Abstract
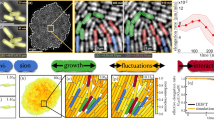
Cells that grow in confined spaces eventually build up mechanical compressive stress. This growth-induced pressure decreases cell growth. Growth-induced pressure is important in a multitude of contexts, including cancer1,2,3, microbial infections4 and biofouling5; yet, our understanding of its origin and molecular consequences remains limited. Here we combine microfluidic confinement of the yeast Saccharomyces cerevisiae6 with rheological measurements using genetically encoded multimeric nanoparticles7 to reveal that growth-induced pressure is accompanied with an increase in a key cellular physical property: macromolecular crowding. We develop a fully calibrated model that predicts how increased macromolecular crowding hinders protein expression and thus diminishes cell growth. This model is sufficient to explain the coupling of growth rate to pressure without the need for specific molecular sensors or signalling cascades. As molecular crowding is similar across all domains of life, this could be a deeply conserved mechanism of biomechanical feedback that allows environmental sensing originating from the fundamental physical properties of cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Source data are available for this paper. All other data that support the plots within this paper and other findings of this study are available from the corresponding authors upon reasonable request.
Code availability
All the codes used in this paper are available from the corresponding authors upon reasonable request.
References
Helmlinger, G., Netti, P. A., Lichtenbeld, H. C., Melder, R. J. & Jain, R. K. Solid stress inhibits the growth of multicellular tumor spheroids. Nat. Biotechnol. 15, 778–783 (1997).
Alessandri, K. et al. Cellular capsules as a tool for multicellular spheroid production and for investigating the mechanics of tumor progression in vitro. Proc. Natl Acad. Sci. USA 110, 14843–14848 (2013).
Rizzuti, I. et al. Mechanical control of cell proliferation increases resistance to chemotherapeutic agents. Phys. Rev. Lett. 125, 128103 (2020).
Bonazzi, D. et al. Intermittent pili-mediated forces fluidize Neisseria meningitidis aggregates promoting vascular colonization. Cell 174, 143–155.e16 (2018).
Delarue, M. et al. Self-driven jamming in growing microbial populations. Nat. Phys. 12, 762–766 (2016).
Holt, L. J., Hallatschek, O. & Delarue, M. Mechano-chemostats to study the effects of compressive stress on yeast. Methods Cell. Biol. 147, 215–231 (2018).
Delarue, M. et al. mTORC1 controls phase separation and the biophysical properties of the cytoplasm by tuning crowding. Cell 174, 338–349.e20 (2018).
Streichan, S. J., Hoerner, C. R., Schneidt, T., Holzer, D. & Hufnagel, L. Spatial constraints control cell proliferation in tissues. Proc. Natl Acad. Sci. USA 111, 5586–5591 (2014).
Bengough, A. G., Croser, C. & Pritchard, J. A biophysical analysis of root growth under mechanical stress. Plant Soil 189, 155–164 (1997).
Fal, K. et al. Tissue folding at the organ–meristem boundary results in nuclear compression and chromatin compaction. Proc. Natl Acad. Sci. USA 118, e2017859118 (2021).
Rivera-Yoshida, N., Arias Del Angel, J. A. & Benítez, M. Microbial multicellular development: mechanical forces in action. Curr. Opin. Genet. Dev. 51, 37–45 (2018).
Asally, M. et al. Localized cell death focuses mechanical forces during 3D patterning in a biofilm. Proc. Natl Acad. Sci. USA 109, 18891–18896 (2012).
Trejo, M. et al. Elasticity and wrinkled morphology of Bacillus subtilis pellicles. Proc. Natl Acad. Sci. USA 110, 2011–2016 (2013).
Stewart, P. S. & Robertson, C. R. Microbial growth in a fixed volume: studies with entrapped Escherichia coli. Appl. Microbiol. Biotechnol. 30, 34–40 (1989).
Chu, E. K., Kilic, O., Cho, H., Groisman, A. & Levchenko, A. Self-induced mechanical stress can trigger biofilm formation in uropathogenic Escherichia coli. Nat. Commun. 9, 4087 (2018).
Nam, S. et al. Cell cycle progression in confining microenvironments is regulated by a growth-responsive TRPV4-PI3K/Akt-p27Kip1 signaling axis. Sci. Adv. 5, eaaw6171 (2019).
Delarue, M. et al. SCWISh network is essential for survival under mechanical pressure. Proc. Natl Acad. Sci. USA 114, 13465–13470 (2017).
Miermont, A. et al. Severe osmotic compression triggers a slowdown of intracellular signaling, which can be explained by molecular crowding. Proc. Natl Acad. Sci. USA 110, 5725–5730 (2013).
Vibhute, M. A. et al. Transcription and translation in cytomimetic protocells perform most efficiently at distinct macromolecular crowding conditions. ACS Synth. Biol. 9, 2797–2807 (2020).
Nettesheim, G. et al. Macromolecular crowding acts as a physical regulator of intracellular transport. Nat. Phys. 16, 1144–1151 (2020).
Fernandez-De-Cossio-DIaz, J. & Vazquez, A. A physical model of cell metabolism. Sci. Rep. 8, 8349 (2018).
Petelenz-Kurdziel, E. et al. Quantification of cell volume changes upon hyperosmotic stress in Saccharomyces cerevisiae. Integr. Biol. 3, 1120–1126 (2011).
Görlich, D. & Kutay, U. Transport between the cell nucleus and the cytoplasm. Annu. Rev. Cell Dev. Biol. 15, 607–660 (2003).
de, I. M., Marechal, P.-A. & Gervais, P. Passive response of Saccharomyces cerevisiae to osmotic shifts: cell volume variations depending on the physiological state. Biochem. Biophys. Res. Commun. 227, 519–523 (1996).
Borodavka, A. et al. Sizes of long RNA molecules are determined by the branching patterns of their secondary structures. Biophys. J. 111, 2077–2085 (2016).
Busti, S., Coccetti, P., Alberghina, L. & Vanoni, M. Glucose signaling-mediated coordination of cell growth and cell cycle in Saccharomyces cerevisiae. Sensors 10, 6195–6240 (2010).
Tamás, M. J. & Hohmann, S. The osmotic stress response of Saccharomyces cerevisiae. in Yeast Stress Responses. Topics in Current Genetics Vol. 1 (eds Hohmann S. & Mager W. H.) 121–200 (Springer, 2003).
Maayan, I. & Engelberg, D. The yeast MAPK Hog1 is not essential for immediate survival under osmostress. FEBS Lett. 583, 2015–2020 (2009).
Rojas, E. R., Huang, K. C. & Theriot, J. A. Homeostatic cell growth is accomplished mechanically through membrane tension inhibition of cell-wall synthesis. Cell Syst. 5, 578–590.e6 (2017).
Puchner, E. M., Walter, J. M., Kasper, R., Huang, B. & Lim, W. A. Counting molecules in single organelles with superresolution microscopy allows tracking of the endosome maturation trajectory. Proc. Natl Acad. Sci. USA 110, 16015–16020 (2013).
Takamori, S. et al. Molecular anatomy of a trafficking organelle. Cell 127, 831–846 (2006).
Vella, D., Ajdari, A., Vaziri, A. & Boudaoud, A. The indentation of pressurized elastic shells: from polymeric capsules to yeast cells. J. R. Soc. Interface 9, 448–455 (2012).
Albersheim, P., Darvill, A., Roberts, K., Sederoff, R. & Staehelin A. Plant Cell Walls (Garland Science, 2010).
Schaber, J. et al. Biophysical properties of Saccharomyces cerevisiae and their relationship with HOG pathway activation. Eur. Biophys. J. 39, 1547–1556 (2010).
Levin, D. E. Cell wall integrity signaling in Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 69, 262–291 (2005).
Ellis, R. J. Macromolecular crowding: obvious but underappreciated. Trends Biochem. Sci 26, 597–604 (2001).
Sbalzarini, I. F. & Koumoutsakos, P. Feature point tracking and trajectory analysis for video imaging in cell biology. J. Struct. Biol. 151, 182–195 (2005).
Griffith, J., Mari, M., De Mazière, A. & Reggiori, F. A cryosectioning procedure for the ultrastructural analysis and the immunogold labelling of yeast Saccharomyces cerevisiae. Traffic 9, 1060–1072 (2008).
Hutter, J. L., & Bechhoefer, J. Calibration of atomic‐force microscope tips. Rev. Sci. Instrum. 64, 1868 (1993).
Acknowledgements
We thank E. Kassianidou for the initial help with the laser ablation experiments. We thank A. Liang, C. Petzold and K. Dancel-Manning at the NYULH DART Microscopy Laboratory for consultation and assistance with the transmission electron microscopy work; this core is partially funded by the NYU Cancer Center Support Grant NIH/NCI P30CA016087. The technological realizations and associated research works were partly supported by the French RENATECH network (M.D.). L.J.H. was funded by NIH grants R01 GM132447 and R37 CA240765, the American Cancer Society, the Pershing Square Sohn Cancer Research Award and Chan Zuckerberg Initiative. We thank E. Rojas for fruitful discussions. L.J.H. and M.D. thank the FACE foundation for travel support.
Author information
Authors and Affiliations
Contributions
B.A. and M.D. designed and performed the experiments and data analysis. C.F.-D. and E.D. performed the AFM experiments. L.J.H. designed the strains used in the study. B.A., L.J.H. and M.D. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Physics thanks Pascal Hersen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Device used in the microfluidic experiments.
a. Cells are loaded in a culture chamber connected on its sides to narrow channels that are used to set the chemical environment. Confined growth lead to the buildup of GIP, which is measured through the deformation of the elastic chamber. b. The culture chamber is, similar to the device presented in a., connected to a set of narrow channels to set the chemical environment. A valve is actuated to confine the cell population and allow it to build up GIP. We estimate GIP by measuring the deformation of the PDMS membrane. Opening of the valve leads to a relaxation of GIP.
Extended Data Fig. 2 Impact of osmotic shock.
Ratio of nucleus and cytoplasm volume under osmotic shock and growth induced pressure.
Extended Data Fig. 3 Linear and exponential fits on the diffusion as a function of GIP data.
The score for each fit is presented. We superimposed the prediction of diffusion as a function of GIP for the 40nm-GEMs, as well as the corresponding score.
Extended Data Fig. 4 40nm-GEMs diffusion as a function of cell volume.
Model fit (Eq. (13)) of the experimental data to extract \(\xi _{40} = 7.4 \pm 2.5\) (r2 = 0.99).
Extended Data Fig. 5 Contribution of density and chamber volume change in the growth rate.
For simplicity, we denoted kg as the growth rate, \(k_\rho = \partial _t\rho /\rho\) as the contribution of cell density ρ to growth rate, and \(k_V = \partial _PV/V\;\partial _tP\) the contribution of the volume of the chamber V.
Extended Data Fig. 6 Induction time of PADH2-mCherry.
The induction time is plotted as a function of time. Inset: induction time plotted as a function of growth rate.
Extended Data Fig. 7 Protein production rate as a function of GIP.
In orange: data from the PADH2-mCherry promoter. In blue: data from the PHIS3-GFP promoter.
Extended Data Fig. 8 Laser ablation experiments and calibration of turgor pressure.
a. We used a high-intensity laser pulse to make a hole in a cell, forcing its deflation. The cell radius changes as a function of turgor pressure, cell wall elasticity, and thickness of the cell wall. b. Cells were punctured with a laser, resulting in a decrease in cell radius proportional to turgor pressure. The similar decrease in radius of WT and hog1Δ cells indicates that, absent osmotic perturbation, these cells develop similar amounts of turgor pressure. The decrease in radius of osmotically compressed (c = 1 M sorbitol) hog1Δ cells indicates that these cells are still pressurized, albeit to a reduced extent. c. We used transmission electron microscopy to measure the cell wall thickness. d. We performed AFM experiments, using small deformations (below 0.2 μm) to extract the effective elasticity of the cell. This elasticity provided a mathematical function of turgor pressure, cell wall elasticity, and cell wall thickness.
Extended Data Fig. 9 Growth induced pressure as a function of time.
In blue, for the WT cells. In orange, for the hog1Δ cells.
Extended Data Fig. 10 Measurement of the fluorescence intensity in the center versus the edge of the chamber prior to induction of PADH2-mCherry.
The data shows an insignificant (p-value = 0.16) 2.3% difference.
Supplementary information
Supplementary Information
Supplementary information containing a table of strains used along with the mathematical modelling.
Source data
Source Data Fig. 1
Raw data for Fig. 1b–e.
Source Data Fig. 2
Raw data for Fig. 2a,c,d.
Source Data Fig. 3
Raw data for Fig. 3a–c.
Source Data Fig. 4
Raw data for Fig. 4b–e.
Source Data Extended Data Fig. 2
Raw data for Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Raw data for Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Raw data for Extended Data Fig. 4.
Source Data Extended Data Fig. 5
Raw data for Extended Data Fig. 5.
Source Data Extended Data Fig. 6
Raw data for Extended Data Fig. 6.
Source Data Extended Data Fig. 7
Raw data for Extended Data Fig. 7.
Source Data Extended Data Fig. 9
Raw data for Extended Data Fig. 9.
Rights and permissions
About this article
Cite this article
Alric, B., Formosa-Dague, C., Dague, E. et al. Macromolecular crowding limits growth under pressure. Nat. Phys. 18, 411–416 (2022). https://doi.org/10.1038/s41567-022-01506-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41567-022-01506-1
This article is cited by
-
Viscosity-dependent control of protein synthesis and degradation
Nature Communications (2024)
-
Collective mechano-response dynamically tunes cell-size distributions in growing bacterial colonies
Communications Physics (2023)
-
Geometric constraint-triggered collagen expression mediates bacterial-host adhesion
Nature Communications (2023)
-
Comparison between carbohydrate and salt-based macromolecular crowders for cell preservation at higher temperatures
3 Biotech (2023)