Abstract
Reconstructing the genomes of bilaterian ancestors is central to our understanding of animal evolution, where knowledge from ancient and/or slow-evolving bilaterian lineages is critical. Here we report a high-quality, chromosome-anchored reference genome for the scallop Patinopecten yessoensis, a bivalve mollusc that has a slow-evolving genome with many ancestral features. Chromosome-based macrosynteny analysis reveals a striking correspondence between the 19 scallop chromosomes and the 17 presumed ancestral bilaterian linkage groups at a level of conservation previously unseen, suggesting that the scallop may have a karyotype close to that of the bilaterian ancestor. Scallop Hox gene expression follows a new mode of subcluster temporal co-linearity that is possibly ancestral and may provide great potential in supporting diverse bilaterian body plans. Transcriptome analysis of scallop mantle eyes finds unexpected diversity in phototransduction cascades and a potentially ancient Pax2/5/8-dependent pathway for noncephalic eyes. The outstanding preservation of ancestral karyotype and developmental control makes the scallop genome a valuable resource for understanding early bilaterian evolution and biology.
Similar content being viewed by others
The nature of Urbilateria, the last common ancestor of all bilaterians, is enigmatic due to the lack of a plausible candidate in the fossil records1. The earliest unambiguous fossil of a bilaterian, Kimberella, shows remarkable resemblance to a mollusc, albeit its relationship with Urbilateria remains uncertain2,3. In the absence of definitive fossil records, genomic reconstruction by comparing extant bilaterian genomes becomes essential to our understanding of early bilaterian ancestors and their subsequent evolution4,5. However, reconstructing the genome of the bilaterian ancestor is challenging due to the paucity of high-order genome assemblies from ancient and/or slow-evolving lineages. Early genome sequencing efforts have mostly focused on two of the three major bilaterian groups, that is, protostome ecdysozoans and deuterostomes. Limited sequencing in the third group of protostome lophotrochozoans, a large superclade that includes molluscs, annelids and brachiopods, has revealed that their genomes are less derived from the ancestral bilaterian state than those of many ecdysozoans5. Unfortunately, none of these less-derived lophotrochozoan genomes were assembled to a degree that permits chromosome-level genome comparison.
Mollusca is the most speciose phylum of Lophotrochozoa and among the first bilaterians to appear in fossil records6. Many molluscan lineages including bivalves showed little change in shell morphology and life style over several hundred million years, and yet extant molluscs are abundant and thriving in diverse marine, freshwater and terrestrial environments, providing key ecological services and significant economic benefits to humans. Molluscs are highly diverse in form, making them excellent subjects to study body plan evolution and in particular its patterning by Hox genes7. Molluscs also have the greatest diversity in eye morphology, ranging from simple cupped to chambered or compound eyes, as well as in the number and placement of their eyes8, providing good subjects to study the origin and evolution of the eye, or Darwin’s ‘organ of extreme perfection’. Despite the great evolutionary and biological significance of molluscs, our sampling of their genomes remains limited to a few species5,9–11 and without high-order assemblies.
Here we report a high-quality, chromosome-anchored reference genome of the scallop Patinopecten yessoensis (Jay, 1857), a bivalve mollusc from the large Pectinidae family that contains ~270 living species and thousands of fossil species (dating back to ~320–340 million years ago, Ma12). Scallops are widely distributed in world oceans. They are mostly free-living and have multiple eyes scattering along the mantle edge. Many scallops are important fishery and aquaculture species. P. yessoensis is a large scallop living on cold and stable ocean bottoms of the northwestern Pacific. It has a conserved 19-chromosome karyotype that is common to diverse bivalves and may represent the ancient karyotype of bivalves13. Analysis of the scallop genome and extensive transcriptomes reveals outstanding preservation of ancestral bilaterian linkage groups, an intact Hox gene cluster under new expression control and diverse phototransduction cascades with a potentially ancient Pax2/5/8-dependent pathway for noncephalic eye formation, providing insights into the evolution of genome organization and developmental control during the emergence of bilaterians.
Results and discussion
Genome sequencing, assembly and characterization
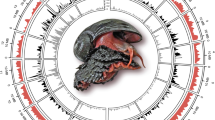
Genomes of marine bivalves are particularly challenging to sequence and assemble with short next-generation sequencing reads due to high polymorphism and repetitive content9,11. To alleviate the polymorphism problem, a highly inbred individual derived from selfing of a hermaphrodite (inbreeding coefficient of 0.5; Supplementary Fig. 1; Fig. 1a) was used for whole-genome shotgun (WGS) sequencing (424.3 Gb data in total; Supplementary Table 1), and an efficient, hybrid-specific SOAPdenovo approach14 was adopted for genome assembly (see Supplementary Text; Supplementary Figs 2–7). The final genome assembly is 988 Mb, with a contig N50 size of 38 kb and a scaffold N50 size of 804 kb (Supplementary Table 2), representing significant improvements over two published bivalve genomes9,11. Our assembly is 442 Mb less than the estimated genome size (~1.43 Gb; Supplementary Figs 8 and 9), probably due to the collapse of repetitive elements (Supplementary Fig. 10). The quality and integrity of the assembly is demonstrated by the mapping of 94.5% paired-end reads, 99.8% of Sanger-sequenced fosmids and 96.0–99.8% of various transcriptomic datasets generated in this and a previous study15 (Supplementary Figs 11 and 12; Supplementary Tables 3–6). With the aid of a high-density linkage map (7,489 markers; Supplementary Table 7) constructed by using the 2b-RAD methodology16, 1,419 scaffolds (covering ~81% of the assembly) are assigned to the 19 haploid chromosomes (Fig. 1a; Supplementary Fig. 13), providing the first chromosome-anchored genome assembly in molluscs or less-derived lophotrochozoans.
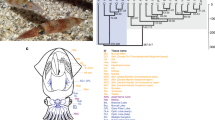
a, From outer to inner circles: a1, marker distribution across 19 chromosomes at megabase scales; a2 and a3, SNP density of the inbred progeny and its hermaphroditic parent, showing global heterozygosity reduction across the genome due to high inbreeding; a4 and a5, sequencing depth and abundance of repetitive sequences across the genome, with high repeat regions generally covered with more reads than low repeat regions (r = 0.58), indicating collapse of repeats in the genome assembly; a6, gene density across the genome. a1–a6 are drawn in non-overlapping 1 Mb sliding windows. b, Venn diagram of shared gene families between the scallop and two bivalve species (Pacific oyster C. gigas and pearl oyster P. fucata). Contrary to expectations, gene family sharing between scallop and each of two oyster species is more than that between the two oyster species, suggesting the less-derived gene repertoire in the scallop. c, Number of gene families shared between each of nine lophotrochozoan species and three major animal groups (deuterostome, ecdysozoan and non-bilaterian). P. yessoensis shared the most gene families with each group, followed by the ‘living fossil’ brachiopod L. anatina. Species names in red refer to bivalve species.
The scallop genome encodes 26,415 protein-coding genes (Supplementary Figs 14 and 15; Supplementary Table 8), of which 91% are annotated based on known proteins in public databases (Supplementary Table 9). The repeat content accounts for 39% (389 Mb) of the assembly (Supplementary Table 10), dominated by tandem repeats (18.4%). Transposable elements, which are usually considered active modulators of genome evolution, are less abundant (8–18% reduction) and less active in the scallop genome than the Pacific oyster and pearl oyster genomes (Supplementary Table 10; Supplementary Fig. 16). Resequencing of the wild hermaphrodite parent provides a genome-wide single nucleotide polymorphism (SNP) and short insertion/deletion (indel) polymorphism level of 1.04% (Supplementary Table 11), which is lower than the 1.30% found in the Pacific oyster Crassostrea gigas but approximately sevenfold higher than that (0.14%) found in humans17. As expected, polymorphism in the inbred scallop is greatly reduced compared to that in its hermaphroditic parent (Supplementary Table 11; Fig. 1a), which may have contributed to our assembly success.
Genome comparison and chromosome evolution
Phylogenetic analysis with 482 highly conserved, single-copy genes show that the scallop lineage diverged around ~425 Ma from the lineage leading to Pacific oyster and pearl oyster (Supplementary Fig. 17). Based on the sister taxon relationship between Bivalvia and Gastropoda18, our phylogenetic analysis gives an estimation of 504 Ma for the appearance of the bivalve lineage or its divergence from the gastropod lineage (Supplementary Fig. 17). P. yessoensis shows relatively slow substitution rate in protein sequences among bilaterians (Supplementary Table 12; Supplementary Fig. 18), supporting the ‘slow-evolving’ feature of scallop coding repertoire.
Gene family analysis of scallop and two other bivalves identifies a core set of 9,365 gene families (Fig. 1b). Comparing with 24 selected animal species (Supplementary Table 13) identified 756 bivalve-specific and 567 expanded gene families with notable enrichment of ion channel- and neurotransmitter-related functions (Supplementary Tables 14 and 15) that may help sessile or less mobile bivalves to cope with environmental changes more efficiently as part of bivalve adaptation. Contrary to expectations, the number of shared gene families between scallop and each of the other two bivalves (C. gigas and Pinctada fucata) is higher than that between C. gigas and P. fucata, which are phylogenetically closer (Supplementary Fig. 17), indicating relatively slower rate of gene divergence or loss in the scallop lineage. This also coincides with the observation of higher polymorphism in the exons of Pacific oyster than those of scallop as noted above. Among lophotrochozoans, bivalves share considerably more gene families with deuterostomes, ecdysozoans and non-bilaterian animals (Fig. 1c; Supplementary Table 16), with the highest values observed for scallop, followed by the brachiopod Lingula anatina, a lophotrochozoan that is commonly considered a ‘living fossil’19. Gene family analysis also identifies 830 scallop-specific and 349 expanded gene families that participate in diverse biological processes (Supplementary Tables 17 and 18) and are probably important for scallop lineage-specific adaptations.
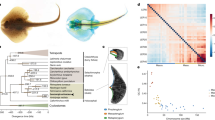
To enable deep phylogenetic comparisons, we conducted macrosynteny analysis of conserved linkage between orthologous genes, which is independent of intra-chromosomal rearrangements4,5. Such analysis has been fruitful in previous studies on lophotrochozoans5 for understanding long-range macrosynteny conservation, but limited in inferring chromosome-scale evolution, as these studies are all based on highly fragmented genome assemblies with the number of scaffolds usually ranging from thousands to tens of thousands. To understand bilaterian chromosome evolution, we generated chromosome-level assemblies not only for scallop but also for Pacific oyster (C. gigas) and pearl oyster (P. fucata) by using two recently published high-density linkage maps20,21, and used them for macrosynteny analysis. Strikingly, our chromosome-based macrosynteny analysis reveals a near-perfect correspondence between the 19 scallop chromosomes and the 17 presumed bilaterian ancestral linkage groups (ALGs; ALGs or proto-chromosomes reconstructed in ref. 5), a level of chromosome preservation that far exceeds other bilaterians with chromosome-level assemblies (Fig. 2; Supplementary Table 19; conservation index: 0.81 for scallop whereas 0–0.42 for other bilaterians), suggesting that scallop has a karyotype highly similar to that of the bilaterian ancestor. Such degree of karyotype preservation is less evident in the two oyster species (Fig. 2), which may attribute to their presumably derived karyotypes (10 chromosomes in C. gigas and 14 chromosomes in P. fucata) in comparison with the highly conserved 19-chromosome karyotype found in scallops and many other bivalves13. To allow more bilaterian genomes (11 additional representative bilaterians) to be included for comparisons, we also performed the conventional scaffold-based macrosynteny analysis4,5, which still shows that scallop has the highest level of macrosynteny conservation, closely followed by amphioxus Branchiostoma floridae (Supplementary Fig. 19). Only two inter-chromosome rearrangements were identified in all three bivalves, including partial translocation of ALG2 and the fusion of ALG5 and ALG16 (Supplementary Table 19) that possibly pre-dates the radiation of bivalves.
Chromosome-based macrosynteny is shown in the form of dot plots with comparisons between the chromosomes of 12 bilaterian genomes (x axis) and the 17 presumed bilaterian ALGs (ALG1–17, y axis) that were previously reconstructed5. The scallop genome anchored in 19 chromosomes shows the highest level of chromosome preservation (conservation index (CI) = 0.81) that far exceeds those of other bilaterians (CI = 0–0.42) with chromosome-level assemblies, suggesting that the scallop has a karyotype closely resembling that of a bilaterian ancestor. The haploid chromosome number (n) is shown for each species, and the CI measures the extent of preservation of bilaterian ALGs in each species. The chromosome-anchored genome assemblies of Pacific oyster (C. gigas) and pearl oyster (P. fucata) were generated in this study by using two recently published high-density linkage maps20,21.
Homeobox clusters and subcluster temporal co-linearity
The homeobox genes of Antennapedia (ANTP)-class are key regulators of development in all animals, which presumably originated from a Mega-cluster that formed by tandem duplications of a Proto-ANTP gene22. They are more or less dispersed in modern bilaterian genomes, but mostly found in four distinct chromosomes in the amphioxus and in the annelid Platynereis, which has led to the hypothesis that the Mega-cluster, if it did exist, had already been broken up onto four chromosomes by the time of the protostome–deuterostome ancestor (PDA)23. Supporting this hypothesis, a similar distribution of ANTP genes on four scallop chromosomes is observed (Supplementary Fig. 20). In particular, it confirms the coexistence of the Hox genes with the NK-linked gene Dlx, providing key support for the ancient linkage of NK-linked and Hox-linked genes in the Mega-cluster hypothesis23.
Contrary to frequent cluster alterations in many animal lineages by gene loss, duplication or physical splits24 (Supplementary Fig. 21), ParaHox and Hox clusters are well-preserved and remain intact in the scallop genome, which enables us to infer the possible ancestral state of these clusters in the lophotrochozoan ancestor or PDA (Fig. 3a). For example, the scallop ParaHox cluster exhibits the same gene order, orientation and relative gene spacing as those found in chordates (Supplementary Fig. 22), strongly supporting the previous speculation of the existence of a typical deuterostome-like cluster in the PDA and lophotrochozoan ancestor25. The scallop Hox cluster contains 11 genes (3 anterior, 6 central and 2 posterior) that largely retain the conserved residues of their homeodomains for each Hox paralogous group (Fig. 3a; Supplementary Fig. 23). Comparison of the scallop Hox cluster with those of other lophotrochozoans suggests that the lophotrochozoan ancestor might already have an 11-gene Hox cluster that resembles the intact Hox clusters of scallop and limpet, with all genes except Post1 arranged in the same orientation.
a, Chromosomal organization of Hox and ParaHox genes of scallop and other lophotrochozoans. Unlike other lophotrochozoans whose Hox and ParaHox clusters are usually fragmented, both clusters are present as single, intact clusters in scallop, possibly representing the ancestral state of these clusters in the lophotrochozoan ancestor (top). Horizontal arrows and triangles denote transcription orientation and external gene insertion, respectively. b, Phylogenetic relationships (in protein sequences) among Hox genes from three lophotrochozoans with high levels of Hox sequence preservation, showing closer relationships between genes within a subcluster than between clusters (S1–S4). Tree topologies obtained from maximum likelihood and neighbour-joining methods are largely consistent and only the maximum likelihood tree is shown. Numbers above branches are supporting percentages of 1,000 bootstrap replicates from maximum likelihood (before slash) and neighbour-joining (after slash) trees. PY, P. yessoensis; CG, C. gigas; CT, C. teleta. c, Temporal expression of scallop Hox and ParaHox cluster genes. Expression of ParaHox cluster follows temporal co-linearity during embryonic and larval stages, whereas expression of Hox cluster exhibits an STC for four subclusters (S1: Hox1–Hox3, S2: Hox4–Hox5, S3: Lox5–Lox2, S4: Post2–Post1), with leading genes generally activated earlier than their followers in each subcluster. Vertical thick arrows indicate co-activation of leading genes of four subclusters, whereas thin arrows indicate gene activation order within each subcluster or the whole cluster. 2-8cell, 2–8 cells; Bla, blastula; Gas, gastrula; Tro, trochophore; Dst, D-stage larva; Ped, pediveliger; Juv, juvenile. d, Spatially co-linear expression of four leading genes (Hox1, Hox4, Lox5, Post2) of each subcluster at the gastrula stage. The expression regions of four leading genes almost span the entire anterior–posterior body axis, suggesting the prominent roles of these leading genes in early body plan determination. Expression of other Hox genes is not detectable at this stage (see Supplementary Fig. 23) except Lox4, which is less likely to be a candidate leading gene as its expression does not conform to the characteristic pattern of leading genes (stronger expression at early developmental stages than latter stages). A, anterior; P, posterior; V, ventral; D, dorsal. Scale bar, 50 μm.
Temporal co-linear activation of homeobox genes for patterning the body plan is well documented in vertebrates and may contribute to the conservation of homeobox clusters25–27. As expected, the expression of scallop ParaHox genes shows temporal co-linearity during embryonic and larval development (Fig. 3c), similar to those found in deuterostomes with intact ParaHox clusters25. The expression of scallop Hox genes, however, does not follow the typical pattern of temporal co-linearity as observed in vertebrates that spans the whole cluster (for example, from Hox1 to Hox13 in the HoxD cluster of mouse28). Instead, their expression exhibits a characteristic subcluster-level temporal co-linearity (STC; Fig. 3c): the expression begins at the gastrula stage with leading genes of four subclusters (Hox1, Hox4, Lox5 and Post2), followed by sequential expression of their subcluster followers. Interestingly, the expression of four leading genes also follows spatial co-linearity at the gastrula stage where ‘followers’ are generally not expressed (Fig. 3d; Supplementary Fig. 24), and their expression regions almost span the entire anterior–posterior body axis, suggesting prominent roles for these leading genes in early body plan determination.
We re-examined published data to determine if STC is present in Hox expression of other bilaterians during development. The oyster Hox expression clearly resembles that of scallop, although the oyster has dispersed Hox subclusters9 (Fig. 4; Supplementary Fig. 25) and STC was not previously recognized. These findings suggest that maintaining STC may depend on the integrity of subclusters but not the whole cluster. We also identified similar/partial STC patterns by analysing published Hox expression data in distantly related bilaterian taxa, including the annelids Nereis virens29 and Platynereis dumerilii (Lophotrochozoa), the shrimp Litopenaeus vannamei30 (Ecdysozoa) and the ascidian Ciona intestinalis31 (Chordata) (Fig. 4; Supplementary Figs 26 and 27), suggesting that STC could be ancestral, although gene regulatory networks underlying these STC patterns may have been substantially modified to support lineage-specific body plans. As genes within each subcluster are preferentially related to each other (Fig. 3b), STC might have been established during the stepwise duplication of primordial Hox genes (represented by three co-activated Hox genes in the basal bilaterian acoels32,33; Fig. 4), and a similar scenario was observed for a newly formed rodent-specific Rhox cluster34. It is also possible, but less likely, that a complete Hox cluster with cluster-wide temporal co-linearity already existed in the bilaterian ancestor, and STC is a derived state that independently occurred in several bilaterian lineages. Interestingly, we found that Hox expression in the annelid Capitella teleta follows an unusual mode of whole-cluster temporal co-linearity (WTC) that is subcluster-based35 (Fig. 4; called S-WTC here), probably representing an intermediate state in evolutionary transition from STC to WTC, or vice versa. Owing to its increased flexibility in developmental patterning, STC may be central to the bilaterian body plan evolution and, if indeed ancestral, would provide the bilaterian ancestor with great potential in generating diverse body plans found in different bilaterian lineages.
The phenomenon of STC exists in an intact Hox cluster of scallop and fragmented clusters of different lophotrochozoan lineages (bivalves and annelids)9,29 and distantly related bilaterian groups (ecdysozoan and deuterostome)30,31, suggesting that STC could be ancestral. The basal bilaterian acoels33 have only three unlinked Hox genes (corresponding to Hox1, Hox4/5 and Post paralogous group46), and it has been proposed that these acoel Hox genes may belong to the ancient Hox cluster of Urbilateria or proto-Urbilateria, duplication of which gives rise to present protostome/deuterostome Hox clusters46. In the acoel Convolutriloba longifissura, all three Hox genes show contemporaneous expression after gastrulation32, which is consistent with the co-activation of subclusters in scallop and other bilaterians, and suggests STC might be established along with the stepwise duplication of primordial Hox genes. Interestingly, the Hox expression of the annelid C. teleta exhibits an unusual mode of WTC that is subcluster-based35 (called S-WTC here), probably representing an intermediate state in evolutionary transition between STC and WTC. STC may be central to the bilaterian body plan evolution and, if indeed ancestral, would provide bilaterian ancestors great potential in generating diverse body plans.
Photoreceptors and the eye regulatory network
Scallops have a large number (~30–100) of noncephalic but complex eyes along the edge of their mantle, which possess double-layered retinas, with the proximal and distal retina comprising rhabdomeric and ciliary photoreceptors, respectively36 (Fig. 5a). Ten full-length opsin genes including four r-opsins, two Go-opsins, two c-opsins and one peropsin are identified in the scallop genome and show primary expression in scallop eyes (Supplementary Figs 28 and 29). R-opsin and Go-opsin are known to mediate rhabdomeric and ciliary phototransduction in scallop eyes, respectively37, and as expected, key genes participating in the two phototransduction cascades show higher expression in scallop eyes than mantle (Fig. 5b). In particular, R-opsin and its associated cascade have the highest expression in scallop eyes, greatly exceeding other opsins (Fig. 5a,b), suggesting that rhabdomeric phototransduction may play a prominent role in scallop eye function. The finding of c-opsin expression in scallop eyes is intriguing (Fig. 5a), as c-opsin has not been identified in scallops before and was once considered a vertebrate-type opsin for ciliary phototransduction38. Further investigation of the scallop genome identified key genes participating in vertebrate canonical (Gi/t) and noncanonical (Gs) c-opsin cascades37, and expression profile of these genes supports the involvement of the c-opsin cascade in scallop eye function (Fig. 5b). The coexistence of r-opsin-, Go-opsin- and c-opsin-mediated phototransduction cascades in scallop eyes is unusual. Considering the differential preservation of rhabdomeric and ciliary photoreceptors for vision in extant animal groups (invertebrates and vertebrates, respectively37,38), scallop eyes provide a unique model to study how multiple phototransduction cascades function and coordinate in a single visual system, which may provide insights into distinctive evolutionary routes of these cascades in invertebrates and vertebrates37,38.
a, Scallop has numerous noncephalic eyes scattered along mantle margins, with two distinct retinal layers consisted of rhabdomeric and ciliary PRCs, respectively (scale bars on the upper and lower images represent 5 cm and 100 μm, respectively). Diverse opsin types (r-opsin, Go-opsin and c-opsin) are identified in the scallop genome, with r-opsins showing the highest expression and probably playing a prominent role in scallop eye function. The finding of c-opsin expression in scallop eyes is intriguing, as c-opsin has not been identified in scallops before and was once considered a vertebrate-type opsin for ciliary phototranduction38. Eye samples from three individuals were used in expression evaluation with standard errors shown for eye and mantle groups. b, Key components of different phototransduction cascades (mediated by r-opsin, Go-opsin and c-opsin) identified in the scallop genome with their gene expression in eyes relative to mantle colour-coded. The coexistence of multiple phototransduction cascades in scallop eyes is unusual and intriguing, as these cascades have been selectively preserved for visual function in invertebrates (r-opsin) and vertebrates (c-opsin)37,38. FC, fold change.
We identified a collection of 825 genes that are significantly up-regulated in scallop eyes relative to mantle (Supplementary Table 20) and enriched for genes of the G-protein-coupled receptors (GPCRs) signalling pathway (Supplementary Table 21). Surprisingly, Pax6, a presumed master control gene for all bilaterian eyes39, is present in the genome but not expressed in the eye and mantle (Fig. 5c). Other genes of the typical invertebrate and vertebrate Pax6 pathways are either not expressed (for example, Six3/6 and Rx) or do not show upregulation in the eye relative to mantle (for example, Six1/2, Eya, Dach) (Fig. 6a). The possibility of transient expression of Pax6 regulatory pathway during early eye development, although not yet investigated, seems unlikely as scallop adult eyes exhibit continuous eye formation and growth (that is, continuous eye morphogenesis) with increasing age40. Our finding therefore suggests that the pax6-dependent pathway may not be involved in scallop eye morphogenesis and function. To understand the gene regulatory network of scallop eyes, we constructed a gene coexpression network using 26 adult transcriptome datasets, and identified M2 as the only eye-related module (Supplementary Figs 30 and 31; Supplementary Table 22). The eye-related transcription factors Pax2/5/8, Brn3, Lmx1b and Six4/5 are members of this module. In particular, Pax2/5/8, Brn3 and Lmx1b are recognized as the most important hub transcription factors in the network (Fig. 6b; Supplementary Table 23), suggesting that they are key regulators of scallop eye development and function. The involvement of Pax2/5/8, Brn3 and Six4/5 in the noncephalic light sensors has been previously reported in Platynereis midventral photoreceptor cells (PRCs)41 and amphioxus Hesse organs42, both of which are also Pax6-independent and have led to the hypothesis that cephalic and noncephalic PRCs may have different evolutionary origins, with the former dependent on Pax6 and the latter on Pax2/5/841. However, previous investigations were all based on simple light sensors, and the possibility that these noncephalic light sensors may represent evolutionary innovations cannot be excluded41. Our finding of Pax2/5/8 as a key regulator in the gene network of scallop mantle eyes provides the first complex eye-based evidence supporting the hypothesis of Pax2/5/8-dependent origin of noncephalic eyes (Fig. 6c), and together with previous studies41–45, argues against Pax6 as the universal master control gene for all bilaterian eyes39.
a, Expression analysis of eye-related transcriptional factors. Pax6 and related transcription factors are either not expressed or do not show up-regulation in scallop eyes, whereas Pax2/5/8 and related transcription factors generally show up-regulation in scallop eyes, with P < 0.05 for Pax2/5/8 and Brn3, and P < 0.1 for Six4/5 based on the one-tailed t-test. This suggests that scallop eyes rely on a regulatory pathway that is independent of Pax6, a presumed master control gene for all bilaterian eyes39. Eye samples from three individuals were used in expression evaluation with standard errors shown for eye and mantle groups. b, Gene coexpression network of an eye-related module (M2, see Supplementary Table 23). The M2 module contains 2,171 genes, of which 170 genes with the highest intramodular connectivity are chosen for network display. Gene names or IDs are shown for the top 30 hub genes with transcription factors labelled in red. Transcriptional factors Pax2/5/8, Brn3 and Lmx1b are the most important hub transcription factors and thus represent key regulators in scallop eye network. Node size represents the intramodular connectivity of a given gene. c, Summary of opsins and Pax genes used in cephalic and noncephalic bilaterian eyes or light sensors from this and previous studies41–43,47–50. The observation of cephalic and noncephalic bilaterian eyes controlled by different Pax regulators (Pax6 for the former and Pax2/5/8 for the latter) across major bilaterian clades, supporting their different evolutionary origins41.
Conclusions
Reconstructing the genomes of ancient bilaterians that pre-dated the split of protostomes and deuterostomes is critical to our understanding of bilaterian evolution, where studying genomes of poorly sampled lophotrochozoans should be particularly informative. Ancient genomes may be reconstructed in both gene repertoire and genome organization through gene family studies and synteny analysis of high-order genome assemblies. In devoting such efforts to the scallop P. yessoensis, we find remarkable conservation of ancestral features in genome organization and gene repertoire that bring us closer to the bilaterian ancestral genome. These include the closest representation of the ancestral bilaterian karyotype to date, intact ParaHox and Hox gene clusters, diverse phototransduction cascades and an ancient regulatory pathway for eye development. The STC that is shared by other bilaterians may be ancestral to whole-cluster co-linearity and central to the great diversity in body plan found in molluscs and other bilaterians. The exceptional conservation of ancestral features suggests that the scallop genome is slow-evolving, probably as a consequence of life on cold and stable deep-ocean bottoms. Similar studies, particularly of chromosome-anchored genomes from basal bilaterians such as monoplacophoran molluscs, annelids and acoels, may identify other genomes more closely related to that of the bilaterian ancestor and lead to the eventual reconstruction of urbilaterian chromosomes, which may greatly improve our understanding of bilaterian evolution.
Methods
Genome sequencing and assembly
A one-year-old male P. yessoensis from a selfing family created with a hermaphroditic individual was used for WGS sequencing and assembly. High-quality genomic DNA was extracted from the adductor muscle of this inbred male using the conventional phenol/chloroform extraction method51. Short-insert (180 bp, 300 bp and 500 bp) paired-end libraries and large-insert (2 kb and 5 kb) mate-pair libraries were prepared using Illumina’s DNA library preparation kits following standard protocols. The 10 kb and 16 kb mate-paired libraries were prepared following the Cre–lox recombination-based protocol52. The libraries were subjected to the paired-end 100 bp/150 bp sequencing on the Illumina HiSeq2000 platform. A modified version of SOAPdenovo was developed for efficient genome assembly to reduce the problem of high genome heterozygosity (see Supplementary Text for methodological details).
Genome size estimation
The genome size of P. yessoensis was estimated using flow cytometry and k-mer analysis. Gills of P. yessoensis were used for flow cytometry analysis as previously described53,54, with Pacific oyster C. gigas (2C = 1.31 pg)9 as an internal reference standard. Briefly, gills were dissected and dissociated into single cells using a 25-gauge syringe needle. Then the cell suspension was filtered through a 20-μm nylon mesh and stained with 10 mg ml−1 4,6-diamidino-2-phenylindole (DAPI). The stained cell suspension was analysed using a flow cytometer (Partec PAII, Germany). The DNA content was then converted to gigabases based on the formula: 1 pg = 0.978 Gb (ref. 55). For k-mer analysis, the genome size was estimated based on the 19-mer frequency distribution using the formula: genome size = (total number of 19-mer)/(position of peak depth).
Quality assessment of genome assembly
The integrity of the final assembly of P. yessoensis genome was examined using three 30–35 kb fosmid sequences, ~45× WGS sequences (from 180 bp library) and three sets of messenger RNA (mRNA) data. Fosmid sequences were aligned to the scallop genome assembly using LASTZ56 with the parameters of ‘M = 254 K = 4,500 L = 3,000 Y = 15,000 --seed = match12 --step = 20 --identity = 85’. Burrows–Wheeler Aligner (BWA)57 was used to align the WGS data with the final assembly with parameters of ‘-n 15 –o 1 –e 10’ by considering high polymorphism between haploids9. Full-length complementary DNA (cDNA) sequences, the assembled transcriptomes generated from 454 sequencing15 and Illumina sequencing (assembled by Trinity58) were mapped to the genome assembly using BLAT59 with default parameters and an identity cutoff of 80%.
Linkage map construction and chromosome anchoring
Three full-sib families each consisting of 38–40 individuals were used for linkage mapping analysis. 2b-RAD libraries were prepared for parents and progenies using the type IIB restriction enzyme BsaXI and following the protocol developed in ref. 16. The adaptors with 5′-NNN-3′ overhangs were used to target all BsaXI fragments in the scallop genome. All libraries were subjected to single-end sequencing (1×50 bp) using the Illumina HiSeq2000 platform. The 2b-RAD reads were preprocessed to remove unreliable ones and then genotyped using the RADtyping program60 under default parameters. The SNP markers that segregated at a 1:1 ratio in each mapping family were obtained and categorized as lm×ll or nn×np. Markers present in both parents that segregated at a 1:2:1 ratio were also retrieved and were categorized as hk×hk. SNP markers that conformed to the expected Mendelian ratios (chi-squared test, P > = 0.01) and could be genotyped in at least 80% of the offspring of each family were used for linkage analysis. Markers were grouped at a logarithm of odds threshold of at least 6.0 and ordered based on the regression mapping algorithm implemented in JoinMap4.0 software61. The recombination frequencies were converted into map distances in centi-Morgan (cM) through the Kosambi mapping function. The consensus map was generated by integrating the linkage maps of three families using the MergeMap software62, with map weight setting as 1.0 for each map.
For chromosome anchoring of scaffolds, marker sequences from the consensus genetic map were aligned back to the genome assembly using BLAST63 with the parameters of ‘-e 1e-4 –F F –G 5 –E 2 –W 7 –r 2 –q −3 –m 8’. Only markers that were mapped to a unique location in the assembly were used for anchoring and orienting scaffolds to corresponding linkage groups (that is, chromosomes) according to the locations of markers in the genetic linkage map. For cases where scaffolds were in conflict with the genetic map (for example, markers from one scaffold assigned to different linkage groups), we manually checked these scaffolds using the 10 kb mate-paired reads and eight scaffolds were broken at points with low-coverage support by mate-paired reads. A similar approach was applied to anchor the existing genome assemblies of Pacific oyster (C. gigas)9 and pearl oyster (P. fucata)11 to linkage groups using two recently published high-density genetic linkage maps20,21.
Transcriptome sequencing and expression profiling
Embryos (two to eight cells, blastulae and gastrulae), larvae (trochophore larvae, D-stage larvae, pedi-veliger larvae and juvenile) and adults of P. yessoensis were collected from the hatchery of Zhangzidao Group Co., Ltd (Dalian, China) in 2013. To obtain embryonic and larval materials, artificial fertilization and larval culture were performed according to the procedure described in ref. 64. The fertilized eggs and larvae were reared at 13–15 °C and more than 1,000 embryos/larvae were sampled for each developmental stage (sampling time is provided in Supplementary Table 24). Nine adult tissues/organs (eye, mantle, gill, gonad, blood, digestive gland, striated muscle, smooth muscle and foot) were dissected from two to three scallop individuals. All the samples were flash frozen in liquid nitrogen and stored at −80 °C until use.
Total mRNA was extracted from each of the seven developmental samples and nine adult tissues/organs following the protocol described in ref. 65. RNA sequencing (RNA-Seq) libraries were constructed using the NEBNext mRNA Library Prep Master Mix Set for Illumina following the manufacturer’s instructions. The libraries were subjected to paired-end 100 bp sequencing on the Illumina HiSeq 2000 platform. Raw reads were first filtered by removing those containing undetermined bases (‘N’) or excessive numbers of low-quality positions (>10 positions with quality scores <10 ). Then the high-quality reads were mapped to the P. yessoensis genome using Tophat (v2.0.9)66 with the parameters of ‘-p 10 -N 3 --read-edit-dist 3 -m 1 -r 0 --coverage-search --microexon-search’. The expression level of all genes was normalized using the trimmed mean of M-values (TMM) method (implemented in the edgeR package67) and represented in the form of reads per kilobase of exon model per million mapped reads (RPKM)68. The RPKM expression values of all genes for all developmental stages and adult tissues/organs are provided in Supplementary Table 25.
Polymorphism analysis
To evaluate polymorphism reduction in the inbred progeny, ~50× genome resequencing was performed for its hermaphroditic parent. Paired-end reads from the inbred progeny (~230×) and its parent (~50×) were aligned onto the final genome assembly for SNP and indel identification using BWA57 with the parameters of ‘-n 15 –o 1 –e 10’. The minimum and maximum read depths for variation calling were set as 0.1 and 2-fold of the average depth of sequencing, respectively. To reduce the false positives, SNPs within 5 bp around a gap were filtered out and adjacent gaps located in 10 bp window size were also removed. The statistical significance of comparison of polymorphism rates between scallop and Pacific oyster (C. gigas) was determined using the two-sided chi-squared test.
Genome annotation
Both homology-based and de novo predictions were used to detect transposable elements in the genome. For homology-based detection, RepeatMasker and RepeatProteinMask (both available from http://www.repeatmasker.org) were used to screen the P. yessoensis genome for known transposable elements (for example, DNA transposon, long terminal repeat, long and short interspersed elements) in the RepBase library (v20140131)69. De novo transposable elements were identified and modelled by RepeatModeler (v1.0.4, http://www.repeatmasker.org). Tandem repeats were identified by searching for two or more contiguous, approximate copies of a pattern of nucleotides using Tandem Repeats Finder (v4.07b)70 under default parameters.
Gene prediction and functional annotation were performed primarily following the procedure described in previous studies71,72. Briefly, three de novo gene prediction tools, Augustus (v2.7)73, GlimmerHMM (v3.02)74 and SNAP (2006-07-28)75, were used to predict genes in the repeat-masked genome sequences. For homology-based gene prediction, protein sequences from C. gigas, Lottia gigantea, Helobdella robusta, Anopheles gambiae, Caenorhabditis elegans, Drosophila melanogaster, Homo sapiens and Strongylocentrotus purpuratus were aligned to the P. yessoensis genome using tblastn (v2.2.26; E-value ≤ 1e−5)76, then the homologous genome sequences were aligned against the matching proteins using GeneWise (v2.4.1)77 for accurate spliced alignments. The RNA-Seq reads from different developmental stages and adult tissues/organs were aligned to the P. yessoensis genome using Tophat (2.0.11)66, and Cufflinks (2.1.1)78 was used to produce assembled transcripts and predict transcript structures. Gene predictions from the de novo approach, homology-based approach and RNA-Seq-based evidence were merged to form a comprehensive consensus gene set using the software EVM79. To obtain gene functional annotations, the predicted protein sequences of P. yessoensis were aligned to public databases including KEGG, SwissProt and TrEMBL using BLASTP with the E-value threshold of 1e-5. InterProScan (v4.8)80 was also used to identify motifs and domains by searching the InterPro and Gene Ontology81 databases.
Gene family analysis
We selected the following 27 representative animal species (Supplementary Table 13) from the sub-kingdom Eumetazoa for gene family analysis: P. yessoensis, C. gigas, P. fucata, L. gigantea, Octopus bimaculoides, L. anatina, C. teleta, H. robusta, Schistosoma mansoni (lophotrochozoan group); C. elegans, D. melanogaster, Tribolium castaneum, A. gambiae, Daphnia pulex, Strigamia maritima, Apis mellifera (ecdysozoan group); H. sapiens, B. floridae, S. purpuratus, Danio rerio, Xenopus tropicalis, Gallus gallus, Mus musculus (deuterostome group); Mnemiopsis leidyi, Nematostella vectensis, Trichoplax adhaerens, Amphimedon queenslandica (non-bilaterian group). We used the OrthoMCL software (version 1.4)82 to define gene family clusters among different species. An all-against-all BLASTP was first applied to determine the similarities between genes in all genomes at the E-value threshold of 1e-7. Then the Markov clustering (MCL) algorithm implemented in OrthoMCL was used to group orthologues and paralogues from all input species with an inflation value (-I) of 1.5. For comparisons of gene families between phylogenetic groups, a shared gene family is required to be present in at least two species within each compared group. Gene families belonging only to P. yessoensis but not to any other species (including other bilaterian and non-bilaterian species) were considered scallop-specific gene families. Within the lophotrochozoan group, the number of P. yessoensis genes from each gene family was compared to those from other lophotrochozoans to detect gene families that were expanded only in P. yessoensis. To compute the statistical significance, Fisher's exact test was applied based on two backgrounds: one is the count of all P. yessoensis genes and the other is the count of genes in other lophotrochozoans. A P value threshold of 0.05 was used to retrieve the gene families that were significantly expanded in scallop. A similar approach was also applied to identify bivalve-specific and expanded gene families.
Phylogeny, divergence time and evolutionary rate estimation
We retrieved protein sequences of all single-copy gene families (that is, only one gene copy for each species in a gene family cluster) from the gene family analysis (see previous section) to constitute a 482-gene dataset for constructing a phylogenetic tree for 14 selected species (P. yessoensis, C. gigas, P. fucata, L. gigantea, O. bimaculoides and C. teleta from the lophotrochozoan group; T. castaneum, D. pulex, S. maritima, A. mellifera and D. melanogaster from the ecdysozoan group; H. sapiens and B. floridae from the deuterostome group; and N. vectensis from the non-bilaterian group). The purpose of our phylogenetic analysis was mainly to infer the phylogenetic relationships and divergence time for the bivalve lineage, and a more comprehensive analysis of Lophotrochozoa phylogeny has been recently provided83. Multiple alignments were performed using MUSCLE84 for each gene family, and gaps were trimmed using Gblocks85. Then the alignments were concatenated to a super alignment matrix. ProtTest86 was used to select the best-fit model (LG+Γ4 model) for amino acid replacement and RA×ML (v8.0.19)87 was used to reconstruct a maximum likelihood tree. Robustness of the maximum likelihood tree was assessed using the bootstrap method (100 pseudo-replicates). Divergence time between species/clade was estimated using mcmctree in PAML88 with the parameters of ‘RootAge = <600 model = REV(GTR) alpha = 0.969 clock = 2’, and the calibration points are provided in Supplementary Table 26.
For substitution rate analysis, the above trimmed multiple protein alignments were first converted into the corresponding codon alignments for each gene family. Then synonymous substitution rate (Ks) and nonsynonymous substitution rate (Ka) were estimated by using the free-ratio model in the PAML87 codeml program for each family and each species, and to be stringent, only Ks values less than five were considered.
Macrosynteny analysis
Based on the phylogenetic positions of the 27 animal species (Supplementary Table 13), a hierarchical clustering method5 was adopted to identify orthologous gene sets. First, two gene clusters from different sides of a branch would be merged when they had mutual best BLASTP hits with each other. Second, clusters of genes within a subtree would be further grouped together if these genes have better hits to each other than to any outgroup genes. Based on these two criteria, genes from different species were clustered starting at the leaves leading to the terminal point at the root. The ancestral bilaterian gene families were determined when they met at least one of the following criteria5: (1) the gene family was present in at least two protostome and two deuterostome species (ingroups); and (2) the gene family was present in at least two protostome or two deuterostome species and in two of the non-bilaterian (outgroup) species.
The conservation of gene macrosynteny between species with chromosome-level assemblies and the 17 presumed bilaterian ALGs was displayed in the form of ‘dot plot’. The 17 bilaterian ALGs (represented by the genes of sea anemone N. vectensis) were retrieved from a previous study5, where ALGs were reconstructed for early bilaterian ancestors based on the chromosome-history-graph approach. Each dot in the dot plot comparison represents a one-to-one orthologous gene pair derived from the same ancestral gene family. For species without chromosome-level assemblies, a heuristic hierarchical method4,5 was adopted to cluster the scaffolds from these draft genomes into corresponding homologous ALGs using the cluster program89 with the tree-cutting threshold of 0.25. For both chromosomal and scaffold-level comparisons, a macrosynteny conservation index5 was calculated as measurement of preservation of ALGs in each species. To be conservative, the number of one-to-one orthologous gene pairs whose genes are located in scaffolds or chromosome segments that were assigned into homologous ALGs was taken as numerator, and the number of one-to-one orthologues where both genes were on a scaffold/chromosome segment that was large enough to be assigned to an ALG was taken as denominator.
Homeobox gene analysis
The homeobox genes were identified in the P. yessoensis genome using BLAST with an E-value threshold of 1e−5 against all homeodomain sequences from the HomeoDB database (http://homeodb.zoo.ox.ac.uk/)90, and were further confirmed by comparing to the Conserved Domains Database (http://www.ncbi.nlm.nih.gov/cdd). Genes were classified based on BLAST results, molecular phylogeny and manual inspection of conserved residues. The same approach was also used to identify homeobox genes in other bilaterian genomes. Phylogenetic analyses were performed using MEGA591 to construct neighbour-joining and maximum likelihood trees. For neighbour-joining analysis, evolutionary distances were computed using the p-distance method. For maximum likelihood analysis, the Poisson correction model was chosen. A discrete gamma distribution was used to model evolutionary rate differences among sites. All positions containing gaps and missing data were eliminated in both analyses, and the robustness of the resulting phylogenies was tested by a reanalysis of 1,000 bootstrap replicates. The heat map of Hox and ParaHox gene expression was drawn using custom R scripts that used the heatmap.2 function of gplots (an R package; http://cran.r-project.org/package=gplots).
Whole mount in situ hybridization
Scallop gastrulas (28 h post-fertilization at 15 °C) were fixed in 4% paraformaldehyde overnight, transferred to methanol and stored at −20 °C. Fragments of Hox genes were amplified from larval cDNA using specific primers (Supplementary Table 27) containing a 5′ T7 promoter sequence (5′-taatacgactcactataggg-3′). Purified polymerase chain reaction products were used as templates in the following in vitro transcription. Digoxigenin-labelled sense and anti-sense probes were synthesized using the DIG RNA Labeling Mix (Roche) and a T7 RNA polymerase (Fermentas). Specimens were serially rehydrated in PBST (PBS plus 0.1% tween-20). Specimens were rinsed twice with each for 5 min in TEA buffer (1% triethanolamine in PBST), transferred to freshly prepared 0.3% acetic anhydride in TEA buffer and incubated for 5 min. Additional acetic anhydride was added to yield a final concentration of 0.6% and specimens were further incubated for 5 min. After rinsing twice with each for 5 min in PBST, specimens were post-fixed in 4% paraformaldehyde for 2 h at room temperature and washed five times with each for 5 min with PBST. Specimens were pre-hybridized in hybridization buffer (50% formamide, 5 × SSC, 50 μg ml−1 heparin, 500 μg ml−1 yeast tRNA, 0.1% tween-20, pH 6.0) at 65 °C for 2 h. For hybridization, specimens were incubated in hybridization buffer containing 0.01–0.1 μg ml−1 of denatured RNA probe overnight at 65 ºC. Specimens were then washed twice in washing solution (50% formamide, 2×SSC, 0.1% tween-20; 30 min each), once in 2×SSCT (2×SSC and 0.1% tween-20; 15 min) and twice in 0.2×SSCT (0.2×SSC and 0.1% tween-20; 30 min each), all of which were conducted at 65 ºC. After washing with PBST for 5 min at room temperature, specimens were incubated in blocking buffer (PBST and 0.5% blocking reagent (Roche)) for 2 h at room temperature and then with 1/5,000 diluted alkaline phosphate-conjugated Fab fragments of a sheep anti-digoxigenin antibody (Roche) overnight at 4 ºC. After extensive washing with PBST, specimens were incubated with Nitro blue tetrazolium/5-Bromo-4-chloro-3-indolyl phosphate (NBT/BCIP) substrate solution to detect signals.
Phototransduction genes and network analysis
Key proteins involved in Homo and Drosophila phototransduction pathways37 were downloaded from the National Center for Biotechnology Information (NCBI) protein database, and homologous proteins were searched against the P. yessoensis genome using BLASTP with the E-value threshold of 1e−5. The obtained candidate genes were further checked by their annotations. Putative opsins were also checked by the presence of common motifs for opsins and GPCRs92, and only those containing all seven transmembrane domains and the lysine residue (296K) were kept for further analysis. Phylogenetic analysis of opsin genes was performed using the program MrBayes (v3.2.2)93 based on the LG+G+F amino-acid model. Differentially expressed (P < 0.05) genes were detected according to the procedure described in the edgeR package67. As scallop eyes are small and reside on the mantle, eye sampling might be contaminated by a minimal amount of mantle tissue. To be stringent, we considered those differentially expressed genes that were significantly up-regulated in the eye relative to mantle as candidate eye-related genes for further analysis. Gene ontology enrichment analysis of the differentially expressed genes was performed using the EnrichPipeline94. A signed coexpression gene network for 26 adult transcriptomic datasets was constructed using the R package WGCNA95, with the parameters of ‘sft = 9, minimum module size = 200 and cutting height = 0.99’. Modules with highly similar expression profiles were merged using the mergedColors function in WGCNA. The hubness of a gene in a given module was measured by its connection strength with other genes in the module, and was determined by intramodular connectivity (Kwithin)95. To identify the eye-related module, over-representation analysis of the eye-related genes (that is, up-regulated differentially expressed genes in the eye relative to mantle) was performed for each module using a hypergeometric test with P values adjusted by the Benjamini–Hochberg method96 for multiple-test correction.
Data availability
The scallop genome project has been deposited at the NCBI under the BioProject number PRJNA259405. The WGS, parental resequencing and 2b-RAD data were deposited in the Sequence Read Archive (SRA) database under the accession numbers SRS788513, SRX1034910 and SRX1027271, respectively. The short-read data of various developmental and adult transcriptomes were deposited in the SRA database under the accession numbers SRX1026991, SRX2238787 to SRX2238809, SRX2250256 to SRX2250259, SRX2251047, SRX2251049, SRX2251056, SRX2251057 and SRX2279546.
Additional information
How to cite this article: Wang, S. et al. Scallop genome provides insights into evolution of bilaterian karyotype and development. Nat. Ecol. Evol. 1, 0120 (2017).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Arthur, W. in Evolving Animals: The Story of our Kingdom Ch. 8, 77–84 (Cambridge Univ. Press, 2014).
Fedonkin, M. A. & Waggoner, B. M. The Late Precambrian fossil Kimberella is a mollusc-like bilaterian organism. Nature 388, 868–871 (1997).
Ivantsov, A. Y. A new reconstruction of Kimberella, a problematic vendian metazoan. Paleontol. J. 43, 601–611 (2009).
Putnam, N. H. et al. The amphioxus genome and the evolution of the chordate karyotype. Nature 453, 1064–1071 (2008).
Simakov, O. et al. Insights into bilaterian evolution from three spiralian genomes. Nature 493, 526–531 (2013).
Budd, G. E. The earliest fossil record of the animals and its significance. Phil. Trans. R. Soc. B. 363, 1425–1434 (2008).
Biscotti, M. A., Canapa, A., Forconi, M. & Barucca, M. Hox and ParaHox genes: a review on molluscs. Genesis 52, 935–945 (2014).
Serb, J. M. & Eernisse, D. J. Charting evolution’s trajectory: using molluscan eye diversity to understand parallel and convergent evolution. Evol. Educ. Outreach 1, 439–447 (2008).
Zhang, G. et al. The oyster genome reveals stress adaptation and complexity of shell formation. Nature 490, 49–54 (2012).
Albertin, C. B. et al. The octopus genome and the evolution of cephalopod neural and morphological novelties. Nature 524, 220–224 (2015).
Takeuchi, T. et al. Bivalve-specific gene expansion in the pearl oyster genome implications of adaptation to a sessile lifestyle. Zool. Lett. 2, 3 (2016).
Mergl, M., Massa, D. & Plauchut, B. Devonian and Carboniferous brachiopods and bivalves of the Djado sub-basin (north Niger, SW Libya). J. Czech. Geol. Soc. 46, 169–188 (2001).
Thiriot-Quievreux, C. Review of the literature on bivalve cytogenetics in the last ten years. Cah. Biol. Mar. 43, 17–26 (2002).
Huang, J. et al. The Jujube genome provides insights into genome evolution and the domestication of sweetness/acidity taste in fruit trees. PLoS Genet. 12, e1006433 (2016).
Hou, R. et al. Transcriptome sequencing and de novo analysis for Yesso scallop (Patinopecten yessoensis) using 454 GS FLX. PLoS ONE 6, e21560 (2011).
Wang, S., Meyer, E., McKay, J. K. & Matz, M. V. 2b-RAD: a simple and flexible method for genome-wide genotyping. Nat. Methods 9, 808–810 (2012).
Guo, X., He, Y., Zhang, L., Lelong, C. & Jouaux, A. Immune and stress responses in oysters with insights on adaptation. Fish Shellfish Immunol. 46, 107–119 (2015).
Kocot, K. M. et al. Phylogenomics reveals deep molluscan relationships. Nature 477, 452–456 (2011).
Luo, Y.-J. et al. The Lingula genome provides insights into brachiopod evolution and the origin of phosphate biomineralization. Nat. Commun. 6, 8301 (2015).
Wang, J., Li, L. & Zhang, G. A high-density SNP genetic linkage map and QTL analysis of growth-related traits in a hybrid family of oysters (Crassostrea gigas x Crassostrea angulata) using genotyping-by-sequencing. G3-Genes Genom. Genet. 6, 1417–1426 (2016).
Shi, Y. et al. High-density single nucleotide polymorphisms linkage and quantitative trait locus mapping of the pearl oyster, Pinctada fucata martensii Dunker. Aquaculture 434, 376–384 (2014).
Garcia-Fernandez, J. The genesis and evolution of homeobox gene clusters. Nat. Rev. Genet. 6, 881–892 (2005).
Hui, J. H. L. et al. Extensive chordate and annelid macrosynteny reveals ancestral homeobox gene organization. Mol. Biol. Evol. 29, 157–165 (2012).
Lemons, D. & McGinnis, W. Genomic evolution of Hox gene clusters. Science 313, 1918–1922 (2006).
Garstang, M. & Ferrier, D. E. K. Time is of the essence for ParaHox homeobox gene clustering. BMC Biol. 11, 72 (2013).
Duboule, D. The rise and fall of Hox gene clusters. Development 134, 2549–2560 (2007).
Ferrier, D. E. K. & Minguillon, C. Evolution of the Hox/ParaHox gene clusters. Int. J. Dev. Biol. 47, 605–611 (2003).
Tschopp, P. & Duboule, D. A regulatory ‘landscape effect’ over the HoxD cluster. Dev. Biol. 351, 288–296 (2011).
Kulakova, M. et al. Hox gene expression in larval development of the polychaetes Nereis virens and Platynereis dumerilii (Annelida, Lophotrochozoa). Dev. Genes Evol. 217, 39–54 (2007).
Sun, X. Q. et al. Hox genes and their expression pattern in early development of Litopenaeus vannamei . Period. Ocean Univ. China 45, 52–62 (2015).
Ikuta, T., Yoshida, N., Satoh, N. & Saiga, H. Ciona intestinalis Hox gene cluster: its dispersed structure and residual colinear expression in development. Proc. Natl Acad. Sci. USA 101, 15118–15123 (2004).
Hejnol, A. & Martindale, M. Q. Coordinated spatial and temporal expression of Hox genes during embryogenesis in the acoel Convolutriloba longifissura . BMC Biol. 7, 65 (2009).
Cannon, J. T. et al. Xenacoelomorpha is the sister group to Nephrozoa. Nature 530, 89–93 (2016).
MacLean J. A. II, et al. Rhox: a new homeobox gene cluster. Cell 120, 369–382 (2005).
Frobius, A. C., Matus, D. Q. & Seaver, E. C. Genomic organization and expression demonstrate spatial and temporal Hox gene colinearity in the Lophotrochozoan Capitella sp. I. PLoS ONE 3, e4004 (2008).
Wilkens, L. A. in Scallops: Biology, Ecology and Aquaculture (eds Shumway, S. E. & Parsons, G. J. ) Ch. 5, 317–356 (Elsevier, 2006).
Yau, K. W. & Hardie, R. C. Phototransduction motifs and variations. Cell 139, 246–264 (2009).
Fernald, R. D. Casting a genetic light on the evolution of eyes. Science 313, 1914–1918 (2006).
Gehring, W. J. & Ikeo, K. Pax6: mastering eye morphogenesis and eye evolution. Trends Genet. 15, 371–377 (1999).
Whoriskey, K., Whoriskey, S. & Whoriskey, F. G. Systematic differences in eye numbers between the left and right valves of the sea scallop Placopecten magellanicus: an evolutionary response for visualizing the water column? J. Shellfish Res. 33, 337–341 (2014).
Backfisch, B. et al. Stable transgenesis in the marine annelid Platynereis dumerilii sheds new light on photoreceptor evolution. Proc. Natl Acad. Sci. USA 110, 193–198 (2013).
Kozmik, Z. The role of Pax genes in eye evolution. Brain Res. Bull. 75, 335–339 (2008).
Glardon, S., Holland, L. Z., Gehring, W. J. & Holland, N. D. Isolation and developmental expression of the amphioxus Pax-6 gene (AmphiPax-6): insights into eye and photoreceptor evolution. Development 125, 2701–2710 (1998).
Pineda, D. et al. The genetic network of prototypic planarian eye regeneration is Pax6 independent. Development 129, 1423–1434 (2002).
Blackburn, D. C. et al. Isolation and expression of Pax6 and atonal homologues in the American horseshoe crab, Limulus polyphemus . Dev. Dynam. 237, 2209–2219 (2008).
Cook, C. E., Jiménez, E., Akam, M. & Saló, E. The Hox gene complement of acoel flatworms, a basal bilaterian clade. Evol. Dev. 6, 154–163 (2004).
Quiring, R., Walldorf, U., Kloter, U. & Gehring, W. J. Homology of the eyeless gene of Drosophila to the small eye gene in mice and aniridia in humans. Science 265, 785–789 (1994).
Tomarev, S. et al. Squid Pax-6 and eye development. Proc. Natl Acad. Sci. USA 94, 2421–2426 (1997).
Arendt, D., Tessmar, K., de Campos-Baptista, M. M., Dorresteijn, A. & Wittbrodt, J. Development of pigment-cup eyes in the polychaete Platynereis dumerilii and evolutionary conservation of larval eyes in Bilateria. Development 129, 1143–1154 (2002).
Vopalensky, P. et al. Molecular analysis of the amphioxus frontal eye unravels the evolutionary origin of the retina and pigment cells of the vertebrate eye. Proc. Natl Acad. Sci. USA 109, 15383–15388 (2012).
Sambrook, J., Fritsch, E. F. & Maniatis, T. Molecular Cloning: A Laboratory Manual (Cold Spring Harbor Lab Press, 1989).
Van Nieuwerburgh, F. et al. Illumina mate-paired DNA sequencing-library preparation using Cre-Lox recombination. Nucleic Acids Res. 40, e24 (2012).
Zhang, L. et al. Fosmid library construction and initial analysis of end sequences in Zhikong scallop (Chlamys farreri). Mar. Biotechnol. 9, 606–612 (2007).
Meng, Q. et al. Growth and reproductive performance of triploid Yesso scallops (Patinopecten yessoensis) induced by hypotonic shock. J. Shellfish Res. 31, 1113–1122 (2012).
Dolezel, J., Bartos, H., Voglmayr, J. & Greilhuber, J. Nuclear DNA content and genome size of trout and human. Cytometry 51, 127–128 (2003).
Harris, R. S. Improved Pairwise Alignment of Genomic DNA PhD thesis, Pennsylvania State Univ. (2007).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Kent, W. J. BLAT - the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Fu, X. et al. RADtyping: an integrated package for accurate de novo codominant and dominant RAD genotyping in mapping populations. PLoS ONE 8, e79960 (2013).
Stam, P. Construction of integrated genetic linkage maps by means of a new computer package: JoinMap. Plant J. 3, 739–744 (1993).
Wu, Y., Close, T. J. & Lonardi, S. Accurate construction of consensus genetic maps via integer linear programming. IEEE ACM Trans. Comput. Biol. Bioinform. 8, 381–394 (2011).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Wang, R. & Wang, Z. Science of Marine Shellfish Culture (Ocean Univ. China Press, 2008).
Hu, X. L. et al. Cloning and characterization of tryptophan 2,3-dioxygenase gene of Zhikong scallop Chlamys farreri (Jones and Preston 1904). Aquac. Res. 37, 1187–1194 (2006).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-Seq data. Genome Biol. 11, R25 (2010).
Jurka, J. Repbase Update: a database and an electronic journal of repetitive elements. Trends Genet. 9, 418–420 (2000).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Xu, P. et al. Genome sequence and genetic diversity of the common carp, Cyprinus carpio. Nat. Genet. 46, 1212–1219 (2014).
Chen S. et al. Whole-genome sequence of a flatfish provides insights into ZW sex chromosome evolution and adaptation to a benthic lifestyle. Nat. Genet. 46, 253–260 (2014).
Stanke, M., Schöffmann, O., Morgenstern, B. & Waack, S. Gene prediction in eukaryotes with a generalized hidden Markov model that uses hints from external sources. BMC Bioinform. 7, 62 (2006).
Majoros, W. H., Pertea, M. & Salzberg, S. L. TigrScan and GlimmerHMM: two open-source ab initio eukaryotic gene-finders. Bioinformatics 20, 2878–2879 (2004).
Korf, I. Gene finding in novel genomes. BMC Bioinform. 5, 59 (2004).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Birney, E., Clamp, M. & Durbin, R. Genewise and genomewise. Genome Res. 14, 988–995 (2004).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using evidence modeler and the program to assemble spliced alignments. Genome Biol. 9, R7 (2008).
Mulder, N. & Apweiler, R. InterPro and InterProScan: tools for protein sequence classification and comparison. Methods Mol. Biol. 396, 59–70 (2007).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Li, L., Stoeckert, C. J. & Roos, D. S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003).
Kocot, K. M. et al. Phylogenomics of Lophotrochozoa with consideration of systematic error. Syst. Biol. 66, 256–282 (2017).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Talavera, G. & Castresana, J. Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst. Biol. 56, 564–577 (2007).
Abascal, F., Zardoya, R. & Posada D. ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21, 2104–2105 (2005).
Stamatakis, A., Ludwig, T. & Meier, H. RAxML-III: a fast program for maximum likelihood-based inference of large phylogenetic trees. Bioinformatics 21, 456–463 (2005).
Yang Z. PAML4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
De Hoon, M. J. et al. Open source clustering software. Bioinformatics 20, 1453–1454 (2004).
Zhong, Y. F., Butts, T. & Holland, P. W. HomeoDB: a database of homeobox gene diversity. Evol. Dev. 10, 516–518 (2008).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Holland, L. Z. et al. The amphioxus genome illuminates vertebrate origins and cephalochordate biology. Genome Res. 18, 1100–1111 (2008).
Huelsenbeck, J. P. & Ronquist, F. MrBayes: Bayesian inference of phylogenetic trees. Bioinformatics 17, 754–755 (2001).
Chen, S. et al. De novo analysis of transcriptome dynamics in the migratory locust during the development of phase traits. PLoS ONE 5, e15633 (2010).
Langfelder, P. & Steve, H. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 9, 559 (2008).
Benjamini, Y. & Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Acknowledgements
We thank P. Holland for helpful comments on the earlier version of the manuscript. We thank X. Zhang, X. Yu, X. Wang, L. Tao, H. Ruan, H. Zhu, J. Wei and J. Lv for assistance with data analysis. We acknowledge grant support from the National Natural Science Foundation of China (31130054, 31322055, 31272656 and 31630081), the National High Technology Research and Development Program of China (863 program; 2012AA92204 and 2012AA10A405), the Taishan Scholar Project Fund of Shandong Province of China, the Natural Science Foundation for Distinguished Young Scholars of Shandong Province of China (JQ201308) and the AoShan Talents Program of Qingdao National Laboratory for Marine Science and Technology (2015ASTP-ES02). X.G. acknowledges support from Taishan Oversea Scholar Program of Shandong and USDA-NIFA/NJAES Project 1004475/NJ32920. We thank Dalian Zhangzidao Group for financial support, as well as providing scallop materials and facilities. We thank G. Jekely for providing access to the Platynereis transcriptome dataset.
Author information
Authors and Affiliations
Contributions
Z.B. conceived the study and initiated the scallop genome project. Z.B. and S.W. designed major scientific objectives and perceived the ‘STC’ phenomenon of Hox expression. S.W., Z.B., D.C. and X.G. directed final data analyses and interpretation. Rq.L. and J.Z. developed the strategy for assembly of highly heterozygous genomes. Rq.L., J.Z., Z.J., Y.H. and Jian L. coordinated genome sequencing, assembly, annotation and comparative genomics. S.W., Wq.J., Z.B. and W.L. coordinated parent resequencing, transcriptomes sequencing and assembly, analyses of homeobox genes and eye-related genes. R.H., Wq.J., D.W., Yc. L. and C.M. conducted DNA extraction and sequencing library preparation. Ll.Z., Y.S. and J.W. conducted mRNA extraction and sequencing library preparation. J.Z., Ji L., X.S. and Wk.J. performed genome assembly and functional annotation. Wq.J., J.D., X.X., Yp.L. and J.M. participated in assembly evaluation. Xl.H., Xue L., Wq.J., Jia L., S.W. and Hd.L. participated in 2b-RAD genotyping and linkage map construction. Ji L., J.Z., X.S. and Y.L. conducted gene family analysis and comparative genomics. S.W., Y.S., Wq.J., Ll.Z., T.L., Y.M. and Rj.L. participated in transcriptome analyses. S.W., X.X. and Yl.L. participated in the analyses of scallop homeobox genes. P.H., B.D. and Z.L. participated in the in situ hybridization experiments. S.W., Y.S. and Ll.Z. participated in the analyses of scallop eye-related genes. Xl.H. and X.N. participated in eye-related experiments and C.Z. provided technical support. Jun L. and Zu.Z. provided scallop materials and facilities for larval and adult rearing. R.H., L.Z., Q.X., H.G., Q.F. and Y.K. participated in scallop culture and sample collection. R.H., H.D., C.M., X.F. and Q.Y. participated in fosmid library construction. Xt.H., H.L. and Xuan L. participated in the cytogenetic analysis of scallop chromosomes. Y.W. participated in construction of genome databases. R.W. and Zh.Z. participated in discussions and provided suggestions for manuscript improvement. S.W., J.Z., X.G., D.C. and Z.B. did most of the writing with input from other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Text; Supplementary Figures 1–31; Supplementary Tables 1–18,21,22,24,26,27 (PDF 7672 kb)
Supplementary Table 19
Supplementary Table 19 (XLSX 64 kb)
Supplementary Tables 20 and 23
Supplementary Tables 20 and 23 (XLSX 207 kb)
Supplementary Table 25
Supplementary Table 25 (XLSX 9495 kb)
Rights and permissions
This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wang, S., Zhang, J., Jiao, W. et al. Scallop genome provides insights into evolution of bilaterian karyotype and development. Nat Ecol Evol 1, 0120 (2017). https://doi.org/10.1038/s41559-017-0120
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41559-017-0120
This article is cited by
-
Chromosome-level genome assembly of Oncomelania hupensis: the intermediate snail host of Schistosoma japonicum
Infectious Diseases of Poverty (2024)
-
A chromosome-level genome for the nudibranch gastropod Berghia stephanieae helps parse clade-specific gene expression in novel and conserved phenotypes
BMC Biology (2024)
-
Genome-wide investigation of the TGF-β superfamily in scallops
BMC Genomics (2024)
-
Pan-evolutionary and regulatory genome architecture delineated by an integrated macro- and microsynteny approach
Nature Protocols (2024)
-
Chromosome-level genome assembly and annotation of rare and endangered tropical bivalve, Tridacna crocea
Scientific Data (2024)