Abstract
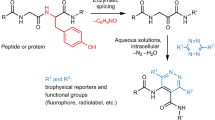
Electrochemistry has recently emerged as a powerful approach in small-molecule synthesis owing to its numerous attractive features, including precise control over the fundamental reaction parameters, mild reaction conditions and innate scalability. Even though these advantages also make it an attractive strategy for chemoselective modification of complex biomolecules such as proteins, such applications remain poorly developed. Here we report an electrochemically promoted coupling reaction between 5-hydroxytryptophan (5HTP) and simple aromatic amines—electrochemical labelling of hydroxyindoles with chemoselectivity (eCLIC)—that enables site-specific labelling of full-length proteins under mild conditions. Using genetic code expansion technology, the 5HTP residue can be incorporated into predefined sites of a recombinant protein expressed in either prokaryotic or eukaryotic hosts for subsequent eCLIC labelling. We used the eCLIC reaction to site-specifically label various recombinant proteins, including a full-length human antibody. Furthermore, we show that eCLIC is compatible with strain-promoted alkyne-azide and alkene-tetrazine click reactions, enabling site-specific modification of proteins at two different sites with distinct labels.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Data supporting the findings of this study are available within the paper and its Supplementary Information. Crystallographic data for the structure reported in this Article have been deposited at the Cambridge Crystallographic Data Centre, under deposition no. CCDC 2179454 (19). Copies of the data can be obtained free of charge via https://www.ccdc.cam.ac.uk/structures/Search?Ccdcid=2179454. Source data are provided with this paper.
References
Faraday, M. Siebente reihe von experimental‐Untersuchungen über Elektricität. Ann. Phys. 109, 481–520 (1834).
Kolbe, H. Beobachtungen über die oxydirende Wirkung des Sauerstoffs, wenn derselbe mit Hülfe einer elektrischen Säule entwickelt wird. J. Prakt. Chem. 41, 137–139 (1847).
Horn, E. J., Rosen, B. R. & Baran, P. S. Synthetic organic electrochemistry: an enabling and innately sustainable method. ACS Central Sci. 2, 302–308 (2016).
Yan, M., Kawamata, Y. & Baran, P. S. Synthetic organic electrochemical methods since 2000: on the verge of a renaissance. Chem. Rev. 117, 13230–13319 (2017).
Little, R. D. A perspective on organic electrochemistry. J. Org. Chem. 85, 13375–13390 (2020).
Zhu, C., Ang, N. W. J., Meyer, T. H., Qiu, Y. & Ackermann, L. Organic electrochemistry: molecular syntheses with potential. ACS Central Sci. 7, 415–431 (2021).
Spicer, C. D. & Davis, B. G. Selective chemical protein modification. Nat. Commun. 5, 4740 (2014).
Stephanopoulos, N. & Francis, M. B. Choosing an effective protein bioconjugation strategy. Nat. Chem. Biol. 7, 876–884 (2011).
Boutureira, O. & Bernardes, Ga. J. Advances in chemical protein modification. Chem. Rev. 115, 2174–2195 (2015).
deGruyter, J. N., Malins, L. R. & Baran, P. S. Residue-specific peptide modification: a chemist’s guide. Biochemistry 56, 3863–3873 (2017).
Lang, K. & Chin, J. W. Bioorthogonal reactions for labeling proteins. ACS Chem. Biol. 9, 16–20 (2014).
McKay, C. S. & Finn, M. G. Click chemistry in complex mixtures: bioorthogonal bioconjugation. Chem. Biol. 21, 1075–1101 (2014).
Shih, H. W., Kamber, D. N. & Prescher, J. A. Building better bioorthogonal reactions. Curr. Opin. Chem. Biol. 21, 103–111 (2014).
Sletten, E. M. & Bertozzi, C. R. Bioorthogonal chemistry: fishing for selectivity in a sea of functionality. Angew. Chem. Int. Ed. 48, 6974–6998 (2009).
Kim, C. H., Axup, J. Y. & Schultz, P. G. Protein conjugation with genetically encoded unnatural amino acids. Curr. Opin. Chem. Biol. 17, 412–419 (2013).
Scinto, S. L. et al. Bioorthogonal chemistry. Nat. Rev. Methods Primers 1, 30 (2021).
Hoyt, E. A., Cal, P. M., Oliveira, B. L. & Bernardes, G. J. Contemporary approaches to site-selective protein modification. Nat. Rev. Chem. 3, 147–171 (2019).
Walsh, S. J. et al. Site-selective modification strategies in antibody-drug conjugates. Chem. Soc. Rev. 50, 1305–1353 (2021).
Bloom, S. et al. Decarboxylative alkylation for site-selective bioconjugation of native proteins via oxidation potentials. Nat. Chem. 10, 205–211 (2018).
Lin, S. et al. Redox-based reagents for chemoselective methionine bioconjugation. Science 355, 597–602 (2017).
Patterson, D. M. & Prescher, J. A. Orthogonal bioorthogonal chemistries. Curr. Opin. Chem. Biol. 28, 141–149 (2015).
Italia, J. S. et al. Mutually orthogonal nonsense-suppression systems and conjugation chemistries for precise protein labeling at up to three distinct sites. J. Am. Chem. Sci. 141, 6204–6212 (2019).
Bednar, R. M., Karplus, P. A. & Mehl, R. A. Site-specific dual encoding and labeling of proteins via genetic code expansion. Cell Chem. Biol. 30, 343–361 (2023).
Osgood, A. O. et al. An efficient opal-suppressor tryptophanyl pair creates new routes for simultaneously incorporating up to three distinct noncanonical amino acids into proteins in mammalian cells. Angew. Chem. Int. Ed. 62, e202219269 (2023).
Mackay, A. S., Payne, R. J. & Malins, L. R. Electrochemistry for the chemoselective modification of peptides and proteins. J. Am. Chem. Soc. 144, 23–41 (2022).
Song, C. et al. Electrochemical oxidation induced selective tyrosine bioconjugation for the modification of biomolecules. Chem. Sci. 10, 7982–7987 (2019).
Alvarez-Dorta, D. et al. Electrochemically promoted tyrosine-click-chemistry for protein labeling. J. Am. Chem. Sci. 140, 17120–17126 (2018).
Sato, S. et al. Site-selective protein chemical modification of exposed tyrosine residues using tyrosine click reaction. Bioconjug. Chem. 31, 1417–1424 (2020).
Depienne, S. et al. Luminol anchors improve the electrochemical-tyrosine-click labelling of proteins. Chem. Sci. 12, 15374–15381 (2021).
Seki, Y. et al. Transition metal-free tryptophan-selective bioconjugation of proteins. J. Am. Chem. Sci. 138, 10798–10801 (2016).
Toyama, E. et al. Electrochemical tryptophan-selective bioconjugation. Preprint at https://chemrxiv.org/engage/chemrxiv/article-details/60c740a4567dfe14a1ec3c1a (2019).
Kendall, G. et al. Specific electrochemical nitration of horse heart myoglobin. Arch. Biochem. Biophys. 392, 169–179 (2001).
Iniesta, J. et al. Specific electrochemical iodination of horse heart myoglobin at tyrosine 103 as determined by Fourier transform ion cyclotron resonance mass spectrometry. Arch. Biochem. Biophys. 474, 1–7 (2008).
Baran, P. S., Guerrero, C. A. & Corey, E. J. The first method for protection−deprotection of the indole 2,3-π bond. Org. Lett. 5, 1999–2001 (2003).
Decoene, K. et al. Protein conjugation with triazolinediones: switching from a general tyrosine-selective labeling method to a highly specific tryptophan bioconjugation strategy. Preprint at https://chemrxiv.org/engage/chemrxiv/article-details/60c754d0842e650404db4222 (2021).
Italia, J. S. et al. An orthogonalized platform for genetic code expansion in both bacteria and eukaryotes. Nat. Chem. Biol. 13, 446–450 (2017).
White, A. M., Palombi, I. R. & Malins, L. R. Umpolung strategies for the functionalization of peptides and proteins. Chem. Sci. 13, 2809–2823 (2022).
Jewett, J. C. & Bertozzi, C. R. Cu-free click cycloaddition reactions in chemical biology. Chem. Soc. Rev. 39, 1272–1279 (2010).
Devaraj, N. K. & Weissleder, R. Biomedical applications of tetrazine cycloadditions. Acc. Chem. Res. 44, 816–827 (2011).
Chin, J. W. Expanding and reprogramming the genetic code. Nature 550, 53–60 (2017).
Italia, J. S. et al. Expanding the genetic code of mammalian cells. Biochem. Soc. Trans. 45, 555–562 (2017).
Young, D. D. & Schultz, P. G. Playing with the molecules of life. ACS Chem. Biol. 13, 854–870 (2018).
Ficaretta, E. D. et al. A robust platform for unnatural amino acid mutagenesis in E. coli using the bacterial tryptophanyl-tRNA synthetase/tRNA pair. J. Mol. Biol. 434, 167304 (2022).
Addy, P. S., Erickson, S. B., Italia, J. S. & Chatterjee, A. A chemoselective rapid azo-coupling reaction (CRACR) for unclickable bioconjugation. J. Am. Chem. Sci. 139, 11670–11673 (2017).
Addy, P. S., Italia, J. S. & Chatterjee, A. An oxidative bioconjugation strategy targeted to a genetically encoded 5-hydroxytryptophan. Chembiochem 19, 1375–1378 (2018).
Addy, P. S., Zheng, Y., Italia, J. S. & Chatterjee, A. A ‘quenchergenic’ chemoselective protein labeling strategy. Chembiochem 20, 1659–1663 (2019).
Singha Roy, S. J. et al. Photoredox-catalyzed labeling of hydroxyindoles with chemoselectivity (PhotoCLIC) for site-specific protein bioconjugation. Angew. Chem. Int. Ed. 62, e202300961 (2023).
Humphries, K. A., Wrona, M. Z. & Dryhurst, G. Electrochemical and enzymatic oxidation of 5-hydroxytryptophan. J. Electroanal. Chem. 346, 377–403 (1993).
Napolitano, A., d’Ischia, M. & Prota, G. A profile of the oxidation chemistry of 5-hydroxyindole under biomimetic conditions. Tetrahedron 44, 7265–7270 (1988).
Wrona, M. Z. & Dryhurst, G. Oxidation of serotonin by superoxide radical: implications to neurodegenerative brain disorders. Chem. Res. Toxicol. 11, 639–650 (1998).
Yamagishi, Y., Ashigai, H., Goto, Y., Murakami, H. & Suga, H. Ribosomal synthesis of cyclic peptides with a fluorogenic oxidative coupling reaction. Chembiochem 10, 1469–1472 (2009).
Ferguson, W. J. et al. Hydrogen ion buffers for biological research. Anal. Biochem. 104, 300–310 (1980).
Grady, J. K., Chasteen, N. D. & Harris, D. C. Radicals from ‘Good’s’ buffers. Anal. Biochem. 173, 111–115 (1988).
Nutting, J. E., Rafiee, M. & Stahl, S. S. Tetramethylpiperidine N-oxyl (TEMPO), phthalimide N-oxyl (PINO), and related N-oxyl species: electrochemical properties and their use in electrocatalytic reactions. Chem. Rev. 118, 4834–4885 (2018).
Sheikholeslami, F. et al. Isolation of a novel nanobody against HER-2/neu using phage displays technology. Lab. Med. 41, 69–76 (2010).
Drago, J. Z., Modi, S. & Chandarlapaty, S. Unlocking the potential of antibody–drug conjugates for cancer therapy. Nat. Rev. Clin. Oncol. 18, 327–344 (2021).
Beck, A., Goetsch, L., Dumontet, C. & Corvaïa, N. Strategies and challenges for the next generation of antibody-drug conjugates. Nat. Rev. Drug Discov. 16, 315–337 (2017).
Axup, J. Y. et al. Synthesis of site-specific antibody-drug conjugates using unnatural amino acids. Proc. Natl Acad. Sci. USA 109, 16101–16106 (2012).
VanBrunt, M. P. et al. Genetically encoded azide containing amino acid in mammalian cells enables site-specific antibody-drug conjugates using click cycloaddition chemistry. Bioconjug. Chem. 26, 2249–2260 (2015).
Agarwal, P. & Bertozzi, C. R. Site-specific antibody–drug conjugates: the nexus of bioorthogonal chemistry, protein engineering and drug development. Bioconjug. Chem. 26, 176–192 (2015).
Tian, F. et al. A general approach to site-specific antibody drug conjugates. Proc. Natl Acad. Sci. USA 111, 1766–1771 (2014).
Chen, Y. et al. Creation of bacterial cells with 5-hydroxytryptophan as a 21st amino acid building block. Chem 6, 2717–2727 (2020).
Roy, G. et al. Development of a high yielding expression platform for the introduction of non-natural amino acids in protein sequences. mAbs 12, 1684749 (2020).
Lang, K. et al. Genetically encoded norbornene directs site-specific cellular protein labelling via a rapid bioorthogonal reaction. Nat. Chem. 4, 298–304 (2012).
Zheng, Y., Addy, P. S., Mukherjee, R. & Chatterjee, A. Defining the current scope and limitations of dual noncanonical amino acid mutagenesis in mammalian cells. Chem. Sci. 8, 7211–7217 (2017).
Backliwal, G. et al. Rational vector design and multi-pathway modulation of HEK 293E cells yield recombinant antibody titers exceeding 1 g/l by transient transfection under serum-free conditions. Nucleic Acids Res. 36, e96 (2008).
Acknowledgements
This work was supported by the National Science Foundation (2128185 to A.C.) and by the NIH (R35GM134964 to E.W., R35GM136437 to A.C.). We thank M. Waegele (Boston College) for insightful discussions, B. Li (Director, XRD facility, B.C.) and T. Jayasundera (Director, NMR facilities, B.C.) for their assistance. The funding agencies had no role in study design.
Author information
Authors and Affiliations
Contributions
A.C., C.L. and S.J.S.R. designed the experiments and interpreted data. C.L. conducted bioconjugation experiments. S.J.S.R. generated the synthetic probes and performed structural characterizations. V.J.O. assisted with CV and earlier electrochemistry experiments. S.E.C. and E.W. performed trypsin digestion/MS and associated analysis. A.M. optimized the eCLIC modification of nanobody 5F7. D.J. assisted with flow cytometry analyses. E.D.F. assisted with bacterial expression of 5HTP-containing proteins. A.C., C.L. and S.J.S.R. prepared the manuscript. A.C. supervised the project.
Corresponding author
Ethics declarations
Competing interests
A patent application on the eCLIC technology reported here has been submitted. A.C. is a cofounder and senior advisor of BrickBio, Inc. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks Sébastien Gouin, Wen-Bin Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Information
Supplementary File 1
Crystal structure of NBoc–OMe–5HTP adduct with N,N-dimethylaniline
Source data
Source Data Fig. 5e
Raw data from MTT assay
Source Data Fig. 5c
Uncropped gels Fig 5c
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Loynd, C., Singha Roy, S.J., Ovalle, V.J. et al. Electrochemical labelling of hydroxyindoles with chemoselectivity for site-specific protein bioconjugation. Nat. Chem. 16, 389–397 (2024). https://doi.org/10.1038/s41557-023-01375-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-023-01375-y