patients with resected glioblastoma. J. Magn. Reson. Imaging 33, 296–305 (2011).
Jang, B. S., Jeon, S. H., Kim, I. H. & Kim, I. A. Prediction of pseudoprogression versus progression using machine learning algorithm in glioblastoma. Sci. Rep. 8, 12516 (2018).
Lohmann, P. et al. FET PET radiomics for differentiating pseudoprogression from early tumor progression in glioma patients post-chemoradiation. Cancers 12, 3835 (2020).
Kickingereder, P. et al. Radiomic profiling of glioblastoma: identifying an imaging predictor of patient survival with improved performance over established clinical and radiologic risk models. Radiology 280, 880–889 (2016).
Macyszyn, L. et al. Imaging patterns predict patient survival and molecular subtype in glioblastoma via machine learning techniques. Neuro Oncol. 18, 417–425 (2016).
Li, Q. et al. A fully-automatic multiparametric radiomics model: towards reproducible and prognostic imaging signature for prediction of overall survival in glioblastoma multiforme. Sci. Rep. 7, 14331 (2017).
Li, Y. et al. Radiomic features predict Ki-67 expression level and survival in lower grade gliomas. J. Neurooncol. 135, 317–324 (2017).
Chang, P. D. et al. A multiparametric model for mapping cellularity in glioblastoma using radiographically localized biopsies. AJNR 38, 890–898 (2017).
Goyal, A. et al. The T2-FLAIR-mismatch sign as an imaging biomarker for IDH and 1p/19q status in diffuse low-grade gliomas: a systematic review with a Bayesian approach to evaluation of diagnostic test performance. Neurosurg. Focus. 47, E13 (2019).
Smits, M. & van den Bent, M. J. Imaging correlates of adult glioma genotypes. Radiology 284, 316–331 (2017).
Chang, P. et al. Deep-learning convolutional neural networks accurately classify genetic mutations in gliomas. AJNR 39, 1201–1207 (2018).
Verma, R. et al. tumor habitat-derived radiomic features at pretreatment MRI that are prognostic for progression-free survival in glioblastoma are associated with key morphologic attributes at histopathologic examination: a feasibility study. Radiol. Artif. Intell. 2, e190168 (2020).
Zhao, J. et al. Diagnostic accuracy and potential covariates for machine learning to identify IDH mutations in glioma patients: evidence from a meta-analysis. Eur. Radiol. 30, 4664–4674 (2020).
Bhandari, A. P., Liong, R., Koppen, J., Murthy, S. V. & Lasocki, A. Noninvasive determination of IDH and 1p19q status of lower-grade gliomas using MRI radiomics: a systematic review. AJNR 42, 94–101 (2021).
van der Voort, S. R. et al. Predicting the 1p/19q codeletion status of presumed low-grade glioma with an externally validated machine learning algorithm. Clin. Cancer Res. 25, 7455–7462 (2019).
Liu, X. et al. A radiomic signature as a non-invasive predictor of progression-free survival in patients with lower-grade gliomas. Neuroimage Clin. 20, 1070–1077 (2018).
Tong, E., McCullagh, K. L. & Iv, M. Advanced imaging of brain metastases: from augmenting visualization and improving diagnosis to evaluating treatment response. Front. Neurol. 11, 270 (2020).
Petrujkic, K. et al. Computational quantitative MR image features – a potential useful tool in differentiating glioblastoma from solitary brain metastasis. Eur. J. Radiol. 119, 108634 (2019).
Ortiz-Ramon, R., Larroza, A., Ruiz-Espana, S., Arana, E. & Moratal, D. Classifying brain metastases by their primary site of origin using a radiomics approach based on texture analysis: a feasibility study. Eur. Radiol. 28, 4514–4523 (2018).
Larroza, A. et al. Support vector machine classification of brain metastasis and radiation necrosis based on texture analysis in MRI. J. Magn. Reson. Imaging 42, 1362–1368 (2015).
Charron, O. et al. Automatic detection and segmentation of brain metastases on multimodal MR images with a deep convolutional neural network. Comput. Biol. Med. 95, 43–54 (2018).
Liu, Y. et al. A deep convolutional neural network-based automatic delineation strategy for multiple brain metastases stereotactic radiosurgery. PLoS ONE 12, e0185844 (2017).
Liu, Y. et al. Automatic metastatic brain tumor segmentation for stereotactic radiosurgery applications. Phys. Med. Biol. 61, 8440–8461 (2016).
Kickingereder, P. et al. Automated quantitative tumour response assessment of MRI in neuro-oncology with artificial neural networks: a multicentre, retrospective study. Lancet Oncol. 20, 728–740 (2019).
Kim, D. W., Jang, H. Y., Kim, K. W., Shin, Y. & Park, S. H. Design characteristics of studies reporting the performance of artificial intelligence algorithms for diagnostic analysis of medical images: results from recently published papers. Korean J. Radiol. 20, 405–410 (2019).
Park, J. E. et al. A systematic review reporting quality of radiomics research in neuro-oncology: toward clinical utility and quality improvement using high-dimensional imaging features. BMC Cancer 20, 29 (2020).
Ma, D. et al. Magnetic resonance fingerprinting. Nature 495, 187–192 (2013).
Zhou, J., Heo, H. Y., Knutsson, L., van Zijl, P. C. M. & Jiang, S. APT-weighted MRI: techniques, current neuro applications, and challenging issues. J. Magn. Reson. Imaging 50, 347–364 (2019).
Ladd, M. E. et al. Pros and cons of ultra-high-field MRI/MRS for human application. Prog. Nucl. Magn. Reson. Spectrosc. 109, 1–50 (2018).
Carre, A. et al. Standardization of brain MR images across machines and protocols: bridging the gap for MRI-based radiomics. Sci. Rep. 10, 12340 (2020).
Ellingson, B. M. et al. Consensus recommendations for a standardized Brain Tumor Imaging Protocol in clinical trials. Neuro Oncol. 17, 1188–1198 (2015).
Kaufmann, T. J. et al. Consensus recommendations for a standardized brain tumor imaging protocol for clinical trials in brain metastases. Neuro Oncol. 22, 757–772 (2020).
Boxerman, J. L. et al. Consensus recommendations for a dynamic susceptibility contrast MRI protocol for use in high-grade gliomas. Neuro Oncol. 22, 1262–1275 (2020).
Zwanenburg, A. et al. The image biomarker standardization initiative: standardized quantitative radiomics for high-throughput image-based phenotyping. Radiology 295, 328–338 (2020).
European Society of Radiology. ESR position paper on imaging biobanks. Insights Imaging 6, 403–410 (2015).
Wilkinson, M. D. et al. The FAIR Guiding Principles for scientific data management and stewardship. Sci. Data 3, 160018 (2016).
Al-Mubarak, H. et al. Stacked in-plane histology for quantitative validation of non-invasive imaging biomarkers: application to an infiltrative brain tumour model. J. Neurosci. Methods 326, 108372 (2019).
Bakas, S. et al. Advancing The Cancer Genome Atlas glioma MRI collections with expert segmentation labels and radiomic features. Sci. Data 4, 170117 (2017).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
M.S. declares speaker fees paid to her institution from GE Healthcare and honorarium paid to her institution from Parexel Ltd. for trial review of EORTC-1410.
Additional information
Peer review information
Nature Reviews Neurology thanks A. Waldman, who co-reviewed with G. Thompson, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
European Imaging Biomarkers Alliance (EIBALL): https://www.myesr.org/research/esr-research-committee
Open Source Initiative for Perfusion Imaging (OSIPI): https://www.osipi.org
Quantitative Imagine Biomarkers Alliance (QIBA): https://www.rsna.org/research/quantitative-imaging-biomarkers-alliance
Quantitative Imaging Biomarkers Alliance (QIBA) wiki: https://qibawiki.rsna.org/index.php/Main_Page
The Cancer Genome Atlas: https://www.cancer.gov/about-nci/organization/ccg/research/structural-genomics/tcga
The Cancer Imaging Archive: https://www.cancerimagingarchive.net
Glossary
- Repeatability
-
The frequency with which the same measurement under the same conditions (for example, same scanner, participant and rater) provides the same result.
- Reproducibility
-
The frequency with which the same measurement performed under different conditions (for example, on a different scanner or by a different rater) provides the same result.
- Phantom
-
An artificial construct, either physical or digital, that provides a reference standard for validation and calibration.
- Sensitivity
-
The proportion results from a given test that are true positives.
- Specificity
-
The proportion results from a given test that are true negatives.
- Brownian motion
-
The random motion of particles within a medium.
- Ki-67 labelling index
-
A marker of cellular proliferation based on immunohistochemical assessment of the expression of the Ki-67 protein.
- Mean kurtosis
-
An estimate of the non-gaussianity of water diffusion resulting from the presence of diffusion barriers and compartments within tissue structure; higher mean kurtosis indicates higher tissue microstructural complexity.
- Deep learning
-
A class of machine learning based on artificial neural networks that are inspired by biological networks of learning and information processing; ‘deep’ refers to the use of multiple layers in the network.
- Overfitting
-
A phenomenon that occurs when the match between the classification model and the data set is too perfect, resulting in a model that cannot be generalized to any other data set.
- Dimensionality
-
The number of dimensions included in a computational model; in radiomics, this term relates primarily to the number of imaging features, each feature being one dimension.
- Noise
-
The unexplained variation or randomness in a computational model.
- Radiomics quality score
-
A score system used to assess the quality of radiomics studies; consists of 16 items covering methodology, reporting, clinical utility and contribution to open science.
Rights and permissions
About this article
Cite this article
Smits, M. MRI biomarkers in neuro-oncology. Nat Rev Neurol 17, 486–500 (2021). https://doi.org/10.1038/s41582-021-00510-y
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41582-021-00510-y
This article is cited by
-
MRIO: the Magnetic Resonance Imaging Acquisition and Analysis Ontology
Neuroinformatics (2024)
-
How to evaluate perfusion imaging in post-treatment glioma: a comparison of three different analysis methods
Neuroradiology (2024)
-
Arterial spin-labeled magnetic resonance perfusion imaging in prediction of pediatric brain tumors grading: inter-observer agreement
Egyptian Journal of Radiology and Nuclear Medicine (2023)
-
Imaging biomarkers for clinical applications in neuro-oncology: current status and future perspectives
Biomarker Research (2023)
-
Nano-imaging agents for brain diseases: Environmentally responsive imaging and therapy
Nano Research (2023)