Abstract
Norovirus infection can cause gastrointestinal disease in humans. Development of therapies and vaccines against norovirus have been limited by the lack of a suitable and reliable animal model. Here we established rhesus macaques as an animal model for human norovirus infection. We show that rhesus macaques are susceptible to oral infection with human noroviruses from two different genogroups. Variation in duration of virus shedding (days to weeks) between animals, evolution of the virus over the time of infection, induction of virus-specific adaptive immune responses, susceptibility to reinfection and preferential replication of norovirus in the jejunum of rhesus macaques was similar to infection reported in humans. We found minor pathological signs and changes in epithelial cell surface glycosylation patterns in the small intestine during infection. Detection of viral protein and RNA in intestinal biopsies confirmed the presence of the virus in chromogranin A-expressing epithelial cells, as it does in humans. Thus, rhesus macaques are a promising non-human primate model to evaluate vaccines and therapeutics against norovirus disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data supporting the findings of this study are available within the paper, its Extended Data or Source Data files. Representative microscopy images are included in the main or extended data figures. Additional microscopy image files are available from the corresponding author upon request. Sequence data for human norovirus isolated from rhesus macaques have been deposited in GenBank with accession numbers OR397766–OR397788. Source data are provided with this paper.
Change history
15 February 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41564-024-01636-7
References
Woodward, J., Gkrania-Klotsas, E. & Kumararatne, D. Chronic norovirus infection and common variable immunodeficiency. Clin. Exp. Immunol. 188, 363–370 (2017).
Lu, M. C., Lin, S. C., Hsu, Y. H. & Chen, S. Y. Epidemiology, clinical features, and unusual complications of norovirus infection in Taiwan: what we know after rotavirus vaccines. Pathogens 11, 451 (2022).
Sanfilippo, E., Habeshian, K., Cotton, C. H. & Kirkorian, A. Y. Severe reactive infectious mucocutaneous eruption mimicking drug-induced epidermal necrolysis triggered by norovirus. Pediatr. Dermatol. https://doi.org/10.1111/pde.15370 (2023).
Yilmaz, N. & Yuksel, S. Hemolytic uremic syndrome associated with norovirus gastroenteritis: case report and literature review. Nephron 146, 489–493 (2022).
Pires, S. M. et al. Aetiology-specific estimates of the global and regional incidence and mortality of diarrhoeal diseases commonly transmitted through food. PLoS ONE 10, e0142927 (2015).
Bartsch, S. M., Lopman, B. A., Ozawa, S., Hall, A. J. & Lee, B. Y. Global economic burden of norovirus gastroenteritis. PLoS ONE 11, e0151219 (2016).
Kumazaki, M. & Usuku, S. Influence of herd immunity on norovirus: a long-term field study of repeated viral gastroenteritis outbreaks at the same facilities. BMC Infect. Dis. 23, 1–15 (2023).
Mattison, C. P. et al. Childcare and school acute gastroenteritis outbreaks: 2009–2020. Pediatrics 150, e2021056002 (2022).
Sosnovtsev, S. V. et al. Cleavage map and proteolytic processing of the murine norovirus nonstructural polyprotein in infected cells. J. Virol. 80, 7816–7831 (2006).
Chhabra, P. et al. Updated classification of norovirus genogroups and genotypes. J. Gen. Virol. 100, 1393–1406 (2019).
Ettayebi, K. et al. Replication of human noroviruses in stem cell-derived human enteroids. Science 353, 1387–1393 (2016).
Jones, M. K. et al. Enteric bacteria promote human and mouse norovirus infection of B cells. Science 346, 755–759 (2014).
Van Dycke, J. et al. A robust human norovirus replication model in zebrafish larvae. PLoS Pathog. 15, e1008009 (2019).
Taube, S. et al. A mouse model for human norovirus. mBiohttps://doi.org/10.1128/mBio.00450-13 (2013).
Cheetham, S. et al. Pathogenesis of a genogroup II human norovirus in gnotobiotic pigs. J. Virol. 80, 10372–10381 (2006).
Souza, M., Azevedo, M. S. P., Jung, K., Cheetham, S. & Saif, L. J. Pathogenesis and immune responses in gnotobiotic calves after infection with the genogroup II.4-HS66 strain of human norovirus. J. Virol. 82, 1777–1786 (2008).
Bok, K. et al. Chimpanzees as an animal model for human norovirus infection and vaccine development. Proc. Natl Acad. Sci. USA 108, 325–330 (2011).
Hutson, A. M., Atmar, R. L., Marcus, D. M. & Estes, M. K. Norwalk virus-like particle hemagglutination by binding to h histo-blood group antigens. J. Virol. 77, 405–415 (2003).
Marionneau, S. et al. Norwalk virus binds to histo-blood group antigens present on gastroduodenal epithelial cells of secretor individuals. Gastroenterology 122, 1967–1977 (2002).
Rockx, B. H., Bogers, W. M., Heeney, J. L., van Amerongen, G. & Koopmans, M. P. Experimental norovirus infections in non-human primates. J. Med. Virol. 75, 313–320 (2005).
Tohma, K., Lepore, C. J., Gao, Y. M., Ford-Siltz, L. A. & Parra, G. I. Population genomics of GII.4 noroviruses reveal complex diversification and new antigenic sites involved in the emergence of pandemic strains. mBio 10, 10–1128 (2019).
Green, K. Y. et al. Human norovirus targets enteroendocrine epithelial cells in the small intestine. Nat. Commun. 11, 2759 (2020).
Kendra, J. A., Tohma, K. & Parra, G. I. Global and regional circulation trends of norovirus genotypes and recombinants, 1995–2019: a comprehensive review of sequences from public databases. Rev. Med. Virol. 32, e2354 (2022).
Lindesmith, L. et al. Human susceptibility and resistance to Norwalk virus infection. Nat. Med. 9, 548–553 (2003).
Hutson, A. M., Airaud, F., LePendu, J., Estes, M. K. & Atmar, R. L. Norwalk virus infection associates with secretor status genotyped from sera. J. Med. Virol. 77, 116–120 (2005).
Tan, M. et al. Outbreak studies of a GII-3 and a GII-4 norovirus revealed an association between HBGA phenotypes and viral infection. J. Med. Virol. 80, 1296–1301 (2008).
Atmar, R. L. et al. Determination of the 50% human infectious dose for Norwalk virus. J. Infect. Dis. 209, 1016–1022 (2014).
Teunis, P. F. et al. Shedding of norovirus in symptomatic and asymptomatic infections. Epidemiol. Infect. 143, 1710–1717 (2015).
Costantini, V. P. et al. Epidemiologic, virologic, and host genetic factors of norovirus outbreaks in long-term care facilities. Clin. Infect. Dis. 62, 1–10 (2016).
Echenique, I. A. et al. Prolonged norovirus infection after pancreas transplantation: a case report and review of chronic norovirus. Transpl. Infect. Dis. 18, 98–104 (2016).
Saif, M. A. et al. Chronic norovirus infection in pediatric hematopoietic stem cell transplant recipients: a cause of prolonged intestinal failure requiring intensive nutritional support. Pediatr. Transpl. 15, 505–509 (2011).
Parrino, T. A., Schreiber, D. S., Trier, J. S., Kapikian, A. Z. & Blacklow, N. R. Clinical immunity in acute gastroenteritis caused by Norwalk agent. N. Engl. J. Med. 297, 86–89 (1977).
Wyatt, R. G. et al. Comparison of three agents of acute infectious nonbacterial gastroenteritis by cross-challenge in volunteers. J. Infect. Dis. 129, 709–714 (1974).
Karandikar, U. C. et al. Detection of human norovirus in intestinal biopsies from immunocompromised transplant patients. J. Gen. Virol. 97, 2291–2300 (2016).
Hutson, A. M., Atmar, R. L., Graham, D. Y. & Estes, M. K. Norwalk virus infection and disease is associated with ABO histo-blood group type. J. Infect. Dis. 185, 1335–1337 (2002).
Meyer, E., Ebner, W., Scholz, R., Dettenkofer, M. & Daschner, F. D. Nosocomial outbreak of norovirus gastroenteritis and investigation of ABO histo-blood group type in infected staff and patients. J. Hosp. Infect. 56, 64–66 (2004).
Huang, P. W. et al. Noroviruses bind to human ABO, Lewis, and secretor histo-blood group antigens: identification of 4 distinct strain-specific patterns. J. Infect. Dis. 188, 19–31 (2003).
Huang, P. W. et al. Norovirus and histo-blood group antigens: demonstration of a wide spectrum of strain specificities and classification of two major binding groups among multiple binding patterns. J. Virol. 79, 6714–6722 (2005).
Brazil, J. C. & Parkos, C. A. Finding the sweet spot: glycosylation mediated regulation of intestinal inflammation. Mucosal Immunol. 15, 211–222 (2022).
Goto, Y., Uematsu, S. & Kiyono, H. Epithelial glycosylation in gut homeostasis and inflammation. Nat. Immunol. 17, 1244–1251 (2016).
Carroll, D. J. et al. Interleukin-22 regulates B3GNT7 expression to induce fucosylation of glycoproteins in intestinal epithelial cells. J. Biol. Chem. 298, 101463 (2022).
Nagao-Kitamoto, H. et al. Interleukin-22-mediated host glycosylation prevents Clostridioides difficile infection by modulating the metabolic activity of the gut microbiota. Nat. Med. 26, 608–617 (2020).
Pham, T. A. et al. Epithelial IL-22RA1-mediated fucosylation promotes intestinal colonization resistance to an opportunistic pathogen. Cell Host Microbe 16, 504–516 (2014).
Pickard, J. M. et al. Rapid fucosylation of intestinal epithelium sustains host-commensal symbiosis in sickness. Nature 514, 638–641 (2014).
Matsuzawa-Ishimoto, Y. et al. The gammadelta IEL effector API5 masks genetic susceptibility to Paneth cell death. Nature 610, 547–554 (2022).
Sugimoto, K. et al. IL-22 ameliorates intestinal inflammation in a mouse model of ulcerative colitis. J. Clin. Invest. 118, 534–544 (2008).
Ouyang, W. Distinct roles of IL-22 in human psoriasis and inflammatory bowel disease. Cytokine Growth Factor Rev. 21, 435–441 (2010).
Yan, J. et al. The pathogenic roles of IL-22 in colitis: its transcription regulation by musculin in T helper subsets and innate lymphoid cells. Front. Immunol. 12, 758730 (2021).
Mathew, S. et al. Mixed viral-bacterial infections and their effects on gut microbiota and clinical illnesses in children. Sci. Rep. 9, 865 (2019).
Shen, X. K., Song, S. L., Li, C. & Zhang, J. Z. Synonymous mutations in representative yeast genes are mostly strongly non-neutral. Nature 606, 725–731 (2022).
Nice, T. J., Strong, D. W., McCune, B. T., Pohl, C. S. & Virgin, H. W. A single-amino-acid change in murine norovirus NS1/2 is sufficient for colonic tropism and persistence. J. Virol. 87, 327–334 (2013).
Bok, K. et al. Evolutionary dynamics of GII.4 noroviruses over a 34-year period. J. Virol. 83, 11890–11901 (2009).
Verardi, R. et al. Disulfide stabilization of human norovirus GI.1 virus-like particles focuses immune response toward blockade epitopes. npj Vaccines 5, 110 (2020).
Parra, G. I. et al. Immunogenicity and specificity of norovirus Consensus GII.4 virus-like particles in monovalent and bivalent vaccine formulations. Vaccine 30, 3580–3586 (2012).
Acknowledgements
This project was funded by the Intramural Research Programs of the VRC and of NIAID, NIH. We acknowledge the outstanding work of veterinary (A. Cook and K. Guerriero) and research technicians at DVR and Bioqual, Inc., as well as animal programme coordinators C. N. Miller Dulan, J. Graves and A. Silvious, and past coordinators E. McCarthy and H. Bao at the VRC, DVR and BioQual, Inc. for expert animal care. We thank VRC Sequencing Core members J. Roberts-Torres and F. Laboune for technical assistance with sequencing procedures. We thank Roederer and Green laboratory members for discussions.
Author information
Authors and Affiliations
Contributions
M.R. and I.R. conceived and designed experiments. I.R. performed experiments, analysed data and prepared figures. K.D.W. assisted with experiments. B.M.N. prepared biopsies for microscopy. D.A.A. evaluated biopsies by microscopy and assisted with necropsy. A.R.H. and D.C.D. assisted with sequencing support. S.D. and S.G. analysed sequencing data. N.C., S.V.S., K.Y.G., A.S.O., R.V. and P.D.K. provided reagents. J.-P.T. and R.W. provided technical support. K.B., K.Y.G. and M.R. provided conceptual advice. I.R., M.R., K.Y.G., B.M.N., D.A.A., A.R.H., S.G. and A.S.O. wrote, and all authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
P.D.K., R.V., M.R. and I.R. have applied for a patent on norovirus stabilized VLPs (application no. PCT/US2021/055018), which were used in ELISA assays of this paper. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Koen Van Rompay and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Serum responses against a mix of norovirus inoculum in monkeys.
IgM (top panel) and IgG (bottom panel) titers in serum of inoculated monkeys were measured by ELISA against GI.1, GII.3, GII.4 and GII.17 VLPs up to eleven months after the challenge. Data points represent a mean of three technical replicates.
Extended Data Fig. 2 Serum IgA responses to GII.1 norovirus challenge in NHPs.
Serum IgA antibody responses measured to norovirus GII.1 Hawaii virus-like particle in ELISA. Data points represent a mean of two technical replicates.
Extended Data Fig. 3 Histo-blood group antigen phenotype in saliva of challenged animals.
ELISA plates were coated with saliva from different monkeys and HBGA phenotype was characterized using HBGA-specific antibodies. Each color represents a single animal. Leb detection was carried out using two clones, SPM194 and T218, where the latter has cross-reactivity to H-1 HBGA. Data points represent three technical replicates.
Extended Data Fig. 4 Necropsy of norovirus challenged animals.
a, Viral titers per g of stool from animals challenged with NVS3 at 1 × 108 genome copies (gc) per animal. On x axis shown days after inoculation. Animals were euthanized on the following day of the first positive stool sample. NHP #12 was euthanized on day 4, and NHP #13 on day 5 after the challenge. Dotted line indicates viral RNA detection limit at 104 gc/g of stool. Results represent a mean of two technical replicates. b, Detection of viral genome per mg of tissue collected from small and large intestines after the necropsy. Each data point summarizes a mean of two technical replicates.
Extended Data Fig. 5 H-1 reduction after norovirus challenge.
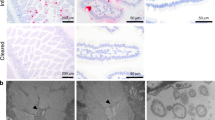
Representative H-1 immunostaining (brown) of jejunal biopsies. Diffuse and strong immunostaining along the brush border of the intestinal mucosal epithelial cells and within the cytoplasm apical to nucleus before the challenge (pre-). At 2 days post-challenge, there is a decrease in H-1 immunostaining intensity along the brush border, but the cytoplasmic immunostaining remains. The immunostaining at 30 days is similar in distribution and intensity to that seen in the pre-challenge biopsy. H-1 IHC, 40x.
Extended Data Fig. 6 Norovirus synonymous mutations in host.
Synonymous mutations found in different regions of human norovirus isolated from several animals during the virus shedding. Each block of rows represents frequency of single nucleotide polymorphisms (snp) found in the virus collected from individual animals. The source of each challenge (ch) is indicated in bold at the top of each block. Isolates that were used in passaging experiments of the shown animals are represented in italic bold. Grey gradient color designates frequency of identified mutations and orange gradient indicates viral titers per g of stool measured at each represented timepoint. Asterisk indicates samples that were amplified before sequencing. Only mutations with >10% detection frequency are shown. ORF1, open reading frame 1; RdRp, RNA-dependent RNA polymerase; VP1/2, viral protein 1/2; nt, nucleotide.
Extended Data Fig. 7 GI.1 norovirus challenge in pre-selected macaques.
a, Norovirus GI.1 genome copies detected in stool collected from animals over the following time after the challenge. Animals received GI.1 norovirus orally at 1 × 108 gc/animal dose. Dotted line indicates viral RNA detection limit at 104 gc/g of stool. Results represent a mean of two technical replicates run twice, and the error bars indicate a standard deviation. b, c, Serum responses of IgM (b) and IgG (c) against GI.1 VLP in ELISA post inoculation (p.i.). Data points represent a mean of two technical replicates. d, e, Chromogenic immunohistochemistry of jejunal biopsies indicating H-1 HBGA presence in brown. Biopsies were collected two weeks before (d) and two days after the challenge (e). f, saliva HBGA phenotype of GI.1 challenged animals. Each color represents a single HBGA. Leb detection was performed using two clones, SPM194 and T218, where the latter has cross-reactivity to H-1 HBGA. Data points represent three technical replicates. Horizontal lines indicate a mean and error bars one standard deviation.
Extended Data Fig. 8 Virus and serum titers in NHP #6.
Data collected from NHP #6 after two challenges (virus stock and dose indicated by orange arrows) is characterized by viral titers found in their stool and by serum antibody titers of IgM and IgG (EP – endpoint). The animal was removed from the study due to observed active H. pylori infection after a challenge. Results represent a mean of three technical replicates, and error bars indicate one standard deviation. Viral RNA detection limit at 104 gc/g of stool.
Supplementary information
Source data
Source Data Fig. 1
Replicate data.
Source Data Fig. 2
Replicate data.
Source Data Fig. 3
Replicate data.
Source Data Extended Data Fig. 1
Replicate data.
Source Data Extended Data Fig. 2
Replicate data.
Source Data Extended Data Fig. 3
Replicate data.
Source Data Extended Data Fig. 4
Replicate data.
Source Data Extended Data Fig. 7
Replicate data.
Source Data Extended Data Fig. 8
Replicate data.
Rights and permissions
About this article
Cite this article
Rimkute, I., Chaimongkol, N., Woods, K.D. et al. A non-human primate model for human norovirus infection. Nat Microbiol 9, 776–786 (2024). https://doi.org/10.1038/s41564-023-01585-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-023-01585-7